Introduction
Sepsis-associated encephalopathy (SAE) is a common
complication of sepsis that is prevalent in hospitalized patients
(sepsis happens in ~2% of all hospitalized patients) (1). Furthermore, it has been estimated
that SAE affects 9-71% of those patients with severe sepsis
(2). The major manifestations of
SAE include emotional abnormalities, impaired memory and cognitive
impairment. Even after the acute phase of sepsis, these mental
manifestations persist, severely affecting the prognosis and
quality of life of patients (3).
To date, numerous biological processes have been reported to be
involved in the pathogenesis of SAE, including inflammatory
response, collapse of the blood-brain barrier (BBB), cerebral
ischemia and alterations in neurotransmitters. Although the
specific mechanism has remained to be established, uncontrolled
neuroinflammation is probably the leading cause of SAE (4).
During the early phase of sepsis, brain microglial
cells that function as peripheral macrophages are activated and
transformed to the M1 cell type (5,6).
Thereafter, A great number of inflammatory cytokines, including
interleukin-1 (IL-1), IL-6 and tumor necrosis factor-α (TNF-α), are
secreted into the brain and lead to uncontrolled neuroinflammation
(7). In addition, the activated
astrocytes are also capable of releasing multiple types of
inflammatory mediators, further facilitating the development of
neuroinflammation (8).
Overactivation of microglia and other immune cells may cause damage
to neuron and endothelial cells, and furthermore deteriorate the
BBB and enhance the release of reactive oxygen species, which
seriously disturb brain function (9). Recent studies have verified that the
suppression of microglial activation and central inflammatory
responses may partially reverse the cognitive impairment caused by
SAE (7,10). However, there is still a lack of
effective treatment for SAE in the clinic, probably due to the
detailed pathology of SAE having remained to be fully
elucidated.
Non-coding (nc) RNAs, including microRNA
(miRNA/miR), long ncRNA (lncRNA) and circular RNA (circRNA), have
pivotal roles in the regulation of cell proliferation (11), autophagy (12) and apoptosis (13), as well as inflammatory responses
(14), ischemia-reperfusion injury
(15) and tumorigenesis (16). Recent studies have indicated that
lncRNAs and circRNAs participate in the process of microglial
activation (17-19).
However, the orchestra of lncRNAs and circRNAs in microglial
activation has remained to be fully determined, particularly during
the early phase of SAE. In the present study, high-throughput
microarray techniques were used to profile the expression of mRNAs,
lncRNAs and circRNAs in BV-2 microglial cells after a short
stimulation with lipopolysaccharide (LPS). Through the analysis of
differentially expressed (DE) RNAs, promising novel signals
involving non-coding RNAs were provided in the present study to
provide clues for future investigation of microglial
activation.
Materials and methods
Cell lines
Murine BV-2 microglial cells were obtained from the
National Infrastructure of Cell Line Resource. The cells were
cultured in high-glucose Dulbecco's modified Eagle's medium (DMEM;
Thermo Fisher Scientific, Inc.) supplemented with 10% fetal bovine
serum (FBS; Biological Industries, Inc.), 1 mM pyruvate, 100 U/ml
penicillin and 100 µg/ml streptomycin in a humidified atmosphere
with 5% CO2 at 37˚C.
Cell treatments
BV-2 microglial cells were seeded in a 96-well plate
at a density of 50,000 cells or in a 6-well plate at a density of
500,000 cells in each well 24 h prior to the experiment. During the
experiment, the culture medium was changed to high-glucose DMEM
supplemented with 0.5% FBS, and then LPS was added into each well
(final concentration of 1.00 µg/ml) for incubation for 2, 4, 8, 16
or 24 h. In the control wells, the same volume of PBS was added and
the cells were cultured for 24 h. A Cell Counting Kit-8 (CCK-8;
BestBio) was used to detect the cell viability. CCK-8 stain (10 µl)
was added into the 100 µl culture medium and the cells were further
cultured for 4 h in the 37˚C incubator prior to the detection of
the optical density at the wavelength of 450 nm on a SpectraMax
multimode microplate (SpectraMax iD3; Molecular Devices).
The morphology of BV-2 cells with the different
incubation times (2, 4, 8, 16 or 24 h) of LPS or incubated with PBS
for 24 h in the 6-well plate was examined under an inverted
microscope (DMi8; Leica Microsystems) in an independent experiment
and the same cells in the 6-well plate were finally collected for
RNA extraction and the detection of the expression levels of TNF-α,
IL-6 and IL-1β. Furthermore, cells seeded in the 6-well plate were
treated with different concentrations of LPS (0, 0.01, 0.10 or 1.00
µg/ml) for 4 h. After washing with PBS, the total RNA in each well
was extracted by RNAiso Plus (Takara Bio, Inc.) for detecting the
mRNA levels of TNF-α, IL-6 and IL-1β.
For the microarray analysis, cells were seeded in a
35-mm dish at a density of 500,000 cells 24 h prior to the
experiment. BV-2 cells were stimulated with 1.00 µg/ml LPS or PBS
for 4 h. Finally, the cells were washed with PBS and collected for
the subsequent treatment.
RNA extraction and microarray
assay
RNAiso Plus (Takara Bio, Inc.) was used to extract
total RNA following the manufacturer's protocol. Total RNA was
quantified by a NanoDrop® 2000 (Thermo Fisher
Scientific, Inc.) and the RNA integrity was assessed using an
Agilent Bioanalyzer 2100 (Agilent Technologies, Inc.). The sample
labeling, microarray hybridization, washing and scanning were
performed based on the manufacturer's standard protocols (Agilent
Technologies, Inc.) by OE Biotech. In brief, total RNA was
reversely transcribed to double-stranded cDNA (The first strand of
cDNA complementary to the RNA was synthesized first and the second
strand of cDNA complementary to the first strand was subsequently
synthesized). Next, cRNA was synthesized using the double-stranded
cDNA as the template and labeled with Cyanine-3-cytidine
triphosphate (CTP). The labeled cRNAs were hybridized onto the
microarray. After washing, the arrays were scanned by the Agilent
Scanner G2505C (Agilent Technologies, Inc). Feature Extraction
software (version 10.7.1.1; Agilent Technologies, Inc.) was used to
analyze array images to obtain raw data. The raw data were
submitted to the Gene Expression Omnibus (GEO) database (dataset
no. GSE171696).
Bioinformatics analysis
DE genes were then identified by performing
Student's t-test to calculate the fold change (FC) and P-value. The
threshold set for upregulated and downregulated genes was FC
>2.0 and P<0.05.
Gene ontology (GO) and kyoto
encyclopedia of genes and genomes (KEGG) enrichment analysis
GO analysis of target genes was performed in the
categories of biological process (BP), cellular component (CC) and
molecular function (MF) (http://www.geneontology.org). KEGG pathway analysis
(www.genome.jp/kegg) was also applied to
analyze the key regulatory pathways involved in the activation of
BV-2 cells. The upregulated or downregulated RNAs were individually
analyzed via an online tool (https://cloud.oebiotech.cn/task/detail/array_enrichment/).
CeRNA network analysis
DE lncRNAs/circRNAs that had a significant positive
correlation (correlation coefficient >0.9) with DE mRNAs were
selected as the targets for the competing endogenous (ce)RNA
analysis. Target prediction of miRNAs to the aforementioned DE
lncRNAs, circRNAs and mRNAs was performed by using all known mouse
miRNAs in the database Mibase22. The ternary regulatory networks of
lncRNA-miRNA-mRNA and circRNA-miRNA-mRNA were established using the
predicted miRNAs as a bridge and the top 100 ceRNA network, which
contained the top 100 links (ranked by the P-value of each link) of
lncRNA-miRNA-mRNA or circRNA-miRNA-mRNA, was drawn by using the R
(version 3.5.1) software (20).
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from BV-2 cells using RNAiso
Plus. For each sample, 1 µg RNA was reverse-transcribed into cDNA
using the Vazyme HiScript III 1st Strand cDNA Synthesis Kit with
random hexamer (Vazyme Biotech Co. Ltd.) according to the
manufacturer's protocols. For qPCR, 5 µl reaction mixture composed
of 5 µl Vazyme SYBR qPCR Master Mix (Vazyme Biotech Co. Ltd.), 3 µl
1:9 diluted cDNA and 2 µl of 10 nmol/l primers were incubated in a
Bio-Rad 96-well PCR plate. The thermocycling conditions in the
RT-qPCR System (CFX Connect; Bio-Rad Laboratories, Inc.) was set to
95˚C for 5 min, followed by 40 cycles of 95˚C for 10 sec and 60˚C
for 30 sec. Data analysis was further performed using Bio-Rad CFX
Maestro software (Bio-Rad Laboratories, Inc.). The relative
expression levels of RNAs were calculated using the
2-ΔΔCq method with β-actin as a reference (21). The primer sequences of lncRNAs,
circRNAs and mRNAs were displayed in Tables SI and SII.
Statistical analysis
Values are expressed as the mean ± standard error of
the mean. An unpaired Student's t-test for parametric data and the
Mann-Whitney U-test for nonparametric data were utilized for
comparisons between two groups. GraphPad Prism 5 (GraphPad
Software, Inc.) was used for all statistical analyses. Pearson
correlation analysis was used to confirm the co-expression of
lncRNAs/circRNAs and mRNA with an online tool (https://cloud.oebiotech.cn/task/detail/correxpress/).
P<0.05 was considered to indicate statistical significance.
Results
Differential expression of mRNAs,
lncRNAs and circRNAs in LPS-treated BV-2 cells
Treatment with LPS (1 µg/ml) for 2-24 h had no
impact on the viability of BV-2 cells (Fig. S1A and B) and 4 h of treatment with LPS
concentration-dependently (0.01, 0.1 and 1 µg/ml) induced the
transcription of TNF-α, IL-6 and IL-1β (Fig. S1C). The expression levels of
TNF-α, IL-6 and IL-1β mRNAs changed after treatment with LPS for
different durations (Fig. S1D).
However, the transcription of these inflammatory factor genes was
already induced after 4 h of treatment. Therefore, the LPS
concentration of 1 µg/ml and the incubation time of 4 h was used in
the subsequent experiments.
The expression profiles of mRNAs, lncRNA and
circRNAs in BV-2 cells were analyzed using microarray technology. A
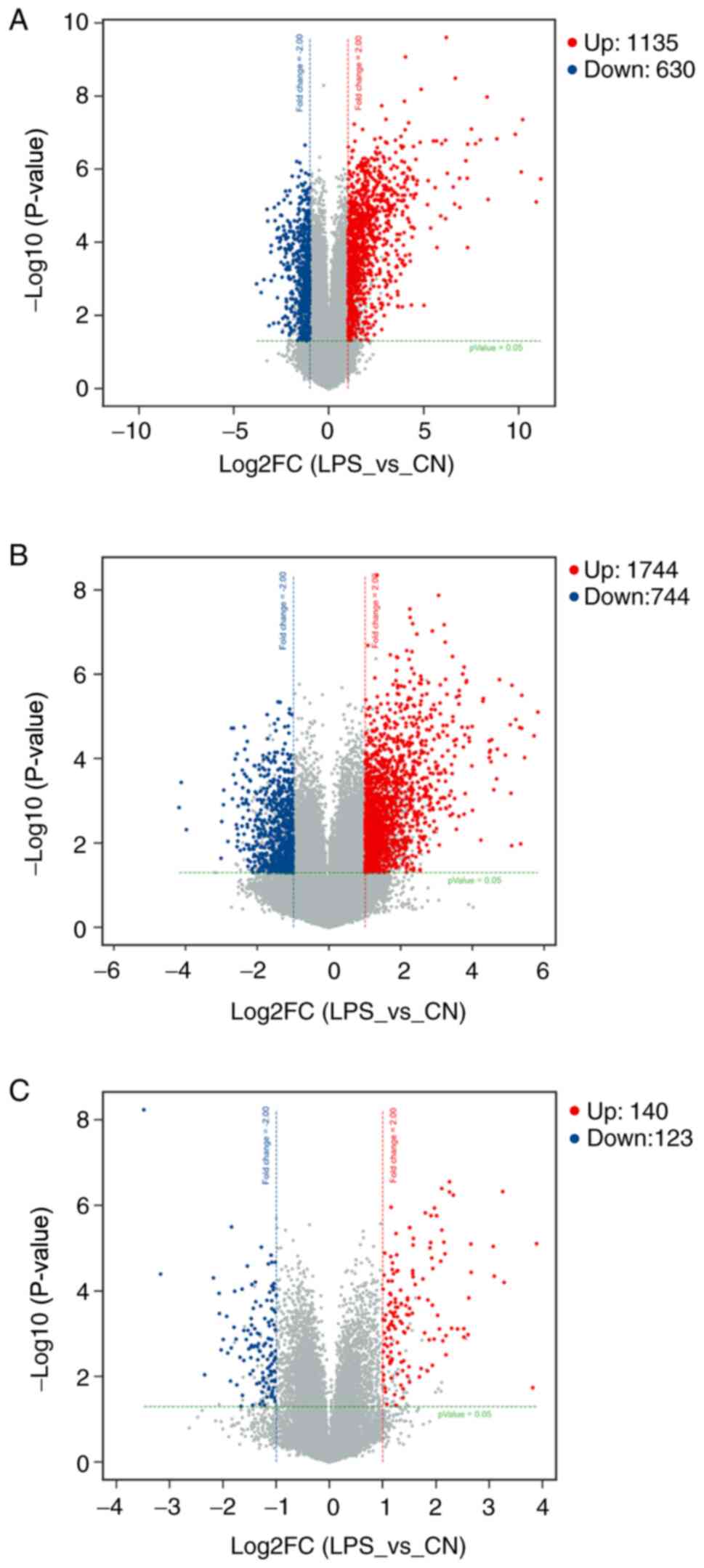
total of 1,135 mRNAs were identified to be upregulated and 630
mRNAs were downregulated. There were 2,488 DE lncRNAs, including
1,744 upregulated and 744 downregulated lncRNAs. Furthermore, 140
circRNAs were upregulated and 123 circRNAs were downregulated. The
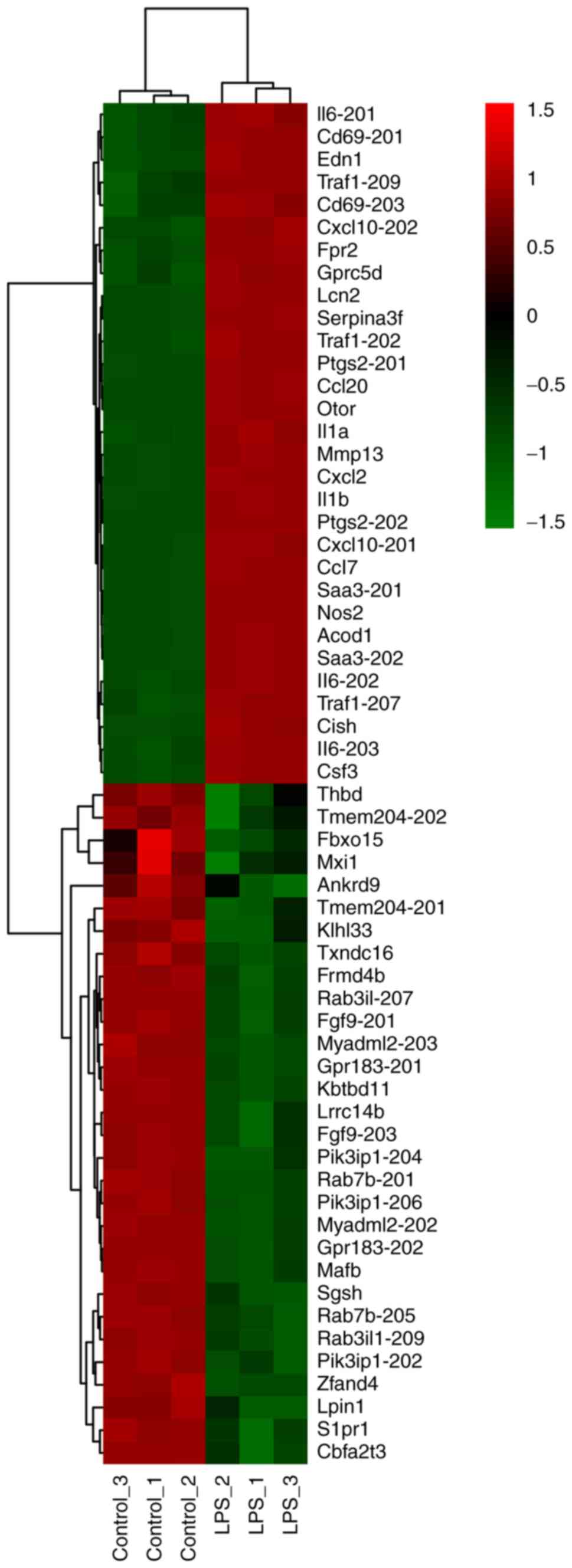
top 30 DE mRNAs ranked by FC, including both upregulated and
downregulated, were present as a heatmap (Fig. 1). Volcano plot analysis was used to
assess the variations in mRNA (Fig.
2A), lncRNA (Fig. 2B) and
circRNA (Fig. 2C) expression
profiles between the two groups.
GO and KEGG analysis of the DE
mRNAs
The results of the GO functional enrichment analysis
indicated that the upregulated DE mRNAs were involved in 585 BP, 54
CC and 112 MF terms. Furthermore, the downregulated mRNAs were
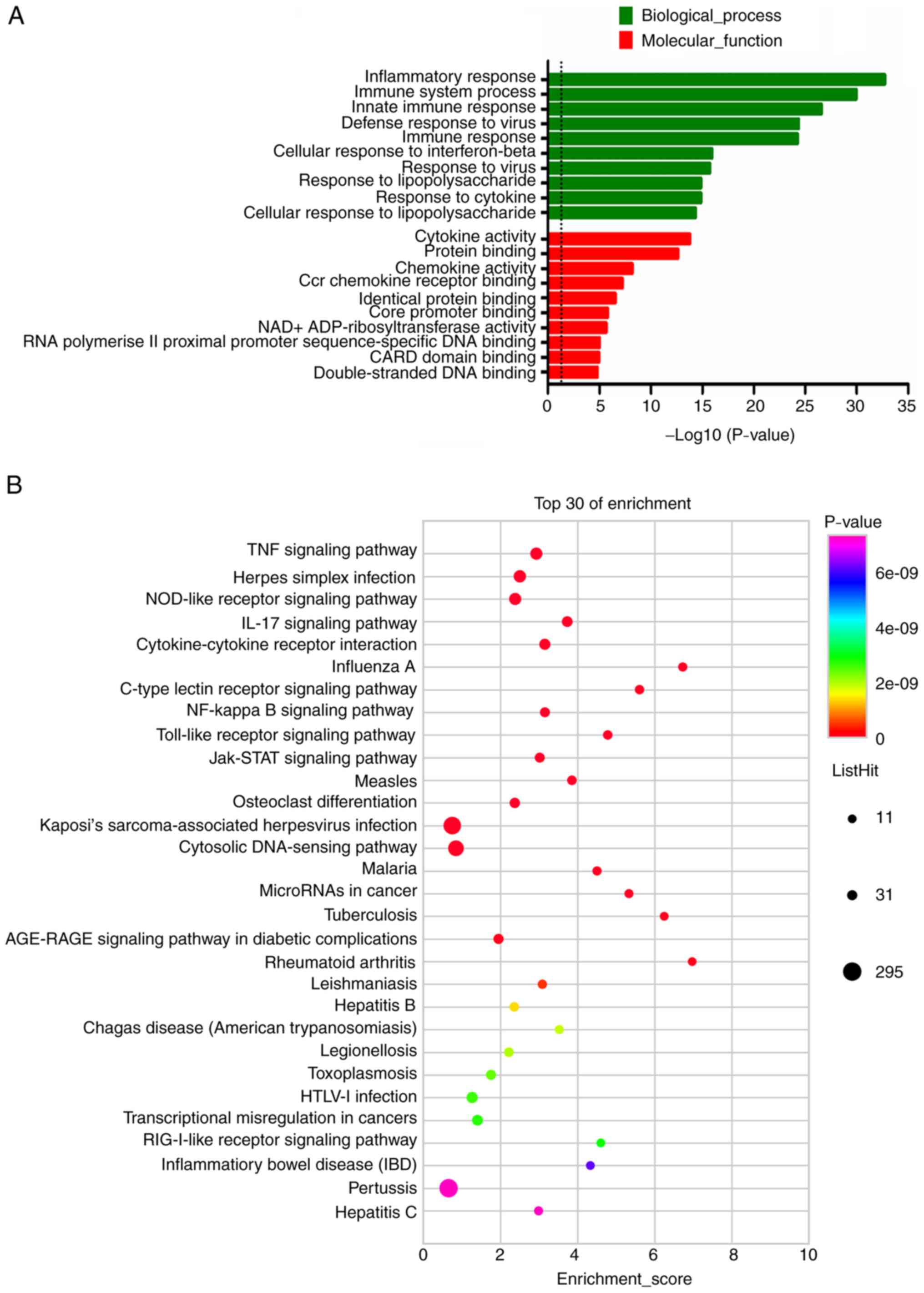
involved in 182 BP, 19 CC and 58 MF terms. The top 10 results with
the most significant P-values of the upregulated DE mRNAs in BP and
MF were presented in the bar blot (Fig. 3A). GO term analysis of upregulated
DE mRNAs revealed that the most enriched GO terms were cytokine
activity (MF) and inflammatory response (BP). The upregulated mRNAs
in BV-2 cells were most significantly enriched in the TNF signaling
pathway, Herpes simplex infection, NOD-like receptor signaling
pathway, IL-17 signaling pathway and cytokine-cytokine receptor
interaction (Fig. 3B). For the
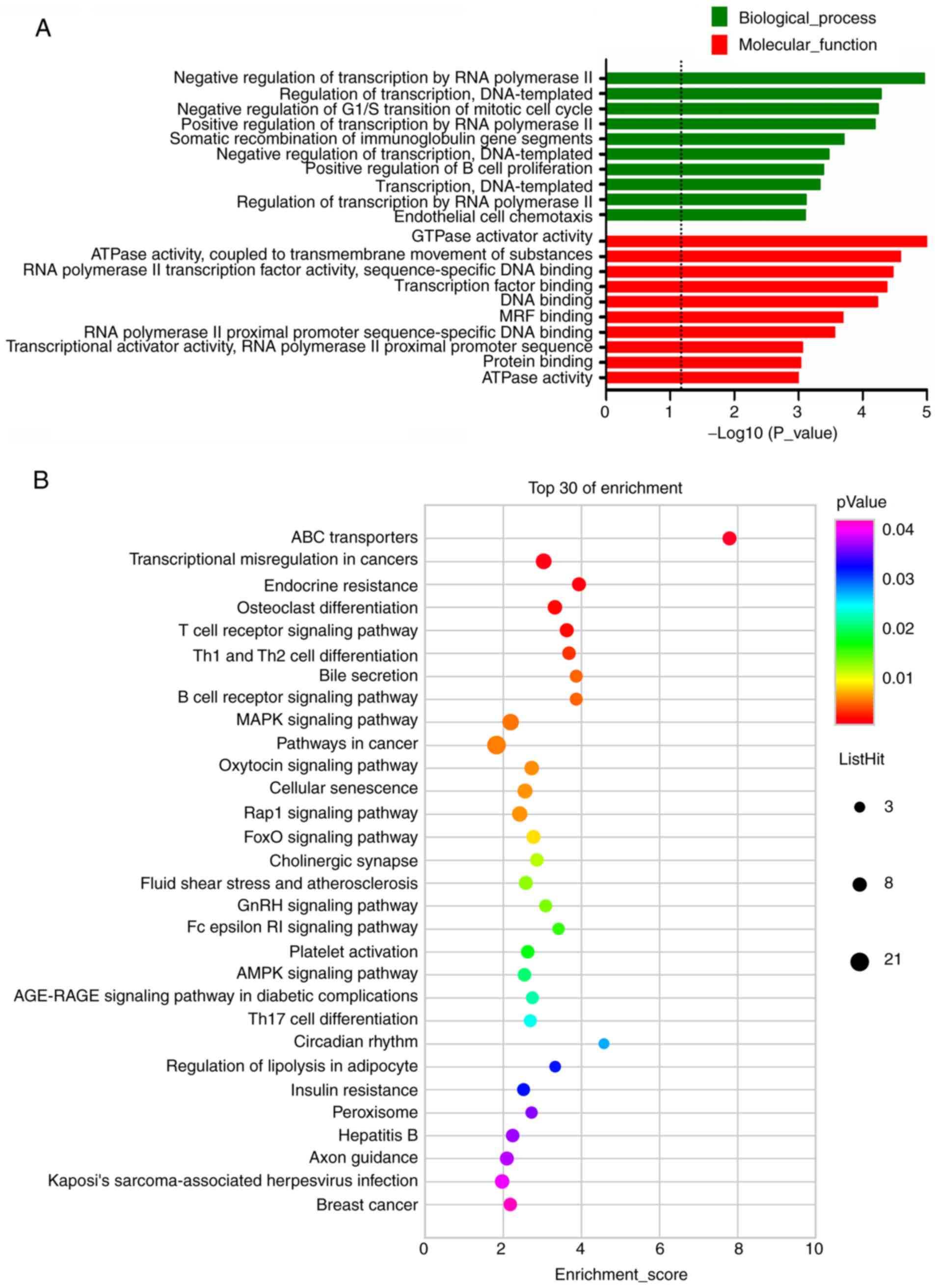
downregulated mRNAs, the most enriched GO terms were GTPase
activator activity (MF) and negative regulation of transcription by
RNA polymerase II (BP), which were shown in Fig. 4A. KEGG pathway analysis of DE mRNAs
determined 79 pathways for upregulated mRNAs and 33 pathways for
downregulated mRNAs. Furthermore, the pathways most significantly
enriched by the downregulated mRNAs were ABC transporters,
transcriptional misregulation in cancers, endocrine resistance,
osteoclast differentiation and T-cell receptor signaling pathway
(Fig. 4B). In addition, pathways
associated with lipid metabolism, including bile secretion,
regulation of lipolysis in adipocyte and insulin resistance, were
among the top 30 pathways enriched by downregulated mRNAs (Fig. 4B).
CeRNA network analysis
Co-expressed lncRNAs/circRNAs and mRNAs were
selected to predict the association with miRNAs. In total,
>300,000 lncRNA-miRNA-mRNA links were predicted. The top 100
links were chosen according to the P-value of their correlation and
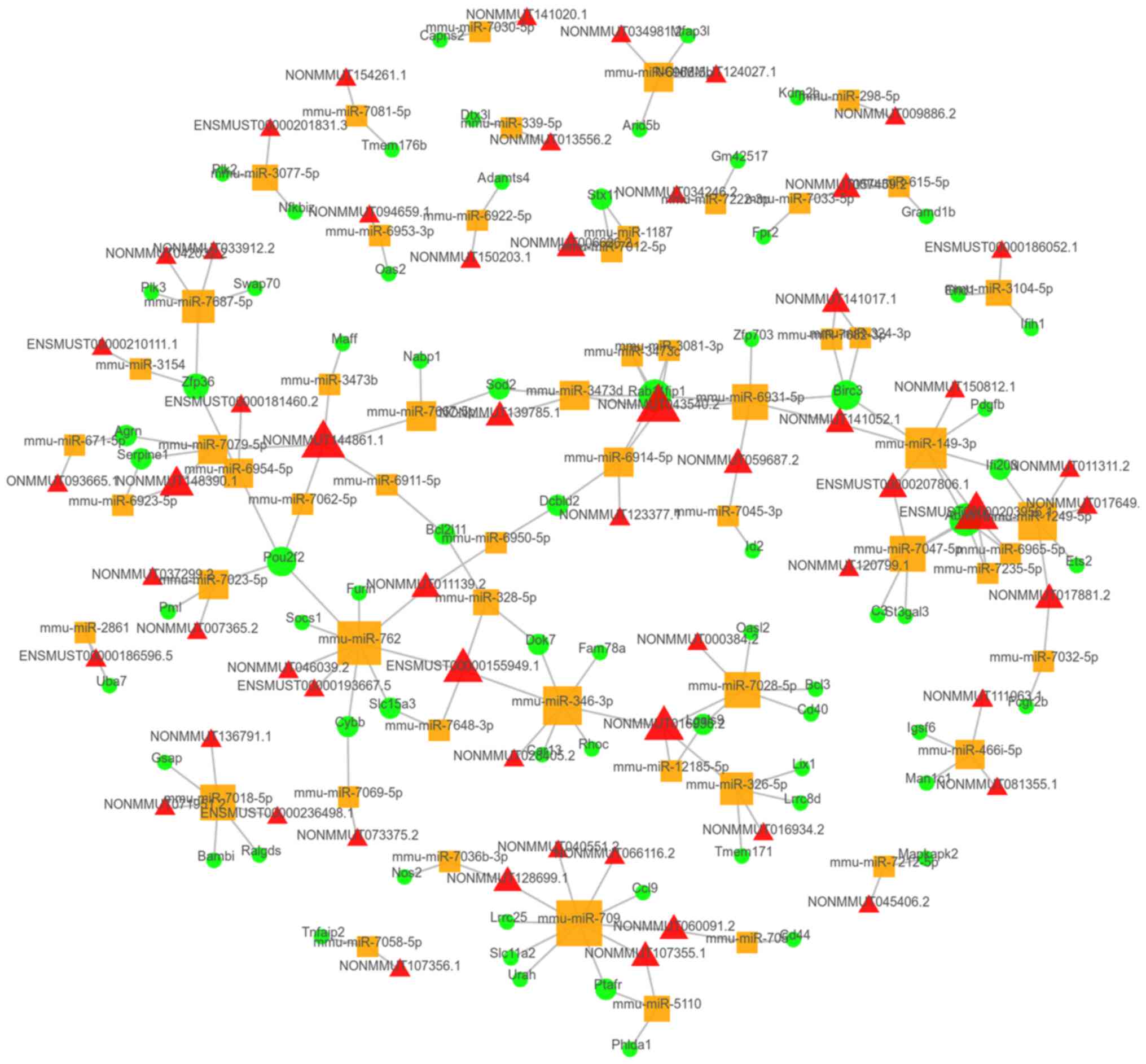
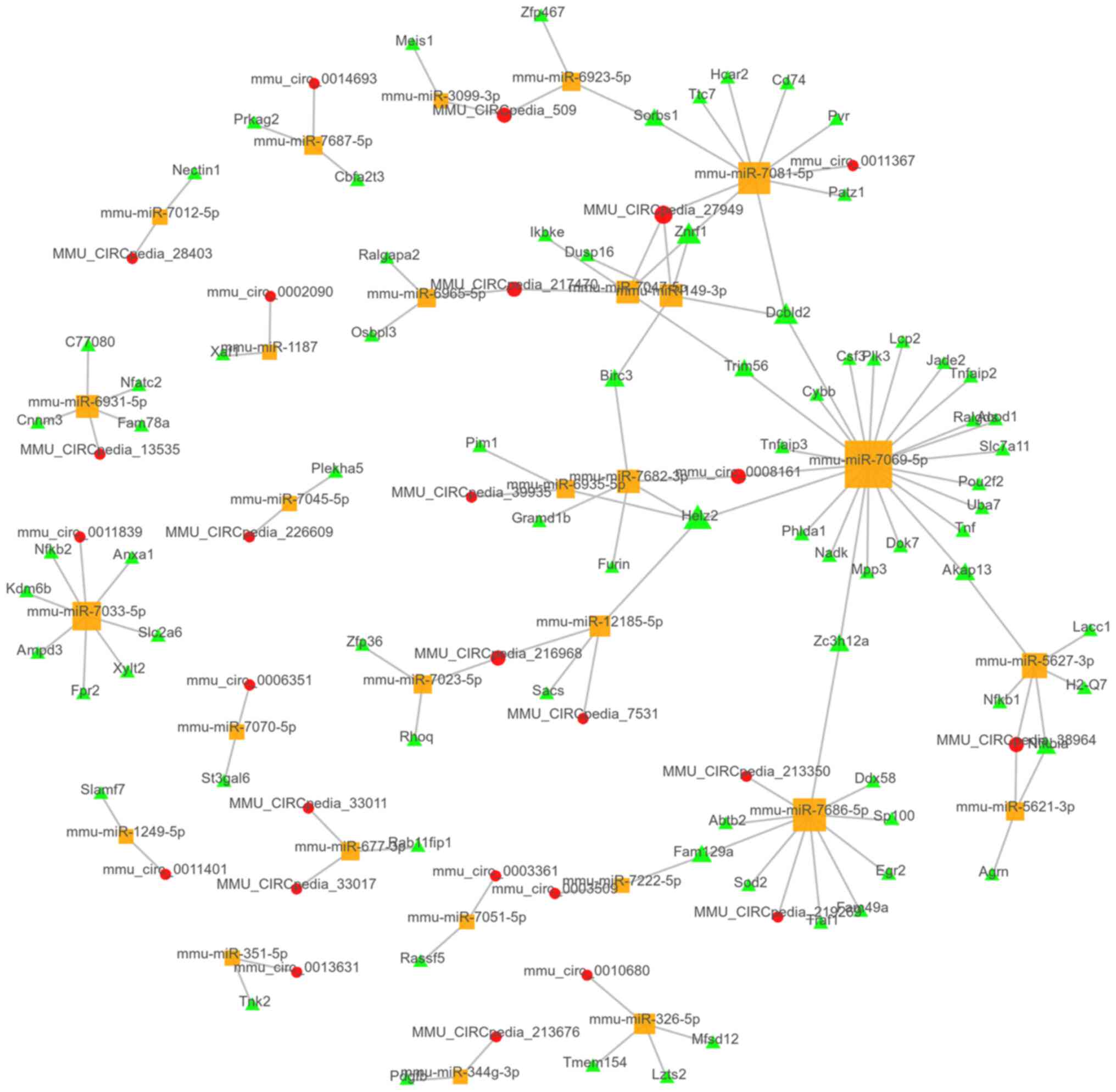
were present in the top 100 ceRNA network (Fig. 5). Furthermore, 4,053
circRNA-miRNA-mRNA links were predicted, with the top 100 ceRNA
network presented in Fig. 6.
Validation of DE mRNAs, lncRNAs and
circRNA
DE mRNAs, lncRNAs and circRNAs from the expression
profile with a high FC or in the hub of the ceRNA networks were
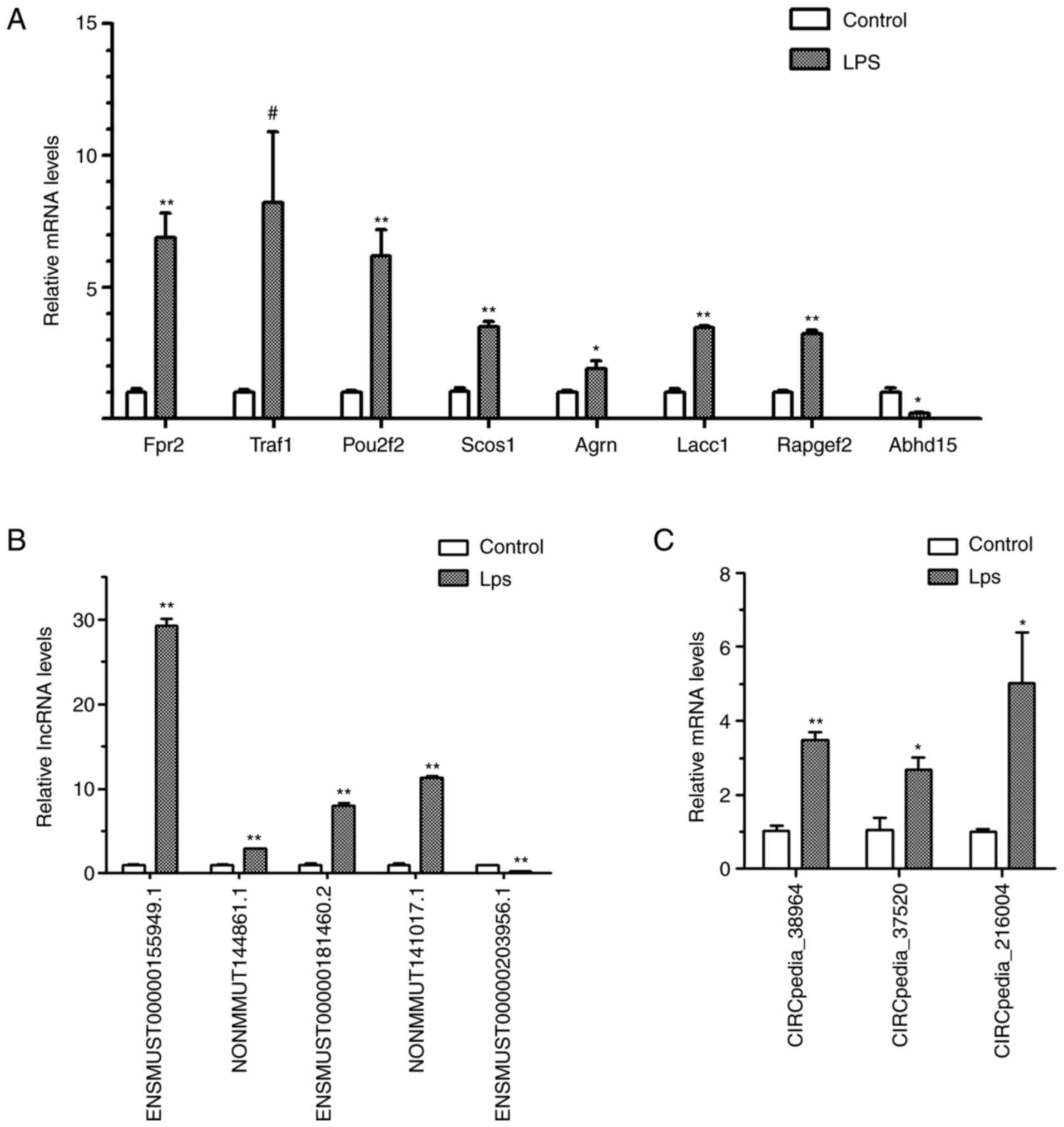
selected for validation (Fig. 7).
RT-qPCR validated that seven selected mRNAs, including TNF receptor
(TNFR)-associated factor 1 (Traf1), formyl peptide receptor 2
(Fpr2), POU class 2 homeobox 2 (Pou2f2), suppressor of cytokine
signaling 1 (Socs1), agrin (Agrn), laccase domain containing 1
(Lacc1) and Rap guanine nucleotide exchange factor 2 (Rapgef2),
were increased after LPS stimulation, and that the mRNA expression
of α/β-hydrolase domain containing protein 15 (Abhd15) was
decreased. Furthermore, four lncRNAs (ENSMUST00000155949.1,
NONMMUT144861.1, ENSMUST00000181460.2 and NONMMUT141017.1) and
three circRNAs (CIRCpedia_38964, CIRCpedia_37520 and
CIRCpedia_216004) were demonstrated to be upregulated and one
lncRNA (ENSMUST00000203956.1) was proven to be downregulated.
Discussion
Neuroinflammation is proposed to have a central role
in the pathogenesis of SAE, which increases the burden on the
health system and may cause major damage to patients' quality of
life (3,22). Microglial cells, which act as
residential macrophages, are activated during the pathological
process of neuroinflammation (5,6). The
modulation of microglial cell activation has become a promising
therapeutic direction in the treatment of SAE, as well as
neuropathic pain (10,23). The murine microglial cell line BV-2
is commonly used for study of neuroinflammation and microglial
activation (24,25). Henn et al (26) indicated that the cell line has a
high transcriptome homology with primary microglial cells and a
similar response to inflammatory challenges. Therefore, in the
present study, BV-2 cells stimulated with LPS were employed as the
model of microglial activation. Microarrays of mRNAs, lncRNAs and
circRNAs of stimulated or vehicle-treated BV-2 cells were performed
and the data were analyzed.
The results of the present study indicated that the
expression profiles of mRNAs and lncRNAs were acutely altered after
4 h of LPS exposure. A total of 1,135 mRNAs were determined to be
significantly upregulated and 630 mRNAs were downregulated. The top
upregulated mRNAs included IL-6, IL-1β, IL-1α, neutrophil
gelatinase-associated lipocalin (Lcn2), C-X-C motif chemokine 2
(Cxcl2), Fpr2, Traf1 and CD69 molecule. Compared with the study
from Li et al (17), who
prolonged the exposure to LPS to 24 h, there were slight variations
in the top changed mRNAs. Traf1, Fpr2 and CD69 were among the top
30 upregulated mRNAs in the early phase of LPS exposure, while
IL-6, IL-1β, Lcn2, serum amyloid A-3 protein and Cxcl2 were the top
upregulated genes in both the early and late phase. Furthermore,
the probes contained in the chip used in the present study were
designed to discern the different transcripts of genes. For
instance, two transcripts of the IL-6 gene (202 and 203), recorded
in the Ensemble Database, were among the top 30 upregulated mRNAs,
while IL6-201 was upregulated to a lesser degree (FC=71.15). The
different expression of the alternative transcripts of the same
gene requires further exploration to elucidate their potential
function.
Recently, Sun et al (27) also employed transcriptome
sequencing approaches to characterize the effects of LPS on lncRNA
and mRNA expression patterns in brain tissue isolated from Sprague
Dawley rats. In this study, Sprague Dawley rats were exposed to LPS
for 6 or 24 h and a large number of lncRNAs and mRNAs were
indicated to be changed. However, no behavioral observation of
these rats was made, which is required to diagnose SAE. The
upregulated gene regulator of G-protein signaling 14 identified by
Sun et al (27) was also
verified in the BV-2 cells of the present study. Furthermore, the
difference in the lncRNA expression signature between the present
study and the study by Sun et al (27) was huge, possibly due to the gap
between different species and different cell types.
GO analysis of these early upregulated DE mRNAs in
the category BP included inflammatory response, immune system
process, response to LPS and response to cytokine, which were all
related to inflammation and with a log10 (P-value) >10. This
differed from downregulated DE mRNAs, which mainly involved the
regulation of transcription, with log10 (P-value) <5. Among the
top downregulated coding RNAs, several had the function of gene
transcription or cooperation with transcription factor. For
instance, MAF bZIP transcription factor B (Mafb), an important
transcription factor gene strongly expressed in macrophages, is a
member of the Maf transcription factor family and regulates gene
transcription via the binding to the Maf recognition element
(28). A study demonstrated that
Mafb determined the anti-inflammatory properties of macrophages
(29). Consistently, in the
present study of BV-2 cells, short-time exposure to LPS markedly
decreased the expression of Mafb.
In the KEGG analysis of the downregulated mRNAs, it
was noted that pathways associated with lipid metabolism were
prevalent. Bile secretion, regulation of lipolysis in adipocyte and
insulin resistance were among the top 30 terms enriched by
downregulated mRNAs. Accumulating evidence has indicated that
different metabolic pathways or energy homeostasis are required for
programming M1 and M2 macrophages (30). Furthermore, it has been indicated
that LPS treatment may induce lipid droplet accumulation and
increased cellular triglyceride content in microglia. Lipid
metabolism was linked with inflammatory signaling and microglial
activation (31). Wang et
al (32) indicated that
exogenous recombinant human fibroblast growth factor 21, which may
suppress lipid accumulation through the activation of adenosine
5'-monophosphate-activated protein kinase (AMPK) (33), ameliorated LPS-induced
depressive-like behavior by inhibiting microglial expression of
proinflammatory cytokines. In addition, it has been indicated that
the activation of AMPK by ENERGI-F704 (a chemical component
identified from bamboo shoot extract) ameliorated the LPS-induced
inflammation in BV-2 cells (34).
Of note, in the present study, Abhd15, which is associated with
glucolipid metabolism commonly expressed in adipose tissue, was
significantly downregulated in LPS-stimulated BV-2 cells.
Furthermore, the expression of Lpin1 that modulates lipid
metabolism and glucose homeostasis was reduced by 14-fold after LPS
stimulation. The present results further substantiated the
association between lipid metabolism and microglial activation. How
these lipid-related genes function in the microglial activation and
whether the modulation of lipid metabolism is able to impact
neuroinflammation remains elusive and further investigation is
required.
Although the validated expression levels of Fpr2 and
TNF receptor-associated factor 1 (Tarf1) after LPS exposure were
not as high as those detected by microarray in the present study,
their functions in inflammation signaling were striking (35). Fpr2, a G-protein-coupled receptor,
has an important role in host defense and inflammation. Fpr2
inhibition is able to markedly reduce the activation of
transforming growth factor beta-activated kinase 1 induced by LPS
and decrease inflammation and oxidative stress in LPS-induced lung
injury (36). Mounting evidence
has indicated that Fpr2 participates in the pathogenesis of sepsis
(37,38). Tarf1 is a signaling adaptor first
identified as part of the TNFR2 signaling complex (39). Tarf1 has a key role in pro-survival
signaling downstream of TNFR superfamily members, such as TNFR2 and
CD40. Moreover, it has been demonstrated that Tarf1 negatively
regulates Toll-like receptor and Nod-like receptor signaling,
through sequestering the linear ubiquitin assembly complex
(40). In a study of acute lung
injury, the interferon of Tarf1 markedly attenuated LPS-induced
mortality, alleviated lung injury and markedly reduced inflammation
(41).
In 2011, Salmena et al (42) proposed the ceRNA hypothesis, which
aimed to explain how RNAs ‘talk’ to each other to establish
interactions to modify functional genetic information, and function
in pathological conditions. Through miRNA response elements, miRNAs
recognize and bind to regions with partial complementary sequences
on target RNA transcripts, resulting in the repression of the
target gene. Thereby, miRNAs mediate regulatory crosstalk between
various components of the transcriptome. On the contrary, lncRNAs
or circRNAs competitively bind and absorb miRNAs via specific
nucleotide elements, acting like a ‘sponge’, and release the coding
mRNAs from repression. Based on the expression profiles and the
ceRNA analysis, several lncRNAs, circRNAs and mRNAs were selected
for validation in the present study. For mRNAs, Traf1, Fpr2,
Pou2f2, Socs1, Agrn, Lacc1, Rapgef and Birc3 were demonstrated to
be upregulated and Abhd15 was downregulated. LncRNA
ENSMUST00000155949.1, also named 6530402F18Rik, was demonstrated to
have high transcript levels. The gene of 6530402F18Rik is located
on chromosome 2 and does not encode any protein. In the ceRNA
network analysis, it was predicted that 6530402F18Rik is able to
combine with miR-762 and miR-7648-3p, with the predicted scores of
750 and 735, respectively. Both miR-762 and miR-7648-3p were
predicted to target Pou2f2, with a predicted score >600. Socs1
was also predicted to be associated with 6530402F18Rik through
miR-762, as well as miR-7648-3p, with high scores. Unlike
6530402F18Rik, ENSMUST00000203956.1 (2010008C14Rik) was
demonstrated to be reduced in expression. In the prediction,
2010008C14Rik was able to ‘sponge’ miR-149-3p. Thus, it is probable
that the downregulated 2010008C14Rik released more free miR-149-3p
that had a predicted domain for Abhd15, leading to the decreased
expression of Abhd15. However, functional research on 6530402F18Rik
and 2010008C14Rik is scarce. LncRNAs functioning in the activation
of BV-2 cells and modulating BV-2 cell activation via ‘sponging’
are worth further investigation.
The present study detected thousands of DE lncRNAs
and mRNAs, while the number of significant DE circRNAs was lower,
with only 140 circRNAs determined to be upregulated and 123
circRNAs were downregulated. The circRNAs with a high FC were not
all included in the circRNA-miRNA-mRNA network. For instance,
CIRCpedia_216004, the top increased circRNA, was determined to be
highly expressed, but was not predicted in the ceRNA network. For
CIRCpedia_37520, there was only one predicted association with
Rapgef2, which was also validated in the present study, likely via
miR-5620-3p. Furthermore, it was predicted that CIRCpedia_38964 was
able to sequester miRNA and lead to the increased translation of
Agrn and Lacc1 that were indicated to moderately increase. It
remains elusive whether these circRNAs are only biomarkers of
inflammation or have other potential functions.
Despite the results obtained, the present study has
certain limitations. First, the pathogenesis of SAE involves not
only microglial cells, but also neurons, endothelial cells and
other cell types. Furthermore, although LPS is the primary
inflammatory substance in sepsis, other components of
microorganisms and the substances of damaged host cells are capable
of inducing inflammatory responses. The in vitro microglial
cells stimulated by LPS cannot completely imitate the pathogenesis
of SAE. Furthermore, the genes and links validated in the present
study require further validation in human microglial cells and
in vivo SAE models and their functions in inflammation
should also be explored in the future.
In conclusion, the present study revealed the ceRNA
networks involved the interrelation of lncRNA-miRNA-mRNA or
circRNA-miRNA-mRNA in the early phase of the microglial activation.
Furthermore, most of the DE genes were mainly involved in
inflammatory responses, immune responses and lipid metabolism.
Although more in vivo and in vitro functional studies
are required, the present study may improve the present knowledge
on neuroinflammation and provide potential therapeutic targets for
SAE.
Supplementary Material
Stimulation of LPS on BV-2 cells. (A)
The morphology of BV-2 cells after 0, 2, 4, 8, 16, 24 incubation of
1 μg/ml LPS (magnification, x200). (B) Viability of BV-2 cells
after 0, 2, 4, 8, 16 and 24 h of incubation with 1.00 μg/ml LPS
determined using the Cell Counting Kit-8. The ordinate value is the
OD at 450 nm in each well. (C) mRNA expression levels of TNF-α,
IL-6 and IL-1β after 4 h of incubation with 0.01, 0.10 or 1.00
μg/ml LPS. (D) Changes in mRNA expression levels of TNF-α, IL-6 and
IL-1β after incubation with LPS for different durations (0, 2, 4,
8, 16, 24 h). **P<0.010, compared with the group at 0
h (two-tailed Student's t-test); ##P<0.05, compared
with the group of 0.10 μg/ml (two-tailed Student's t-test). LPS,
lipopolysac-charide; OD, optical density; RQ, relative quantity;
(relative quantity were calculated according to the standard
2-ΔΔCq method using the Cq of β-actin to
normalize).
Primers used for the validation of
mRNAs.
Primers used for the validation of
lncRNAs and circRNAs.
Acknowledgements
The authors acknowledge the support of Shanghai OE
Biomedical Technology Co., Ltd. for the microarray of RNAs.
Funding
This work was supported by the Natural Science Foundation of
Anhui Province (grant no. 1808085MH308) and the School Research
Fund Project of Anhui Medical University (grant no.
2019xkj178).
Availability of data and materials
The datasets of the microarray used in the present
study are available from the GEO database under accession no.
GSE171696 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE171696).
Other datasets used and/or analyzed during the current study are
available from the corresponding author upon reasonable
request.
Authors' contributions
JW and QX conceived and designed the experiments. YL
and SC contributed to the cell culture and treatments. JW, JG, ZZ
and BH analyzed the data and drafted the manuscript. JG and QX
contributed to manuscript editing. JW, JG and QX confirm the
authenticity of the raw data. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Huang M, Cai SL and Su JQ: The
pathogenesis of sepsis and potential therapeutic targets. Int J Mol
Sci. 20(5376)2019.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Sprung CL, Peduzzi PN, Shatney CH, Schein
RM, Wilson MF, Sheagren JN and Hinshaw LB: Impact of encephalopathy
on mortality in the sepsis syndrome. The veterans administration
systemic sepsis cooperative study group. Crit Care Med. 18:801–806.
1990.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Iwashyna TJ, Ely EW, Smith DM and Langa
KM: Long-term cognitive impairment and functional disability among
survivors of severe sepsis. JAMA. 304:1787–1794. 2010.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Ren C, Yao RQ, Zhang H, Feng YW and Yao
YM: Sepsis-associated encephalopathy: A vicious cycle of
immunosuppression. J Neuroinflammation. 17(14)2020.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Block ML, Zecca L and Hong JS:
Microglia-mediated neurotoxicity: Uncovering the molecular
mechanisms. Nat Rev Neurosci. 8:57–69. 2007.PubMed/NCBI View
Article : Google Scholar
|
|
6
|
Lee E, Eo JC, Lee C and Yu JW: Distinct
features of brain-resident macrophages: Microglia and
non-parenchymal brain macrophages. Mol Cells. 44:281–291.
2021.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Li Y, Yin L, Fan Z, Su B, Chen Y, Ma Y,
Zhong Y, Hou W, Fang Z and Zhang X: Microglia: A potential
therapeutic target for sepsis-associated encephalopathy and
sepsis-associated chronic pain. Front Pharmacol.
11(600421)2020.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Shulyatnikova T and Verkhratsky A:
Astroglia in sepsis associated encephalopathy. Neurochem Res.
45:83–99. 2020.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Catarina AV, Branchini G, Bettoni L, De
Oliveira JR and Nunes FB: Sepsis-associated encephalopathy: From
pathophysiology to progress in experimental studies. Mol Neurobiol.
58:2770–2779. 2021.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Ye B, Tao T, Zhao A, Wen L, He X, Liu Y,
Fu Q, Mi W and Lou J: Blockade of IL-17A/IL-17R pathway protected
mice from sepsis-associated encephalopathy by inhibition of
microglia activation. Mediators Inflamm.
2019(8461725)2019.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Fatica A and Bozzoni I: Long non-coding
RNAs: New players in cell differentiation and development. Nat Rev
Genet. 15:7–21. 2014.PubMed/NCBI View
Article : Google Scholar
|
|
12
|
Yao H, Han B, Zhang Y, Shen L and Huang R:
Non-coding RNAs and autophagy. Adv Exp Med Biol. 1206:199–220.
2019.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Su Z, Yang Z, Xu Y, Chen Y and Yu Q:
MicroRNAs in apoptosis, autophagy and necroptosis. Oncotarget.
6:8474–8490. 2015.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Wang J, Yan S, Yang J, Lu H, Xu D and Wang
Z: Non-coding RNAs in rheumatoid arthritis: From bench to bedside.
Front Immunol. 10(3129)2020.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Ghafouri-Fard S, Shoorei H and Taheri M:
Non-coding RNAs participate in the ischemia-reperfusion injury.
Biomed Pharmacother. 129(110419)2020.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Kondo Y, Shinjo K and Katsushima K: Long
non-coding RNAs as an epigenetic regulator in human cancers. Cancer
Sci. 108:1927–1933. 2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Li Y, Li Q, Wang C, Li S and Yu L: Long
noncoding RNA expression profile in BV2 microglial cells exposed to
lipopolysaccharide. Biomed Res Int. 2019(5387407)2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Xiaoying G, Guo M, Jie L, Yanmei Z, Ying
C, Shengjie S, Haiyan G, Feixiang S, Sihua Q and Jiahang S:
CircHivep2 contributes to microglia activation and inflammation via
miR-181a-5p/SOCS2 signalling in mice with kainic acid-induced
epileptic seizures. J Cell Mol Med. 24:12980–12993. 2020.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Wu T, Li Y, Liang X, Liu X and Tang M:
Identification of potential circRNA-miRNA-mRNA regulatory networks
in response to graphene quantum dots in microglia by microarray
analysis. Ecotoxicol Environ Saf. 208(111672)2021.PubMed/NCBI View Article : Google Scholar
|
|
20
|
R Development Core Team. R: A language and
environment for statistical computing. R Foundation for Statistical
Computing. Vienna, Austria, 2018. URL https://www.R-project.org/.
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Widmann CN and Heneka MT: Long-term
cerebral consequences of sepsis. Lancet Neurol. 13:630–636.
2014.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Huang CT, Lue JH, Cheng TH and Tsai YJ:
Glycemic control with insulin attenuates sepsis-associated
encephalopathy by inhibiting glial activation via the suppression
of the nuclear factor kappa B and mitogen-activated protein kinase
signaling pathways in septic rats. Brain Res.
1738(146822)2020.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Kimura T, Toriuchi K, Kakita H, Tamura T,
Takeshita S, Yamada Y and Aoyama M: Hypothermia attenuates neuronal
damage via inhibition of microglial activation, including
suppression of microglial cytokine production and phagocytosis.
Cell Mol Neurobiol. 41:459–468. 2021.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Weis GCC, Assmann CE, Mostardeiro VB,
Alves AO, da Rosa JR, Pillat MM, de Andrade CM, Schetinger MRC,
Morsch VMM, da Cruz IBM and Costabeber IH: Chlorpyrifos pesticide
promotes oxidative stress and increases inflammatory states in BV-2
microglial cells: A role in neuroinflammation. Chemosphere.
278(130417)2021.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Henn A, Lund S, Hedtjärn M, Schrattenholz
A, Pörzgen P and Leist M: The suitability of BV2 cells as
alternative model system for primary microglia cultures or for
animal experiments examining brain inflammation. ALTEX. 26:83–94.
2009.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Sun W, Pei L and Liang Z: mRNA and long
non-coding RNA expression profiles in rats reveal inflammatory
features in sepsis-associated encephalopathy. Neurochem Res.
42:3199–3219. 2017.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Gautier EL, Shay T, Miller J, Greter M,
Jakubzick C, Ivanov S, Helft J, Chow A, Elpek KG, Gordonov S, et
al: Gene-expression profiles and transcriptional regulatory
pathways that underlie the identity and diversity of mouse tissue
macrophages. Nat Immunol. 13:1118–1128. 2012.PubMed/NCBI View
Article : Google Scholar
|
|
29
|
Cuevas VD, Anta L, Samaniego R,
Orta-Zavalza E, Vladimir de la Rosa J, Baujat G, Dominguez-Soto Á,
Sanchez-Mateos P, Escribese MM, Castrillo A, et al: MAFB determines
human macrophage anti-inflammatory polarization: Relevance for the
pathogenic mechanisms operating in multicentric carpotarsal
osteolysis. J Immunol. 198:2070–2081. 2017.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Remmerie A and Scott CL: Macrophages and
lipid metabolism. Cell Immunol. 330:27–42. 2018.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Chausse B, Kakimoto PA and Kann O:
Microglia and lipids: How metabolism controls brain innate
immunity. Semin Cell Dev Biol. 112:137–144. 2021.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Wang X, Zhu L, Hu J, Guo R, Ye S, Liu F,
Wang D, Zhao Y, Hu A, Wang X, et al: FGF21 attenuated LPS-induced
depressive-like behavior via inhibiting the inflammatory pathway.
Front Pharmacol. 11(154)2020.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Salminen A, Kauppinen A and Kaarniranta K:
FGF21 activates AMPK signaling: Impact on metabolic regulation and
the aging process. J Mol Med (Berl). 95:123–131. 2017.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Chen CC, Lin JT, Cheng YF, Kuo CY, Huang
CF, Kao SH, Liang YJ, Cheng CY and Chen HM: Amelioration of
LPS-induced inflammation response in microglia by AMPK activation.
Biomed Res Int. 2014(692061)2014.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Maciuszek M, Cacace A, Brennan E, Godson C
and Chapman TM: Recent advances in the design and development of
formyl peptide receptor 2 (FPR2/ALX) agonists as pro-resolving
agents with diverse therapeutic potential. Eur J Med Chem.
213(113167)2021.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Liu H, Lin Z and Ma Y: Suppression of Fpr2
expression protects against endotoxin-induced acute lung injury by
interacting with Nrf2-regulated TAK1 activation. Biomed
Pharmacother. 125(109943)2020.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Zhou Y, Lei J, Xie Q, Wu L, Jin S, Guo B,
Wang X, Yan G, Zhang Q, Zhao H, et al: Fibrinogen-like protein 2
controls sepsis catabasis by interacting with resolvin Dp5. Sci
Adv. 5(eaax0629)2019.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Gobbetti T, Coldewey SM, Chen J, McArthur
S, le Faouder P, Cenac N, Flower RJ, Thiemermann C and Perretti M:
Nonredundant protective properties of FPR2/ALX in polymicrobial
murine sepsis. Proc Natl Acad Sci USA. 111:18685–18690.
2014.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Abdul-Sater AA, Edilova MI, Clouthier DL,
Mbanwi A, Kremmer E and Watts TH: The signaling adaptor TRAF1
negatively regulates toll-like receptor signaling and this
underlies its role in rheumatic disease. Nat Immunol. 18:26–35.
2017.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Edilova MI, Abdul-Sate AA and Watts TH:
TRAF1 signaling in human health and disease. Front Immunol.
9(2969)2018.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Bin W, Ming X and Wen-Xia C: TRAF1
meditates lipopolysaccharide-induced acute lung injury by up
regulating JNK activation. Biochem Biophys Res Commun. 511:49–56.
2019.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Salmena L, Poliseno L, Tay Y, Kats L and
Pandolfi PPA: A ceRNA hypothesis: The rosetta stone of a hidden RNA
language? Cell. 146:353–358. 2011.PubMed/NCBI View Article : Google Scholar
|