Introduction
Pancreatic cancer is a malignant tumor type that
shows one of the highest mortality rates; it ranks as the eighth
leading cause of cancer-related deaths worldwide (1). The poor prognosis of pancreatic
cancer is attributed to its late manifestation, lack of accurate
biomarkers for early diagnosis and assessment of curative resection
options, its propensity for early metastasis, as well as the
limited effects of standard chemotherapeutic agents and
radiotherapy (2). Therefore, novel
diagnostic modalities for early diagnosis and new therapeutic
strategies are urgently needed for the treatment of pancreatic
cancer.
microRNAs (miRNAs) are a class of small,
endogenously expressed, well-conserved, non-coding RNA molecules,
18–25 nucleotides long. They play important regulatory roles on
their gene targets, by degrading their mRNA or inhibiting their
translation (3,4). Growing evidence suggests that miRNAs
play an important role in various biological processes, including
cell proliferation, development, and differentiation (5,6).
Furthermore, emerging evidence suggests that miRNAs play essential
roles in tumorigenesis, and thus may function as promising targets
for the treatment of cancer (7,8).
Recent studies have demonstrated that miR-193b is downregulated in
a variety of cancers, and that it regulates cancer cell
proliferation, migration, invasion, and metastasis (9–13).
Although Ikeda et al (14)
reported that the expression of miR-193b is altered by the
mitogen-activated protein kinase (MAPK) in pancreatic cancer cells
and that its exogenous overexpression markedly inhibits cell
proliferation, the exact role of miR-193b in pancreatic cancer
remains unclear.
In the present study, we not only investigated the
expression of miR-193b in pancreatic cancer tissue, but also
studied the regulatory effects of miR-193b on panceatic cancer cell
proliferation, migration and invasion in vitro, as well as
the underlying molecular mechanisms, which may help develop
potential diagnostic and therapeutic tools for pancreatic cancer.
Our data indicated that miR-193b may be downregulated in pancreatic
cancer and affects the behavior of Panc-1 cells, namely their
proliferation, apoptosis, migration and invasion. We also
demonstrated that the stathmin 1 (STMN1) and urokinase-type
plasminogen activator (uPA) genes are negatively regulated
by miR-193b.
Materials and methods
Patients and tumor samples
Pancreatic cancer and adjacent tissues were obtained
from 27 patients (17 males and 10 females, median age 63 years,
range 38–79 years) undergoing pancreatic cancer surgery at the
Xiangya Hospital at the Central South University, Changsha, China
(Table I). Six healthy pancreatic
samples were collected at surgery from patients with acute
pancreatic injury. The samples were immediately snap-frozen in
liquid nitrogen and stored at −80°C for RNA extraction. Informed
consent was obtained from all patients. Both tumor and
non-cancerous samples were histologically confirmed. The study was
approved by the Ethics Committee of the Central South
University.
 | Table IClinical features and miR-193b
expression profiles in the studied patients. |
Table I
Clinical features and miR-193b
expression profiles in the studied patients.
| Patient | Age (years) | Gender | Tumor size
(cm) | TNM stage | Normalized miR-193b
level |
|---|
| 1 | 60 | F | 3.6×3.9×4.5 | II | 0.1869 |
| 2 | 38 | M | 2.8×3.6×4.4 | IV | 0.0409 |
| 3 | 63 | M | 2.7×2.8×2.6 | I | 2.1585 |
| 4 | 58 | M | 1.9×1.8×2.3 | I | 0.2517 |
| 5 | 75 | F | 2.2×2.8×3.6 | III | 0.0797 |
| 6 | 66 | M | 2.5×3.5×4.4 | II | 0.1111 |
| 7 | 63 | M | 3.0×3.0×3.1 | II | 0.2432 |
| 8 | 73 | F | 2.5×3.2×3.3 | I | 0.1267 |
| 9 | 61 | M | 4.0×5.0×3.5 | II | 0.0896 |
| 10 | 70 | F | 2.0×3.0×2.2 | I | 1.5369 |
| 11 | 62 | M | 2.1×2.6×3.1 | I | 0.1989 |
| 12 | 48 | M | 3.4×3.8×4.5 | III | 0.0813 |
| 13 | 57 | F | 3.8.×4.0×3.6 | III | 0.4323 |
| 14 | 73 | M | 2.5×2.9×3.6 | II | 0.0981 |
| 15 | 71 | M | 1.7×2.4×2.6 | I | 0.2717 |
| 16 | 56 | F | 2.3×3.0×2.2 | II | 0.1869 |
| 17 | 77 | M | 3.2×4.2×4.1 | II | 0.0608 |
| 18 | 59 | F | 4.0×4.2×4.6 | III | 0.0836 |
| 19 | 79 | M | 6.0×5.0×4.2 | IV | 0.0902 |
| 20 | 77 | M | 3.0×4.0×4.0 | III | 0.1387 |
| 21 | 69 | F | 3.1×4.3×3.7 | III | 0.4234 |
| 22 | 65 | F | 2.7×3.5×3.0 | III | 0.0703 |
| 23 | 56 | M | 3.8×4.5×3.3 | II | 0.1285 |
| 24 | 61 | M | 3.3×2.9×2.7 | II | 0.3635 |
| 25 | 64 | F | 4.2×4.5×4.4 | III | 0.1975 |
| 26 | 39 | M | 2.6×1.9×3.0 | II | 0.0915 |
| 27 | 57 | F | 2.5×3.8×3.6 | I | 0.1368 |
RNA extraction and reverse
transcription-quantitative PCR (RT-qPCR) analysis
Total RNA was extracted from the 27 pancreatic
cancer tissues and their adjacent non-carcinoma tissues, the six
healthy pancreatic tissues and the cultured cells. For miR-193b
expression analysis, total RNA was poly-adenylated using a Poly (A)
Tailing kit, according to the manufacturer’s instructions
(GeneCopoeia, Guanzhou, China) and 10 ng of poly(A) mRNA were
converted to cDNA using miR-193b-specific primers of U6, which were
synthesized forward 5′-CTCGCTTCGGCAGCACA-3′ and reverse
5′-AACGCTTCACGAATTTGCGT-3′ (cat.no HmiRQP0278; GeneCopeia), and an
Applied Biosystems® TaqMan® MicroRNA Reverse
Transcription kit (Thermo Fisher Scientific, Waltham, MA, USA).
Following reverse transcription, qPCR was performed on an ABI 7500
thermocycler (Thermo Fisher Scientific) with the following cycling
conditions: 95°C, 5 min, 1 cycle, 95°C, 10 sec, 65°C, 20 sec, 72°C,
10 sec, 40 cycles. The U6 gene was used as a normalization control.
Each sample was analyzed in triplicate.
Cell culture
The human pancreatic cancer cell line Panc-1
(Institute of Biochemistry and Cell Biology, Shanghai, China) was
maintained in our laboratory and cultured in Hepes-buffered
Dulbecco’s modified Eagle’s medium (H-DMEM) supplemented with 10%
Gibco® fetal bovine serum (FBS) (Thermo Fisher
Scientific). Cells were cultured at 37°C in 5% CO2.
Cell transfection
Panc-1 cells were transiently transfected for 48 h
with chemically synthesized miR-193b and the negative control miRNA
(miR-NC) (GenePharma, Shanghai, China) (Table II). Transfection was performed
with Invitrogen™ Lipofectamine® 2000 (Thermo Fisher
Scientific) according to the manufacturer’s recommendations.
 | Table IISequences of the chemically
synthesized miR-193b, miR-NC, miR-193b inhibitor and NC
inbibitors. |
Table II
Sequences of the chemically
synthesized miR-193b, miR-NC, miR-193b inhibitor and NC
inbibitors.
| Synthesized
miRNA | Sequence 5′-3′ |
|---|
| miR-193b |
AACUGGCCCUCAAAGUCCCGCU |
| miR-NC |
UUCUCCGAACGUGUCACGU |
| miR-193b
inhibitor |
AGCGGGACUUUGAGGGCCAGUU |
| NC inbibitor |
CAGUACUUUUGUGUAGUACAA |
Cell proliferation assay
The MTT assay was used to measure cell
proliferation. At 48 h post-transfection, the transfection medium
in each well was replaced by 100 μl of fresh serum-free medium
supplemented with 100 μl 0.5 g/l MTT solution. After incubation at
37°C for 4 h, the MTT medium was removed by aspiration and 50 μl of
dimethyl sulfoxide were added to each well. After incubation at
37°C for an additional 10 min, the absorbance of the sample at 570
nm was measured using a plate reader (cat.no. AD 340C; Beckman
Coulter, Miami, FL, USA).
Cell apoptosis assay
Cells were collected and washed with
phosphate-buffered saline (PBS) mixed with 2% ethylene diamine
tetraacetic acid. Then, each sample, containing
105–106 cells/ml, was stained with 5 μl
Annexin V-fluorescein isothiocyanate (FITC) and 10 μl propidium
iodide (PI) (Annexin V-FITC/PI Apoptosis Detection kit; KeyGen
Biotech Co., Ltd., Nanjing, China) for 15 min. Afterwards, the
cells were diluted using 400 μl Annexin V binding buffer (Haoran
Biological Technology Co, Ltd., Shanghai, China) and analyzed on a
FACSAria flow cytometer (BD Biosciences, Bedford, MA, USA).
Cell migration and invasion assays
Invasion assays were performed in a 24-well
transwell chamber purchased from Corning (Cambridge, MA, USA). The
chamber contained an 8 μm-pore size polycarbonate membrane filter
and was precoated with 100 μg of Matrigel (BD Biosciences). Panc-1
cells transfected with miR-193b or miR-NC were collected and
resuspended in serum-free H-DMEM medium at a concentration of
1×105 cells/ml. Then, the cell suspensions were added
into the top chambers (200 ml/well) and the bottom chambers were
filled with H-DMEM medium containing 10% FBS (500 ml/well),
followed by a 24-h incubation at 37°C. The cells that did not
penetrate the polycarbonate membrane were swabbed using a cotton
bud. The cells that had passed through the membrane and adhered to
the bottom of the polycarbonate membrane were stained for 20 min
with a solution containing 0.1% crystal violet and 20% methanol,
and were photographed and manually counted under an inverted
fluorescence microscope (Olympus, IX70; Olympus, Tokyo, Japan). The
migration assay was performed following similar procedures except
for the use of Matrigel coating on the filters. Two independent
experiments were performed for each assay. The average of cell
counts in five randomly selected fields (x400) was recorded as the
value of each chamber. The experiment was repeated twice, with
triplicate measurements in each experiment.
Bioinformatics analysis
To examine the potential downstream target genes of
miR-193b, the TargetScan (http://www.targetscan.org) (15), miRBase (http://www.mirbase.org) (16) and PicTar (http://www.mirbase.org) (17) were used. The results revealed that
a series of 3′ UTR of human genes contained potential
miR-193b-binding sequences. Amonth these, the oncogenes, STMN1 and
uPA were of interest. The 3′ UTR of STMN1 (59–65 nt, Genebank
accession no. NM_005563) gene and the uPA (777–783 ntl Genebank
accesion no. NM_00145031) gene contained the miRb-binding site.
Dual luciferase reporter assay
Construction of the luciferase report vectors were
as described by Chen et al (18). Briefly, DNA fragments were
amplified from human genomic DNA and cloned into the multiple
cloning sites (XhoI and NotI) distal to the Renilla luciferase
coding region of the psiCHECK2 vector (Promega, Madison, WI, USA).
PCR was performed using KOD hot start DNA polymerase (Novagen,
Madison, WI, USA). The primer sequences used to construct the
psiCHECK-STMN1 vector containing the wild type STMNI 3′ UTR were as
follows: XhoI-STMN1, forward
5′-ACGCCTCGAGTTGTTCTGAGAACTGACTTTCTC-3′ and NotI-STMN1 reverse
5′-ATAAGAATGCGGCCGCATATTCTGATTCTCGTGTCATAGC-3′. Overlapping PCR was
used to generate psiCHECK-STMN1-mutant (M), which contained a
deletion at the miR-193b seed binding site. The following two
primers were also used: STMN1-M, forward
5′-ATATCCAAAGACTGTACTTCATTTTATTTTTTCCCTG-3 and reverse
5′-AGTACAGTCTTTGGATAT-3′. The wild-type and deletion mutant of the
uPA 3′ UTR were generated using the same technique and were cloned
into multiple cloning sites (XhoI and NotI) distal to the Renilla
luciferase coding region of the psiCHECK vector (Promega). The
primer sequences used to construct the psiCHECK-STMN1 containing
wild type and mutant uPA 3′ UTR were as follows: XhoI-uPA, forward
5′-CGTCTAGAGGGTCCCCAGGGAGGAAAC-3′ and NotI-uPA, reverse
5′-CGCATATGTCATCAGAAAAATCACATT-3′, uPA-M forward
5′-CAGTTTCACTTTCACATATCCCTTCCTTTTAGC-3′ and reverse
5′-GCTAAAAGGAAGGGATATGTGAAAGTGAAACTG-3′. The following cycling
conditions were used: pre-denaturation at 95°C for 5 min;
denaturation at 95°C for 30 sec, annealing at 60°C for 30 sec and
elongation at 72°C for 30 sec for a total of 30 cycles, followed by
elongation at 72°C for 10 min. They were then inserted into
multiple cloning sites of the psiCHECK™-2 vector (Promega, Madison,
WI, USA) to obtain the pMIR-STMN1, -mut STMN1, -uPA, and -mut uPA
vectors. For the luciferase assay, Panc-1 cells were transfected in
24-well plates using the Lipofectamine® 2000
transfection reagent. Each well was transfected with 100 ng of
either pMIR-STMN1/uPA or pMIR-mut STMN1/mut uPA vector, along with
10 pmol of hsa-miR-193b or miR-NC. The vector pRL-TK (Promega) was
also transfected as a control. Forty-eight hours after
transfection, the luciferase activity was measured using the Dual
Luciferase® Reporter Assay system (Promega).
Western blot analysis
Cells were solubilized in cold RIPA lysis buffer
(Santa Cruz Biotechnology, Inc.). Proteins were extracted from the
cell lysate using the Total Protein Extraction kit (BestBio,
Shanghai, China) and were quantified using the BCA Protein assay
kit (Santa Cruz Biotechnology, Inc.). The proteins were separated
by 5% SDS-PAGE, and then transferred onto a polyvinylidene
difluoride membrane. The membrane was blocked in 5% non-fat dried
milk in PBS with Tween-20 (PBST) for 3 h, and then incubated
overnight with the primary rabbit anti-STMN1 polyclonal (1:500;
Abcam, Cambridge, UK) and anti-uPA polyclonal antibodies (1:1,000;
Santa Cruz Biotechnology, Inc.). GAPDH was used as the loading
control. After incubation for 1 h at room temperature with the
secondary antibodies, goat ant-rabbit immunoglobulin (Ig)
G/horseradish peroxidase (HRP) at a dilution of 1:30,000 (Santa
Cruz Biotechnology, Inc.) and goat anti-mouse IgG/HRP (1:50,000;
Santa Cruz Biotechnology, Inc.). The immune complexes were detected
using an enhanced chemiluminescence (ECL) kit (Beyotime Institute
of Biotechnology, Shanghai, China).
Statistical analysis
The data were expressed as mean ± standard deviation
and were analyzed using the SPSS 13.0 software (SPSS Inc., Chicago,
IL, USA). Comparisons of the STMN1 mRNA expression data were
performed with the Mann-Whitney U-test. The statistical
significance of differences among groups was analyzed by
paired-sample t-tests or a one-way analysis of variance (ANOVA),
followed by post-hoc least significant difference (LSD) multiple
testing correction, when appropriate. P<0.05 was considered to
indicate statistically significant differences.
Results
miR-193b expression is decreased in
pancreatic cancer tissues
To investigate the roles of miR-193b in pancreatic
cancer, we examined its expression in 27 primary pancreatic cancer
and adjacent matched non-tumorous tissues, and in 6 healthy
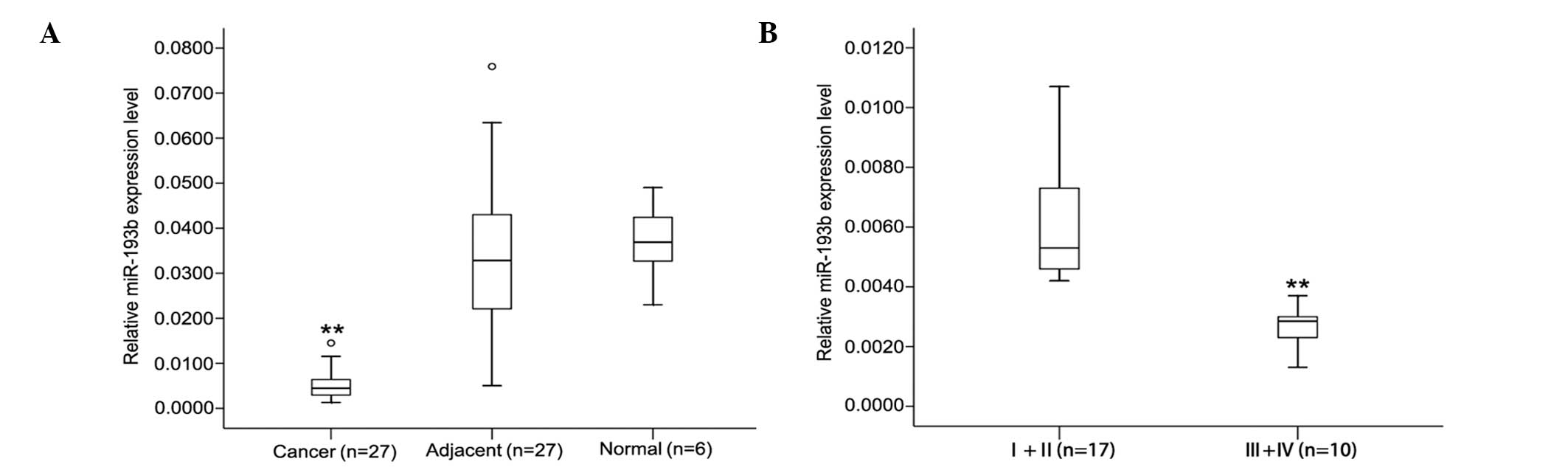
pancreatic tissues. Table I shows
the miR-193b relative expression data in the 27 samples; the
expression of miR-193b was decreased (from 2.3- to 24.4-fold) in
>90% of the pancreatic cancer samples (25 out of 27 patients).
The level of miR-193b in the pancreatic cancer tissues was lower
compared to the adjacent matched non-tumorous tissues
(0.0084±0.0094 vs. 0.0341±0.0217, P<0.01; Fig. 1A); however, no significant
differences were found between adjacent matched non-tumorous
tissues and healthy tissues (0.0341±0.0217 vs. 0.0368±0.0089,
P>0.05). Moreover, analysis of expression data in the pancreatic
cancer samples of different TNM stages showed that there is a
difference between early- and advanced-stage miR-193b expression
levels. As shown in Fig. 1B, in
early pancreatic cancer at stages I and II (n=17), miR-193b
expression was significantly higher than that observed in the
advanced stages III and IV (n=10). These results suggested that
downregulation of miR-193b may be related to the development of
human pancreatic cancer.
Effects of miR-193b transfection on
pancreatic cancer cell proliferation
Since miR-193b may be related to the development of
human pancreatic cancer, we further investigated its effects on
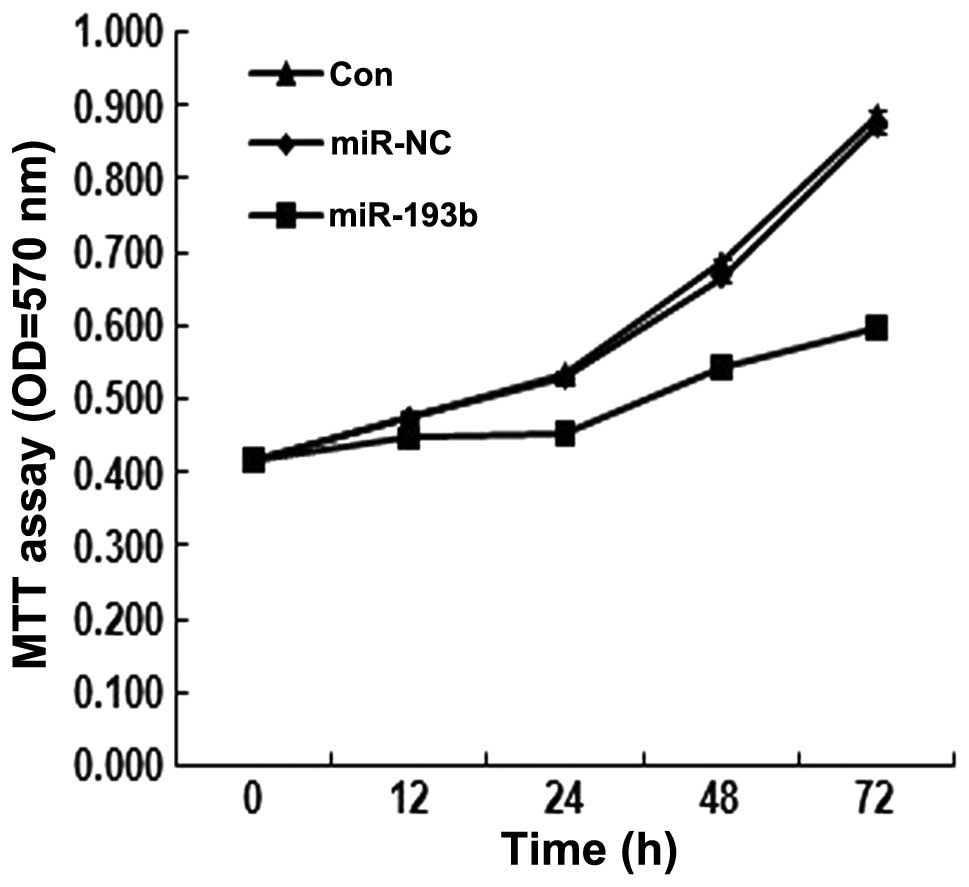
in vitro proliferation. The MTT assay was performed in
Panc-1 cells transfected with the miR-193b, in which the miR is
expected to be overexpressed. As shown in Fig. 2, the assay showed that in the
miR-193b-transfected Panc-1 cells, the proliferation rate was lower
than that observed in the control groups (P<0.05). These results
suggested that miR-193b inhibits Panc-1 cell proliferation.
Effects of miR-193b transfection on cell
apoptosis
Immortality is a common characteristic of cancer
cells. To further explore the role of miR-193b on the apoptosis of
pancreatic cells, we performed Annexin V/PI flow cytometry analysis
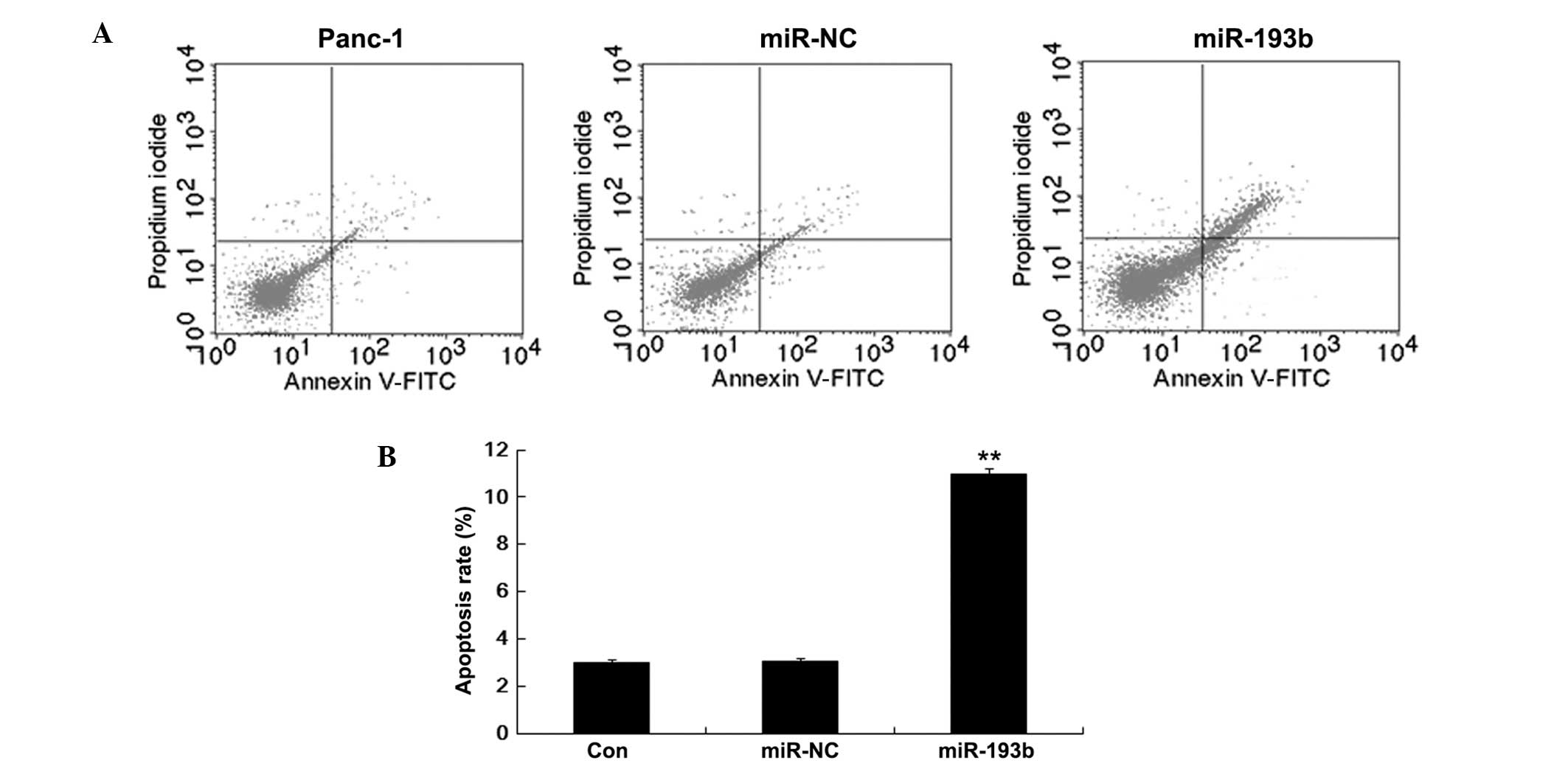
on Panc-1 cells transfected with miR-193b and miR-NC. Fig. 3 shows that the percentage of
apoptotic Panc-1 cells was higher in the miR-193b (10.97±0.22%)
than in the miR-NC (2.98±0.12%; P<0.01) group. Thus, these
results suggested that miR-193b enhances Panc-1 cell apoptosis.
Effects of miR-193b transfection on cell
migration and invasion
To address the potential effects of miR-193b on the
migration and invasion of pancreatic cancer cells in vitro,
we performed cell migration and invasion assays in the different
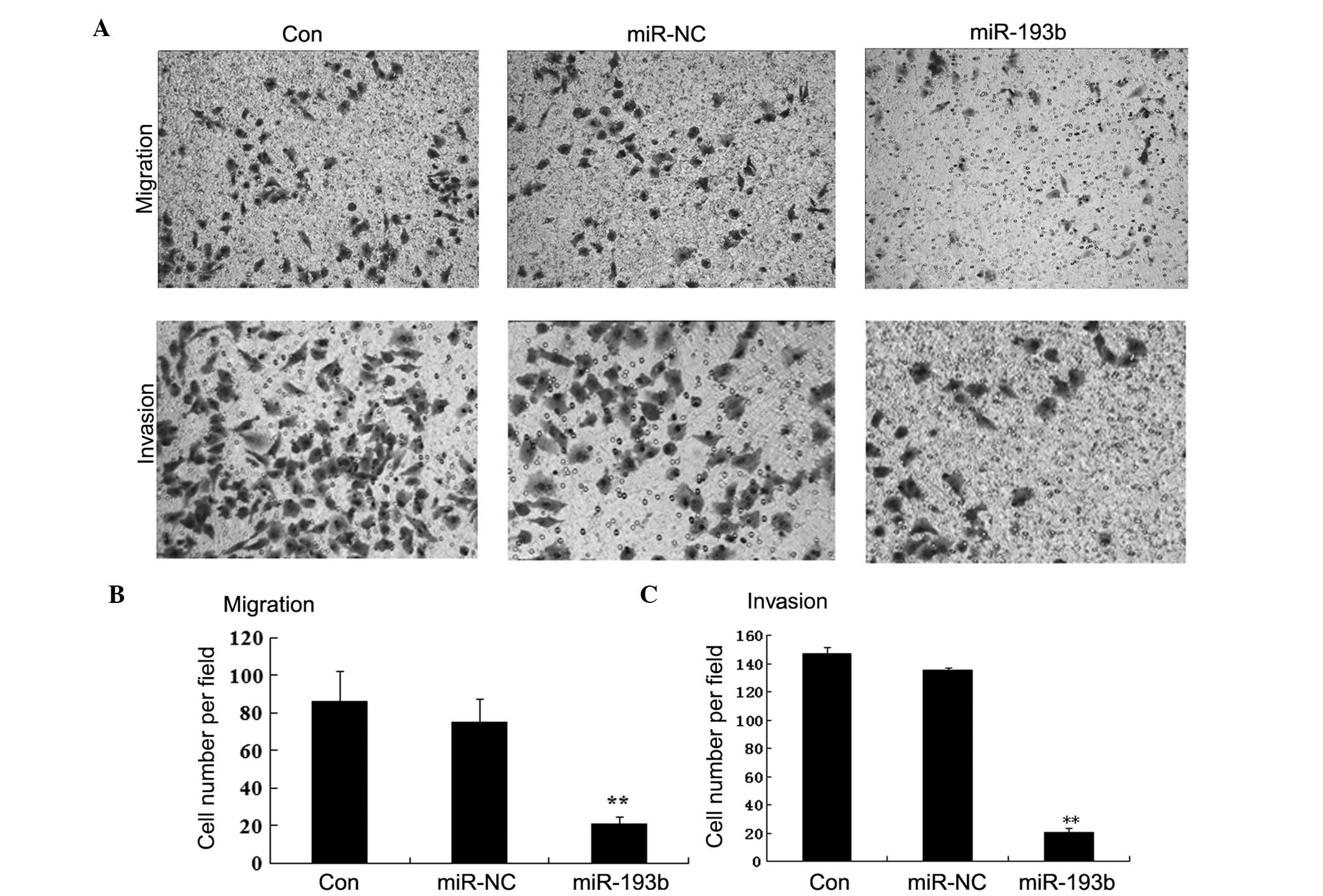
cell lines. As shown in Fig. 4,
Panc-1 cells transfected with miR-193b showed markedly reduced
migratory and invasive activity than the miR-NC or the
untransfected Panc-1 cells (P<0.01). Taken together, these
results suggest that miR-193b may act as a potent suppressor of
pancreatic cancer cell migration and invasion.
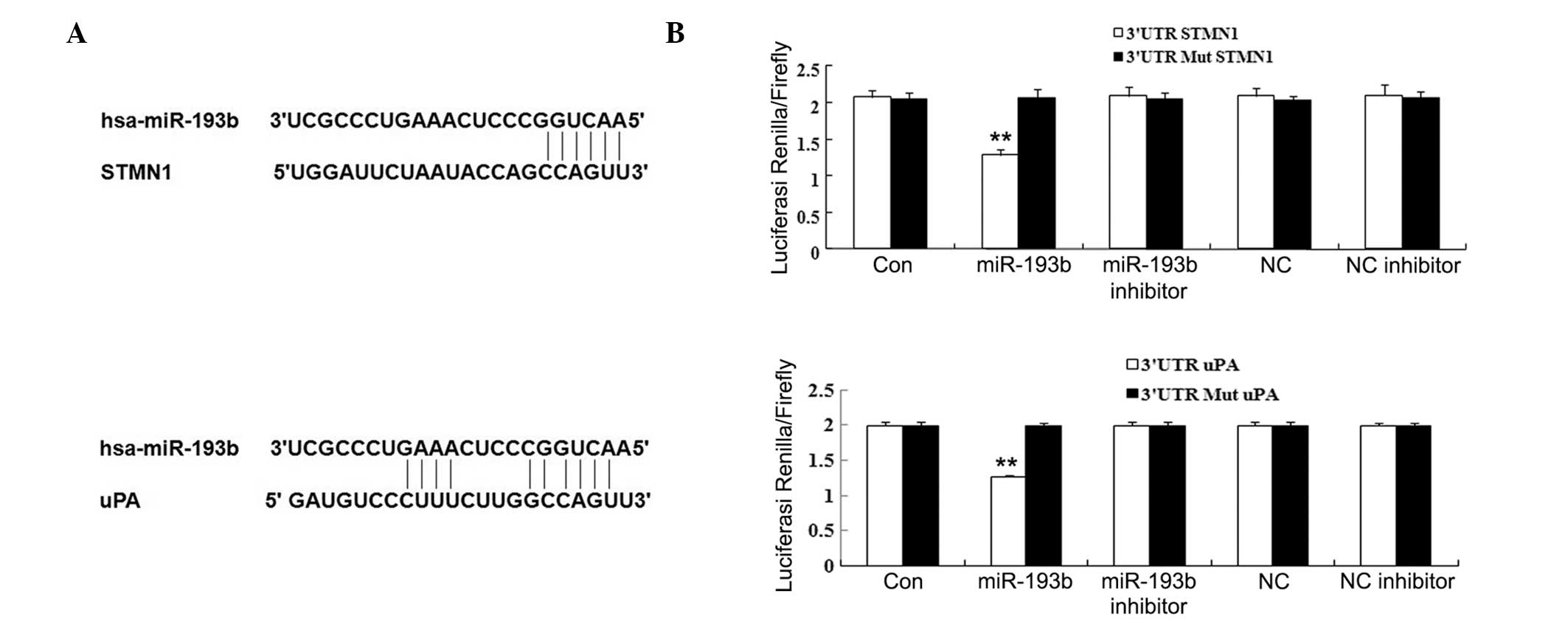
miR-193b directly represses the
expression of STMN1 and uPA through binding to their 3′-UTRs
To investigate the mechanisms by which miR-193b
inhibits pancreatic cancer growth and metastasis, we performed a
bioinformatic analysis to identify the potential target genes of
miR-193b. We found a number of human genes that contain potential
miR-193b-binding sequences in their 3′-UTR regions. Among these, we
focused on two oncogenes, STMN1 and uPA. To confirm
that these genes are direct targets of miR-193b, luciferase assays
were used. The 3′-UTRs of the STMN1 and uPA genes
(Fig. 5A) were cloned downstream
of the coding sequence of luciferase. The resulting constructs were
cotransfected with miR-193b or miR-NC (Table II) into Panc-1 cells. The results
showed that miR-193b, but not miR-NC, specifically decreases the
luciferase reporter levels (Fig.
5B). Moreover, the inhibitory effects of miR-193b were
eliminated upon seed-sequence-deletion mutation of the
miR-193b-binding site within the 3′-UTRs of STMN1 and
uPA. These results indicated that STMN1 and
uPA are the direct downstream targets of miR-193b in
pancreatic cancer.
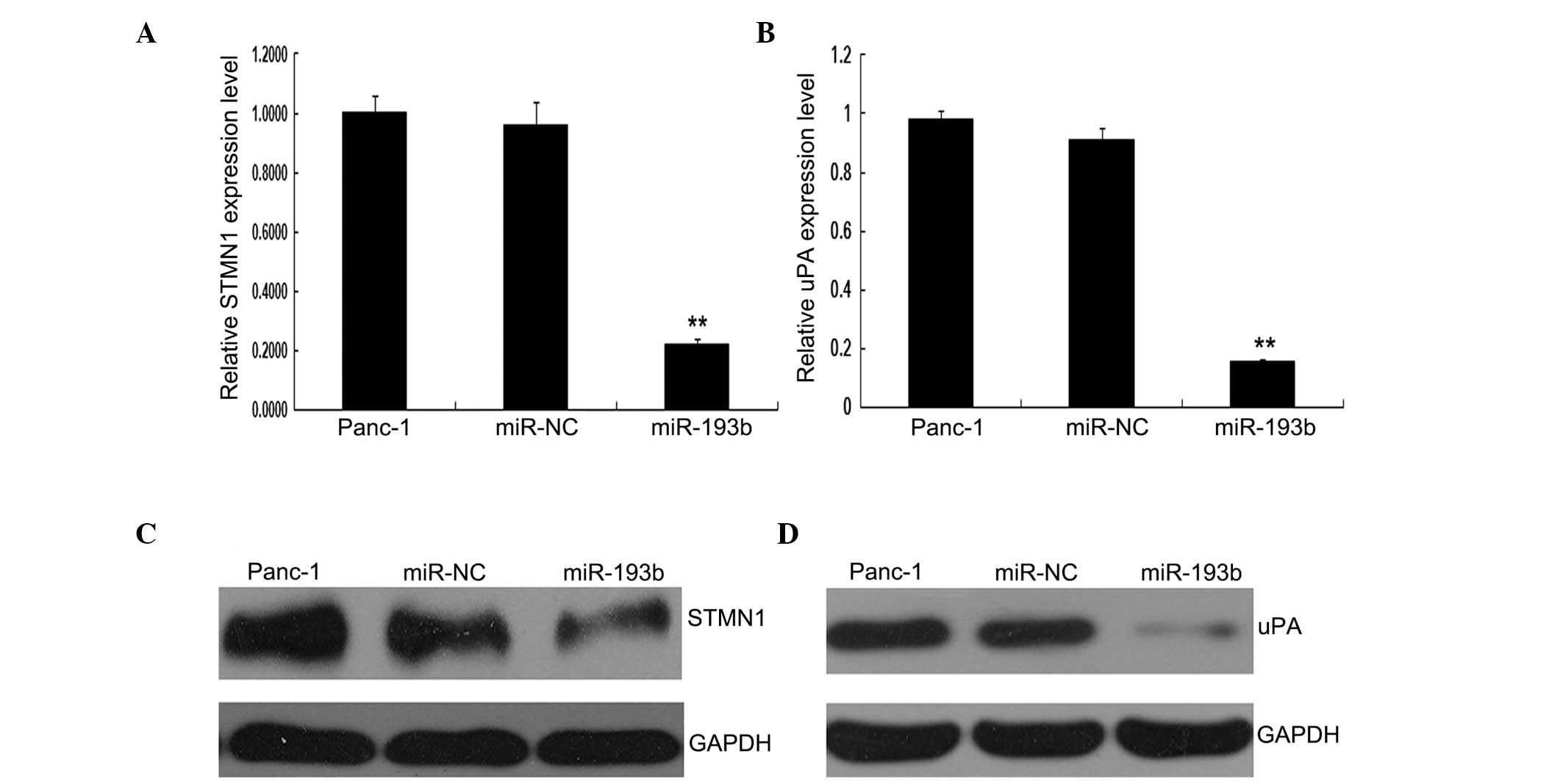
To further investigate the regulatory roles of
miR-193b on the STMN1 and uPA genes, we transfected
Panc-1 cells with miR-193b or miR-NC, and detected the expression
of STMN1 and uPA using RT-qPCR and western blotting
assays. As shown in Fig. 6, both
mRNA and protein expression levels of STMN1 and uPA were
significantly decreased after miR-193b transfection, as compared to
the miR-NC group or the untransfected cells (P<0.05). These data
suggest that STMN1 and uPA expression is negatively regulated by
miR-193b.
Discussion
The present study revealed, for the first time to
the best of our knowledge, the roles of miR-193b in pancreatic
cancer, as well as two of its target genes, STMN1 and
uPA. We found that the expression level of miR-193b is
reduced in pancreatic caner tissues, as compared to the level of
adjacent matched non-tumorous tissues and healthy pancreatic
tissues. Additional assays demonstrated that transfection of cells
with miR-193b, which is expected to increase its expression level,
significantly inhibited cellular proliferation, migration and
invasion, and markedly promoted apoptosis in pancreatic cancer
cells. In addition, STMN1 and uPA were identified as
direct targets of miR-193b, and their expression in pancreatic
cancer cells was found to be negatively regulated by miR-193b.
Accumulating evidence has suggested that miR-193b
acts as a tumor suppressor in various types of cancer, such as
breast, gastric, cervical, prostate cancer, etc. (19–22).
For instance, Wu and colleagues reported that the expression level
of miR-193b is significantly downregulated in endometrioid
adenocarcinoma (9). Rauhala et
al found that miR-193b is an epigenetically regulated tumor
suppressor in prostate cancer (10). Furthermore, miR-193b was found to
be significantly downregulated in melanoma tissues, and
overexpression of miR-193b in melanoma cell lines repressed cell
proliferation, probably through regulating cyclin D1 (23). However, the detailed role of
miR-193b in pancreatic cancer remains unclear. In our study, we
report that miR-193b is systematically downregulated in pancreatic
cancer tissues. Moreover, we found that the expression level of
miR-193b negatively correlates to the TNM stages of pancreatic
cancer, indicating that downregulation of miR-193b may promote the
development and progression of pancreatic cancer.
Recently, Ikeda and colleagues suggested that
activation of the MAPK may play a role in aberrant expression of
miR-193b, which is associated with pancreatic cancer (14). They used RT-qPCR to identify the
MAPK-associated miRNAs in pancreatic cancer cells, and further
found that overexpression of the MAPK-associated miR-193b exerts
the most notable inhibitory effect on proliferation of cultured
pancreatic cancer cells (14).
Consistent with their findings, we also found that transfection of
miR-193b inhibits pancreatic cancer cell proliferation. We further
demonstrated that transfection of cells with miR-193b effectively
promoted apoptosis and suppressed migration and invasion in
pancreatic cancer cells. Considering published and our present
findings, we suggest that miR-193b plays a tumor-suppressive role
in pancreatic cancer.
To further investigate the molecular mechanism
underlying the involvement of miR-193b in pancreatic cancer, we
performed a luciferase reporter assay, and demonstrated that the
genes STMN1 and uPA are the direct targets of
miR-193b, their expression levels being negatively regulated by
this miR in pancreatic cancer cells. A number of studies have
identified miR-193b targets in human malignancies, such as the
genes CCND1, NT5E, PLAU, STARD7,
STMN1, uPA, and YWHAZ, and showed that the
tumor-suppressive role of miR-193b mainly involves inhibition of
the expression of its targets in human cancer (12,14).
STMN1 encodes a cytosolic phosphoprotein,
which plays crucial roles in the formation and the function of the
mitotic spindle. By acting as a microtubule destabilizer, STMN1
participates in cellular biological processes such as cell
division, motility, and differentiation (24). Recent studies have demonstrated
that STMN1 is upregulated in multiple types of malignant tumors,
including sarcoma, hepatocellular carcinoma, gastric, breast and
prostate cancer, oral squamous cell carcinoma, and lung
adenocarcinomas (25–28). Furthermore, STMN1 has been
suggested to act as an oncogene in human malignancies. For
instance, STMN1 was recently reported to upregulate ovarian clear
cell adenocarcinoma via the regulation of HIF-1α through the
PI3K/Akt/mTOR pathway (28).
Moreover, the oncogenic role of STMN1 has been revealed in
pancreatic cancer. Jiang et al found that ectopic
overexpression of STMN1 prevents transforming growth factor-β
inducible early gene 1 (TIEG1)-mediated growth inhibition of
pancreatic cancer cells, while small interfering RNAs targeting
STMN1 inhibited pancreatic cancer cell growth (26). Wang and colleagues suggested that
the therapeutic effect of gemcitabine in pancreatic cancer may be
associated with the inhibition of STMN1 (30). In this study, we showed that
transfection of miR-193b significantly reduced the expression of
STMN1 in pancreatic cancer cells. Based on these findings and ours,
we suggest that overexpression of miR-193b effectively inhibits
pancreatic cancer growth in vitro, at least in part through
the inhibition of STMN1 expression.
The second miR-193 target studied herein is the gene
encoding uPA, which has been implicated in different physiological
and pathophysiological processes, including cell adhesion and
migration (31). In fact, the
oncogenic role of uPA in pancreatic cancer is well established.
Cantero et al demonstrated that the expression of the uPA
protein, as well as of its receptor, is significantly increased in
pancreatic cancer, which correlates to shorter postoperative
survival (32). Moreover,
upregulation of uPA was demonstrated to induce pancreatic cancer
cell invasion (33). In this
study, we found that miR-193b transfection markedly reduced uPA
expression in pancreatic cancer cells, suggesting that the
inhibition of pancreatic cancer growth by miR-193b in vitro
may involve direct modulation of uPA expression. In addition, uPA
was suggested to participate in the post-translational modification
of STMN1 (34). Therefore, future
studies need to focus on the molecular mechanism underlying
miR-193b effects on pancreatic cancer, and specifically on the
relationship among miR-193b, uPA and STMN1.
In conclusion, our study revealed that the
expression of miR-193b is reduced in pancreatic cancer tissues, and
that transfection with miR-193b has an inhibitory effect on
pancreatic cancer cell proliferation in vitro, potentially
via regulating the expression of the STMN1 and uPA
genes, which were identified as direct targets of miR-193b.
Therefore, our study expands the current understanding on the
molecular mechanism underlying the effects of miR-193b on
pancreatic cancer cells, and suggests that miR-193 may constitute a
promising therapeutic agent for suppressing pancreatic cancer
growth and metastasis.
Acknowledgements
This study was supported by the Fundamental Research
Funds for the Central Universities of the Central South
University.
References
|
1
|
Arora S, Bhardwaj A, Srivastava SK, Singh
S, McClellan S, Wang B and Singh AP: Honokiol arrests cell cycle,
induces apoptosis, and potentiates the cytotoxic effect of
gemcitabine in human pancreatic cancer cells. PLoS One.
6:e215732011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Niedergethmann M, Alves F, Neff JK, et al:
Gene expression profiling of liver metastases and tumour invasion
in pancreatic cancer using an orthotopic SCID mouse model. Br J
Cancer. 97:1432–1440. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Garzon R, Calin GA and Croce CM: MicroRNAs
in cancer. Annu Rev Med. 60:167–179. 2009. View Article : Google Scholar
|
|
4
|
Iorio MV and Croce CM: MicroRNAs in
cancer: small molecules with a huge impact. J Clin Oncol.
27:5848–5856. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Croce CM and Calin GA: miRNAs, cancer, and
stem cell division. Cell. 122:6–7. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Gregory RI and Shiekhattar R: MicroRNA
biogenesis and cancer. Cancer Res. 65:3509–3512. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Shen J, Stass SA and Jiang F: MicroRNAs as
potential biomarkers in human solid tumors. Cancer Lett.
329:125–136. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Nana-Sinkam SP and Croce CM: Clinical
applications for microRNAs in cancer. Clin Pharmacol Ther.
93:98–104. 2013. View Article : Google Scholar
|
|
9
|
Wu W, Lin Z, Zhuang Z and Liang X:
Expression profile of mammalian microRNAs in endometrioid
adenocarcinoma. Eur J Cancer Prev. 18:50–55. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Rauhala HE, Jalava SE, Isotalo J, et al:
miR-193b is an epigenetically regulated putative tumor suppressor
in prostate cancer. Int J Cancer. 127:1363–1372. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Xu C, Liu S, Fu H, et al: MicroRNA-193b
regulates proliferation, migration and invasion in human
hepatocellular carcinoma cells. Eur J Cancer. 46:2828–2836. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Li XF, Yan PJ and Shao ZM: Downregulation
of miR-193b contributes to enhance urokinase-type plasminogen
activator (uPA) expression and tumor progression and invasion in
human breast cancer. Oncogene. 28:3937–3948. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Hu H, Li S, Liu J and Ni B: MicroRNA-193b
modulates proliferation, migration, and invasion of non-small cell
lung cancer cells. Acta Biochim Biophys Sin (Shanghai). 44:424–430.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ikeda Y, Tanji E, Makino N, Kawata S and
Furukawa T: MicroRNAs associated with mitogen-activated protein
kinase in human pancreatic cancer. Mol Cancer Res. 10:259–269.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lewis BP, Shih IH, Jones-Rhoades MW,
Bartel DP and Burge CB: Prediction of mammalian microRNA targets.
Cell. 115:787–98. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Griffiths-Jones S, Grocock RJ, van Dongen
S, Bateman A and Enright AJ: MiR-Base: microRNA sequences, targets
and gene nomenclature. Nucleic Acids Res. 34:D140–D144. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Krek A, Grün D, Poy MN, Wolf R, Rosenberg
L, Epstein EJ, MacMenamin P, da Piedade I, Gunsalus KC, Stoffel M
and Rajewsky N: Combinatorial microRNA target predictions. Nat
Genet. 37:495–500. 2005. View
Article : Google Scholar
|
|
18
|
Chen J, Zhang X, Lentz C, Abi-Daoud M,
Pare GC, Yang X, et al: miR-193b regulates mcl-1 in melanoma. Am J
Pathol. 176:2162–2168. 2011. View Article : Google Scholar
|
|
19
|
Tahiri A, Leivonen SK, Lüders T, et al:
Deregulation of cancer-related miRNAs is a common event in both
benign and malignant human breast tumors. Carcinogenesis. 35:76–85.
2014. View Article : Google Scholar
|
|
20
|
Zhou H, Wang K, Hu Z and Wen J: TGF-β1
alters microRNA profile in human gastric cancer cells. Chin J
Cancer Res. 25:102–111. 2013.
|
|
21
|
Cheung TH, Man KN, Yu MY, et al:
Dysregulated microRNAs in the pathogenesis and progression of
cervical neoplasm. Cell Cycle. 11:2876–2884. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Xie C, Jiang XH, Zhang JT, et al: CFTR
suppresses tumor progression through miR-193b targeting urokinase
plasminogen activator (uPA) in prostate cancer. Oncogene.
32:2282–2291. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Chen J, Feilotter HE, Pare GC, et al:
MicroRNA-193b represses cell proliferation and regulates cyclin D1
in melanoma. Am J Pathol. 176:2520–2529. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Rana S, Maples PB, Senzer N and Nemunaitis
J: Stathmin 1: a novel therapeutic target for anticancer activity.
Expert Rev Anticancer Ther. 8:1461–1470. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Belletti B and Baldassarre G: Stathmin: a
protein with many tasks. New biomarker and potential target in
cancer. Expert Opin Ther Targets. 15:1249–1266. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Mistry SJ and Atweh GF: Role of stathmin
in the regulation of the mitotic spindle: potential applications in
cancer therapy. Mt Sinai J Med. 69:299–304. 2002.PubMed/NCBI
|
|
27
|
Ke B, Wu LL, Liu N, Zhang RP, Wang CL and
Liang H: Overexpression of stathmin 1 is associated with poor
prognosis of patients with gastric cancer. Tumour Biol.
34:3137–3145. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Tamura K, Yoshie M, Miyajima E, Kano M and
Tachikawa E: Stathmin regulates hypoxia-inducible factor-1α
expression through the mammalian target of rapamycin pathway in
ovarian clear cell adenocarcinoma. ISRN Pharmacol. 2795932013.
View Article : Google Scholar
|
|
29
|
Jiang L, Chen Y, Chan CY, et al:
Down-regulation of stathmin is required for TGF-beta inducible
early gene 1 induced growth inhibition of pancreatic cancer cells.
Cancer Lett. 274:101–108. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Wang Y, Kuramitsu Y, Ueno T, et al:
Proteomic differential display identifies upregulated vinculin as a
possible biomarker of pancreatic cancer. Oncol Rep. 28:1845–1850.
2012.PubMed/NCBI
|
|
31
|
Reichel CA, Kanse SM and Krombach F: At
the interface of fibrinolysis and inflammation: the role of
urokinase-type plasminogen activator in the leukocyte extravasation
cascade. Trends Cardiovasc Med. 22:192–196. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Cantero D, Friess H, Deflorin J, et al:
Enhanced expression of urokinase plasminogen activator and its
receptor in pancreatic carcinoma. Br J Cancer. 75:388–395. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
He X, Zheng Z, Li J, et al: DJ-1 promotes
invasion and metastasis of pancreatic cancer cells by activating
SRC/ERK/uPA. Carcinogenesis. 33:555–562. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Saldanha RG, Xu N, Molloy MP, Veal DA and
Baker MS: Differential proteome expression associated with
urokinase plasminogen activator receptor (uPAR) suppression in
malignant epithelial cancer. J Proteome Res. 7:4792–4806. 2008.
View Article : Google Scholar
|