Introduction
Glioblastoma (GBM) is the most common and most
lethal type of brain tumor in adults (1). The current treatment for GBM involves
resection, followed by radiation and chemotherapy. Therefore, there
is an urgent requirement to elucidate the underlying molecular
mechanisms of GBM and identify novel molecular targets for the
treatment of this disease.
MicroRNAs (miRs) are endogenous non-coding small
RNAs, which suppress genes by triggering either mRNA degradation or
translational repression by binding to the 3′-untranslated region
(3′-UTR) of target mRNA (2–4).
miRNAs were confirmed to be involved in several processes,
including cell proliferation, apoptosis, invasion and
differentiation in various types of cancer (4,5). The
Wnt signaling pathway is important in tumorigenesis. The
low-density lipoprotein (LDL) receptor-related protein-6 (LRP6), a
novel member of the expanding LDL receptor family, is a cell
surface receptor and is critical in the activation of the canonical
Wnt/β-catenin signaling pathway (6–8). The
underlying mechanism of the tumor suppressive function of miR-513
as well as its target genes have remained to be elucidated. The
present study identified and investigated the targeted regulation
of LRP6 by miR-513c in GBM cells and assessed its function in cell
proliferation.
Materials and methods
Clinical specimens
Human GBM tissues (n=8) and normal brain tissues
were obtained from patients with GBM at Shandong Provincial
Hospital affiliated to Shandong University (Jinan, China). The
present study was approved by the ethics committee of the Shandong
Provincial Hospital Affiliated to Shandong University, Shandong
University (Jinan, China) and written informed consent was obtained
from all patients. Tissue samples were collected at surgery,
immediately frozen in liquid nitrogen and stored until the total
RNAs or proteins were extracted.
Cell culture
The human GBM cell lines LN319, SNB19, LN18, U251MG,
U118MG, T98G, U373MG and LN382T were purchased from the American
Type Culture Collection (Manassas, VA, USA) and grown in Dulbecco's
modified Eagle's medium (Gibco-BRL, Invitrogen Life Technologies,
Carlsbad, CA, USA), supplemented with 10% fetal bovine serum (FBS;
Sigma-Alrich, St. Louis, MO, USA) and 100 units/ml
penicillin-streptomycin (Invitrogen Life Technologies). Normal
human astrocytes (NHAs) were obtained from Lonza (Walkersville, MD,
USA) and cultured in the provided astrocyte growth media,
supplemented with recombinant human epidermal growth factor
(Robustnique Corporation Ltd., Tianjin, China), insulin
(Sigma-Aldrich), ascorbic acid (Sigma-Aldrich), GA-1000
(Sigma-Aldrich), L-glutamine (Sigma-Aldrich) and 5% FBS. All cell
lines were cultured in a humidified incubator at 37°C in an
atmosphere of 5% CO2 and 95% air.
Plasmids, small interfering (si)RNA and
transfection
For ectopic expression of LRP6, the LRP6 open
reading frame with a 3′-UTR was amplified using polymerase chain
reaction (PCR) and subcloned into p-enhanced green fluorescent
protein-N3 (Invitrogen Life Technologies). To construct a
luciferase reporter vector, the LRP6 3′-UTR fragment, containing
putative binding sites for miR-513c, was amplified using PCR and
cloned downstream of the luciferase gene in the pGL3-luciferase
reporter plasmid (Promega Corporation, Madison, WI, USA). The
constructs were verified by sequencing. The miR-513c mimics
(HmiR0671), negative control (NC) and miR-513c inhibitor
(HmiR-AN0568) were purchased from Genecopoeia Co. (Rockville, MD,
USA) and transfected into glioblastoma cells using Lipofectamine
2000 reagent (Invitrogen Life Technologies) according to the
manufacturer's instructions.
For LRP6 depletion, small interfering (si)RNA was
synthesized and purified by RiboBio Co., Ltd. (Guangzhou,
Guangdong, China). The LRP6-siRNA sequences used were as follows:
LRP6-siRNA#1, 5′-CCGATGCAATGGAGATGCAAA-3′ and
LRP6-siRNA#2, 5′-CGCACTACATTAGTTCCAAAT-3′. The
transfection of siRNAs was performed using Lipofectamine 2000
(Invitrogen Life Technologies) according to the manufacturer's
instructions.
RNA extraction and reverse transcription
quantitative (RT-q)PCR
For miRNA quantification, the total RNA, including
miR, was extracted from cultured cells and patient samples using
the mirVana miRNA Isolation kit (Ambion, Austin, TX, USA),
according to the manufacturer's instructions. The cDNA was
synthesized from 5 ng total RNA using the Taqman miRNA reverse
transcription kit (Applied Biosystems, Thermo Fisher Scientific,
Waltham, MA, USA). The expression levels of miR-513c were
quantified using the miRNA-specific TaqMan miRNA Assay kit (Applied
Biosystems). The relative expression levels of miR-513c following
normalization to U6 small nuclear RNA were calculated using
2−[Ct(miR-513c)−Ct(U6)]. qPCR was performed using an
SYBR kit (Qiagen, Shanghai, China) on a Light Cycler system
(LightCycler480; Roche Diagnostics, Basel, Switzerland). The
primers selected were as follows: LRP6 forward,
5′-CCTCAATGCGATTTGTTCC-3′ and reverse, 5′-GGTGTCAAAGAAGCCTCTGC-3′;
cyclin D1 forward, 5′-TCCTCTCCAAAATGCCAGAG-3′ and reverse,
5′-GGCGGATTGGAAATGAACTT-3′; Myc forward, 5′-TCAAGAGGCGAACACACAAC-3′
and reverse, 5′-GGCCTTTTCATTGTTTTCCA-3′; and LEF1 forward,
5′-CACTGTAAGTGATGAGGGGG-3′ and reverse,
5′-TGGATCTCTTTCTCCACCCA-3′.
The expression was normalized against the geometric
mean of GAPDH to control the variability in expression levels
(GAPDH forward, 5′-GACTCATGACCACAGTCCATGC-3′ and reverse,
3′-AGAGGCAGGGATGATGTTCTG-5′) and was calculated as
2−[Ct(LRP6/cyclinD1/Myc/LEF1)−Ct(GAPDH)].
MTT and colony formation assays
Cell growth was measured using an MTT assay. The
SNB19 cells were seeded into 96-well plates in medium containing
10% FBS at 3,000 cells/well. For quantitation of cell viability,
the cultures were stained after 1, 2, 3 and 4 days with MTT.
Briefly, 20 μl 5 mg/ml MTT solution (Sigma-Aldrich) was
added to each well followed by incubation at 37°C for 4 h. The
medium was then removed from each well and the resulting MTT
formazan was solubilized in 150 μl dimethylsulfoxide
(Sigma-Aldrich). The absorbance at 490 nm was measured using a
Multiskan plate reader (Thermo Fisher Scientific).
For colony formation assays, the SNB19 cells were
seeded into three 6-cm cell culture dishes (1×103
cells/well) and incubated for 10 days in medium containing 10% FBS.
The colonies were washed with phosphate-buffered saline and stained
with 1% crystal violet for 30 sec following fixation with 10%
formaldehyde (Beyotime Institute of Biotechnology, Shanghai, China)
for 15 min. The number of colonies, defined as >50 cells/colony,
were counted. Three independent experiments were performed and the
results were statistically analyzed using a paired t-test.
Anchorage-independent growth assay
The cells were trypsinized and 1,000 cells were
re-suspended in 2 ml complete medium containing 0.3% agar
(Sigma-Aldrich). The agar-cell mixture was seeded on top of a
bottom layer, consisting of 1% agar in complete medium. The plates
were incubated at 37°C in a humid atmosphere of 5% CO2
for 2 weeks until colony formation. The colonies were stained with
0.5% crystal violet for counting colony number under an inverted
fluorescence microscope (Motic AE30; Microscope Systems Ltd.,
Glasgow, UK) and images of cell colonies were captured at
magnification of x100. Only cell colonies containing >50 cells
were counted. The experiment was performed three independent times
for each cell line.
Luciferase assays
The cells were seeded in triplicate in 24-well
plates (5×104/well) and cultured for 24 h. The
pGL3-luciferase reporter gene plasmid, pGL3-LRP6-3′-UTR, or the
control-luciferase plasmid were co-transfected into the cells with
the control pRL-TK Renilla plasmid (Promega Corporation) using
Lipofectamine 2000 reagent (Invitrogen Life Technologies).
Luciferase and Renilla activities were assayed 48 h following the
transfection by lysing the cells with Cell Lysis solution (Beyotime
Institute of Biotechnology) and detecting the fluorescence
intensity using the dual luciferase assay kit.
Western blot analysis
The protein lysates were prepared and equal
quantities (40 μg) of protein were subjected to 10%
SDS-PAGE. The proteins were transferred onto polyvinylidene
fluoride membranes for 2 h at 4°C, at a current of 125 mA. The
membranes were blocked with 5% non-fat milk (Beyotime Institute of
Biotechnology) for 2 h and were subsequently incubated overnight
with anti-LRP6 (#3395), anti-cyclin D1 (#2978), anti-C-Myc (#5605),
anti-LEF1 (#2230) (1:1,000; Cell Signaling Technology, Inc.,
Danvers, MA, USA), anti-p84 (ab487; 1:2,000; Abcam, Cambridge, UK)
and α-tubulin (1:2,000; Sigma-Aldrich), which was used as a
reference protein. Following antibody incubation, the membranes
were washed with Tris-buffered saline containing Tween-20 (TBST;
Sigma-Aldrich) three times for 5 min each time. Following washing
with TBST and incubation with either anti-rabbit horseradish
peroxidase-conjugated secondary antibody (A0545; 1:5,000;
Sigma-Aldrich) for 2 h at room temperature, immunocomplexes were
visualized using the ECL Advanced Western Blotting Detection kit
(GE Healthcare, Little Chalfont, UK) according to the
manufacturer's instructions.
Statistical analysis
All statistical analyses were performed using SPSS
19.0 (International Business Machines, Armonk, NY, USA). Student's
t-test was used to determine the significance of the differences
between the two groups of data in all the pertinent experiments.
P<0.05 was considered to indicate a statistically significantly
difference.
Results
Expression of miR-513c is downregulated
in GBM tissues and cell lines
To investigate the role of miR-513c in the
development of GBM, the expression levels of miR-513c in GBM
tissues and GBM cell lines were assessed by RT-qPCR (Fig. 1A). The results demonstrated that
the expression levels of miR-513c were consistently lower in the
GBM tissues compared with those in the normal brain tissues. In
addition, in all eight GBM cell lines, significantly lower
expression levels of miR-513c were observed compared with those in
the NHAs. These results suggested that miR-513c is significantly
reduced in GBM and may serve as a prognostic marker for patients
with GBM.
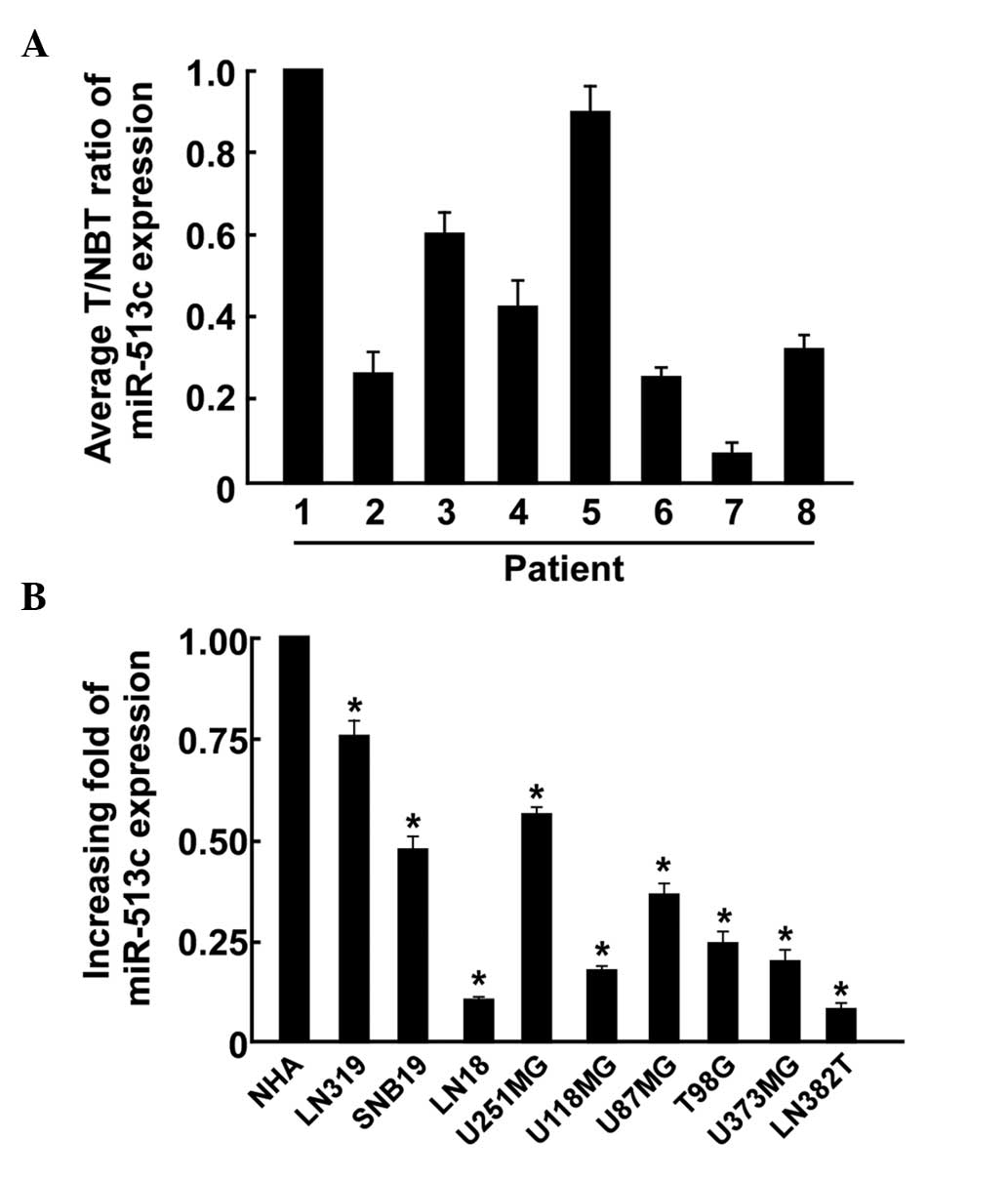 | Figure 1Expression of miR-513c in human GBM
tissues and cell lines. (A) Relative expression levels of miR-513c
in eight paired primary GBM tissues and the matched normal brain
tissue from patients were detected by RT-qPCR analysis. (B) RT-qPCR
analysis of the expression of miR-513c in GBM cell lines, including
LN319, SNB19, LN18, U251MG, U118MG, T98G, U373MG and LN382T. All
experiments were repeated at least three times and values are
expressed as the mean of three independent experiments
(*P<0.05 vs. NHA). GBM, glioblastoma; RT-qPCR,
reverse transcription quantitative polymerase chain reaction;
T/NBT, tumor/normal brain tissue; miR, microRNA; NHA, normal human
astrocytes. |
miR-513c inhibits GBM cell
proliferation
Since miR-513c was significantly downregulated in
GBM tissues and cell lines, the present study investigated whether
miR-513c exhibits a tumor suppressive role in the cell
proliferation of GBM. SNB19 cells were transfected with miR-513c
mimics, miR-513c inhibitor or the respective controls. The relative
expression levels of miR-513c were confirmed using RT-qPCR
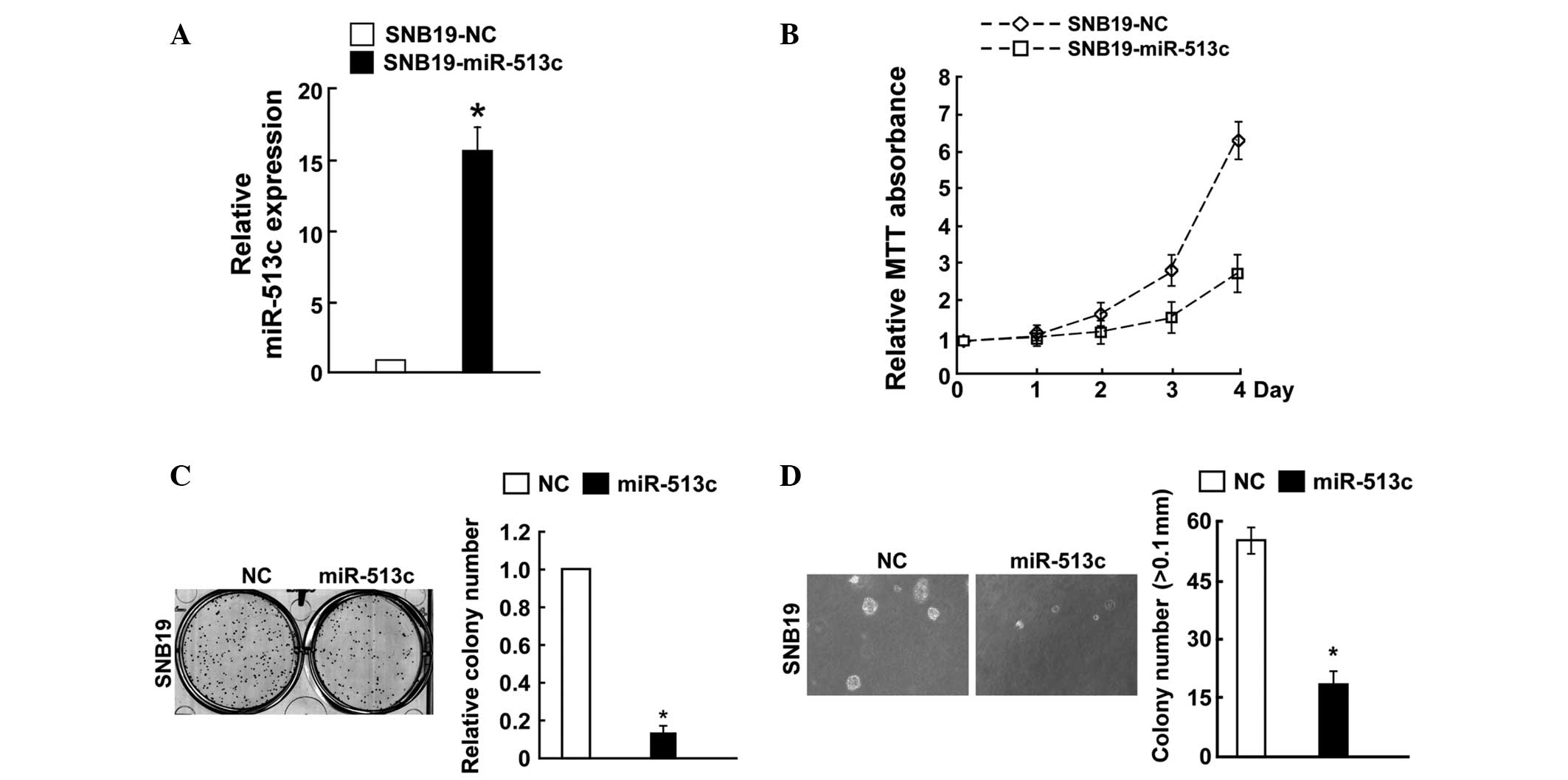
(Figs. 2A and 3A). The effect of miR-513c on cell
proliferation was determined using MTT, colony formation and
anchorage-independent growth assays. To investigate the role of
upregulation of miR-513c in the development and progression of GBM,
its effect on cellular proliferation was assessed. An MTT assay
demonstrated that miR-513c-transduced SNB19 cells exhibited
significantly lower growth rates compared with those in the control
cells at day four after seeding (Fig.
2B). The colony formation assays consistently revealed that
enforced expression of miR-513c significantly reduced the number of
colonies of SNB19 cells following 14 days of culture compared with
that in the control group (Fig.
2C). Of note, it was demonstrated that enforced expression of
miR-513c in SNB19 cells significantly decreased their
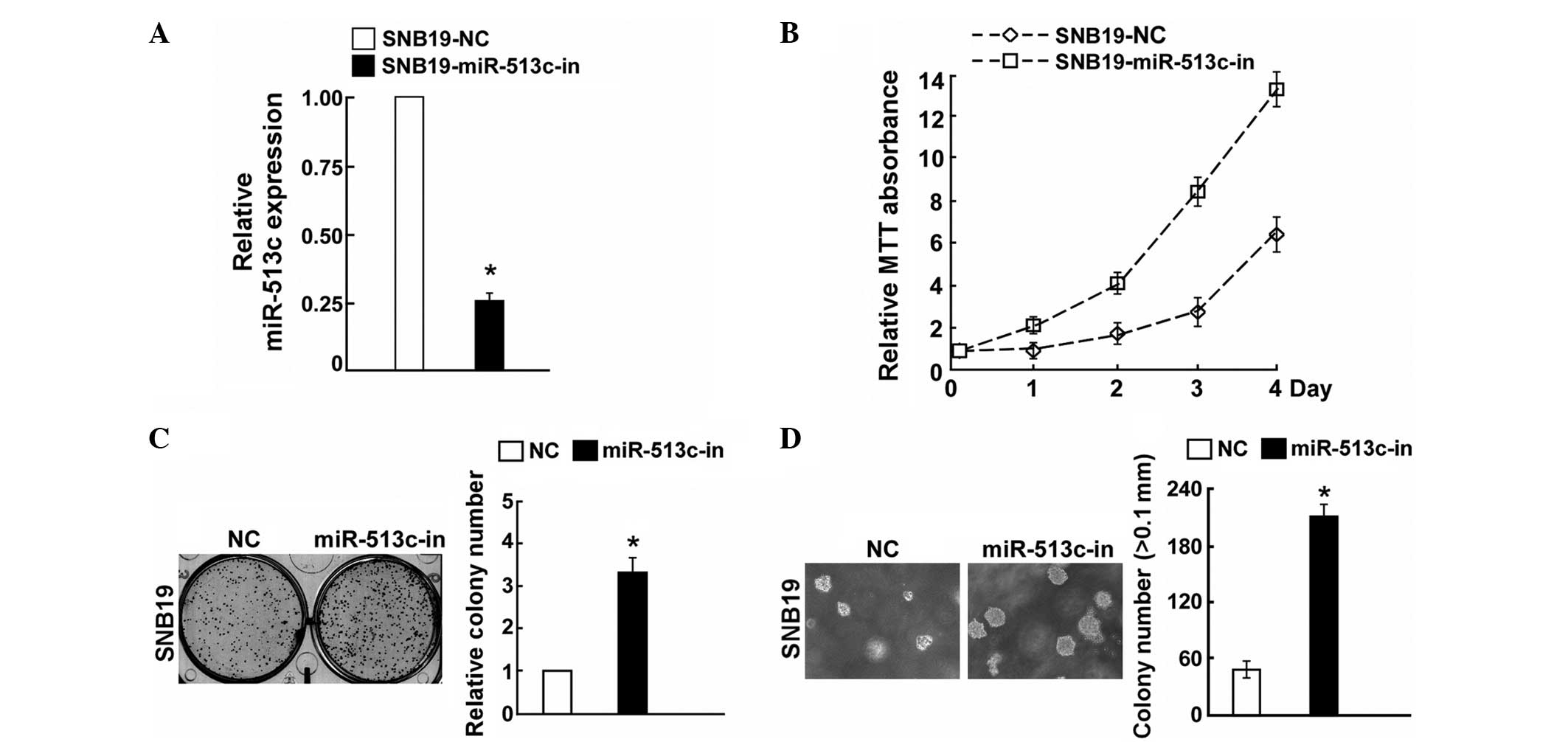
anchorage-independent growth ability (Fig. 2D). By contrast, the cell growth
rates and colony numbers of SNB19 cells transfected with the
miR-513c inhibitor (miR-513c-in) were significantly higher compared
with those transfected with the NC (Fig. 3B and C). In addition, the
anchorage-independent growth ability of SNB19 cells was
significantly increased in following transfection with miR-513c-in
(Fig. 3D). These results
demonstrated that miR-513c reduced GBM cell tumorigenicity in
vitro.
miR-513c directly targets LRP6 by binding
to its 3′-UTR
It is generally accepted that miRNAs exert their
functions by regulating the gene expression of their downstream
target genes (2–4). Bioinformatic analysis was used to
search for the potential regulatory targets of miR-513c. LRP6, a
tumor suppressor gene containing a binding site for miR-513c, was
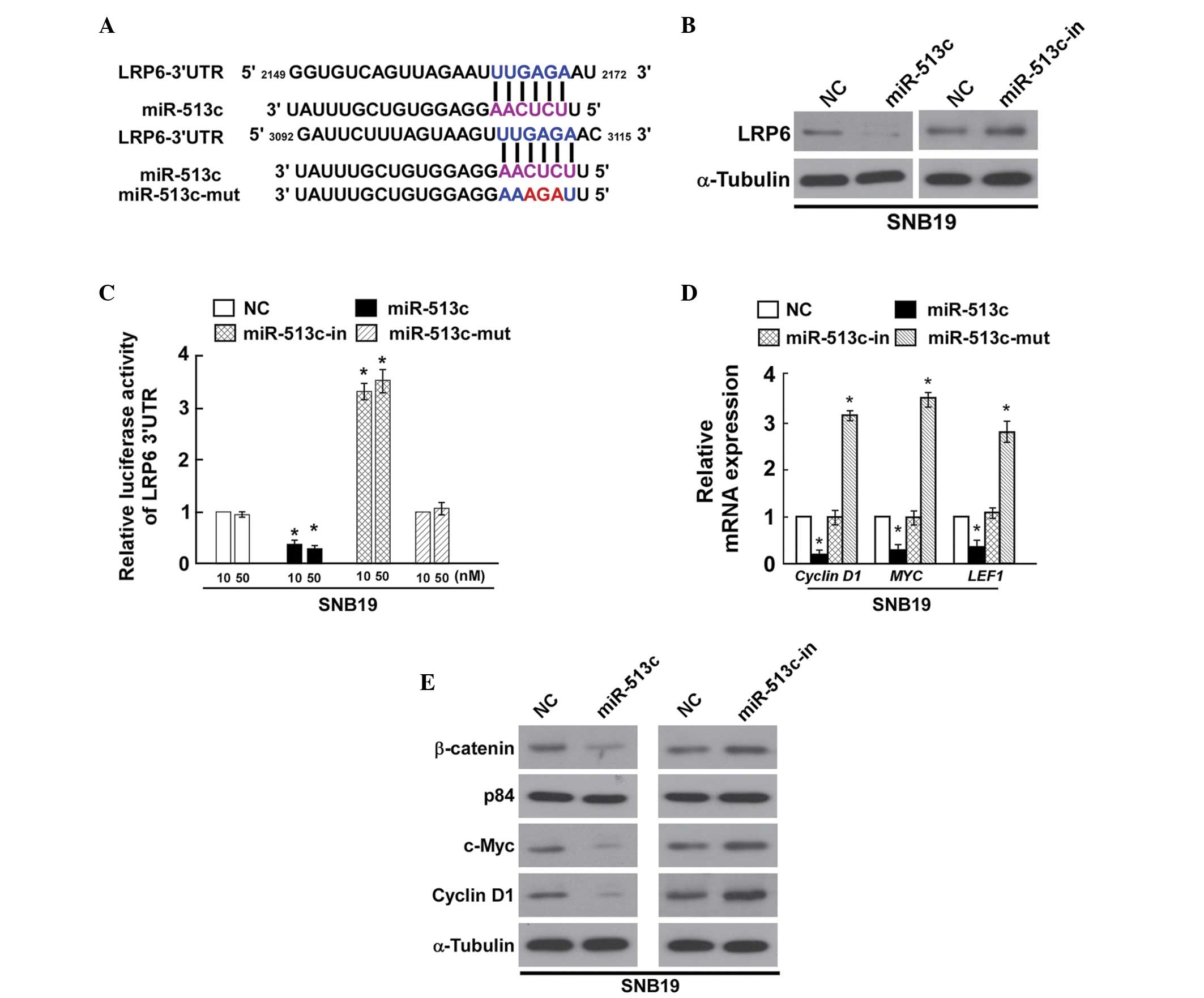
selected as the target gene for further analysis (Fig. 4A).
To determine whether miR-513c affected the
expression of LRP6, the expression levels of LRP6 were detected in
the SNB19 cells, which were transfected with miR-513c mimics,
miR-513c-in or the respective controls. Western blot analysis
demonstrated that the miR-513c mimics markedly suppressed the
protein expression of LRP6 in SNB19 cells (Fig. 4B), while miR-513c-in clearly
promoted the protein expression of LRP6. To confirm the effect of
miR-513c on the inhibition of the expression of LRP6, the present
study assessed whether LRP6 was regulated by miR-513c through
direct binding to its 3′UTR. A LRP6 3′-UTR wild-type vector was
co-transfected into SNB19 cells with either the miR-513c mimic,
miR-513c-in, miR-513c-mut or miR-NC, followed by measurement of
luciferase activity. As shown in Fig.
4C, a consistent and dose-dependent reduction in the luciferase
activity was observed in SNB19 cells transfected with the miR-513c
mimic, whereas LRP6 luciferase activity was increased in the
presence of miR-513c inhibitor. In addition, overexpression of
miR-513c-mut had no significant effect on the luciferase activity
of the LRP6 3′-UTR. These results demonstrated that LRP6 is a bona
fide target of miR-513c.
miR-513c alters the levels of proteins
associated with proliferation in GBM cells
The expression levels of a number of critical
cell-proliferation regulators were also detected. It was previously
reported that LRP6 is closely correlated with Wnt/β catenin
signaling activity (6–8). The present study assessed the mRNA
expression levels of the downstream genes in the Wnt/β-catenin
signaling pathway. The downstream genes regulated by Wnt/β-catenin,
cyclin D1, Myc and LEF1 were all significantly downregulated by
miR-513c and upregulated by the miR-513c inhibitor (Fig. 4D). Furthermore, the protein
expression levels of β-catenin, c-Myc and cyclin D1 were reduced in
SNB19 cells transfected with the miR-513c mimic, however, their
expression was upregulated in the cells transfected with
miR-513c-in compared with that in the control cells (Fig. 4E). These findings provided further
evidence that miR-513c is important in GBM cell proliferation.
These results indicated that miR-513c functionally modulates the
cellular proliferation regulators c-Myc, LEF-1, cyclin D1 and
β-catenin, and is therefore relevant to cell proliferation.
LRP6 upregulation is required for
miR-513c-in-induced proliferation of GBM cells
To further investigate the role of LRP6 promotion in
miR-513c-in-induced GBM proliferation, the effects of
downregulation of LRP6 on GBM cell proliferation were assessed. As
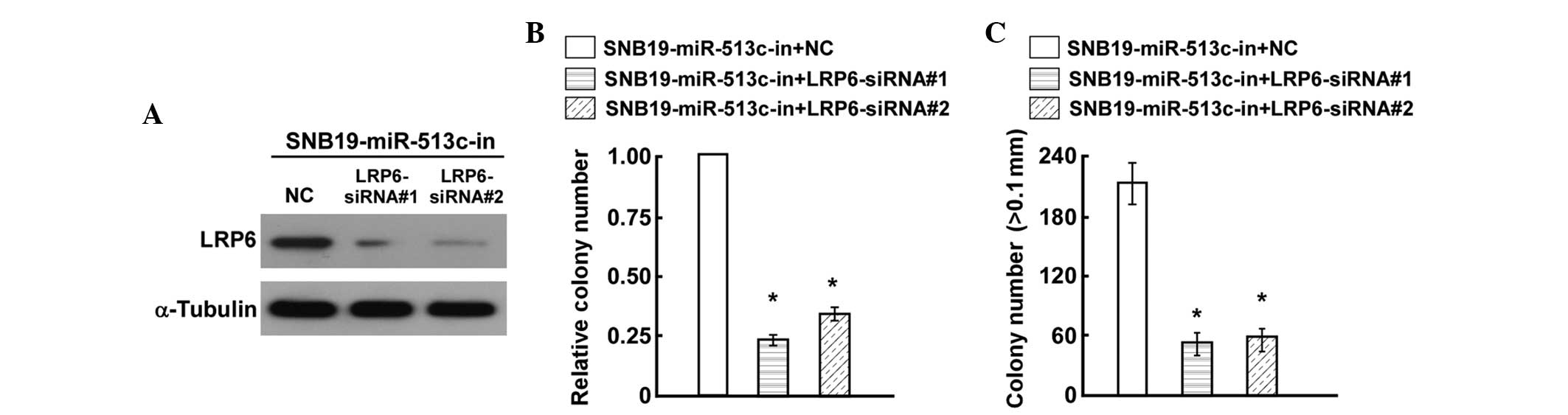
predicted, western blot analysis confirmed that
LRP6-siRNA#1 and LRP6-siRNA#2 effectively
decreased the expression levels of LRP6 in miR-513c-in-transfected
SNB19 cells (Fig. 5A). Colony
formation and anchorage-independent growth assays revealed that
miR-513c-in in the LRP6-silenced cells had no additive effect on
cell proliferation (Fig. 5B and
C). These results demonstrated that direct upregulation of LRP6
was required for miR-513-in-induced GBM cell proliferation.
Discussion
miRNAs are a class of small (22 nucleotides),
non-coding, highly conserved single-stranded RNAs with
post-transcriptional regulatory features, and are important in
multiple biological processes, including the regulation of cell
proliferation, differentiation, survival and apoptosis (3,9,10).
The present study demonstrated that the expression levels of
miR-513c were significantly downregulated in GBM tissues and GBM
cells compared with those in the NHAs and normal brain tissues.
Furthermore, ectopic expression of miR-513c decreased the growth
rate and anchorage-independent growth of GBM cells, whereas
miR-513c-in caused the opposite effect, indicating that miRNA-513c
may function as a tumor suppressor miRNA. Through bioinformatic
analysis, the LRP6 tumor promoter gene was identified as a
theoretical miRNA-513c target gene. Western blot analysis
demonstrated that ectopic miR-513c expression markedly reduced the
protein expression levels of LRP6, whereas miR-513c-in clearly
promoted the protein expression of LRP6. In addition, a luciferase
reporter assay demonstrated that the downregulation of LRP6 was
mediated by miR-513c through the LRP6 3′-UTR. These results
demonstrated that LRP6 is a bona fide target of miR-513c.
LRP6 is a novel member of the expanding LDL receptor
family, functions as an indispensable co-receptor and is critical
in the activation of the canonical Wnt/β-catenin signaling pathway
(11–15). Wnt/β-catenin signaling controls
fundamental cellular processes and aberrant activation of this
pathway is implicated in several types of human cancer (16–18).
β-catenin is one of the major cellular effectors of the Wnt
signaling pathway, which is essential in development and tissue
maintenance (19). It is well
known that the β-catenin signaling pathway is essential in the
pathogenesis of various types of human cancer (16,20,21).
The present study aimed to further investigate the molecular
mechanisms of growth inhibition induced by miR-513c. The expression
of β-catenin was decreased in SNB19 cells transfected with the
miR-513c mimic; however, was upregulated in the cells transfected
with miR-513c-in. Furthermore, the expression levels of β-catenin
downstream genes, including cyclin D1, Myc and LEF-1, were
determined. CyclinD1, Myc and LEF-1 have been identified as target
genes of the Wnt/β-catenin signaling pathway, exerting a pivotal
role in human cancer carcinogenesis (22–26).
The present study demonstrated that the mRNA expression levels of
these molecules were all significantly suppressed, suggesting that
miR-513c may be an important regulator of this signaling pathway.
Consistent with these observations, the protein expression levels
of cyclin D1 and c-Myc were markedly upregulated following
transfection with the miR-513c inhibitor and downregulated
following overexpression of miR-513c. Furthermore, inhibition of
miR-513c in the LRP6-silenced cells revealed no additive effect on
cell proliferation, suggesting that direct upregulation of LRP6 was
required for miR-513c-in-induced GBM cell proliferation.
In conclusion, the present study revealed that
miR-513c is a tumor suppressor miRNA in GBM cells and that LRP6 is
a direct target of miRNA-513c. Therefore, these findings indicated
that miR-513c may serve as a potential therapeutic target for the
treatment of GBM.
Acknowledgments
This study was supported by the Shandong Province
Young and Middle-aged Scientists Research Awards Fund (no.
BS200403017).
References
|
1
|
Wen PY and Kesari S: Malignant gliomas in
adults. N Engl J Med. 359:492–507. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Baek D, Villén J, Shin C, Camargo FD, Gygi
SP and Bartel DP: The impact of microRNAs on protein output.
Nature. 455:64–71. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Miska EA: How microRNAs control cell
division, differentiation and death. Curr Opin Genet Dev.
15:563–568. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ambros V: The functions of animal
microRNAs. Nature. 431:350–355. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Hsu SY, Kudo M, Chen T, et al: The three
subfamilies of leucine-rich repeat-containing G protein-coupled
receptors (LGR): Identification of LGR6 and LGR7 and the signaling
mechanism for LGR7. Mol Endocrinol. 14:1257–1271. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Li Y, Lu W, He X, Schwartz AL and Bu G:
LRP6 expression promotes cancer cell proliferation and
tumorigenesis by altering beta-catenin subcellular distribution.
Oncogene. 23:9129–9135. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lu W, Lin C, King TD, Chen H, Reynolds RC
and Li Y: Silibinin inhibits Wnt/beta-catenin signaling by
suppressing Wnt coreceptor LRP6 expression in human prostate and
breast cancer cells. Cell Signal. 24:2291–2296. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Formosa A, Markert EK, Lena AM, et al:
MicroRNAs, miR-154, miR-299-5p, miR-376a, miR-376c, miR-377,
miR-381, miR-487b, miR-485-3p, miR-495 and miR-654-3p, mapped to
the 14q32.31 locus, regulate proliferation, apoptosis, migration
and invasion in metastatic prostate cancer cells. Oncogene.
33:5173–5182. 2014. View Article : Google Scholar
|
|
10
|
Lehmann U, Streichert T, Otto B, et al:
Identification of differentially expressed microRNAs in human male
breast cancer. BMC Cancer. 10:1092010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Gong X, Carmon KS, Lin Q, Thomas A, Yi J
and Liu Q: LGR6 is a high affinity receptor of R-spondins and
potentially functions as a tumor suppressor. PLoS One.
7:e371372012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gray JD, Kholmanskikh S, Castaldo BS, et
al: LRP6 exerts non-canonical effects on Wnt signaling during
neural tube closure. Hum Mol Genet. 22:4267–4281. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Joiner DM, Ke J, Zhong Z, Xu HE and
Williams BO: LRP5 and LRP6 in development and disease. Trends
Endocrinol Metab. 24:31–39. 2013. View Article : Google Scholar :
|
|
14
|
Maletínská L, Blakely EA, Bjornstad KA,
Deen DF, Knoff LJ and Forte TM: Human glioblastoma cell lines:
levels of low-density lipoprotein receptor and low-density
lipoprotein receptor-related protein. Cancer Res. 60:2300–2303.
2000.PubMed/NCBI
|
|
15
|
Semënov MV, Zhang X and He X: DKK1
antagonizes Wnt signaling without promotion of LRP6 internalization
and degradation. J Biol Chem. 283:21427–21432. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ochoa-Hernández AB, Juárez-Vázquez CI,
Rosales-Reynoso MA and Barros-Núñez P: WNT-β-catenin signaling
pathway and its relationship with cancer. Cir Cir. 80:389–398.
2012.In Spanish.
|
|
17
|
Kim KH, Seol HJ, Kim EH, et al:
Wnt/β-catenin signaling is a key downstream mediator of MET
signaling in glioblastoma stem cells. Neuro Oncol. 15:161–171.
2013. View Article : Google Scholar :
|
|
18
|
Hsieh IS, Chang KC, Tsai YT, et al:
MicroRNA-320 suppresses the stem cell-like characteristics of
prostate cancer cells by downregulating the Wnt/beta-catenin
signaling pathway. Carcinogenesis. 34:530–538. 2013. View Article : Google Scholar
|
|
19
|
Fu Y, Zheng S, An N, et al: β-catenin as a
potential key target for tumor suppression. Int J Cancer.
129:1541–1551. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chocarro-Calvo A, García-Martínez JM,
Ardila-González S, De la Vieja A and García-Jiménez C:
Glucose-induced β-catenin acetylation enhances Wnt signaling in
cancer. Mol Cell. 49:474–486. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang D, Wise ML, Li F and Dey M:
Phytochemicals attenuating aberrant activation of β-catenin in
cancer cells. PLoS One. 7:e505082012. View Article : Google Scholar
|
|
22
|
He TC, Sparks AB, Rago C, et al:
Identification of c-MYC as a target of the APC pathway. Science.
281:1509–1512. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kolligs FT, Bommer G and Göke B:
Wnt/beta-catenin/tcf signaling: a critical pathway in
gastrointestinal tumorigenesis. Digestion. 66:131–144. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Nagayama M, Iwamoto M, Hargett A, et al:
Wnt/beta-catenin signaling regulates cranial base development and
growth. J Dent Res. 87:244–249. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Huang FI, Chen YL, Chang CN, Yuan RH and
Jeng YM: Hepatocyte growth factor activates Wnt pathway by
transcriptional activation of LEF1 to facilitate tumor invasion.
Carcinogenesis. 33:1142–1148. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Planutiene M, Planutis K and Holcombe RF:
Lymphoid enhancer-binding factor 1, a representative of
vertebrate-specific Lef1/Tcf1 sub-family, is a Wnt-beta-catenin
pathway target gene in human endothelial cells which regulates
matrix metal-loproteinase-2 expression and promotes endothelial
cell invasion. Vasc Cell. 3:282011. View Article : Google Scholar
|