Introduction
Autophagy is a conserved and important metabolic
pathway in eukaryotic cells. Under starvation or stress conditions,
harmful and unnecessary intracellular macromolecules are degraded
and recycled through autophagy, which thereby promotes cell
survival and growth (1). In
populations of undifferentiated cells as well as cells undergoing
self-renewal or differentiation into mature cells, the cells
exhibit protein renewal and organelle degradation (2). Autophagy, which is stimulated by the
macroenvironment or cytokines, has an important role in remodeling
stem cells, thus regulating cell differentiation and self-renewal.
However, the role of autophagy in bone marrow-derived mesenchymal
stem cell (BMSC) proliferation and differentiation has remained
elusive.
Notch1 signaling is a regulator which is widely
distributed in various organs of developing embryos (3), a variety of adult organs as well as
undifferentiated cells with the ability to proliferate, and which
has important roles in the development, proliferation and
differentiation of various cell types (4–6).
Notch1 signaling has been shown to have important roles in BMSC
proliferation and differentiation, where it exerts an activating
and inhibitory function, respectively (7). Furthermore, induction of autophagy is
associated with the Notch1 signaling pathway. In order to
investigate the role of autophagy in BMSC proliferation and
differentiation, as well as the underlying molecular mechanisms,
the present study used autophagy inducer rapamycin and autophagy
inhibitors 3-methyladenine (3-MA) and chloroquine, to modulate
autophagic activity in BMSCs. The effects of the modified
autophagic activity on BMSC proliferation, apoptosis and
differentiation into neurons were examined, and the role of the
Notch1 signaling pathway in this process was investigated.
Materials and methods
Animals
Adult male Sprague-Dawley rats (n=30; age, four
weeks, weight, 160–200 g) were obtained from the Animal Center of
Zhengzhou University (Zhengzhou, China) and housed in a specific
pathogen-free room. Animal procedures were approved by the Ethics
Committee of Zhengzhou University (Zhengzhou, China) and performed
according to the 'Guidance On the Care and Use of Laboratory
Animals' (People's Republic of China National Science and
Technology Committee; 2006). BCMCs were obtained three times from
10 rats.
Reagents
The reagents used in the present study were as
follows: 0.25% Trypsin-EDTA (1X), Dulbecco's modified Eagle's
medium (DMEM), fetal bovine serum (FBS) (all from Gibco; Thermo
Fisher Scientific, Waltham, MA, USA), radioimmunoprecipitation
assay (RIPA) lysis buffer, phenylmethanesulfonyl fluoride (PMSF),
bicinchoninic acid (BCA) protein assay kit (cat. no. P0010s), and
Electro-Chemi-Luminescence Plus kit (cat. no. P0018; all from
Beyotime Institute of Biotechnology, Haimen, China), Tris, glycine
and SDS (all from Beijing Solarbio Science & Technology Co.,
Ltd., Beijing, China), Temed (Shanghai Huashuo Fine Chemical Co.,
Ltd., Shanghai, China), ammonium persulfate (Tianjin Kemiou
Chemical Reagent Co., Ltd., Tianjin, China), rapamycin, 3-MA,
chloroquine and dimethylsulfoxide (DMSO) (all from Sigma-Aldrich,
St. Louis, MO, USA), rabbit polyclonal anti-Notch1, rabbit
polyclonal anti-Hes1, rabbit monoclonal anti-P62, and rabbit
polyclonal anti-microtubule-associated protein 2 (MAP2) antibodies
(All from Abcam, Cambridge, MA, USA; cat. nos. ab33932, ab49170,
ab109012, and ab32454; 1:1,000), rabbit-anti polyclonal
microtubule-associated protein 1 light chain 3B (LC3B), and
rabbit-anti polyclonal β-actin antibodies (Cell Signaling
Technology, Inc., Danvers, MA, USA; cat. nos. 3868 and 4967;
1:1,000), and horseradish peroxidase-labeled goat anti-rabbit
secondary antibodies (Santa Cruz Biotechnology, Inc.; cat. no.
SC-2004; 1:2,000).
Isolation, purification and induction of
BMSCs
Following sacrificing the rats by cervical
dislocation, bone marrow cells were collected by repeatedly
flushing both femurs and tibias with culture medium using a 5-ml
syringe under the sterile/cell culture hood. Cells at
1×106/ml were seeded into culture dishes (75
cm2) and cultured in a humidified incubator containing
5% CO2 at 37°C, and the medium was changed every 3–4
days. 6–7 days later, cells were re-seeded at the ratio of 1:2.
Cells in the third passage were randomly divided
into four groups: The control group and three groups treated with
rapamycin (10 mmol/l), 3-MA (3 mmol/l) and chloroquine (25 mmol/l),
respectively, for 12 h. Experiments were performed in triplicate.
For the experiments on cell differentiation, cells were divided
into the same groups as mentioned above, but cultured with
induction medium [low-glucose DMEM (Gibco; Thermo Fisher
Scientific, Inc.) containing 2% DMSO and 200 µmol/l
butylated Hydroxyanisole (BHA; Sigma-Aldrich)] for 12 h (8).
Flow cytometry
Cell cycle analysis
After trypsinization and washing in
phosphate-buffered saline (PBS) twice, cells were re-suspended with
200 µl cold PBS and fixed with 800 µl chilled
ethanol, which was added drop wise with agitation. The suspensions
were stored at −20°C overnight, followed by centrifugation (179 ×
g, 5 min, room temperature), re-suspension of the cell pellet in
propidium iodide (PI; BD Biosciences, Erembodegem, Belgium)/RNase
staining buffer (200 µl; BD Biosciences, Franklin Lakes, NJ,
USA) and incubation in the dark for 15 min at room temperature. The
cell cycle was then analyzed by flow cytometry.
Assessment of apoptosis
Cell treatments were conducted according to the
protocol of the fluorescein isothiocyanate (FITC)-Annexin V
Apoptosis Detection kit (BD Biosciences; cat. no. 556547). After
trypsinization and washing in PBS twice, cells were re-suspended in
100 µl 1X Annexin V binding buffer, 5 µl FITC-Annexin
V and 5 µl PI (all from BD Biosciences), and incubated in
the dark for 15 min after gentle mixing. Following addition of 200
µl 1X Annexin V binding buffer, apoptosis was analyzed by
flow cytometry (BD FACSCanto II; BD Biosciences) within 1 h.
Analysis of neuron-specific markers
neuron-specific enolase (NSE) and Notch1
Cells were fixed and permeabilized using the BD
cytofix/cytoperm kit (cat. no. 554714; BD Biosciences, Franklin
Lakes, NJ, USA) according to the manufacturer's instructions. In
brief, 500 µl Fixation/Permeabilization solution was added
and cells were incubated in the dark for 20 min at room
temperature. After centrifugation (179 × g, 5 min, room
temperature), cell pellets were re-suspended in 2 ml BD Perm/wash
buffer and incubated in the dark for 10 min at room temperature.
Subsequently, phycoerythrin (PE)-conjugated mouse IgG anti-Notch1
antibodies (1 µl; BD Biosciences; cat. no. 552768; 1:100) or
rabbit polyclonal anti-NSE (Abcam; cat. no. ab53025; 1:500) were
added, followed by incubation for 30 min at room temperature in the
dark or 2 h on ice in the dark, respectively. After the cells were
washed using BD Perm/wash buffer, the cells incubated with
anti-Notch1 antibodies were re-suspended in 300 µl PBS
containing 1% paraformaldehyde and stored at 4°C, whereas those
incubated with anti-NSE antibodies were treated with PE-conjugated
rabbit IgG antibodies (BD Biosciences; cat. no. 558416; 1:1,000)
for 1 h on ice in the dark. The cells were then washed and
re-suspended as above. The ratio of NSE-positive cells and the mean
fluorescence intensity (MFI) of Notch1 were analyzed using flow
cytometry (BD FACSCanto II) with a Data-Interpolating Vibrational
Analysis software version 5.0 (BD Biosciences) for data acquisition
and analysis.
Western blot analysis
Following cell lysis and protein extraction using
RIPA lysis buffer, the protein concentration was measured using a
BCA assay according to the instructions of the BCA kit. Protein in
loading buffer (40 µg) was resolved by 10% sodium dodecyl
polyacrylamide electrophoresis (using 80-V constant-voltage
electrophoresis until the bromophenol blue entered the gel,
followed by 120-V constant-voltage electrophoresis for 90 min;
Beijing Dingguo Changsheng Biotechnology Co., Ltd., Beijing, China)
and transferred onto a polyvinylidene difluoride membrane (250 mA
for 1 h; Beijing Dingguo Changsheng Biotechnology Co., Ltd.). After
blocking with 5% skimmed milk for 1 h at room temperature,
membranes were incubated with rabbit-anti LC3B, rabbit-anti MAP2,
rabbit-anti P62, rabbit anti-Notch1, rabbit anti-Hes1 (1:1,000) or
β-actin antibodies (1:2,000) overnight at 4°C. After washing with
Tris-buffered saline containing Tween 20 three times, membranes
were then incubated with horseradish peroxidase-conjugated goat
anti-rabbit antibody (1:2,000) for 1 h at room temperature. The
membranes were further washed three times for 5 min. Bands were
detected using the Electro-Chemi-Luminescence Plus kit. Grey value
analysis was performed using Quantity One software version 4.6.2
(Bio-Rad Laboratories, Inc., Hercules, CA, USA). β-actin was used
as a loading control, and the expression levels of the proteins
were normalized to β-actin.
Statistical analysis
Values are expressed as the mean ± standard
deviation. Statistical analysis was performed using the Student's
t-test with SPSS 13.0 software (SPSS, Inc., Chicago, IL, USA).
P<0.05 was considered to indicate a significant difference
between values.
Results
Pharmacological modulation of autophagic
activity in BMSCs
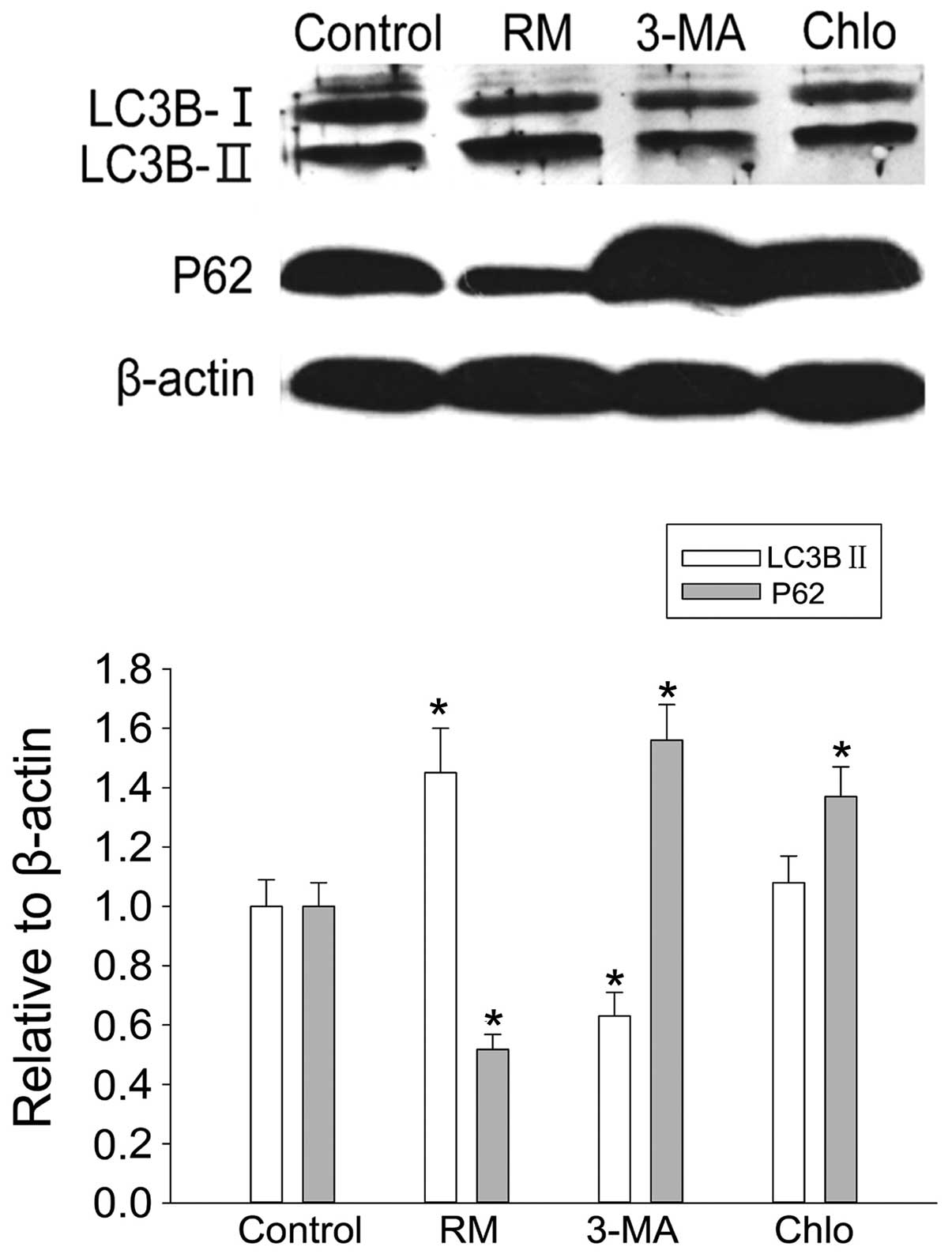
Treatment with autophagy inducer rapamycin for 12 h
promoted the expression of LC3B-II and reduced P62 expression in
BMSCs, while autophagy inhibitors 3-MA and chloroquine produced
opposite effects, except for a slight enhancement of LC3B-II
expression following chloroquine treatment (Fig. 1). These results indicated that
rapamycin was able to enhance autophagy, while 3-MA and chloroquine
inhibited autophagy in BMSCs.
Autophagy inhibits the cell cycle of
BMSCs
Cell cycle analysis BMSCs showed that induction of
autophagy by rapamycin inhibited the cell proliferation, as
indicated by a reduced S-phase cell population (11.04±0.86%) when
compared to that in the control group (14.18±0.76%; P<0.05)
(Fig. 2). Conversely, treatment
with 3-MA and chloroquine marginally increased the S-phase cell
population to 15.13±0.17% and 15.56±0.29%, respectively; however,
differences from the control group were not statistically
significant (Fig. 2).
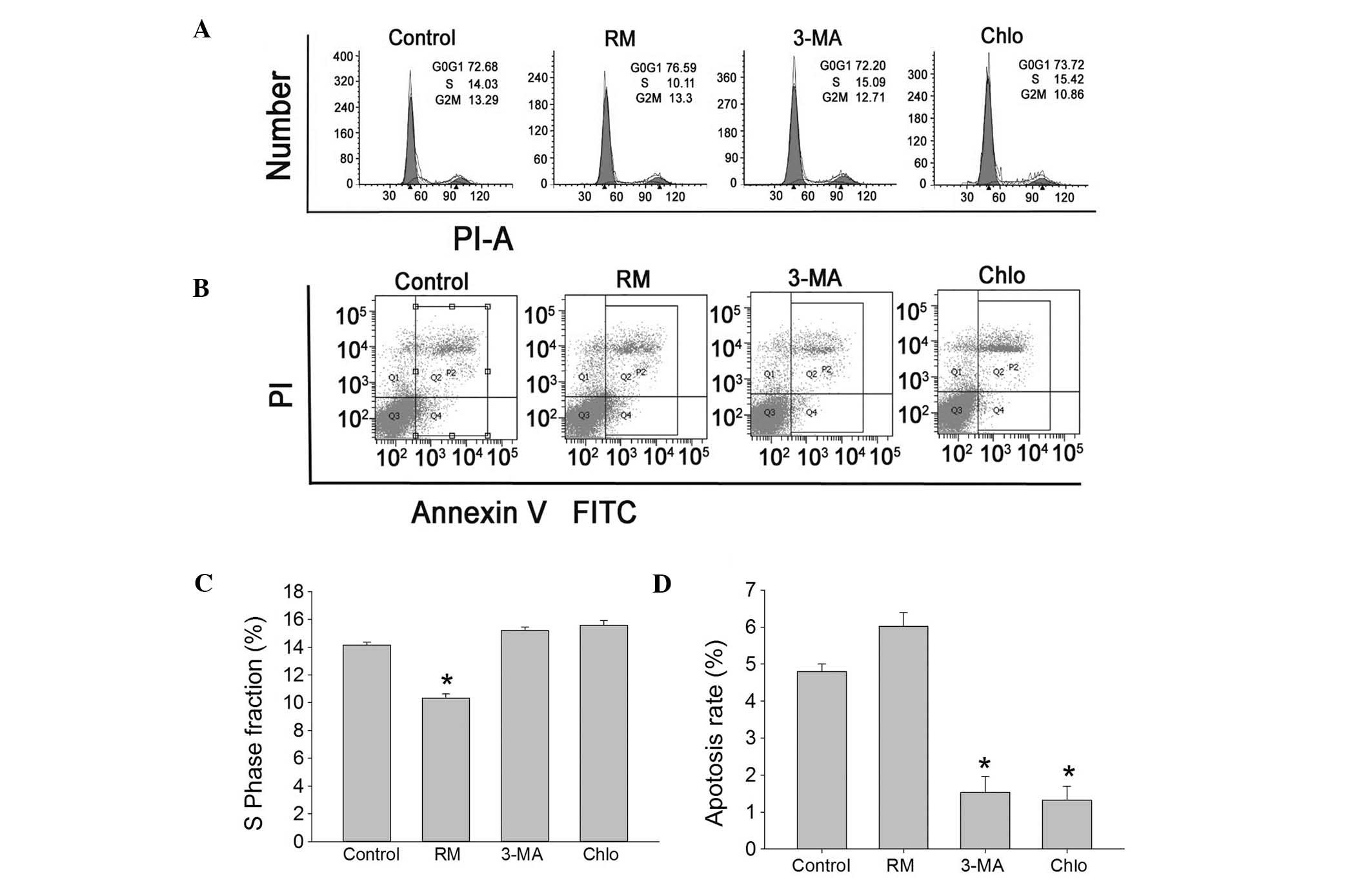 | Figure 2Effects of autophagy on cell cycle
and apoptosis of BMSCs. After incubation of BMSCs with various
drugs for 12 h, (A) cell cycle and (B) apoptosis were detected by
flow cytometry. (C) The S-phase population in the control,
RM-treated, 3-MA-treated and Chlo-treated groups was 14.03, 10.11,
15.09 and 15.42%, respectively. (D) The percentage of apoptotic
cells in the control, RM-treated, 3-MA-treated and Chlo-treated
groups was 4.7, 5.9, 1.5 and 1.3%, respectively. Values are
expressed as the mean ± standard deviation. *P<0.05
vs. control group. RM, rapamycin; Chlo, chloroquine. BMSC, bone
marrow mesenchymal stem cell; 3-MA, 3-methyladenine; PI, propidium
iodide; FITC, fluorescein isothiocyanate. |
Autophagy promotes BMSC apoptosis
Flow cytometric analysis showed that autophagy
inducer rapamycin slightly enhanced the apoptotic rate of BMSCs
(5.8±0.42%), which was, however, not significantly different from
that of the control group (4.6±0.55%). Following treatment with
autophagy inhibitors 3-MA or chloroquine, the apoptotic rate of
BMSCs (1.5±0.1 and 1.23±0.25, respectively) was significantly
reduced when compared to that in the control group (P<0.05)
(Fig. 2). These results indicated
that autophagy promotes apoptosis, while inhibition of autophagy
prevents apoptosis of BMSCs.
Autophagy enhances the differentiation of
BMSCs into neurons
BMSCs at passage 3 were spindle-shaped and had a
fibroblast-like morphology. Following incubation in induction
medium (low-glucose DMEM with 2% DMSO and 200 µmol/l BHA)
for 12 h, cells exhibited neurite extensions and their shape was
changed to resemble neurons. Flow-cytometric analysis showed that
rapamycin-induced autophagy promoted BMSC differentiation into
neurons, as demonstrated by a significantly increased number of
NSE-positive cells (78.2±2.6) compared to that in the control group
(36.7±2.8; P<0.05) (Fig. 3A and
B). Conversely, treatment with autophagy inhibitors 3-MA or
chloroquine reduced the percentage of NSE-positive cells (32.8±3.1
and 29.9±1.56, respectively) compared that in the control group
(P<0.05) (Fig. 3A and B).
Furthermore, western blot analysis showed that upon differentiation
into neurons, the expression of microtubule-associated protein 2
(MAP2) was increased in the rapamycin-treated and reduced in the
3-MA- or chloroquine-treated groups compared to that in the control
group (P<0.05 for all) (Fig. 3C and
D).
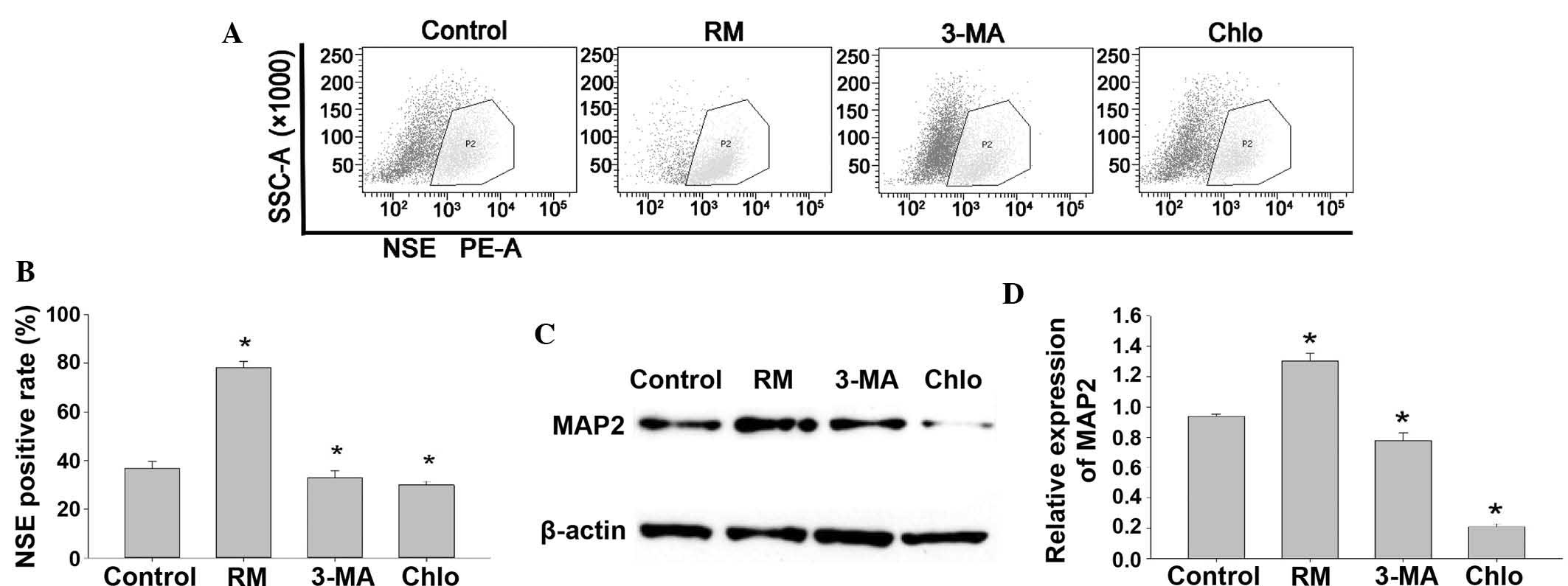 | Figure 3Effects of autophagy on the
differentiation of BMSCs into neurons. (A and B) Following
induction of BMSCs to differentiate into neurons for 12 h, flow
cytometric analysis revealed that the percentage of NSE-positive
cells in the control, RM-treated, 3-MA-treated and Chlo-treated
groups was 38.2, 80.1, 33.7 and 29.7%, respectively. (C and D)
Western blot analysis showed that RM treatment promoted the
expression of MAP2, while treatment with 3-MA or Chlo inhibited
MAP2 expression. Values are expressed as the mean ± standard
deviation. *P<0.05 vs. control group. RM, rapamycin;
Chlo, chloroquine; MA, methyladenine; BMSC, bone marrow mesenchymal
stem cell; SSC, side scatter; MAP2, microtubule-associated protein
2; NSE, neuron-specific enolase; PE-A; absorption of
phycoerythrin. |
The effect of autophagy on Notch1
signaling pathway in BMSCs
To investigate the role of the Notch1 signaling
pathway in the observed autophagy-mediated enhancement of neuronal
differentiation of BMSCs, flow cytometry was used to detect Notch1
MFI after modulation of autophagy. The results showed that the
Notch1 MFI was significantly reduced after treatment with rapamycin
(410±19), when compared to that in the control group (552±23;
P<0.05) (Fig. 4A and B).
However, treatment with 3-MA or chloroquine did not change the MFI
of Notch1 (560±39 and 570±30, respectively) compared to that in the
control group. Furthermore, western blot analysis showed that
treatment with autophagy inducer rapamycin reduced the expression
of Notch1 and Hes1, whereas 3-MA and chloroquine did not affected
the expression of Notch1, but both increased Hes1 expression levels
(Fig. 4C and D).
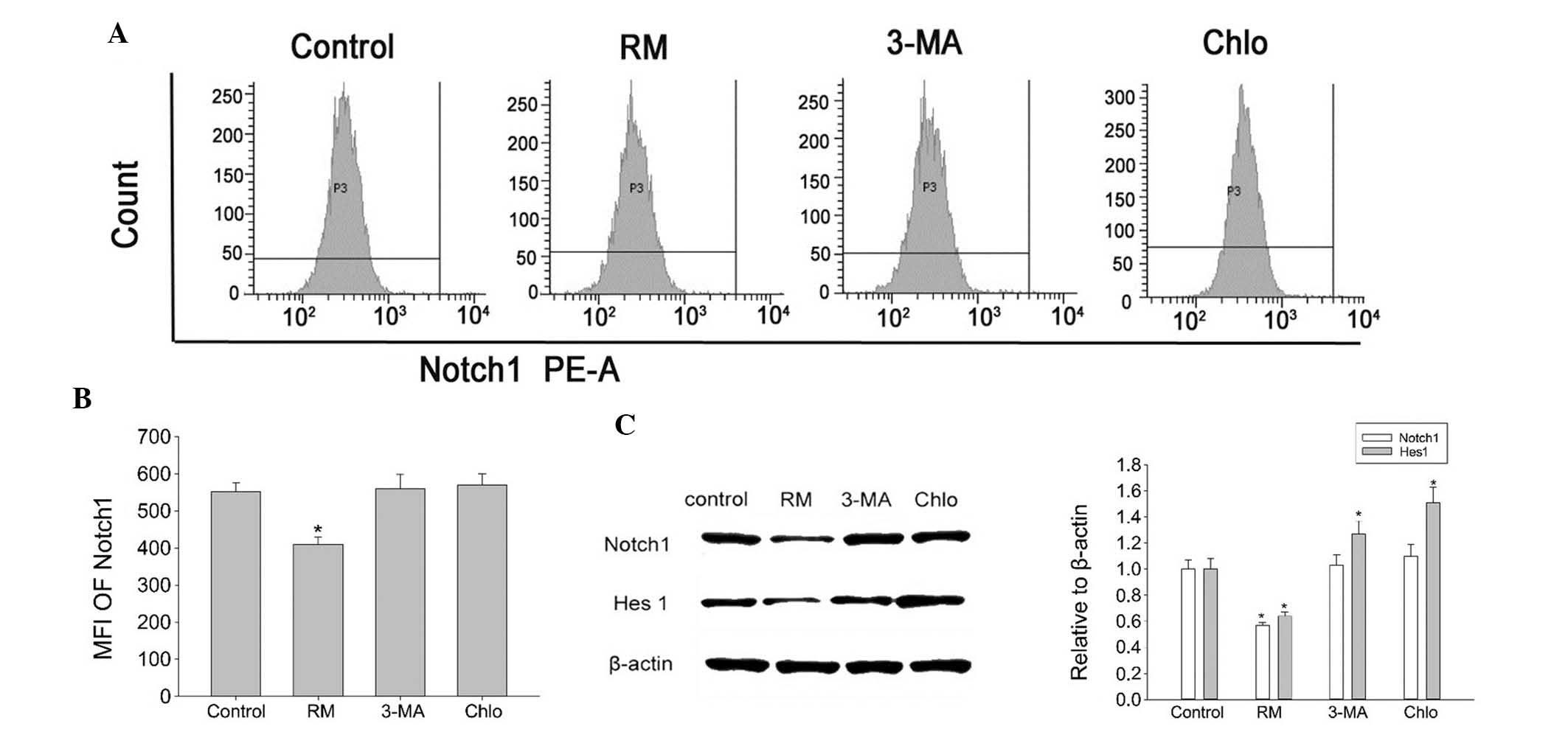 | Figure 4Effects of autophagy on Notch1
signaling in BMSCs. (A and B) Following incubation of BMSCs with
various drugs for 12 h, flow cytometric analysis revealed that the
MFI of Notch1 was 562, 415, 587 and 582 in the control, RM-treated,
3-MA-treated and Chlo-treated group, respectively. Values are
expressed as the mean ± standard deviation. *P<0.05
vs. control group. (C) Western blot analysis showed that RM
treatment reduced the expression of Notch1 and Hes1, while
treatment with 3-MA or Chlo did not affect the expression of Notch1
and Hes1. RM, rapamycin; Chol, chloroquine; MFI, mean fluorescence
intensity; MA, methyladenine; BMSC, bone marrow mesenchymal stem
cell; PE-A; absorption of phycoerythrin. |
Discussion
Transdifferentiation of BMSCs into neurons provides
a novel approach for the treatment of neurological diseases by cell
transplantation (9,10). However, in vitro, the
efficacy of BMSC proliferation and differentiation is low and the
function of differentiated mature cells is also limited;
furthermore, the underlying mechanisms of these processes have
largely remained elusive (11–13).
Autophagy is a process during which cells digest their own
long-lived proteins and organelles under conditions of nutritional
starvation and stress (14–16).
Autophagy has an important role in cell proliferation,
differentiation and apoptosis (17–20),
and can rapidly and efficiently degrade transcription factors,
enzymes, adhesion molecules and certain secreted proteins (21,22),
which are important for stem-cell proliferation and differentiation
(23–25). Further studies focusing on
autophagy in BMSCs may reveal the underlying mechanisms of
proliferation and differentiation, and thereby provide potential
strategies for clinical treatment.
The LC3 protein is localized in the membrane surface
of autophagosomes. Upon the induction of autophagy, LC3-I binds to
phosphatidylethanolamine on the membrane surface of the
autophagosome and is then modulated by autophagy-related 7 (Atg7)
to form LC3-II. As the levels of LC3-II are positively correlated
with the number of autophagic vacuoles, it is considered to be a
marker for autophagy (26). P62 is
able to bind with LC3 and targeting proteins, which are then
degraded in lysosomes. P62 is extensively degraded when the
activity of autophagy is increased, while it accumulates in the
cytoplasm when autophagy is attenuated, suggesting that P62 levels
reflect the level of autophagic activity (27). Mammalian target of rapamycin (mTOR)
and the phosphoinositide-3 kinase (PI3K)/AKT pathway are the most
important signaling pathways involved in the regulation of
autophagy (28). Rapamycin can
enhance autophagic activity by inhibiting the mTOR signaling
pathway (28), which explains for
the result of the present study that rapamycin treatment enhanced
LC3-II levels and reduced P62 levels in BMSCs. 3-MA inhibits
autophagy by decreasing the activity of type III PI3K signaling
(29). Thus, treatment of BMSCs
with 3-MA reduced the expression of LC3-II and enhanced P62 in the
present study. Chloroquine is able to inhibit autophagy at the late
stages by preventing the fusion of autophagosomes and lysosomes
(30). Thus, in the present study,
treatment with chloroquine slightly increased LC3-II, while it
markedly enhanced the expression of P62 in BMSCs.
Deficiency of Atg7 in hematopoietic stem cells has
been reported to promote the amplification of bone marrow precursor
cells, thus inducing invasive bone marrow hyperplasia and
presenting the phenotype of acute myeloid leukemia (25). Autophagy has been suggested to
occur in BMSCs (31,32), and is enhanced under stress
conditions, such as hypoxia (33).
However, the role of autophagy in BMSC apoptosis has remained to be
elucidated. Autophagy has been shown to exert cytoprotective
effects under stress conditions (34,35).
Hypoxia and starvation are known to promote autophagy and decrease
apoptosis in BMSC, while 3-MA treatment increases BMSC apoptosis
(36). Ugland et al
(37) showed that autophagy can be
activated via the cyclic adenosine monophosphate (cAMP) signaling
pathway in BMSCs, and that the activated cAMP is able to inhibit
cell proliferation though modulation of extracellular
signal-regulated kinase-mediated activation of cyclin E, thus
recruiting autophagy-associated protein Beclin1 to surround the
nucleus, which resulted in the formation of autophagic bodies.
In the present study, rapamycin was used to induce
autophagy in BMSCs, which was demonstrated to decrease the S-phase
population of the cell cycle and promote apoptosis, while treatment
with 3-MA and chloroquine exerted the opposite effects, suggesting
that activation of autophagy inhibits BMSC proliferation. Autophagy
may be a double-edged sword, which can have pro-survival effects
under stress conditions to promote cell growth and proliferation,
while inhibiting cell proliferation and promoting apoptosis upon
excessive activation by degrading proteins and organelles essential
for cell proliferation (38).
The present study revealed that rapamycin-induced
autophagy enhanced the expression of neuron-specific markers NSE
and MAP2. These findings suggested that autophagy is involved in
BMSC differentiation. Zeng and Zhou (2) found that autophagy is enhanced during
the process of retinoic acid-induced differentiation of N2a
neuroblastoma cells. Furthermore, the differentiation ability was
inhibited by treatment with autophagy inhibitors 3-MA and LY294002
and by RNA interference-mediated suppression of Beclin1. In
addition, the mTOR pathway was shown to be inhibited during N2a
neuroblastoma cell differentiation, and treatment with mTOR
inhibitor rapamycin promoted neurite extension, increased the cell
size and enhanced the expression of neuron-specific markers
(1). Furthermore, deficiency of
Atg7 and Atg5 as well as treatment with 3-MA has been shown to
inhibit BMSC differentiation into adipocytes, suggesting that
autophagy has an important role in BMSC differentiation into
adipocytes (39). Li et al
(40) have shown that autophagy is
activated during the process of BMSC differentiation into neurons,
and that treatment with rapamycin increased the expression levels
of Tau and MAP2, while 3-MA had the opposite effect.
The present study provided evidence that the
degradation of 'stemness'-associated proteins and the production of
neuron-specific proteins is part of the process of BMSC
differentiation into neurons. Induction of autophagy was indicated
to accelerate this process and thus represents an approach for
stimulating cell differentiation, increasing the expression of
neuron-specific markers, and promoting BMSC differentiation into
neurons.
Notch, a regulatory factor and an important
signaling pathway controlling cell fate, has an important role in
growth, development, proliferation, differentiation and apoptosis
of various cell types (41–43).
A previous study by our group demonstrated that the expression of
Notch1 and Jag1 genes and their downstream targeted genes PS1 and
Hes1 were significantly reduced during the process of human
mesenchymal stem cell differentiation into neurons in vitro
(44), suggesting that Notch1 is
inhibited during the process of neuronal differentiation. Thus, the
Notch1 signaling pathway is considered an important regulator for
BMSC differentiation into neurons (4).
The present study examined the association between
autophagy and the Notch1 signaling pathway in BMSCs and found that
induction of autophagy inhibits Notch1. This affects the ability of
BMSCs to differentiate into neurons, which is supported by the
evidence that the Notch1 signaling pathway and autophagy are linked
(45–47). Thus, autophagy may inhibit
proliferation and promote differentiation by inhibiting the Notch1
signaling pathway in BMSCs.
The results of the present study demonstrated that
induction of autophagy decreased the S-phase population and
promoted neuronal differentiation of BMSCs, while inhibition of
autophagy reduced apoptosis and neuronal differentiation. These
processed were regulated by modulation of the Notch1 signaling
pathway. The present study suggested that autophagy has an
important role in BMSC proliferation and differentiation. Further
study is required to clarify the underlying mechanisms of BMSCs
differentiation in vivo.
Acknowledgments
The present study was supported by National Nature
Science Foundation of China (grant nos. 81171250 and 31100790).
References
|
1
|
Lapierre LR, Gelino S, Meléndez A and
Hansen M: Autophagy and lipid metabolism coordinately modulate life
span in germline-less C. elegans. Curr Biol. 21:1507–1514. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Zeng M and Zhou JN: Roles of autophagy and
mTOR signaling in neuronal differentiation of mouse neuroblastoma
cells. Cell Signal. 20:659–665. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Greenwald I and Kovall R: Notch signaling:
Genetics and structure. WormBook. 17:1–28. 2013. View Article : Google Scholar
|
|
4
|
Chiba S: Notch signaling in stem cell
systems. Stem cells. 24:2437–2447. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Bray SJ: Notch signalling: A simple
pathway becomes complex. Nat Rev Mol Cell Biol. 7:678–689. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Louvi A and Artavanis-Tsakonas S: Notch
and disease: A growing field. Semin Cell Dev Biol. 23:473–480.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Chen BY, Wang X, Chen LW and Luo ZJ:
Molecular targeting regulation of proliferation and differentiation
of the bone marrow-derived mesenchymal stem cells or mesenchymal
stromal cells. Curr Drug Targets. 13:561–571. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Woodbury D, Schwarz EJ, Prockop DJ and
Black IB: Adult rat and human bone marrow stromal cells
differentiate into neurons. J Neurosci Res. 61:364–370. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhang R, Li J and Xie J: Efficient in
vitro labeling rabbit bone marrow-derived mesenchymal stem cells
with SPIO and differentiating into neural-like cells. Mol Cells.
37:650–655. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Caplan AI: Why are MSCs therapeutic? New
data: New insight. J Pathol. 217:318–324. 2009. View Article : Google Scholar
|
|
11
|
Anbari F, Khalili MA, Bahrami AR,
Khoradmehr A, Sadeghian F, Fesahat F and Nabi A: Intravenous
transplantation of bone marrow mesenchymal stem cells promotes
neural regeneration after traumatic brain injury. Neural Regen Res.
9:919–923. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Jing L and Jia Y, Lu J, Han R, Li J, Wang
S, Peng T and Jia Y: MicroRNA-9 promotes differentiation of mouse
bone mesenchymal stem cells into neurons by Notch signaling.
Neuroreport. 22:206–211. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Hayashi T, Wakao S, Kitada M, Ose T,
Watabe H, Kuroda Y, Mitsunaga K, Matsuse D, Shigemoto T, Ito A, et
al: Autologous mesenchymal stem cell-derived dopaminergic neurons
function in parkinsonian macaques. J Clin Invest. 123:272–284.
2013. View
Article : Google Scholar :
|
|
14
|
Todd LR, Gomathinayagam R and Sankar U: A
novel Gfer-Drp1 link in preserving mitochondrial dynamics and
function in pluripotent stem cells. Autophagy. 6:821–822. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Mukhopadhyay S, Panda PK, Sinha N, Das DN
and Bhutia SK: Autophagy and apoptosis: Where do they meet?
Apoptosis. 19:555–566. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Meléndez A and Neufeld TP: The cell
biology of autophagy in metazoans: A developing story. Development.
135:2347–2360. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Mizushima N, Levine B, Cuervo AM and
Klionsky DJ: Autophagy fights disease through cellular
self-digestion. Nature. 451:1069–1075. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yang Z and Klionsky DJ: Mammalian
autophagy: Core molecular machinery and signaling regulation. Curr
Opin Cell Biol. 22:124–131. 2010. View Article : Google Scholar :
|
|
19
|
Yorimitsu T and Klionsky DJ: Autophagy:
Molecular machinery for self-eating. Cell Death Differ. 12(Suppl
2): S1542–S1552. 2005. View Article : Google Scholar
|
|
20
|
Guan JL, Simon AK, Prescott M, Menendez
JA, Liu F, Wang F, Wang C, Wolvetang E, Vazquez-Martin A and Zhang
J: Autophagy in stem cells. Autophagy. 9:830–849. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zirin J and Perrimon N: Drosophila as a
model system to study autophagy. Semin Immunopathol. 32:363–372.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Rubinsztein DC, Gestwicki JE, Murphy LO
and Klionsky DJ: Potential therapeutic applications of autophagy.
Nat Rev Drug Discov. 6:304–312. 2007. View
Article : Google Scholar : PubMed/NCBI
|
|
23
|
Coller HA, Sang L and Roberts JM: A new
description of cellular quiescence. PLoS Biol. 4:e832006.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Phadwal K, Watson AS and Simon AK:
Tightrope act: Autophagy in stem cell renewal, differentiation,
proliferation and aging. Cell Mol Life Sci. 70:89–103. 2013.
View Article : Google Scholar :
|
|
25
|
Mortensen M, Watson AS and Simon AK: Lack
of autophagy in the hematopoietic system leads to loss of
hematopoietic stem cell function and dysregulated myeloid
proliferation. Autophagy. 7:1069–1070. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Kabeya Y, Mizushima N, Yamamoto A,
Oshitani-Okamoto S, Ohsumi Y and Yoshimori T: LC3, GABARAP and
GATE16 localize to autophagosomal membrane depending on form-II
formation. J Cell Sci. 117:2805–2812. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Komatsu M, Kurokawa H, Waguri S, Taguchi
K, Kobayashi A, Ichimura Y, Sou YS, Ueno I, Sakamoto A, Tong KI, et
al: The selective autophagy substrate p62 activates the stress
responsive transcription factor Nrf2 through inactivation of Keap1.
Nat Cell Biol. 12:213–223. 2010.PubMed/NCBI
|
|
28
|
Martelli AM, Evangelisti C, Chiarini F,
Grimaldi C, Cappellini A, Ognibene A and McCubrey JA: The emerging
role of the phosphatidylinositol 3-kinase/Akt/mammalian target of
rapamycin signaling network in normal myelopoiesis and
leukemogenesis. Biochim Biophys Acta. 1803:991–1002. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wu YT, Tan HL, Shui G, Bauvy C, Huang Q,
Wenk MR, Ong CN, Codogno P and Shen HM: Dual role of
3-methyladenine in modulation of autophagy via different temporal
patterns of inhibition on class I and III phosphoinositide
3-kinase. J Biol Chem. 285:10850–10861. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Yoon YH, Cho KS, Hwang JJ, Lee SJ, Choi JA
and Koh JY: Induction of lysosomal dilatation, arrested autophagy
and cell death by chloroquine in cultured ARPE-19 cells. Invest
Ophthalmol Vis Sci. 51:6030–6037. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Pantovic A, Krstic A, Janjetovic K, Kocic
J, Harhaji-Trajkovic L, Bugarski D and Trajkovic V: Coordinated
time-dependent modulation of AMPK/Akt/mTOR signaling and autophagy
controls osteogenic differentiation of human mesenchymal stem
cells. Bone. 52:524–531. 2013. View Article : Google Scholar
|
|
32
|
Lee Y, Jung J, Cho KJ, Lee SK, Park JW, Oh
IH and Kim GJ: Increased SCF/c-kit by hypoxia promotes autophagy of
human placental chorionic plate-derived mesenchymal stem cells via
regulating the phosphorylation of mTOR. J Cell Biochem. 114:79–88.
2013. View Article : Google Scholar
|
|
33
|
Wu J, Niu J, Li X, Li Y, Wang X, Lin J and
Zhang F: Hypoxia induces autophagy of bone marrow-derived
mesenchymal stem cells via activation of ERK1/2. Cell Physiol
Biochem. 33:1467–1474. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Li N, Zhang Q, Qian HY, Jin C, Yang Y and
Gao R: Atorvastatin induces autophagy of mesenchymal stem cells
under hypoxia and serum deprivation conditions by activating the
mitogen-activated protein kinase/extracellular signal-regulated
kinase pathway. Chin Med J (Engl). 127:1046–1051. 2014.
|
|
35
|
Song C, Song C and Tong F: Autophagy
induction is a survival response against oxidative stress in bone
marrow-derived mesenchymal stromal cells. Cytotherapy.
26:1361–1370. 2014. View Article : Google Scholar
|
|
36
|
Wang L, Hu X, Zhu W, Jiang Z, Zhou Y, Chen
P and Wang J: Increased leptin by hypoxic-preconditioning promotes
autophagy of mesenchymal stem cells and protects them from
apoptosis. Sci China Life Sci. 57:171–180. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Ugland H, Naderi S, Brech A, Collas P and
Blomhoff HK: CAMP induces autophagy via a novel pathway involving
ERK, cyclin E and Beclin 1. Autophagy. 7:1199–1211. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Shintani T and Klionsky DJ: Autophagy in
health and disease: A double-edged sword. Science. 306:990–995.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Singh R, Xiang Y, Wang Y, Baikati K,
Cuervo AM, Luu YK, Tang Y, Pessin JE, Schwartz GJ and Czaja MJ:
Autophagy regulates adipose mass and differentiation in mice. J
Clin Invest. 119:3329–3339. 2009.PubMed/NCBI
|
|
40
|
Li Y, Wang C, Zhang G, Wang X, Duan R, Gao
H, Peng T, Teng J and Jia Y: Role of autophagy and mTOR signaling
in neural differentiation of bone marrow mesenchymal stem cells.
Cell Biol Int. 38:1337–1343. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Patel PN, Yu XM, Jaskula-Sztul R and Chen
H: Hesperetin activates the Notch1 signaling cascade, causes
apoptosis, and induces cellular differentiation in anaplastic
thyroid cancer. Ann Surg Oncol. 21(Suppl 4): S497–S504. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Su BH, Qu J, Song M, Huang XY, Hu XM, Xie
J, Zhao Y, Ding LC, She L, Chen J, et al: NOTCH1 signaling
contributes to cell growth, anti-apoptosis and metastasis in
salivary adenoid cystic carcinoma. Oncotarget. 5:6885–6895. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Zhao C, Guo H, Li J, Myint T, Pittman W,
Yang L, Zhong W, Schwartz RJ, Schwarz JJ, et al: Numb family
proteins are essential for cardiac morphogenesis and progenitor
differentiation. Development. 141:281–295. 2014. View Article : Google Scholar :
|
|
44
|
Xing Y, Bai RY, Yan WH, Han XF, Duan P, Xu
Y and Fan ZG: Expression changes of Notch-related genes during the
differentiation of human mesenchymal stem cells into neurons. Sheng
Li Xue Bao. 59:267–272. 2007.In Chinese. PubMed/NCBI
|
|
45
|
Barth JM and Köhler K: How to take
autophagy and endocytosis up a notch. Biomed Res Int.
2014:9608032014. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Kwon MH, Callaway H, Zhong J and
Yedvobnick B: A targeted genetic modifier screen links the
SWI2/SNF2 protein domino to growth and autophagy genes in
Drosophila melanogaster. G3 (Bethesda). 3:815–825. 2013. View Article : Google Scholar
|
|
47
|
Yamamoto S, Charng WL and Bellen HJ:
Endocytosis and intracellular trafficking of Notch and its ligands.
Curr Top Dev Biol. 92:165–200. 2010. View Article : Google Scholar : PubMed/NCBI
|