Introduction
Ovarian cancer is the second the most common
gynecological malignancy, and is the seventh most common cancer in
women worldwide, with 239,000 new cases diagnosed in 2012 (1). Epithelial ovarian cancer (EOC) is the
major pathological type of ovarian cancer, and accounts for
approximately 90% cases, including serous adenocarcinoma,
endometrial adenocarcinoma and clear cell carcinoma (2). Despite advances in the treatments by
combining surgical resection, chemotherapy and radiotherapy, the
prognosis for patients with EOC remains unsatisfactory, with a
five-year overall survival rate of only 30% (3,4). The
poor prognosis of EOC has been demonstrated to be associated with
the occurrence of tumor metastasis, and recurrence (5). Therefore, it is of great clinical
significance to elucidate the molecular mechanisms, which
contribute to the tumorigenesis and tumor development of EOC, and
investigate novel therapeutic targets for this disease.
Previous studies have reported the importance of the
regulatory roles of microRNAs (miRNAs) in the carcinogenesis and
progression of EOC (6–8). miRNAs are a major group of
endogenous, non-protein-coding and short RNA molecules
approximately 22 nucleotides in length (9). At present, more than 1,000 miRNAs
have been validated, which only represent ~1% of the predicted
genes in the genome, and are estimated to regulate the expression
of more than 60% of protein-coding genes (10,11).
miRNAs negatively modulate their target genes through binding to
the 3′untranslated region (UTR) of target genes for the
post-transcriptional regulation, and therefore to participate in a
great deal of biological processes, including cell proliferation,
cell cycle, apoptosis, invasion, metastasis, glucose and lipid
metabolism and infection and immune responses (12–14).
Studies have indicated that the abnormal expression of miRNAs serve
significant roles in the occurrence and development of tumors,
including bladder cancer (15),
renal cell carcinoma (16),
colorectal cancer (17), breast
cancer (18), hepatocellular
carcinoma (19) and EOC (20). Therefore, exploring the expression
patterns and roles of miRNAs in EOC may provide potential
diagnostic and therapeutic targets for the treatments of EOC.
Thus, the current study demonstrated that the
expression level of miR-139 was markedly reduced in EOC, and
restoration of miR-139 repressed cellular proliferation, migration
and invasion. In addition, it was identified that miR-139 directly
targeted and downregulated HDGF through binding to its 3′UTR.
Finally, it was demonstrated that HDGF overexpression may rescue
the inhibitory effects mediated by miR-139 in EOC.
Materials and methods
Tissue specimens
The current study was approved by the Ethics
Committee of Changzhi Peace Hospital Affiliated to Changzhi Medical
College (Changzhi, China), and informed consent was also obtained
from each patient in accordance with the guidelines of Changzhi
Peace Hospital Affiliated to Changzhi Medical College. A total of
22 primary EOC tissues and matched adjacent normal tissues were
collected from patients with EOC (age range, 41–71 years; median
age, 56) who underwent surgical resection at the Department of
Gynaecology and Obstetrics, Changzhi Peace Hospital Affiliated to
Changzhi Medical College between June 2014 and October 2015. All
tissues were immediately snap-frozen in liquid nitrogen following
surgical resection and stored at −80°C in a refrigerator until
required for RNA extraction.
Cell lines and oligonucleotides
transfection
Four EOC cell lines (OVCAR3, CAOV3, SKOV3 and ES-2)
and the human normal ovarian epithelial cell line (NOEC) were
obtained from the American Type Culture Collection (ATCC; Manassas,
VA, USA). All cell lines were cultured in RPMI-1640 medium or Ham's
F-12 medium containing 10% FBS, 100 mg/ml penicillin and 100 mg/ml
streptomycin (all from Gibco; Thermo Fisher Scientific, Inc.,
Waltham, MA, USA) in a humidified 5% CO2 at 37°C.
The miR-139 mimics and negative control mimics (NC)
were purchased from Shanghai GenePharma Co., Ltd. (Shanghai,
China). HDGF-overexpressed plasmid (pcDNA3.1-HDGF) and the blank
vector pcDNA3.1 (pcDNA3.1) were synthesized by the Chinese Academy
of Sciences (Changchun, China). Transient transfections of the
miRNA mimics or plasmid were conducted with Lipofectamine 2000
(Invitrogen; Thermo Fisher Scientific, Inc.) and OPTI-MEM reduced
serum medium (Gibco; Thermo Fisher Scientific, Inc.), following the
manufacturer's instructions.
Total RNA extraction and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
Total RNA and miRNAs were extracted from tissues or
cells using TRIzol (Invitrogen; Thermo Fisher Scientific, Inc.) and
mirVana miRNA isolation kit (Ambion; Thermo Fisher Scientific,
Inc.), respectively. TaqMan microRNA assay (Applied Biosystems;
Thermo Fisher Scientific, Inc.) was adopted to measure miR-139
expression. The reaction mixture contained 1.0 µl TaqMan miRNA
assay (20X), 10.0 µl TaqMan 2X Universal PCR Master mix (Thermo
Fisher Scientific, Inc.), 1.33 µl cDNA, 1 µl forward primer and 1
µl reverse primer and 5.67 µl double distilled water. For HDGF mRNA
expression, reverse transcription was performed with PrimeScript™
RT Master mix (Takara Bio, Inc., Otsu, Japan). The SYBR-Green PCR
master mixture (Takara Bio, Inc.) was used to determine HDGF mRNA
expression. The reaction system for qPCR consisted of 10 µl
SYBR-Green PCR Master mix, 2 µl forward primer, 2 µl reverse
primer, 2 µl cDNA and 4 µl double distilled water. The thermocycler
conditions were as follows: 95°C for 10 min; 40 cycles of 95°C for
15 sec and 60°C for 1 min. RT-qPCR was performed in triplicate
using ABI Prism 7500 Sequence Detection system (Applied Biosystems;
Thermo Fisher Scientific, Inc.), the 2−ΔΔCq method was
used to determine the relative gene expression (21). Primers are presented in Table I. U6 snRNA and GAPDH mRNA were used
as endogenous controls for miR-139 and HDGF mRNA, respectively.
 | Table I.Primer sequences for reverse
transcription-quantitative polymerase chain reaction. |
Table I.
Primer sequences for reverse
transcription-quantitative polymerase chain reaction.
| Gene | Sequences
(5′→3′) |
|---|
| miR-139 |
|
|
Forward |
CGACGCGTCCCTCTTCCCATTCCTTC |
|
Reverse |
CCGGAATTCCGAGACCCACTGACACTATCT |
| U6 |
|
|
Forward |
CTCGCTTCGGCAGCACATATACT |
|
Reverse |
ACGCTTCACGAATTTGCGTGTC |
| HDGF |
|
|
Forward |
ATCAACAGCCAACAAATACC |
|
Reverse |
TTCTTATCACCGTCACCCT |
| GAPDH |
|
|
Forward |
CATCACCATCTTCCAGGAGCG |
|
Reverse |
TGACCTTGCCCACAGCCTTG |
MTT assay
The proliferative ability of EOC cells was
determined using the
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)
assay (Sigma-Aldrich; Merck Millipore, Darmstadt, Germany).
Briefly, transfected cells were collected, counted and seeded in
96-well plates at a density of 2,000 cells/well. Cells were then
incubated in a humidified 5% CO2 at 37°C for continual
24–96 h. The MTT assay was performed at specified time points, 20
µl MTT solution (5 mg/ml) was added into each well and incubated at
37°C for 4 h. Subsequently, the culture medium containing MTT
solution was removed and replaced with 100 µl DMSO (Sigma-Aldrich;
Merck Millipore). Finally, the optical density at 490 nm (OD) was
detected with a microplate reader (ELx800; Bio-Tek Instruments,
Inc., Winooski, VT, USA).
Cell migration and invasion assay
The cell migration and invasion assay was performed
in triplicate using Transwell chambers (8-µm pore size; EMD
Millipore, Billerica, MA, USA) and Matrigel (BD Biosciences, San
Jose, CA, USA)-coated Transwell chambers. In brief, transfected
cells were collected at 48 h post-transfection, counted, and
resuspended in FBS-free culture medium. A total of 4×104
cells in 100 µl FBS-free culture medium were plated into the upper
chamber, and 500 µl culture medium containing 20% FBS was added to
the lower chamber. Cells were incubated in a humidified 5%
CO2 at 37°C for 48 h. Cells that remained on top of the
filter were carefully removed with cotton swabs, and those that
migrated or invaded through the membranes were fixed with 95%
methanol (Beyotime Institute of Biotechnology, Haimen, China),
stained with 0.5% crystal violet (Beyotime Institute of
Biotechnology) and photographed under a microscope (magnification,
×200; Olympus Corporation, Tokyo, Japan).
Bioinformatics analysis
The target genes information of miR-139 were
analyzed using TargetScan (www.targetscan.org) and miRanda (www.microrna.org).
Luciferase report assay
For the luciferase reporter assay, the wild type
(Wt) 3′UTR and mutant (Mut) 3′UTR of HDGF into the pMIR-promotor
vector was synthesized by Shanghai GenePharma Co., Ltd. HEK293T
cells (ATCC) were seeded into 24-well plates at a density of 50–60%
influence. Subsequent to incubation overnight, cells were
co-transfected with miR-139 mimics or NC, along with
pMIR-HDGF-3′UTR Wt and pMIR-HDGF-3′UTR Mut using Lipofectamine
2000. Cells were then incubated in a humidified 5% CO2
at 37°C for 48 h, and luciferase activities were determined with
the Dual-luciferase assay system (Promega Corporation, Madison, WI,
USA), following to the manufacturer's instructions.
Western blotting
A total of 72 h post-transfection, cells were washed
with ice-cold PBS (Gibco; Thermo Fisher Scientific, Inc.) three
times and lysed in RIPA lysis buffer (Beyotime Institute of
Biotechnology) supplemented with proteinase/phosphatase inhibitors
(Thermo Fisher Scientific, Inc.). The concentration of total
protein was detected by using a BCA assay kit (Beyotime Institute
of Biotechnology). Equal amounts protein were subjected to 10% SDS
polyacrylamide gel electrophoresis and transferred onto the
polyvinylidene difluoride membranes (EMD Millipore). The membranes
were then blocked with 3% skimmed milk in Tris-buffered saline/0.1%
Tween (TBST) at room temperature for 1 h, and then incubated with
primary antibodies at 4°C overnight. Subsequent to washing with
TBST for three times, the membranes were incubated with goat
anti-mouse horseradish peroxidase-conjugated secondary antibodies
(cat. no. sc-2005; 1:5,000 dilution; Santa Cruz Biotechnology,
Inc., Santa Cruz, CA, USA), and then visualized with enhanced
chemiluminescence (Pierce; Thermo Fisher Scientific, Inc.). The
primary antibodies used in the present study were mouse anti-human
monoclonal HDGF (1:1,000 dilution; sc-271344; Santa Cruz
Biotechnology, Inc.) and mouse anti-human monoclonal GAPDH (1:1,000
dilution; sc-51907; Santa Cruz Biotechnology, Inc.). GAPDH was used
as an internal control for HDGF.
Statistical analysis
The results were presented as the mean ± standard
deviation. The differences between groups were compared using SPSS
software, version 17 (SPSS, Inc., Chicago, IL, USA). Two-tailed
P<0.05 was considered to indicate a statistically significant
difference.
Results
miR-139 expression in EOC tissues and
cell lines
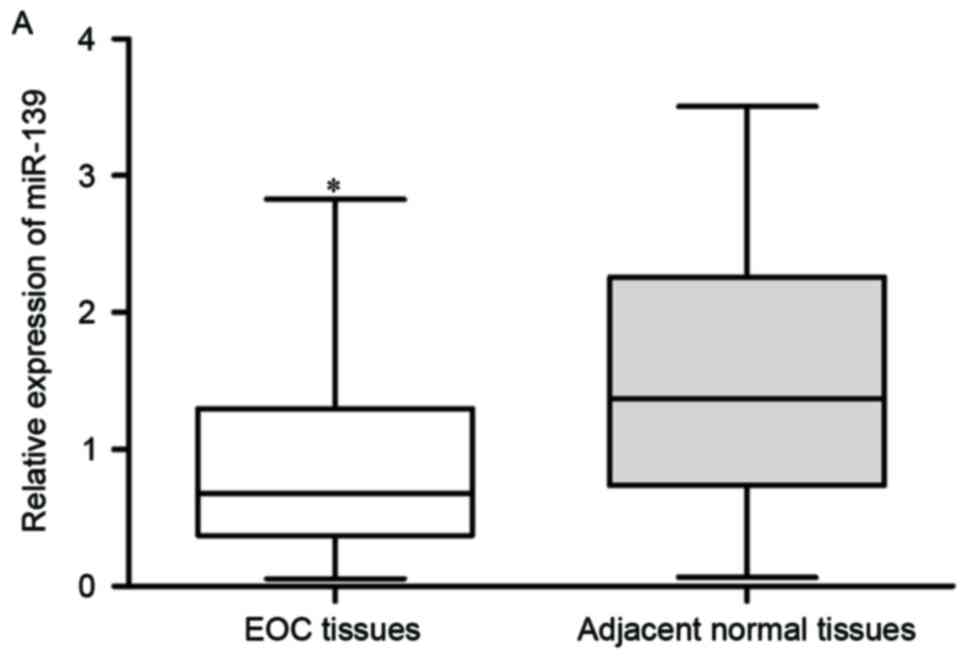
Firstly, miR-139 expression was assayed in EOC
tissues and matched adjacent normal tissues using RT-qPCR. As
presented in Fig. 1A, the
expression level of miR-139 was significantly declined in EOC
tissues compared with those in matched adjacent normal tissues
(P<0.05). miR-139 expression in EOC cell lines (OVCAR3, CAOV3,
SKOV3 and ES-2) in addition to the human NOEC was also determined.
As predicted, miR-139 was significantly downregulated in EOC cell
lines compared with NOEC (Fig. 1B;
P<0.05).
miR-139 suppressed the proliferation
and motility of EOC cells
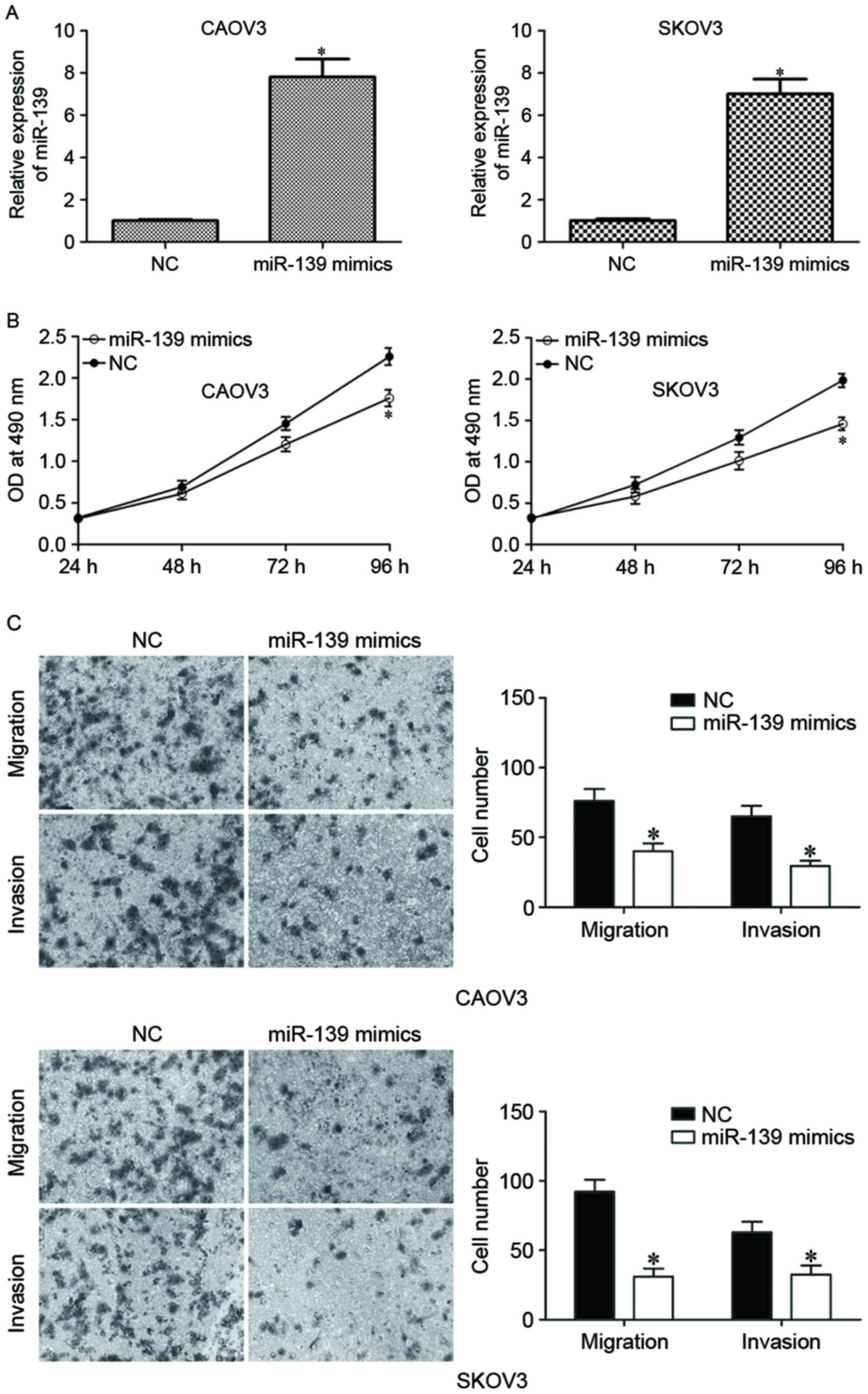
The effects of miR-139 were further investigated on
EOC carcinogenesis and progression. CAOV3 and SKOV3 were
transfected with miR-139 mimics or NC. The upregulation of miR-139
on miR-139 mimics-transfected CAOV3 and SKOV3 was confirmed by
RT-qPCR (Fig. 2A; P<0.05). MTT
assay was performed to measure proliferation of miR-139
mimics-transfected CAOV3 and SKOV3 cells, and the results indicated
that miR-139 overexpression inhibited proliferation of CAOV3 and
SKOV3 cells (Fig. 2B; P<0.05).
The effect of miR-139 on the motility of EOC cells was determined
using cell migration and invasion assay. As presented in Fig. 2C, upregulation of miR-139
substantially suppressed migration and invasion capacities of CAOV3
and SKOV3 cells (Fig. 2C;
P<0.05). These data suggested that miR-139 acted as a tumor
suppressor in EOC.
HDGF was a direct target gene of
miR-139
In order to explore the molecular mechanism of
miR-139 in EOC, the predicted targets of miR-139 were analyzed
using bioinformatics analysis with two publicly available databases
(TargetScan and miRanda). As presented in Fig. 3A, the 3′UTR of HDGF contained
potential binding sites of miR-139. To confirm HDGF as a direct
target gene of miR-139, the luciferase reporter assay was
performed. HEK293T cells were transfected with pMIR-HDGF-3′UTR Wt
or pMIR-HDGF-3′UTR Mut, and miR-139 mimics or NC. As presented in
Fig. 3B, results of luciferase
reporter assay indicated that upregulation of miR-139 significantly
reduced luciferase activities of pMIR-HDGF-3′UTR Wt (P<0.05),
however not luciferase activities of pMIR-HDGF-3′UTR Mut.
miR-139 negatively regulated HDGF
expression in EOC cells
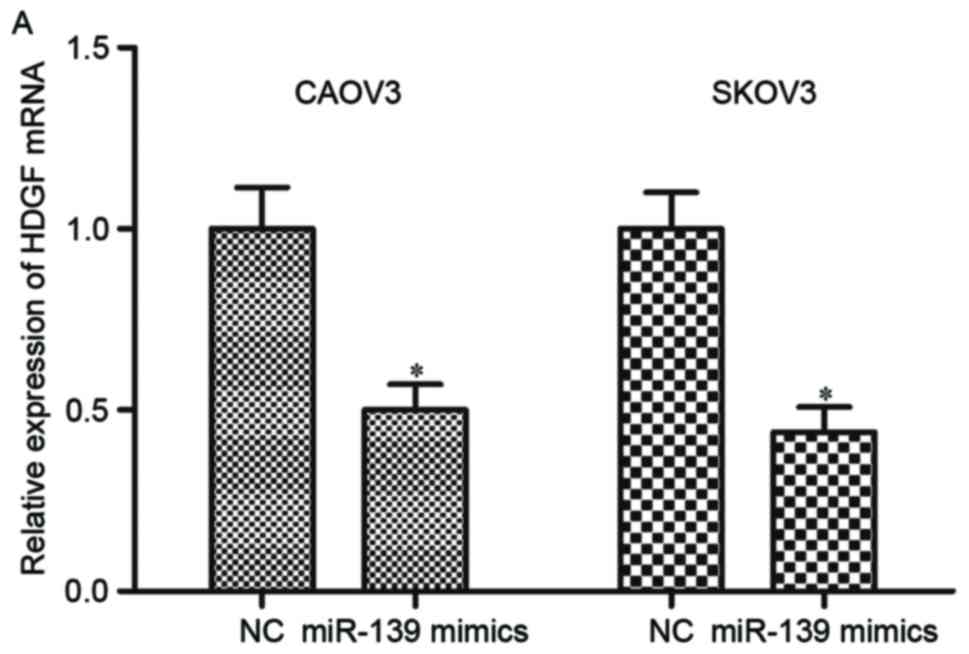
To investigate the regulation roles of miR-139 on
HDGF expression, CAOV3 and SKOV3 cells were transfected with
miR-139 mimics or NC, and measured HDGF expression by RT-qPCR and
western blotting. The results of RT-qPCR indicated that restoration
of miR-139 expression suppressed HDGF mRNA expression in CAOV3 and
SKOV3 cells (Fig. 4A; P<0.05).
In addition, similar to the mRNA changes, miR-139 overexpression
additionally reduced HDGF protein expression level in CAOV3 and
SKOV3 cells compared with NC groups (Fig. 4B; P<0.05). These observations
demonstrated that HDGF was a direct target gene of miR-139, and
could negatively regulate HDGF expression through binding to its
3′UTR.
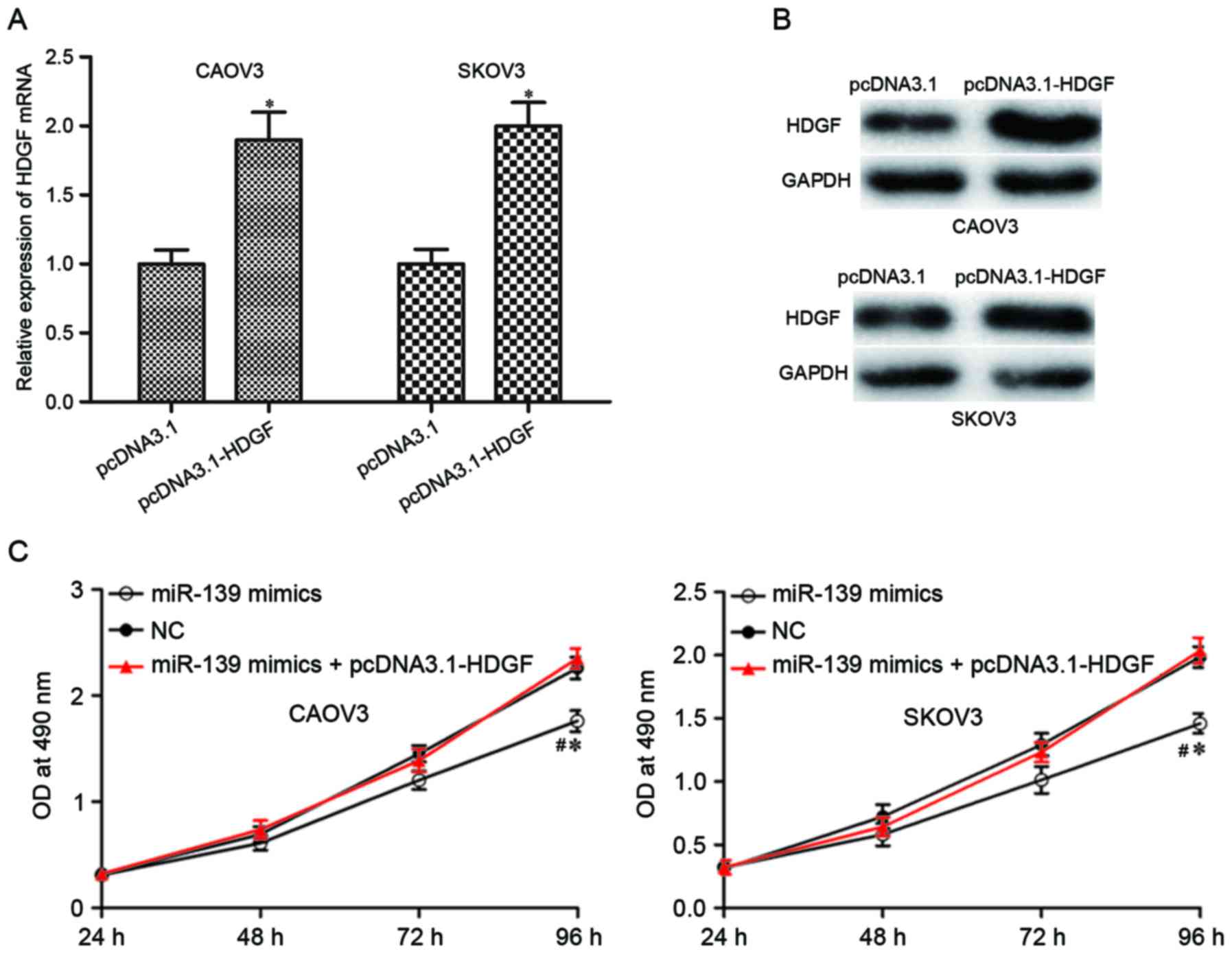
Upregulation of HDGF could rescue the
inhibitory effects of miR-139 on EOC cells
HDGF was identified as a direct target gene of
miR-139 in EOC. If these tumor suppressive roles of miR-139 on EOC
were mediated by HDGF, HDGF overexpression could rescue these
effects induced by miR-139. pcDNA3.1-HDGF or pcDNA3.1 was injected
into CAOV3 and SKOV3 cells, and upregulation of HDGF was determined
by RT-qPCR (Fig. 5A; P<0.05)
and western blotting (Fig. 5B;
P<0.05). Reintroduction of HDGF in miR-139 mimics-transfected
CAOV3 and SKOV3 cells rescued the inhibitory effects on cell
proliferation (Fig. 5C; P<0.05)
and motility (Fig. 5D and E;
P<0.05) induced by miR-139 overexpression. These results further
demonstrated that HDGF was a direct target of miR-139 in EOC.
Discussion
The dysregulation of miR-139 was a frequent event
and it is involved in the carcinogenesis and progression of various
kinds of human cancer. For example, in hepatocellular carcinoma,
miR-139 expression was downregulated, and reduced miR-139 levels
were correlated with clinicopathological features, including venous
invasion, microsatellite formation, absence of tumor encapsulation
and reduced differentiation (22).
In colon cancer, expression levels of miR-139 were reduced in tumor
tissues, and its low expression was associated with age.
Furthermore, miR-139 underexpression was correlated with poor
overall survival, particularly in patients with TNM stages I and II
(23). In esophageal squamous cell
carcinoma, reduced miR-139 expression was associated with lymph
node metastases (24). The
downregulation of miR-139 was also observed to correlate with
gastric cancer (25), parathyroid
carcinoma (26), and basal cell
carcinoma (27). However, the
expression pattern of miR-139 in EOC was not investigated. In the
present study, it was observed that miR-139 was significantly
downregulated in EOC tissues and cell lines using RT-qPCR. These
observations suggested that miR-139 may serve important roles in
cancer.
miR-139 has been subject to various studies and
serves significant roles in numerous biological functions. In
hepatocellular carcinoma, upregulation of miR-139 clearly
attenuated cell motility in vitro and the incidence and
severity of lung metastasis from orthotopic liver tumors in
vivo through negative regulation of rho associated coiled-coil
containing protein kinase 2 (22).
Gu et al (28) reported
that miR-139 overexpression suppressed hepatocellular carcinoma
cells growth, migration, invasion and enhanced apoptosis via the
WNT/TCF-4 pathway. In glioma, miR-139 inhibited cell proliferation
and invasion both in vitro and in vivo through
directly targeting IGF-1R, AMY-1 and PGC-1β (29). In addition, restoration of miR-139
repressed glioma cells migration and invasion by targeting ZEB1 and
ZEB2 (30), and improved
temozolomide-induced apoptosis via blockade of MCl-1 (31). Ren et al (32) demonstrated that miR-139 decreased
cell growth and induced cell apoptosis via the protein kinase B
signaling pathway. Zhang et al (33) observed that miR-139 overexpression
inhibited breast cancer cell growth, motility, enhanced cell
apoptosis, caused cell cycle arrest in S phase and improved
chemosensitivity to docetaxel by targeting Notch1. However, the
roles of miR-139 on EOC cells remained to be investigated. The
current study observed that ectopic of miR-139 expression
significantly inhibited proliferation, migration and invasion of
EOC cells. These results suggested that miR-139 acted as a tumor
suppressor in cancer, and could be investigated as a therapeutic
target for the therapy of these types of cancer.
Regarding the molecular mechanism underlying the
regulation effect of miRNA on cancers, it is crucial to explore
their target genes. Firstly, bioinformatics analysis indicated the
presence of miR-139 binding site on the 3′UTR of HDGF. In addition,
luciferase reporter assays indicated that miR-139 overexpression
decreased the luciferase activities of Wt 3′UTR of HDGF, whereas
there was no effect on the Mut 3′UTR of HDGF, suggesting that HDGF
was a direct target gene of miR-139. In addition, results of
RT-qPCR and western blotting indicated that enforced miR-139
expression suppressed HDGF expression at both mRNA and protein
levels in EOC cells. Finally, upregulation of HDGF in miR-139
mimics-transfected cells could rescue the tumor suppressive roles
induced by miR-139 overexpression on cell proliferation, migration
and invasion, further demonstrating that miR-139 inhibited EOC cell
proliferation, migration and invasion through directly targeting
HDGF.
HDGF, located on chromosome 1, region q21-q23
(34), is a heparin-binding growth
factor and firstly purified from culture medium conditioned with
hepatoma-cell line HuH7 (35). In
EOC, HDGF was significantly upregulated in tumor tissues and higher
expression of HDGF was correlated with lymphatic metastasis. In
addition, patients with EOC with higher HDGF levels had a poorer
five-year overall survival rate, and multivariate analysis
identified HDGF as an independent prognostic factor for patients
with EOC (36). Therefore,
regarding its cancer-associated functions, HDGF is worthwhile to be
investigated as a novel therapeutic target in EOC. Increasing
studies indicated that HDGF may be regulated by multiple miRNAs in
various types of cancer, including miR-610 in colorectal cancer
(37), miR-497 in prostate cancer
(38), miR-195 in non-small cell
lung cancer (39), miR-141 in
gastric cancer (40), and miR-214
in hepatocellular carcinoma (41).
In the present study, to the best of our knowledge for the first
time, it was observed that HDGF may be negatively regulated by
miR-139 in EOC, and therefore to inhibit cell growth and
metastasis. Collectively, miR-139 could be investigated as a
targeted therapy to against HDGF and to block EOC rapidly growth
and metastasis.
In conclusion, the present study indicated that
miR-139 exhibited tumor suppressive roles against EOC through
directly targeting HDGF. This newly identified miR-139/HDGF
association provided potential novel therapeutic targets for
patients with EOC.
References
|
1
|
Ferlay J, Soerjomataram I, Dikshit R, Eser
S, Mathers C, Rebelo M, Parkin DM, Forman D and Bray F: Cancer
incidence and mortality worldwide: Sources, methods and major
patterns in GLOBOCAN 2012. Int J Cancer. 136:E359–E386. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Suh DH, Kim JW, Kim K, Kim HJ and Lee KH:
Major clinical research advances in gynecologic cancer in 2012. J
Gynecol Oncol. 24:66–82. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Maldonado L and Hoque MO: Epigenomics and
ovarian carcinoma. Biomark Med. 4:543–570. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Dong R, Liu X, Zhang Q, Jiang Z, Li Y, Wei
Y, Li Y, Yang Q, Liu J, Wei JJ, et al: miR-145 inhibits tumor
growth and metastasis by targeting metadherin in high-grade serous
ovarian carcinoma. Oncotarget. 5:10816–10829. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yiwei T, Hua H, Hui G, Mao M and Xiang L:
HOTAIR Interacting with MAPK1 regulates ovarian cancer skov3 cell
proliferation, migration, and invasion. Med Sci Monit.
21:1856–1863. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Dwivedi SK, Mustafi SB, Mangala LS, Jiang
D, Pradeep S, Rodriguez-Aguayo C, Ling H, Ivan C, Mukherjee P,
Calin GA, et al: Therapeutic evaluation of microRNA-15a and
microRNA-16 in ovarian cancer. Oncotarget. 7:15093–15104. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Chen X, Dong C, Law PT, Chan MT, Su Z,
Wang S, Wu WK and Xu H: MicroRNA-145 targets TRIM2 and exerts
tumor-suppressing functions in epithelial ovarian cancer. Gynecol
Oncol. 139:513–519. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Luo J, Zhou J, Cheng Q, Zhou C and Ding Z:
Role of microRNA-133a in epithelial ovarian cancer pathogenesis and
progression. Oncol Lett. 7:1043–1048. 2014.PubMed/NCBI
|
|
9
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Esquela-Kerscher A and Slack FJ:
Oncomirs-microRNAs with a role in cancer. Nat Rev Cancer.
6:259–269. 2006. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Schmiedel JM, Klemm SL, Zheng Y, Sahay A,
Bluthgen N, Marks DS and van Oudenaarden A: Gene expression.
MicroRNA control of protein expression noise. Science. 348:128–132.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ebert MS and Sharp PA: Roles for microRNAs
in conferring robustness to biological processes. Cell.
149:515–524. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Bartel DP: MicroRNAs: Target recognition
and regulatory functions. Cell. 136:215–233. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Friedman RC, Farh KK, Burge CB and Bartel
DP: Most mammalian mRNAs are conserved targets of microRNAs. Genome
Res. 19:92–105. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Sasaki H, Yoshiike M, Nozawa S, Usuba W,
Katsuoka Y, Aida K, Kitajima K, Kudo H, Hoshikawa M, Yoshioka Y, et
al: Expression level of urinary microRNA-146a-5p is increased in
patients with bladder cancer and decreased in those after
transurethral resection. Clin Genitourin Cancer. 14:e493–e499.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Li Y, Jin L, Chen D, Liu J, Su Z, Yang S,
Gui Y, Mao X, Nie G and Lai Y: Tumor suppressive miR-196a is
associated with cellular migration, proliferation and apoptosis in
renal cell carcinoma. Mol Med Rep. 14:560–566. 2016.PubMed/NCBI
|
|
17
|
Chen Z, Liu H, Jin W, Ding Z, Zheng S and
Yu Y: Tissue microRNA-21 expression predicted recurrence and poor
survival in patients with colorectal cancer-a meta-analysis. Onco
Targets Ther. 9:2615–2624. 2016.PubMed/NCBI
|
|
18
|
Tao S, Liu YB, Zhou ZW, Lian B, Li H, Li
JP and Zhou SF: miR-3646 promotes cell proliferation, migration,
and invasion via regulating G2/M transition in human breast cancer
cells. Am J Transl Res. 8:1659–1677. 2016.PubMed/NCBI
|
|
19
|
Wang N, Wang Q, Shen D, Sun X, Cao X and
Wu D: Downregulation of microRNA-122 promotes proliferation,
migration, and invasion of human hepatocellular carcinoma cells by
activating epithelial-mesenchymal transition. Onco Targets Ther.
9:2035–2047. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen H, Zhang L, Zhang L, Du J, Wang H and
Wang B: MicroRNA-183 correlates cancer prognosis, regulates cancer
proliferation and bufalin sensitivity in epithelial ovarian caner.
Am J Transl Res. 8:1748–1755. 2016.PubMed/NCBI
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wong CC, Wong CM, Tung EK, Au SL, Lee JM,
Poon RT, Man K and Ng IO: The microRNA miR-139 suppresses
metastasis and progression of hepatocellular carcinoma by
down-regulating Rho-kinase 2. Gastroenterology. 140:322–331. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Liu X, Duan B, Dong Y, He C, Zhou H, Sheng
H, Gao H and Zhang X: MicroRNA-139-3p indicates a poor prognosis of
colon cancer. Int J Clin Exp Pathol. 7:8046–8052. 2014.PubMed/NCBI
|
|
24
|
Liu R, Yang M, Meng Y, Liao J, Sheng J, Pu
Y, Yin L and Kim SJ: Tumor-suppressive function of miR-139-5p in
esophageal squamous cell carcinoma. PLoS One. 8:e770682013.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Guo J, Miao Y, Xiao B, Huan R, Jiang Z,
Meng D and Wang Y: Differential expression of microRNA species in
human gastric cancer versus non-tumorous tissues. J Gastroenterol
Hepatol. 24:652–657. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Corbetta S, Vaira V, Guarnieri V,
Scillitani A, Eller-Vainicher C, Ferrero S, Vicentini L, Chiodini
I, Bisceglia M, Beck-Peccoz P, et al: Differential expression of
microRNAs in human parathyroid carcinomas compared with normal
parathyroid tissue. Endocr Relat Cancer. 17:135–146. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Sand M, Skrygan M, Sand D, Georgas D, Hahn
SA, Gambichler T, Altmeyer P and Bechara FG: Expression of
microRNAs in basal cell carcinoma. Br J Dermatol. 167:847–855.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gu W, Li X and Wang J: miR-139 regulates
the proliferation and invasion of hepatocellular carcinoma through
the WNT/TCF-4 pathway. Oncol Rep. 31:397–404. 2014.PubMed/NCBI
|
|
29
|
Wang H, Yan X, Ji LY, Ji XT, Wang P, Guo
SW and Li SZ: miR-139 functions as an antioncomir to repress glioma
progression through targeting IGF-1 R, AMY-1, and PGC-1β. Technol
Cancer Res Treat. Feb 10–2016.(Epub ahead of print).
|
|
30
|
Yue S, Wang L, Zhang H, Min Y, Lou Y, Sun
H, Jiang Y, Zhang W, Liang A, Guo Y, et al: miR-139-5p suppresses
cancer cell migration and invasion through targeting ZEB1 and ZEB2
in GBM. Tumour Biol. 36:6741–6749. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Li RY, Chen LC, Zhang HY, Du WZ, Feng Y,
Wang HB, Wen JQ, Liu X, Li XF, Sun Y, et al: MiR-139 inhibits Mcl-1
expression and potentiates TMZ-induced apoptosis in glioma. CNS
Neurosci Ther. 19:477–483. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Ren Y, Zhu H, Chi C, Yang F and Xu X:
MiRNA-139 regulates oral cancer Tca8113 cells apoptosis through Akt
signaling pathway. Int J Clin Exp Pathol. 8:4588–4594.
2015.PubMed/NCBI
|
|
33
|
Zhang HD, Sun DW, Mao L, Zhang J, Jiang
LH, Li J, Wu Y, Ji H, Chen W, Wang J, et al: MiR-139-5p inhibits
the biological function of breast cancer cells by targeting Notch1
and mediates chemosensitivity to docetaxel. Biochem Biophys Res
Commun. 465:702–713. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Bao C, Wang J, Ma W, Wang X and Cheng Y:
HDGF: A novel jack-of-all-trades in cancer. Future Oncol.
10:2675–2685. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Huang JS, Chao CC, Su TL, Yeh SH, Chen DS,
Chen CT, Chen PJ and Jou YS: Diverse cellular transformation
capability of overexpressed genes in human hepatocellular
carcinoma. Biochem Biophys Res Commun. 315:950–958. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Liu XJ, Liu WL, Yang FM, Yang XQ and Lu
XF: Hepatoma-derived growth factor predicts unfavorable prognosis
of epithelial ovarian cancer. Onco Targets Ther. 8:2101–2109.
2015.PubMed/NCBI
|
|
37
|
Sun B, Gu X, Chen Z and Xiang J: MiR-610
inhibits cell proliferation and invasion in colorectal cancer by
repressing hepatoma-derived growth factor. Am J Cancer Res.
5:3635–3644. 2015.PubMed/NCBI
|
|
38
|
Wu D, Niu X, Pan H, Zhang Z, Zhou Y, Qu P
and Zhou J: MicroRNA-497 targets hepatoma-derived growth factor and
suppresses human prostate cancer cell motility. Mol Med Rep.
13:2287–2292. 2016.PubMed/NCBI
|
|
39
|
Guo H, Li W, Zheng T and Liu Z: MiR-195
targets HDGF to inhibit proliferation and invasion of NSCLC cells.
Tumour Biol. 35:8861–8866. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Chen B, Huang T, Jiang J, Lv L, Li H and
Xia S: miR-141 suppresses proliferation and motility of gastric
cancer cells by targeting HDGF. Mol Cell Biochem. 388:211–218.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Shih TC, Tien YJ, Wen CJ, Yeh TS, Yu MC,
Huang CH, Lee YS, Yen TC and Hsieh SY: MicroRNA-214 downregulation
contributes to tumor angiogenesis by inducing secretion of the
hepatoma-derived growth factor in human hepatoma. J Hepatol.
57:584–591. 2012. View Article : Google Scholar : PubMed/NCBI
|