Introduction
Glioblastoma multiforme (GBM) develops from the
lineage of star-shaped glial cells (1). GBM is a highly prevalent cancer,
accounting for 12–15% of all brain tumors and ~70% of all diagnosed
gliomas (2,3). GBM is defined by the World Health
Organization as the most aggressive form of grade IV glioma,
characterized by unrestrained cellular proliferation, strong
resistance to apoptosis, diffuse infiltration, vigorous
angiogenesis and widespread genomic instability (4). The annual incidence of malignant
glioma is >5/100,000 population (5). Currently, the clinical outcome of
glioma has improved (6–8); however, the median survival rate of
patients with GBM remains at only 12–15 months (9), thus, novel therapeutic targets are
required.
Increasing evidence has confirmed that microRNAs
(miRNAs/miRs) serve essential roles in the pathogenesis of various
tumors. miRNAs are a class of small RNAs, ~18–22 nucleotides in
length (10–12). miRNAs suppress protein expression
through the inhibition of translation or the induction of mRNA
degradation, by binding to the 3′ untranslated region (3′UTR) of
target mRNAs (13). miRNAs are
involved in a diverse range of biological processes, including
cellular proliferation, migration, apoptosis and differentiation
(14–18).
The insulin-like growth factor (IGF) signaling axis
is critical to the growth, development and maintenance of a number
of tissue types (19,20). A previous study demonstrated that
the IGF 1 receptor (IGF1R) is an independent prognostic factor
associated with decreased survival time in patients with GBM
(21). The present study
investigated the role of miR-194-5p in GBM and identified IGF1R as
a target of miR-194-5p. IGF1R may warrant further investigation as
a potential novel therapeutic target.
Materials and methods
Patients and tissue samples
A total of 10 GBM tissues and 10 corresponding
adjacent normal tissues were collected from the Department of
Neurosurgery, The People's Hospital of Leshan City (Leshan, China).
The use of human tissues in the present study was evaluated and
approved by the Ethic Committee of The People's Hospital of Leshan
City. All patients enrolled in the present study provided written
informed consent, and all specimens were handled and made anonymous
according to the ethical and legal standards of China. All GBM
tissues were confirmed and evaluated by a pathologist, and were
frozen at −80°C immediately following surgery.
Cell culture
The normal human astrocytes (NHA; cat. no. 43923),
and human glioblastoma cell lines U373 and U118, were purchased
from the Type Culture Collection of the Chinese Academy of Sciences
(Shanghai, China). All cells were cultured in Dulbecco's modified
Eagle's medium (Gibco; Thermo Fisher Scientific, Inc., Waltham, MA,
USA) with 10% fetal bovine serum (Gibco; Thermo Fisher Scientific,
Inc.) at 37°C with 5% CO2. The U373 cell line is known
to be contaminated with the U251 astrocytoma cell line and the U118
cell line is known to be contaminated with the U138 astrocytoma
cell line (22).
Analysis of miR-194-5p in tissues and
cell lines
Total RNA was extracted from cells using TRIzol
reagent (Invitrogen; Thermo Fisher Scientific, Inc.), according to
the manufacturer's protocol. The level of miR-194-5p was evaluated
using a TaqMan MiRNA Assay (Applied Biosystems; Thermo Fisher
Scientific, Inc.), according to the manufacturer's protocol. U6
small nuclear RNA was used as an internal loading control. Reverse
transcription-quantitative polymerase chain reaction was performed
on an ABI 7900HT instrument (Applied Biosystems; Thermo Fisher
Scientific, Inc.). The primers were synthesized and tested by
Sangon Biotech Co., Ltd. (Shanghai, China). The primer sequences
were as follows: U6 forward, GTG GAC CGC ACA AGC TCG CT, and
reverse, TTG TTG AAC GGC ACT GTG TAT AGC A; miR-194-5p forward, AGT
GTG ACG TTG ACA TCC GT, and reverse, GCA GCT CAG TAA CAG TCC GC.
Relative miR-194-5p expression was calculated according to the
comparative Cq method (23) and
normalized to the expression of U6.
Overexpression and inhibition of
miR-194-5p in vitro
miR-194-5p was overexpressed and inhibited in
vitro by transfection with miR-194-5p mimics and an miR-194-5p
antisense oligonucleotide (ASO), respectively. The miR-194-5p
mimics and miR-194-5p ASO were purchased from Sangon Biotech Co.,
Ltd. The miRNA sequences were as follows: miR-194-5p mimic,
5′-AAGGCAGGGCCCCCGCUCCCC-3′; miR-194-5p ASO,
5′-GGAGCGGGGGCCCUGCCUUUU-3; scrambled miRNA control,
5′-CAGUACUUUUGUGUAGUACAA-3. The cells were seeded (1×106
cells/well) and cultured overnight. Transfection was performed for
30 min using Lipofectamine 2000 (Thermo Fisher Scientific, Inc.),
according to the manufacturer's protocol.
Cellular proliferation assay
After 24, 48 and 72 h of transfection, cellular
proliferation was assessed by the MTT assay. Cells were seeded into
96-well plates at a density of 5×105 cells/well. MTT
reagent was added into the medium at a final concentration of 0.1
mg/ml. A total of 100 µl DMSO was added. Optical density was
measured by a microplate reader at 570 nm.
Cellular apoptosis analysis
A cell suspension (5×105 cells/ml) was
prepared in annexin V-fluorescein isothiocyanate (FITC; Abcam,
Shanghai, China) binding buffer and Annexin V-FITC was added and
incubated for 15 min at room temperature. Subsequently, propidium
iodide (PI; Abcam, Shanghai, China) was added, samples were
analyzed by flow cytometry using an LSR II flow cytometer (BD
Biosciences, Franklin Lakes, NJ, USA) at an excitation wavelength
of 488 nm (argon-ion laser or solid state laser), and an emission
wavelength of 530 nm for FITC and 590 nm for PI (24). Data analysis was performed using
the BD FACSDiva software version 8.0.1 (BD Biosciences).
Prediction of the possible targets of
miR-149-5p
Targetscan software (www.targetscan.org) was used to predict the potential
targets of miR-149-5p.
Dual-luciferase reporter assays
Cells were seeded at 1×105 cells/well and
were serum starved for 6 h prior to transfection. The wild-type
(WT) 3′UTR of IGF1R and mutated controls were cloned and inserted
into the dual-luciferase-reporter plasmid (500 ng; Shengong
Biotechnology Company) and the pGL3-control (100 ng; Promega
Corporation, Madison, WI, USA). miR-194-5p mimics and controls were
subsequently transfected into U373 cells containing wild-type or
mutant 3′UTR plasmids with Lipofectamine 2000 (Invitrogen; Thermo
Fisher Scientific, Inc.). Cells were harvested and luciferase
activities were analyzed after 24 h using the Dual-Luciferase
Reporter assay system (Promega Corporation). The mutant IGF1R
3′UTRs were generated using a Site-Directed Mutagenesis kit
(Promega Corporation). The primers used to perform mutagenesis were
as follows: IGF1R forward,
5′-ctgtccctgatgctgaaggcaggcagaatgacttc-3′, and reverse,
5′-gaagtcattctgctctgccttcagctcagggacag-3′. The kit was used
according to the manufacturer's protocol.
Western blot analysis
Whole-cell lysates were prepared as previously
described (25). The cell lysates
were centrifuged at 4°C, 12,000 × g for 10 min, extracted and the
proteins were quantified using a bicinchoninic acid protein
quantification kit (cat. no. ab102536; Abcam, Cambridge, UK).
Proteins (30 µg) were separated via 4–10% SDS-PAGE. Membranes were
electrotransferred to nitrocellulose membranes (EMD Millipore,
Billerica, MA, USA). The membranes were blocked with 5% non-fat
milk overnight at 4°C and incubated with the anti-IGF1R antibody
(cat. no. 3027; 1:1,000; Cell Signaling Technology, Inc., Danvers,
MA, USA) overnight at 4°C. β-actin (cat. no. 4967; 1:1,000; Cell
Signaling Technology, Inc.) was used as the endogenous control,
incubated overnight at 4°C. Then the membranes were incubated with
an anti-rabbit horseradish peroxidase-conjugated secondary antibody
(cat. no. 7074; 1:1,000; Cell Signaling Technology, Inc.) for 2 h
at room temperature. The bands were visualized using enhanced
chemiluminescence (GE Healthcare Life Sciences, Little Chalfont,
UK).
Statistical analysis
Data are presented as the mean ± standard deviation
of 3 independent experiments. Statistical analysis was performed
using SPSS software (version 10.0; SPSS, Inc., Chicago, IL, USA).
The difference between two groups was analyzed using a two-tailed
Student's t-test. One-way analysis of variance was used to analyze
the difference between multiple groups, followed by the
Student-Newman-Keuls method. P<0.05 was considered to indicate a
statistically significant difference.
Results
miR-194-5p targets IGF1R
IGF1R is an independent prognostic factor associated
with decreased survival of patients with GBM (21). Therefore, miR-194-5p was selected
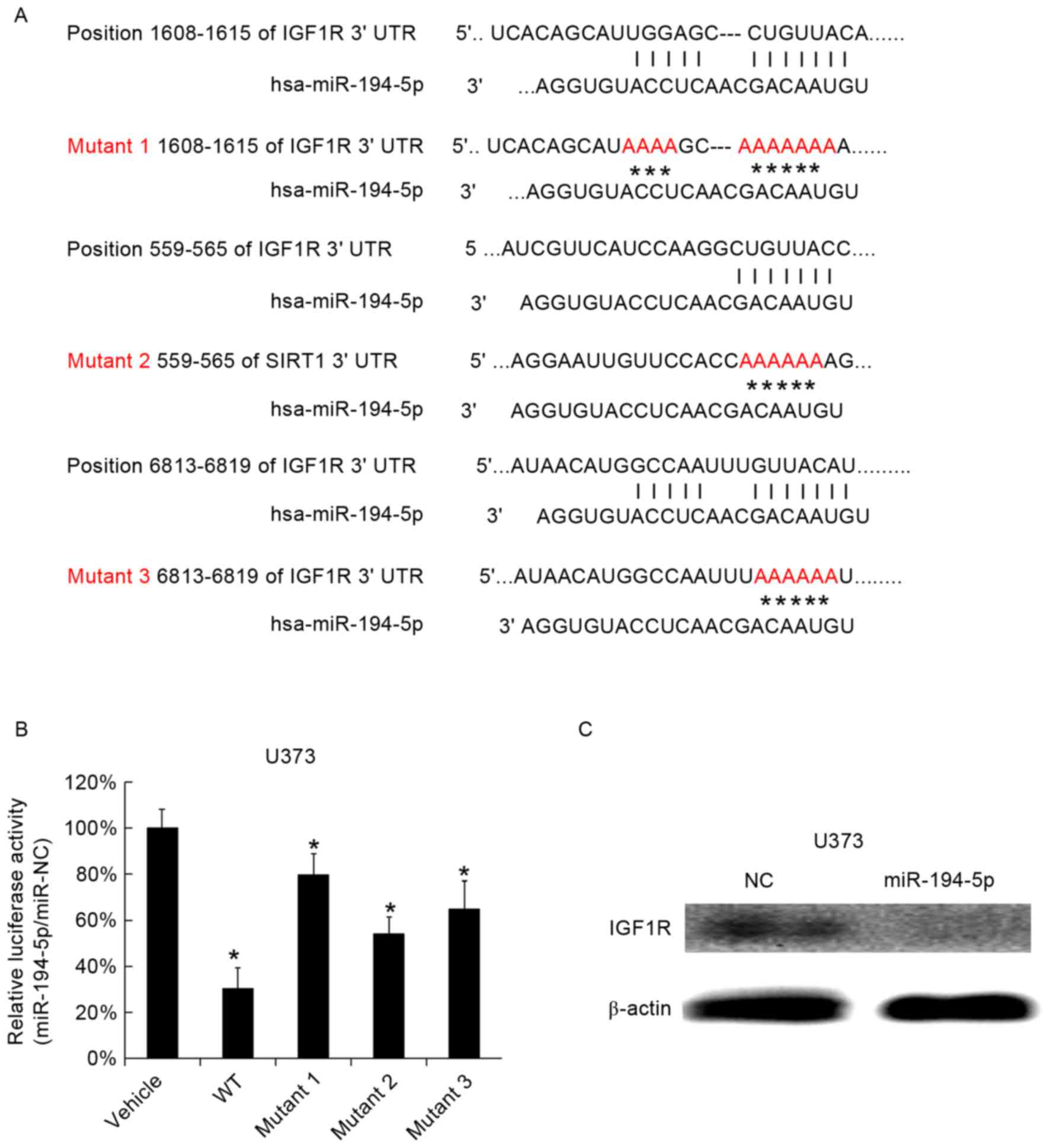
for investigation, as it was identified to target IGF1R. A total of
three miR-194-5p binding sites were identified in the 3′UTR of the
IGF1R gene and three mutants were subsequently generated (Fig. 1A). The three mutated versions of
the IGF1R 3′UTRs were cloned into luciferase reporter plasmids.
miR-194-5p mimics and the mutant 3′UTRs were cotransfected into
U373 cells, and it was observed that the miR-194-5p mimics reduced
the luciferase activity of the WT 3′UTR, while the mutated 3′UTRs
partially reduced the inhibitory function of miR-194-5p, scrambled
miRNA was used as a vehicle (Fig.
1B). miR-194-5p mimics were subsequently transfected into U373
cells and the IGF-1R protein level was observed to be reduced 48 h
following transfection (Fig. 1C).
The results indicated that miR-194-5p may serve a role in GBM via
IGF1R, thus miR-194-5p was selected for further study.
Decreased miR-194-5p in GBM
tissues
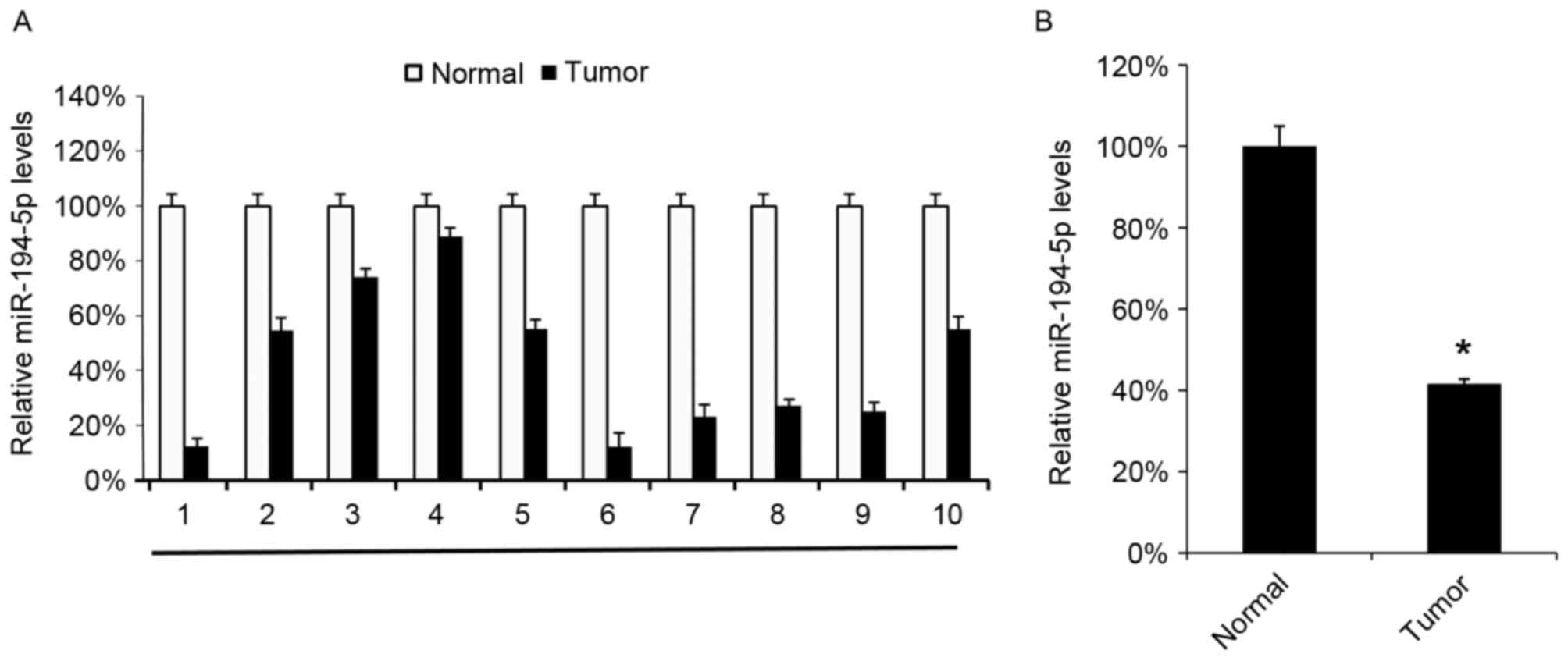
miR-194-5p levels were evaluated in 10 GBM tissues
and compared with the miR-194-5p levels in the 10 corresponding
adjacent normal tissues. All the GBM tissues were observed to
express decreased miR-194-5p levels compared with the corresponding
adjacent normal tissues (Fig. 2A).
The mean levels of the 10 GBM tissues also exhibited a decreased
level of miR-194-5p compared with the normal tissues (Fig. 2B).
Effect of miR-194-5p mimics on GBM
cell growth and apoptosis
miR-194-5p was overexpressed in GBM cells to test
the function of miR-194-5p. The miR-194-5p levels in U373, U118 and
NHA cells were analyzed by RT-qPCR, and it was observed that U373
and U118 cells expressed decreased levels of miR-194-5p compared
with NHA cells (Fig. 3A).
miR-194-5p was successfully overexpressed in U373 and U118 cells by
miR-194-5p mimic transfection (Fig.
3B). MTT analysis revealed that the miR-194-5p mimics inhibited
the growth of U373 and U118 cells (Fig. 3C). The cellular apoptotic rate was
analyzed 48 h following miR-194-5p mimic transfection, and it was
observed that miR-194-5p upregulated the apoptotic rate of U373 and
U118 cells (Fig. 3D).
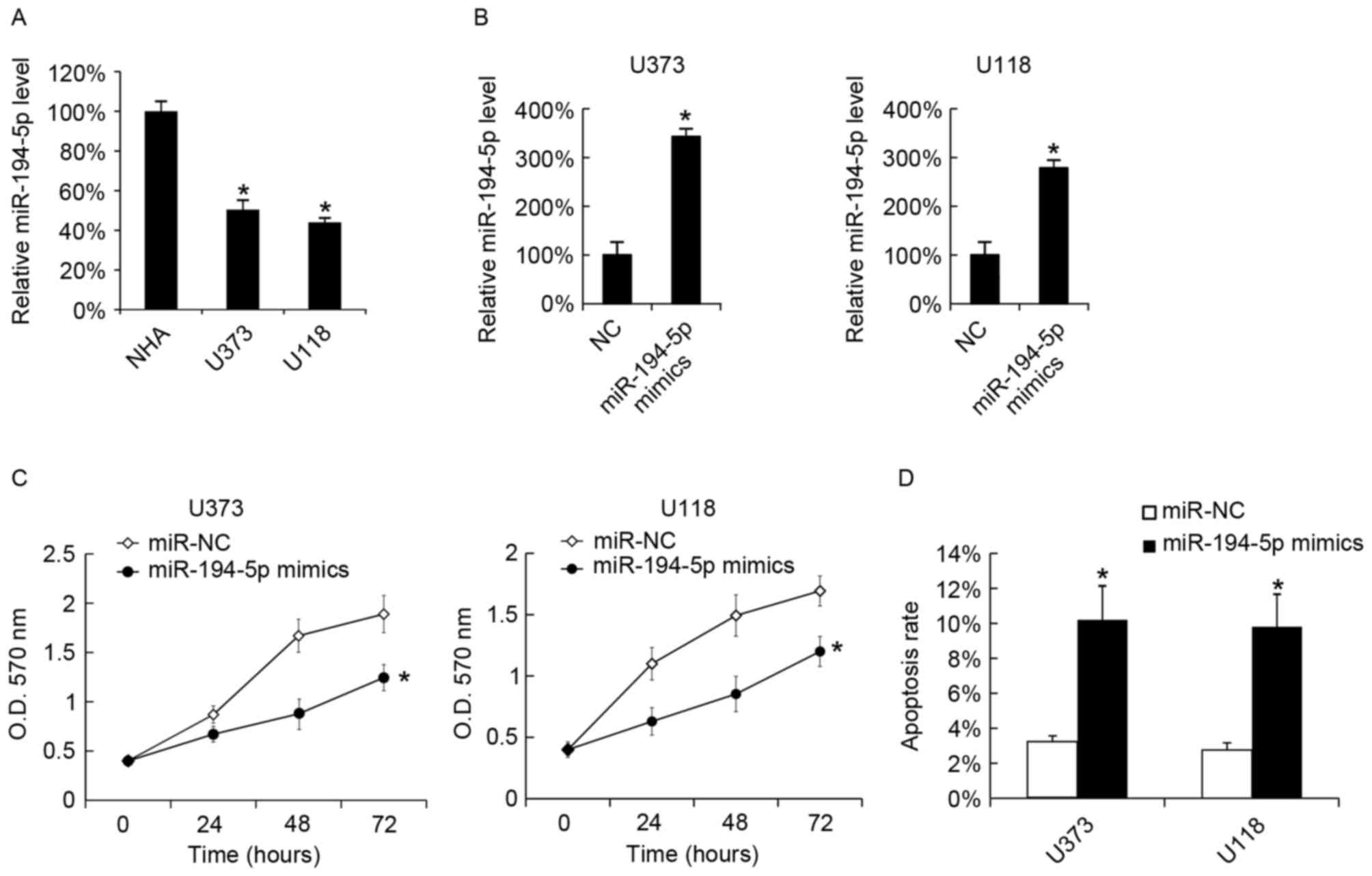 | Figure 3.Transfection with miR-194-5p mimics
inhibits U373 and U118 cellular proliferation, and promotes
cellular apoptosis. (A) miR-194-5p levels in NHA, U373 and U118
cells were analyzed by RT-qPCR. miR-194-5p mimics were transfected
into (B) U373 and (C) U118 cells, and the miR-194-5p levels were
evaluated by RT-qPCR 48 h following transfection. (C) After 24, 48
and 72 h of transfection, cellular proliferation was analyzed using
the MTT assay. (D) A total of 48 h following miR-194-5p mimic
transfection, cellular apoptosis was measured. The data are
presented as the mean ± standard deviation. Experiments were
repeated at least three times. *P<0.05 vs. the miR-NC. miR,
microRNA; NHA, normal human astrocyte; RT-qPCR, reverse
transcription-quantitative polymerase chain reaction; NC, negative
control; O.D., optical density. |
Effect of miR-194-5p ASO on cell
growth
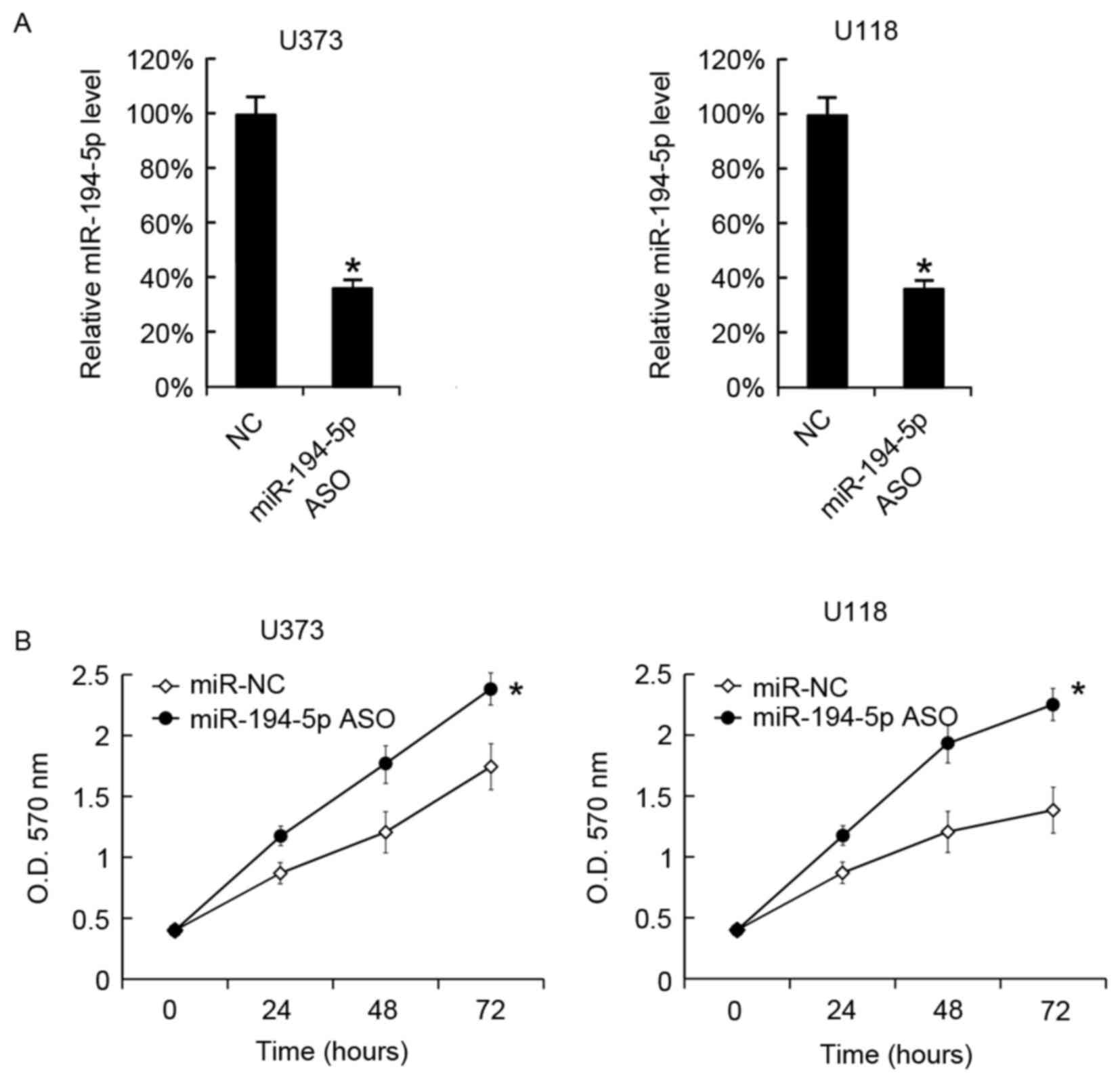
miR-194-5p was successfully downregulated in U373
and U118 cells by miR-194-5p ASO transfection. After 48 h, the
miR-194-5p levels in cells were analyzed by RT-qPCR (Fig. 4A). Cellular proliferation in the
two cell lines was analyzed using the MTT assay, and it was
observed that miR-194-5p ASO promoted the cellular proliferation of
U373 and U118 cells (Fig. 4B).
Discussion
miR-194-5p inhibited cellular proliferation,
invasion and migration in non-small cell lung cancer by targeting
forkhead box A1 (26). In the
present study, miR-194-5p was demonstrated to inhibit cellular
proliferation and apoptosis in GBM by targeting IGF1R.
GBM immunotherapy studies observed that macrophages
accumulate during GBM progression and may be targeted by inhibition
of colony-stimulating factor-1 receptor (CSF-1R) (27,28).
In clinical trials, several approaches to inhibit CSF-1R are
currently being used, including antibodies and small molecules
(29–31). Quail et al (32) studied acquired resistance to CSF-1R
inhibition immunotherapy in glioma in 2016 and demonstrated that
the phosphatidylinositol 3-kinase pathway activity was elevated in
recurrent GBM, driven by macrophage-derived IGF1 and IGF1R. IGF1R
inhibition with CSF-1R inhibition in recurrent tumors significantly
prolonged overall survival in mice. In the present study,
miR-194-5p was observed to target IGF1R in GBM cell lines.
Therefore, it was hypothesized that miR-194-5p may be useful in GBM
therapy. It was hypothesized that overexpression of miR-194-5p may
inhibit tumor cell growth and promote cell apoptosis, and that
inhibition of IGF1R may promote the effect of the
macrophage-targeted immunotherapy. The role of miR-194-5p in
macrophage-targeted immunotherapy requires further
investigation.
In conclusion, the results of the present study
demonstrated that miR-194-5p inhibited cellular proliferation and
promoted cell apoptosis. The target gene of miR-194-5p is IGF1R.
The present study demonstrated that miR-194-5p inhibited GBM cell
growth and promoted apoptosis via IGF1R.
References
|
1
|
Parsons DW, Jones S, Zhang X, Lin JC,
Leary RJ, Angenendt P, Mankoo P, Carter H, Siu IM, Gallia GL, et
al: An integrated genomic analysis of human glioblastoma
multiforme. Science. 321:1807–1812. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ohgaki H and Kleihues P: Genetic pathways
to primary and secondary glioblastoma. Am J Pathol. 170:1445–1453.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ohgaki H and Kleihues P: Population-based
studies on incidence, survival rates, and genetic alterations in
astrocytic and oligodendroglial gliomas. J Neuropathol Exp Neurol.
64:479–489. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ramakrishna R and Pisapia D: Recent
molecular advances in our understanding of glioma. Cureus.
7:e2872015.PubMed/NCBI
|
|
5
|
Fisher JL, Schwartzbaum JA, Wrensch M and
Wiemels JL: Epidemiology of brain tumors. Neurol Clin. 25:867–90,
vii. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Stupp R, Hegi ME, van den Bent MJ, Mason
WP, Weller M, Mirimanoff RO and Cairncross JG; European
Organisation for Research and Treatment of Cancer Brain Tumor and
Radiotherapy Groups, ; National Cancer Institute of Canada Clinical
Trials Group, : Changing paradigms-an update on the
multidisciplinary management of malignant glioma. Oncologist.
11:165–180. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Oermann E, Collins BT, Erickson KT, Yu X,
Lei S, Suy S, Hanscom HN, Kim J, Park HU, Eldabh A, et al:
CyberKnife® enhanced conventionally fractionated
chemoradiation for high grade glioma in close proximity to critical
structures. J Hematol Oncol. 3:222010. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ottenhausen M, Krieg SM, Meyer B and
Ringel F: Functional preoperative and intraoperative mapping and
monitoring: Increasing safety and efficacy in glioma surgery.
Neurosurg Focus. 38:E32015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Paw I, Carpenter RC, Watabe K, Debinski W
and Lo HW: Mechanisms regulating glioma invasion. Cancer Lett.
362:1–7. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Xie B, Ding Q, Han H and Wu D: miRCancer:
A microRNA-cancer association database constructed by text mining
on literature. Bioinformatics. 29:638–644. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Mitchell PS, Parkin RK, Kroh EM, Fritz BR,
Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O'Briant
KC, Allen A, et al: Circulating microRNAs as stable blood-based
markers for cancer detection. Proc Natl Acad Sci USA. 105:pp.
10513–10518. 2008; View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Shimono Y, Zabala M, Cho RW, Lobo N,
Dalerba P, Qian D, Diehn M, Liu H, Panula SP, Chiao E, et al:
Downregulation of miRNA-200c links breast cancer stem cells with
normal stem cells. Cell. 138:592–603. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ambros V: The functions of animal
microRNAs. Nature. 431:350–355. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Babae N, Bourajjaj M, Liu Y, Van Beijnum
JR, Cerisoli F, Scaria PV, Verheul M, Van Berkel MP, Pieters EH,
Van Haastert RJ, et al: Systemic miRNA-7 delivery inhibits tumor
angiogenesis and growth in murine xenograft glioblastoma.
Oncotarget. 5:6687–6700. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hwang H and Mendell J: microRNAs in cell
proliferation, cell death, and tumorigenesis. Br J Cancer.
94:776–780. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Frankel LB, Wen J, Lees M, Høyer-Hansen M,
Farkas T, Krogh A, Jäättelä M and Lund AH: microRNA-101 is a potent
inhibitor of autophagy. EMBO J. 30:4628–4641. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhang H, Li Y and Lai M: The microRNA
network and tumor metastasis. Oncogene. 29:937–948. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Dontula R, Dinasarapu A, Chetty C, Pannuru
P, Herbert E, Ozer H and Lakka SS: microRNA 203 modulates glioma
cell migration via Robo1/ERK/MMP-9 signaling. Genes cancer.
4:285–296. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
LeRoith D and Roberts CT Jr: The
insulin-like growth factor system and cancer. Cancer Lett.
195:127–137. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Denduluri SK, Idowu O, Wang Z, Liao Z, Yan
Z, Mohammed MK, Ye J, Wei Q, Wang J, Zhao L and Luu HH:
Insulin-like growth factor (IGF) signaling in tumorigenesis and the
development of cancer drug resistance. Genes Dis. 2:13–25. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Maris C, D'Haene N, Trépant AL, Le Mercier
M, Sauvage S, Allard J, Rorive S, Demetter P, Decaestecker C and
Salmon I: IGF-IR: A new prognostic biomarker for human
glioblastoma. Br J Cancer. 113:729–737. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Capes-Davis A, Theodosopoulos G, Atkin I,
Drexler HG, Kohara A, MacLeod RA, Masters JR, Nakamura Y, Reid YA,
Reddel RR and Freshney RI: Check your cultures! A list of
cross-contaminated or misidentified cell lines. Int J Cancer.
127:1–8. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Vermes I, Haanen C, Steffens-Nakken H and
Reutellingsperger C: A novel assay for apoptosis. Flow cytometric
detection of phosphatidylserine expression on early apoptotic cells
using fluorescein labelled Annexin V. J Immunol Methods. 184:39–51.
1995. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Song B, Zhang C, Li G, Jin G and Liu C:
miR-940 inhibited pancreatic ductal adenocarcinoma growth by
targeting MyD88. Cell Physiol Biochem. 35:1167–1177. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhu X, Li D, Yu F, Jia C, Xie J, Ma Y, Fan
S, Cai H, Luo Q, Lv Z and Fan L: miR-194 inhibits the
proliferation, invasion, migration, and enhances the
chemosensitivity of non-small cell lung cancer cells by targeting
forkhead box A1 protein. Oncotarget. 7:13139–13152. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Josephs DH, Bax HJ and Karagiannis SN:
Tumour-associated macrophage polarisation and re-education with
immunotherapy. Front Biosci (Elite Ed). 7:293–308. 2015.PubMed/NCBI
|
|
28
|
Seton-Rogers S: Tumour microenvironment:
Teaching old macrophages new tricks. Nat Rev Cancer. 13:7532013.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ruffell B and Coussens LM: Macrophages and
therapeutic resistance in cancer. Cancer Cell. 27:462–472. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ries CH, Cannarile MA, Hoves S, Benz J,
Wartha K, Runza V, Rey-Giraud F, Pradel LP, Feuerhake F, Klaman I,
et al: Targeting tumor-associated macrophages with anti-CSF-1R
antibody reveals a strategy for cancer therapy. Cancer Cell.
25:846–859. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Tap WD, Wainberg ZA, Anthony SP, Ibrahim
PN, Zhang C, Healey JH, Chmielowski B, Staddon AP, Cohn AL, Shapiro
GI, et al: Structure-guided blockade of CSF1R kinase in
tenosynovial giant-cell tumor. N Engl J Med. 373:428–437. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Quail DF, Bowman RL, Akkari L, Quick ML,
Schuhmacher AJ, Huse JT, Holland EC, Sutton JC and Joyce JA: The
tumor microenvironment underlies acquired resistance to CSF-1R
inhibition in gliomas. Science. 352:aad30182016. View Article : Google Scholar : PubMed/NCBI
|