|
1
|
Stewart B and Wild CP: World Cancer Report
2014. IARC press; Lyon: 2015
|
|
2
|
Lauren P: The two histological main types
of gastric carcinoma: Diffuse and so-called intestinal-type
carcinoma. An attempt at a histo-clinical classification. Acta
Pathol Microbiol Scand. 64:31–49. 1965. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wadhwa R, Taketa T, Sudo K, Blum MA and
Ajani JA: Modern oncological approaches to gastric adenocarcinoma.
Gastroenterol Clin North Am. 42:359–369. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wagner AD, Grothe W, Haerting J, Kleber G,
Grothey A and Fleig WE: Chemotherapy in advanced gastric cancer: A
systematic review and meta-analysis based on aggregate data. J Clin
Oncol. 24:2903–2909. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chen X, Leung SY, Yuen ST, Chu KM, Ji J,
Li R, Chan AS, Law S, Troyanskaya OG, Wong J, et al: Variation in
gene expression patterns in human gastric cancers. Mol Biol Cell.
14:3208–3215. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Boussioutas A, Li H, Liu J, Waring P, Lade
S, Holloway AJ, Taupin D, Gorringe K, Haviv I, Desmond PV and
Bowtell DD: Distinctive patterns of gene expression in premalignant
gastric mucosa and gastric cancer. Cancer Res. 63:2569–2577.
2003.PubMed/NCBI
|
|
7
|
Tan P and Yeoh KG: Genetics and molecular
pathogenesis of gastric adenocarcinoma. Gastroenterology.
149:1153–1162.e3. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Pharoah PD, Guilford P and Caldas C;
International Gastric Cancer Linkage Consortium, : Incidence of
gastric cancer and breast cancer in CDH1 (E-cadherin) mutation
carriers from hereditary diffuse gastric cancer families.
Gastroenterology. 121:1348–1353. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Chang HR, Nam S, Kook MC, Kim KT, Liu X,
Yao H, Jung HR, Lemos R Jr, Seo HH, Park HS, et al: HNF4α is a
therapeutic target that links AMPK to WNT signalling in early-stage
gastric cancer. Gut. 1–32. 2014.
|
|
10
|
Kim YH, Liang H, Liu X, Lee JS, Cho JY,
Cheong JH, Kim H, Li M, Downey TJ, Dyer MD, et al: AMPKα modulation
in cancer progression: Multilayer integrative analysis of the whole
transcriptome in Asian gastric cancer. Cancer Res. 72:2512–2521.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Mäder U, Nicolas P, Richard H, Bessières P
and Aymerich S: Comprehensive identification and quantification of
microbial transcriptomes by genome-wide unbiased methods. Curr Opin
Biotechnol. 22:32–41. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhang P, Li C, Zhu L, Su X, Li Y, Jin C
and Li T: De novo assembly of the sea cucumber Apostichopus
japonicus hemocytes transcriptome to identify miRNA targets
associated with skin ulceration syndrome. PLoS One. 8:1254–1256.
2013.
|
|
13
|
Schmieder R and Edwards R: Quality control
and preprocessing of metagenomic datasets. Bioinformatics.
27:863–864. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kim D, Pertea G, Trapnell C, Pimentel H,
Kelley R and Salzberg SL: TopHat2: Accurate alignment of
transcriptomes in the presence of insertions, deletions and gene
fusions. Genome Biol. 14:R362013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Anders S, Pyl PT and Huber W: HTSeq-a
Python framework to work with high-throughput sequencing data.
Bioinformatics. 31:166–169. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Robinson MD, McCarthy DJ and Smyth GK:
edgeR: A Bioconductor package for differential expression analysis
of digital gene expression data. Bioinformatics. 26:139–140. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
McCarthy DJ, Chen Y and Smyth GK:
Differential expression analysis of multifactor RNA-Seq experiments
with respect to biological variation. Nucleic Acids Res.
40:4288–4297. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Law CW, Chen Y, Shi W and Smyth GK: voom:
Precision weights unlock linear model analysis tools for RNA-seq
read counts. Genome biol. 15:R292014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Smyth GK: Limma: Linear models for
microarray dataBioinformatics and computational biology solutions
using R and Bioconductor. Springer; New York, NY: pp. 397–420.
2005, View Article : Google Scholar
|
|
20
|
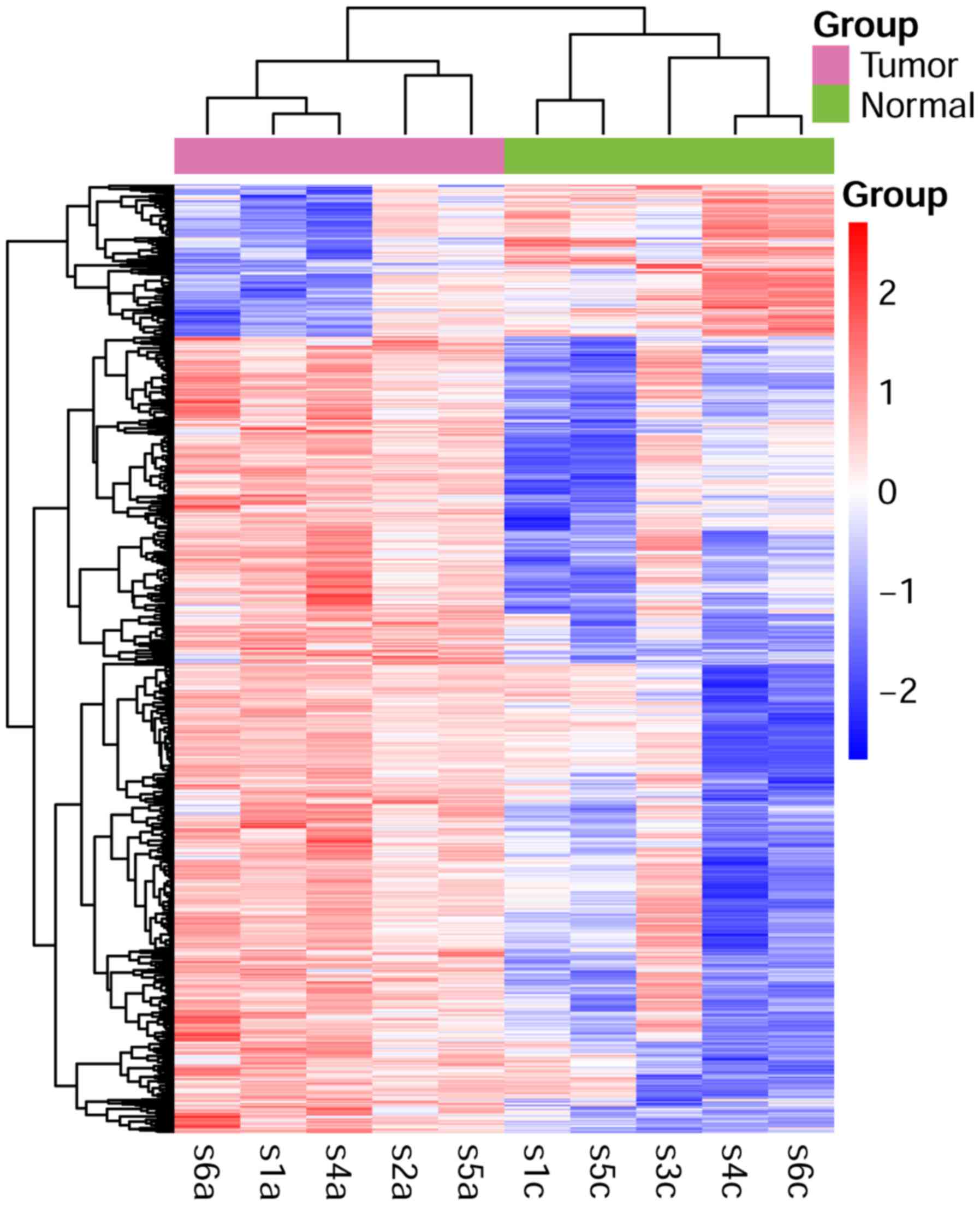
Kolde R: pheatmap: Pretty heatmaps. R
package version 1.0. 8. 2015.
|
|
21
|
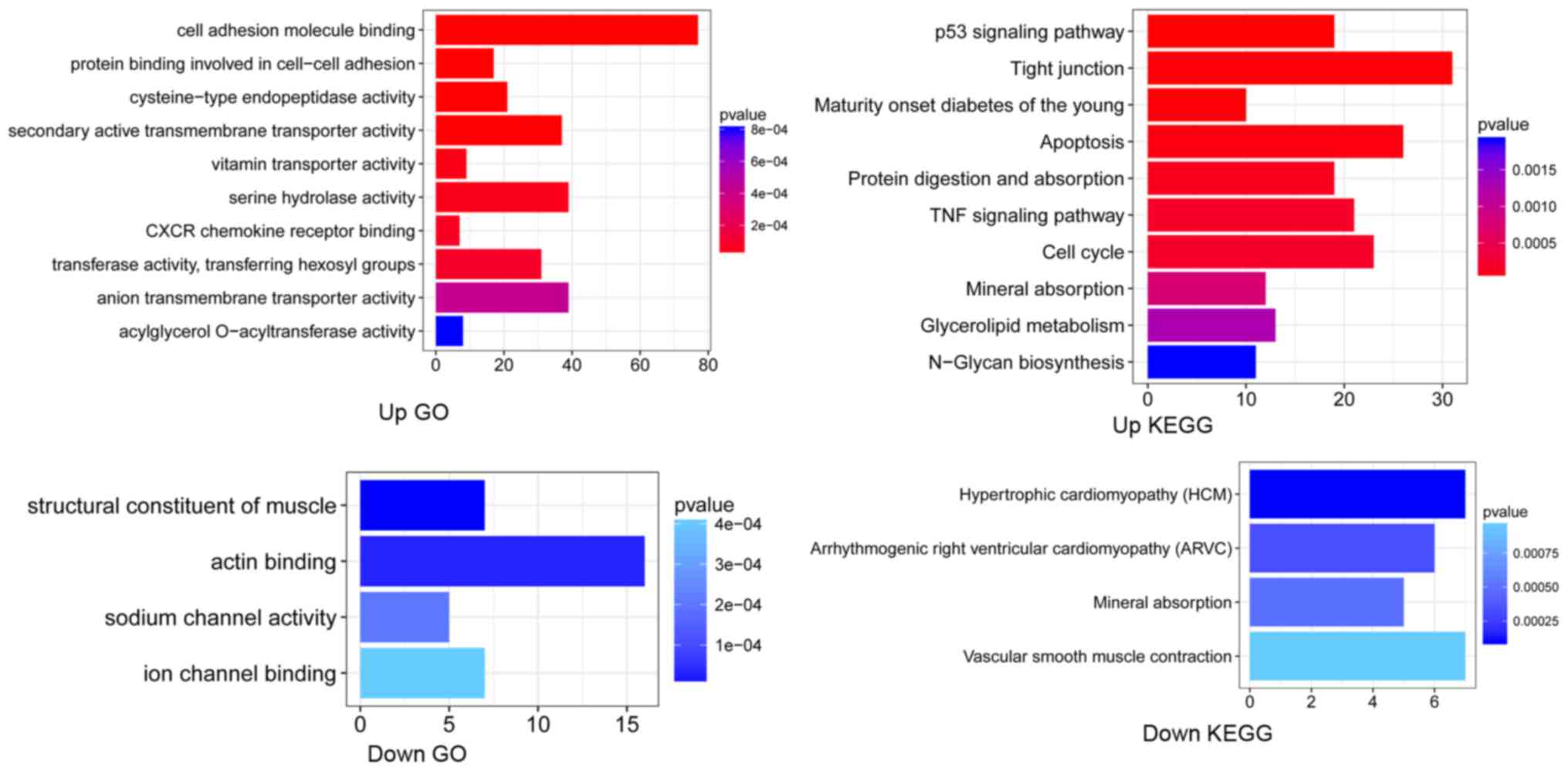
Ashburner M, Ball CA, Blake JA, Botstein
D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT,
et al: Gene ontology: Tool for the unification of biology. The gene
ontology consortium. Nat Genet. 25:25–29. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kanehisa M and Goto S: KEGG: Kyoto
encyclopedia of genes and genomes. Nucleic Res. 28:27–30. 2000.
View Article : Google Scholar
|
|
23
|
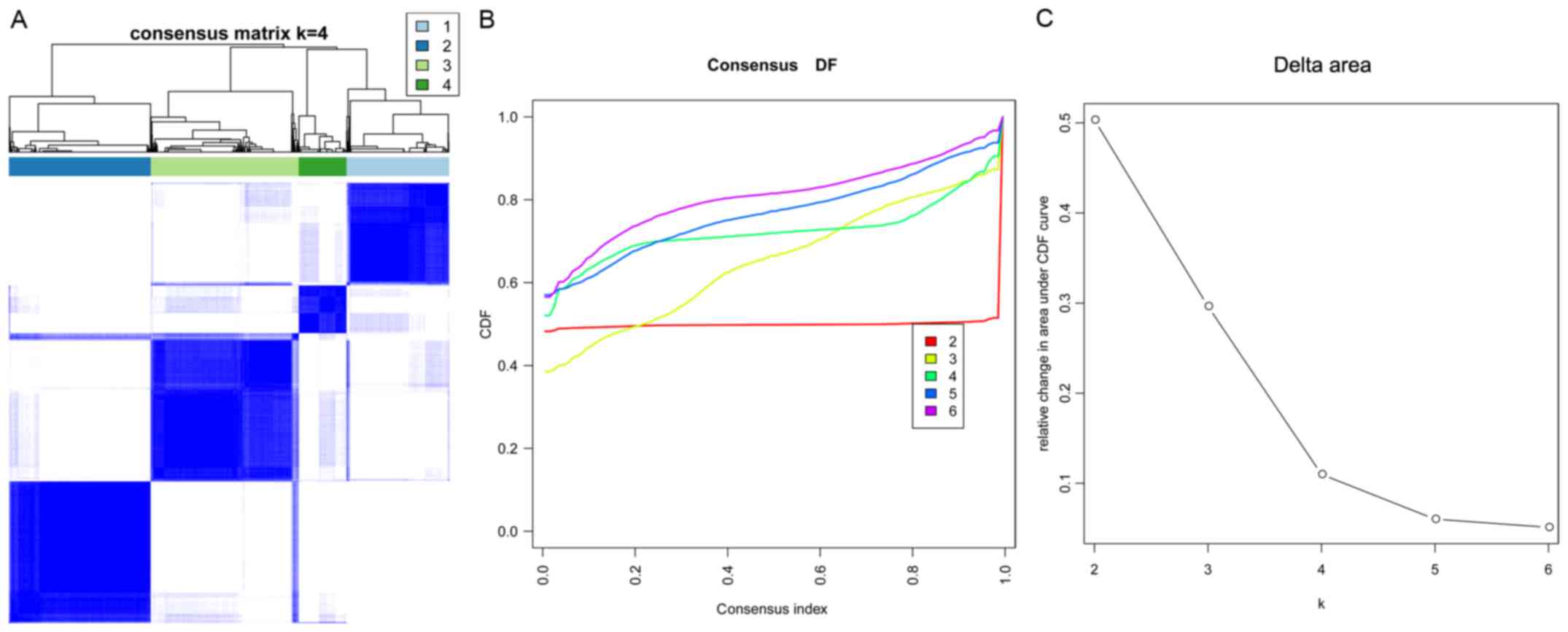
Wilkerson MD and Hayes DN:
ConsensusClusterPlus: A class discovery tool with confidence
assessments and item tracking. Bioinformatics. 26:1572–1573. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Xue B, Oldfield CJ, Dunker AK and Uversky
VN: CDF it all: Consensus prediction of intrinsically disordered
proteins based on various cumulative distribution functions. FEBS
Lett. 583:1469–1474. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
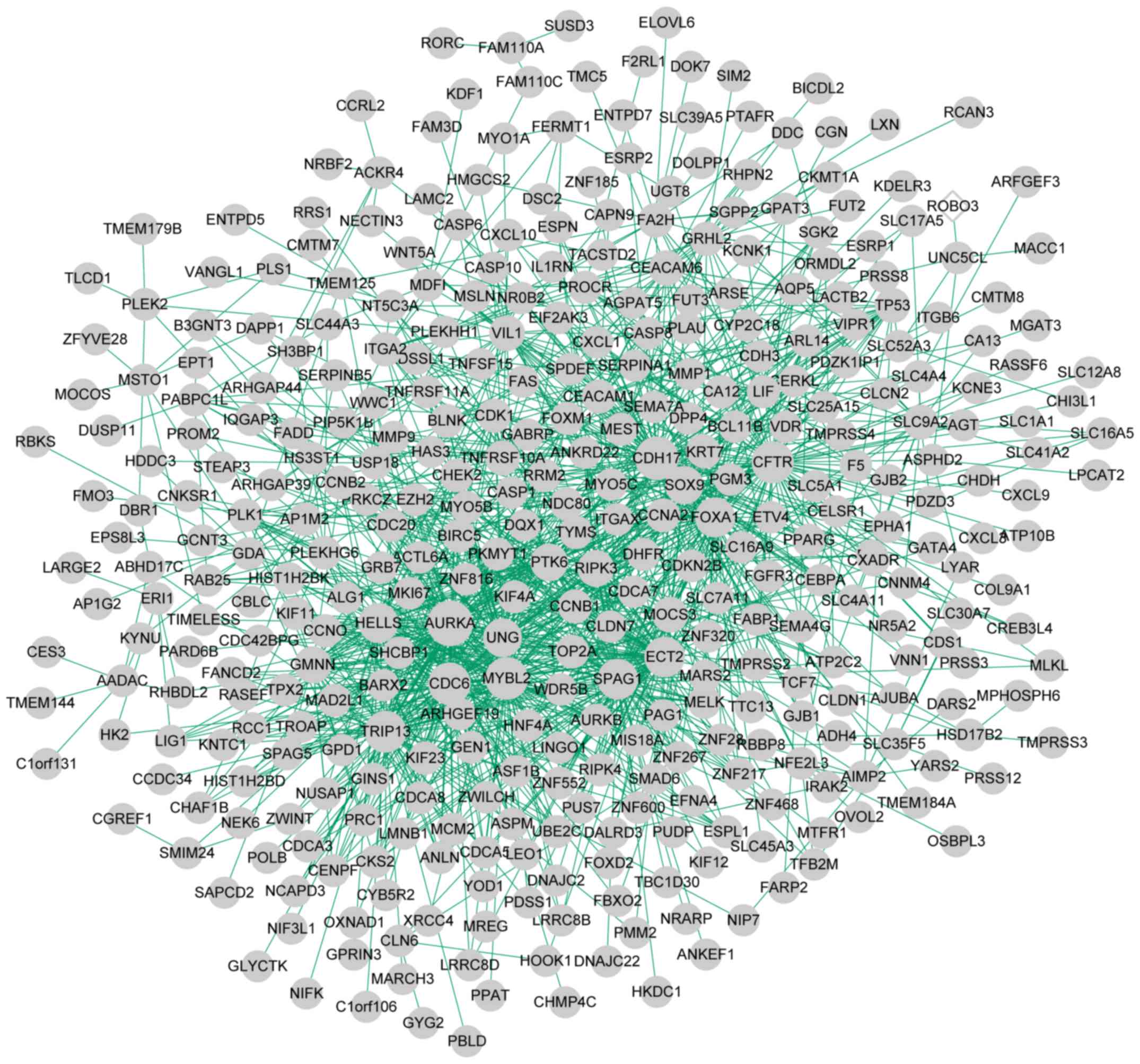
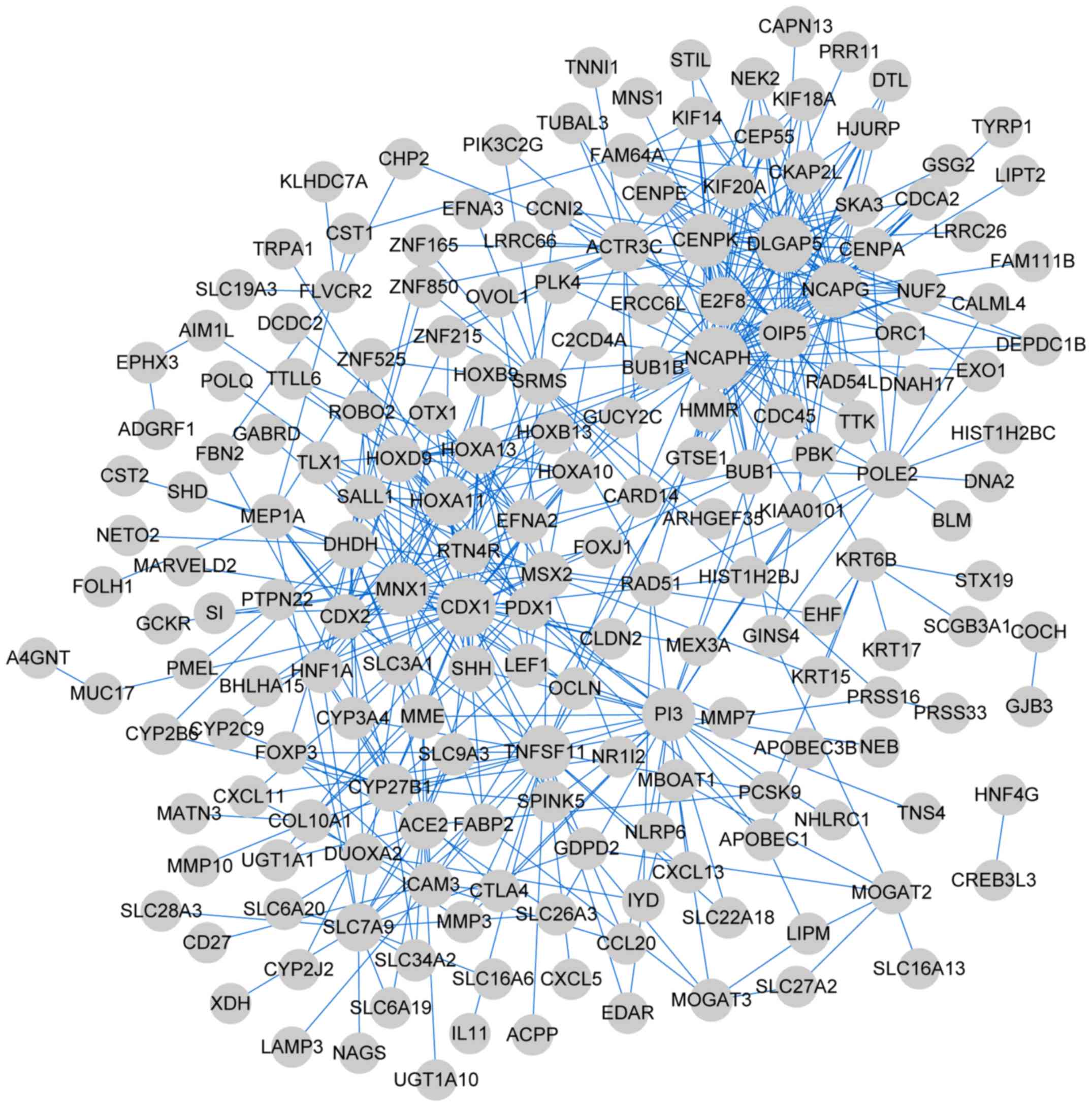
Szklarczyk D, Franceschini A, Wyder S,
Forslund K, Heller D, Huerta-Cepas J, Simonovic M, Roth A, Santos
A, Tsafou KP, et al: STRING v10: Protein-protein interaction
networks, integrated over the tree of life. Nucleic Acids Res.
43:D447–D452. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yu Tang ML, Jianxin, Wang Yi and Pan
Fang-Xiang Wu: CytoNCA: A cytoscape plugin for centrality analysis
and evaluation of biological networks. Biosystems. 127:67–72. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
He X and Zhang J: Why do hubs tend to be
essential in protein networks? PLoS Genet. 2:e882006. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Nebert DW and Dalton TP: The role of
cytochrome P450 enzymes in endogenous signalling pathways and
environmental carcinogenesis. Nat Rev Cancer. 6:9472006. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Nebert DW and Russell DW: Clinical
importance of the cytochromes P450. Lancet. 360:1155–1162. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Tsukino H, Kuroda Y, Qiu D, Nakao H, Imai
H and Katoh T: Effects of cytochrome P450 (CYP) 2A6 gene deletion
and CYP2E1 genotypes on gastric adenocarcinoma. Int J Cancer.
100:425–428. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Lee SK, Moon J, Park SW, Song SY, Chung JB
and Kang JK: Loss of the tight junction protein claudin 4
correlates with histological growth-pattern and differentiation in
advanced gastric adenocarcinoma. Oncol Rep. 13:193–199.
2005.PubMed/NCBI
|
|
33
|
Johnson AH, Frierson HF, Zaika A, Powell
SM, Roche J, Crowe S, Moskaluk CA and El-Rifai W: Expression of
tight-junction protein claudin-7 is an early event in gastric
tumorigenesis. Am J Pathol. 167:577–584. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Yasuda K, Hirohashi Y, Kuroda T, Takaya A,
Kubo T, Kanaseki T, Tsukahara T, Hasegawa T, Saito T, Sato N and
Torigoe T: MAPK13 is preferentially expressed in gynecological
cancer stem cells and has a role in the tumor-initiation. Biochem
Biophys Res Commun. 472:643–647. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Franco AT, Corken A and Ware J: Platelets
at the interface of thrombosis, inflammation and cancer. Blood.
126:582–588. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Nash GF, Turner LF, Scully MF and Kakkar
AK: Platelets and cancer. Lancet Oncol. 3:4252002. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Taucher S, Salat A, Gnant M, Kwasny W,
Mlineritsch B, Menzel RC, Schmid M, Smola MG, Stierer M, Tausch C,
et al: Impact of pretreatment thrombocytosis on survival in primary
breast cancer. Thromb Haemost. 89:1098–1106. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Gasic GJ, Gasic TB and Stewart CC:
Antimetastatic effects associated with platelet reduction. Proc
Natl Acad Sci USA. 61:pp. 46–52. 1968; View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Pyagay P, Heroult M, Wang Q, Lehnert W,
Belden J, Liaw L, Friesel RE and Lindner V: Collagen triple helix
repeat containing 1, a novel secreted protein in injured and
diseased arteries, inhibits collagen expression and promotes cell
migration. Circ Res. 96:261–268. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Wang P, Wang YC, Chen XY, Shen ZY, Cao H,
Zhang YJ, Yu J, Zhu JD, Lu YY and Fang JY: CTHRC1 is upregulated by
promoter demethylation and transforming growth factor-β1 and may be
associated with metastasis in human gastric cancer. Cancer Sci.
103:1327–1333. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Tang L, Dai DL, Su M, Martinka M, Li G and
Zhou Y: Aberrant expression of collagen triple helix repeat
containing 1 in human solid cancers. Clin Cancer Res. 12:3716–3722.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Ebinu JO, Bottorff DA, Chan EY, Stang SL,
Dunn RJ and Stone JC: RasGRP, a Ras guanyl nucleotide-releasing
protein with calcium-and diacylglycerol-binding motifs. Science.
280:1082–1086. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Lauchle JO, Kim D, Le DT, Akagi K, Crone
M, Krisman K, Warner K, Bonifas JM, Li Q, Coakley KM, et al:
Response and resistance to MEK inhibition in leukaemias initiated
by hyperactive Ras. Nature. 461:411–414. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Oki-Idouchi CE and Lorenzo PS: Transgenic
overexpression of RasGRP1 in mouse epidermis results in spontaneous
tumors of the skin. Cancer Res. 67:276–280. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Yang D, Kedei N, Li L, Tao J, Velasquez
JF, Michalowski AM, Tóth BI, Marincsák R, Varga A, Bíró T, et al:
RasGRP3 contributes to formation and maintenance of the prostate
cancer phenotype. Cancer Res. 70:7905–7917. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Djiogue S, Kamdje AH, Vecchio L, Kipanyula
MJ, Farahna M, Aldebasi Y and Seke Etet PF: Insulin resistance and
cancer: the role of insulin and IGFs. Endocr Relat Cancer.
20:R1–R17. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Bruce W and Corpet D: The colonic protein
fermentation and insulin resistance hypotheses for colon cancer
etiology: Experimental tests using precursor lesions. Eur J Cancer
Prev. 5:41–47. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Trevisan M, Liu J, Muti P, Misciagna G and
Menotti A; Risk Factors and Life Expectancy Research Group, :
Markers of insulin resistance and colorectal cancer mortality.
Cancer Epidemiol Biomarkers Prev. 10:937–941. 2001.PubMed/NCBI
|
|
49
|
Mu N, Zhu Y, Wang Y, Zhang H and Xue F:
Insulin resistance: A significant risk factor of endometrial
cancer. Gynecol Oncol. 125:751–757. 2012. View Article : Google Scholar : PubMed/NCBI
|