Introduction
Lung cancer is an aggressive malignant tumor with
poor therapeutic outcomes (1).
Patients with non-small cell lung cancer (NSCLC) account for the
majority of lung cancer cases and the incidence of NSCLC is
increasing annually (2–4). Thus, developing effective and novel
biomarkers and targets for lung cancer diagnosis and therapy is
urgent.
Transcription factor II B (TFIIB)-related factor 2
(BRF2) is a gene located on chromosome 8p12 (5). As a subunit, BRF2 protein, which is
located on TFIIB and participates in small RNA production, is
catalyzed under RNA polymerase III (pol III) (6–9).
During cell cycle, the transcription of RNA pol III is regulated to
ensure normal cellular growth (10,11).
Previous studies reported that RNA pol III activity plays a key
role in the deregulation of a variety of cancers, regardless of
tissue types (12,13). Deregulation of TFIIIB-mediated
transcription is an important factor in tumor development (14). In addition, TBP expression is found
increased in a large number of cancers including in human kidney,
colon, melanoma, and gastric cancers (15,16).
BRF2 has a pivotal role in proliferation,
metastasis, angiogenesis, and tumorigenesis by acting as an
oncogene (2). In addition,
overexpression of BRF2 is associated with a higher risk of cancer
recurrence (5,17). Lockwood et al (18) demonstrated that genetic activation
of BRF2 is a special mechanism of squamous cell carcinoma
tumorigenesis, and this finding is the first clinical evidence to
suggest that BRF2 is a novel oncogene in lung cancer. Previous
studies suggest that BRF2 expression is increased in breast
cancers, suggesting that it is potentially a candidate oncogene
(19,20). However, the mechanism of action of
BRF2 still remains to be elucidated, especially the relationship
between BRF2 and lung cancer.
By using immunofluorescence staining and
immunohistochemistry methods, the present study detected BRF2
protein expression levels in lung cancer tissues, and analyzed the
function of the BRF2 gene in the metastasis of lung cancer
cells.
Materials and methods
Clinical samples
The clinical samples of lung cancer tissue and
paired normal adjacent tissues were collected from lung cancer
patients at Zhuji Affiliated Hospital of Shaoxing University in
2019 for treatment. A total of 72 patients with lung cancer (age,
35–70 years) were enrolled between December 2018 and December 2019.
The male to female ratio was 3:1. Patients were divided into three
groups according to the age of onset as follows: Group 1, 35–50;
group 2, 50–62; group 3, 62–70. Cancer and normal adjacent tissues
of all patients were obtained and used to detect the gene
expression level of BRF2 in the tissues. The patients had no
history of chemotherapy and did not have other types of cancer,
infectious diseases, or autoimmune diseases. All patients signed
informed consent and agreed that their tissues would be used for
clinical research. The study was approved by the Ethics Committee
of Zhuji Affiliated Hospital of Shaoxing University (approval no.
ZLK20181124). All clinical samples were obtained at the time of
initial resection, and stored at −80°C.
Bioinformatics analysis
BRF2 mRNA expression levels in lung cancer and
normal tissues were compared using a bioinformatics website
(gepia.cancer-pku.cn).
Cell culture
Normal human lung epithelial cells (BEAS-2B; cat.
no. CRL-9609) and human lung cancer cells (A549, cat. no. CCL-185;
H292, cat. no. CRL-1848) were purchased from the American Type
Culture Collection. The cells were cultured in Dulbecco's modified
Eagle's medium (DMEM, Sigma-Aldrich; Merck KGaA) supplemented with
10% fetal bovine serum (FBS, Thermo Fisher Scientific, Inc.), 100
U/ml penicillin and 100 µg/ml streptomycin at 37°C in 5%
CO2.
Immunofluorescence staining
The cells (2×104/ml) were grown on
coverslips and after the liquid had been aspirated, the cells were
covered 2–3 mm under 4% formaldehyde diluted in warm
phosphate-buffered saline (PBS) to be fixed for 15 min at room
temperature. Following removal of the fixative, the cells were
rinsed in PBS with for 5 min. Then the cells were blocked with
immunostaining blocking buffer (cat. no. P0102; Beyotime Institute
of Biotechnology) at 37°C for 60 min, after aspirating the
solution, primary antibody anti-BRF2 antibody (mouse, cat. no.
ab194442, 1:500, Abcam) was added to the cells. Next, the cells
were incubated overnight at 4°C. The cells were rinsed 3 times in
PBS for 5 min at room temperature in the dark, and then incubated
with fluorochrome-conjugated secondary antibody [horseradish
peroxidase (HRP)-conjugated goat anti-mouse IgG H&L (1:100;
cat. no. ab6789; Abcam)] at 37°C for 1 h. DAPI (cat. no. D1306,
Thermo Fisher Scientific, Inc.) was added to the cells and
incubated together for 5 min in the dark. Following aspiration of
the liquid, the cells were observed under a fluorescence microscope
(magnification, ×200; cat. no. BX43; Olympus Corporation).
Immunohistochemistry
Following fixation in 4% formaldehyde at 25°C for 24
h, the tissues were dehydrated and made transparent by gradient
alcohol, then paraffin-embedded, and sectioned (section thickness,
5 µm). The sections were soaked in citrate buffer solution (pH 6.0)
and heated in a 850 W power microwave oven for 10 min to conduct
antigen repair. The endogenous peroxidase enzyme activity was
minimized by rinsing the sections in 3% H2O2
in methanol for 20 min at room temperature. Then, the tissues were
incubated with primary rabbit anti-BRF2 polyclonal antibody (1:400,
cat. no. ab194442, Abcam) at 4°C overnight. Next the sections were
incubated with secondary antibody [HRP-conjugated goat anti-rabbit
immunoglobulin (IgG) H&L (1:2,000; cat. no. ab150113; Abcam)
and streptavidin-peroxidase complex for 30 min at 37°C. The
sections were stained with hematoxylin, dried in an oven at 65°C,
rinsed in water, then mixed with alcohol and xylene and naturally
dried. Finally, the sections were sealed and observed under an
optical microscope (magnification, ×100; BX53M, Olympus
Corporation).
Cell transfection
BRF2 small interfering (si)RNA (cat. no. 11968S,
Cell Signaling Technology, Inc.) was transfected into the A549
cells using Lipofectamine® 2000 Transfection reagent
(Invitrogen; Thermo Fisher Scientific, Inc.). A549 cells were
seeded into 6-well plates at 1×105 cells per well and
incubated at 37°C for 24 h. Then 1.5 ml medium without serum or
antibiotics was added into each well of the plate, with the mixture
of 100 pmol BRF2 siRNA and Lipofectamine® 2000 to
incubate for 4–6 h at 37°C with 5% CO2. The siRNA
sequences were as follows: BRF2 siRNA sense,
5′-GGUGGGAAAUAAUUCCUUATT-3′; si-negative control (siNC) sense,
5′-UUCUCCGAACGUGUCACGUTT-3′. After 72 h of transfection, the cells
were collected for later analysis.
Reverse transcription-quantitative
(RT-q)PCR
Total RNAs were extracted from tissues or cells
(2×104/ml) by TRIzol® reagent (Invitrogen;
Thermo Fisher Scientific, Inc.) according to the manufacturer's
protocols. Total RNAs were placed in a refrigerator at 4°C, and
quantified by a biological spectrometer (Nano Drop 2000C; Thermo
Fisher Scientific, Inc.). The extracted RNAs were
reverse-transcribed to cDNA using a Prime Script RT reagent kit
(Takara Bio, Inc.) according to the manufacturer's instructions.
The results were normalized to β-actin. Next, SYBR®
Green PCR Master Mix (cat. no. 4312704, Applied Biosystems, Thermo
Fisher Scientific, Inc.) and Bio-Rad CFX 96 Touch Real-Time PCR
Detection system (cat. no. 1855196, Bio-Rad Laboratories, Inc.)
were used for RT-qPCR, according to the manufacturer's
instructions. The thermocycling conditions were as follows: 95°C
for 5 min, 40 cycles of 95°C for 15 sec, 60°C for 30 sec and 70°C
for 10 sec. The primers for β-actin and BRF2 were: β-actin forward,
5′-ATTGGCAATGAGCGGTTC-3′ and reverse: 5′-GGATGCCACAGGACTCCA-3′;
BRF2 forward, 5′-CACAGGGGAAAACGAACAAG-3′ and reverse,
5′-TCGACAAAGGTCTCTCACTCG-3′. The gene expression was calculated by
the 2−ΔΔCq method (21). Each experiment was performed 3
times.
Western blotting
Following transfection for 48 h, A549 cells were
collected for western blot analysis as previously described
(22). Total proteins were
extracted from the cells by RIPA lysis buffer (Thermo Fisher
Scientific, Inc.), and protein concentration was determined by
bicinchoninic protein assay kit (Sigma-Aldrich; Merck KGaA). The
proteins (10 µg) were separated by 10% SDS-PAGE (cat. no. P0012A,
Beyotime Institute of Biotechnology) and transferred onto
polyvinylidene fluoride membranes (cat. no. FFP28, Beyotime
Institute of Biotechnology), which were blocked with 5% fat-free
milk for 1 h at room temperature. The membranes were then incubated
overnight at 4°C with the primary antibodies: anti-Akt antibody
(rabbit, cat. no. ab8805, 1:500, Abcam), anti-p-Akt antibody
(rabbit, cat. no. ab38449, 1:500, Abcam), anti-Bax antibody
(rabbit, cat. no. ab32503, 1:1,000, Abcam), anti-Bcl-2 antibody
(rabbit, cat. no. ab59348, 1:500, Abcam), anti- E-cadherin antibody
(rabbit, cat. no. ab40772, 1:1,000, Abcam), anti-N-cadherin
(rabbit, cat. no. ab18203, 1 µg/ml, Abcam), anti-Snail antibody
(rabbit, cat. no. ab229701, 1:1,000, Abcam), anti-EGFR antibody
(rabbit, cat. no. ab52849, 1:1,000, Abcam) and anti-β-actin
antibody (mouse, cat. no. ab8226, 1:500, Abcam). β-actin served as
an internal reference. The membranes were then incubated with
secondary HRP-conjugated antibodies [goat anti-rabbit IgG H&L
(HRP) (1:2,000; cat. no. ab150113; Abcam) and goat anti-mouse IgG
H&L (HRP) (1:2,000; cat. no. ab6789; Abcam] at 37°C for 1 h and
washed 3 times with 0.9% TBST. The protein bands were developed by
an enhanced chemiluminescence (ECL) kit (Millipore), and the grey
values of the strips were calculated by ImageJ (version 5.0;
National Institutes of Health) (23).
MTT assay
The A549 cells were incubated in 96-well plates at a
density of 5×103 cells/well. Following transfection, the
cells were cultured for 48 h with 5% CO2 at 37°C. Next,
20 µl MTT (Promega Corp.) was added to each well. The formazan
products were dissolved in 100 µl dimethylsulfoxide (DMSO), and the
absorbance was detected at 540 nm using a microplate reader (PR3100
TSC, Bio-Rad Laboratories, Inc.). The cells were subjected to a
multifunctional enzyme-linked analyzer (Attune NxT; Thermo Fisher
Scientific, Inc.) for 4 h, and the absorbance value of each hole
was measured at 490 nm.
Cell Counting Kit- 8 (CCK-8)
The cells (5×103 cells/well) from the
different treatment groups were seeded in a 96-well plate, and
cultured in an incubator with 5% CO2 at 37°C for 48 h.
Next, 100 µl CCK-8 and serum-free basic medium (DMEM; Thermo Fisher
Scientific, Inc.) were mixed at a 1:10 ratio and added to a cell
culture plate and cultured for 4 h. Finally, the absorbance at 450
nm wavelength was determined using a microplate reader (RNE90002,
Reagen Biology LLC).
Apoptosis assay
Following transfection, the A549 cells were washed
with cold PBS twice. Annexin V/Dead Cell Apoptosis kit (cat. no.
V13242, Thermo Fisher Scientific, Inc.) was used to identify
apoptotic cells. Briefly, the cells were re-suspended in Annexin V
binding buffer, added with Annexin V-FITC and propidium iodide (PI)
buffer, and incubated at room temperature for 15 min in the dark.
Cell apoptosis was analyzed using a flow cytometer (FACSCalibur™;
version 2.0; BD Biosciences).
Wound-healing assay
Following transfection, the cells were subcultured
in plates at a density of 1×105 cells/well and incubated
in serum without FBS at 37°C for 24 h. The wounds were created
using a 200-µl yellow sterile pipette tip on the monolayer, and
detached cells were removed by washing the cells 3 times. The
healing process was observed under a 600 Autobiochemical Analyzer
(Olympus Corporation), and ImageJ software (Image Pro Plus; version
6.0; National Institutes of Health) was used to calculate the
average distance between cells.
Transwell invation assay
Matrigel (BD Biosciences) diluted at a 1:8 ratio was
used to cover the upper surface of the membrane of the bottom of
Transwell chambers, and the chambers were incubated at 37°C for 30
min in order to polymerize the Matrigel into gel. The cells were
resuspended in the upper chambers of the Transwell (8-µm pore size,
Corning Inc.), which contained serum-free DMEM medium, while the
lower chambers were supplemented with DMEM containing 10% FBS. The
Transwell chambers were all incubated for 48 h. Next, the cells
remaining in the upper chamber were gently removed using a cotton
swab, while the invaded cells were fixed in 4% paraformaldehyde for
15 min and stained by 0.1% crystal violet staining at 37°C for 20
min. The cells were randomly counted under an inverted microscope
(Eclipse Ti2, Nikon Corporation) and images were captured.
Statistical analysis
All data were statistically analyzed by SPSS version
18.0 (SPSS, Inc.). The data are presented as means ± standard
deviation, and all experiments were repeated 3 times. Two-group
comparisons were conducted by the Student's t-test, and multi-group
comparisons were conducted by one-way analysis of variance (ANOVA)
followed by Dunnett's post hoc test. P<0.05 was considered to
indicate a statistically significant difference.
Results
BRF2 expression levels are increased
in lung cancer tissues and cells
To investigate whether BRF2 is involved in tumor
migration and invasion, its protein expression in normal human lung
epithelial BEAS-2B cells and lung cancer cells A549 and H292 were
detected by immunofluorescence staining. As shown in Fig. 1A, marker protein BRF2 was
positively expressed in A549 and H292 cells, especially in the A549
cells, while in BEAS-2B cells, marker protein BRF2 demonstrated a
low expression. Meanwhile, BRF2 expression was detected by western
blotting, as shown in Fig. 1B and
C; BRF2 expression was significantly increased in A549 and H292
cells compared to that noted in the BEAS-2B cells. In addition,
according to Fig. 1D, BRF2 protein
staining was detected in cancer tissues by immunohistochemistry;
however, BRF2 was barely observed in normal lung tissues. As shown
in Fig. 1E, the BRF2 mRNA
expression levels in lung cancer tissues and normal tissues were
compared using a bioinformatics website (gepia.cancer-pku.cn), and
the result demonstrated that the BRF2 mRNA expression level in the
tumor tissues was notably higher compared with normal tissues. The
BRF2 mRNA expression in lung cancer A549 and H292 cells and normal
lung epithelial BEAS-2B cells was detected by RT-qPCR. Relative
BRF2 mRNA expression level was the highest in A549 cells (Fig. 1F, P<0.001) and its protein
expression was also high in H292 cells, while the mRNA expression
level of BRF2 in BEAS-2B cells was low. In addition, normal
adjacent and cancer tissues were divided into groups 1–3. Relative
BRF2 mRNA expression level in lung cancer cells was clearly higher
than normal cells (Fig. 1G,
P<0.001).
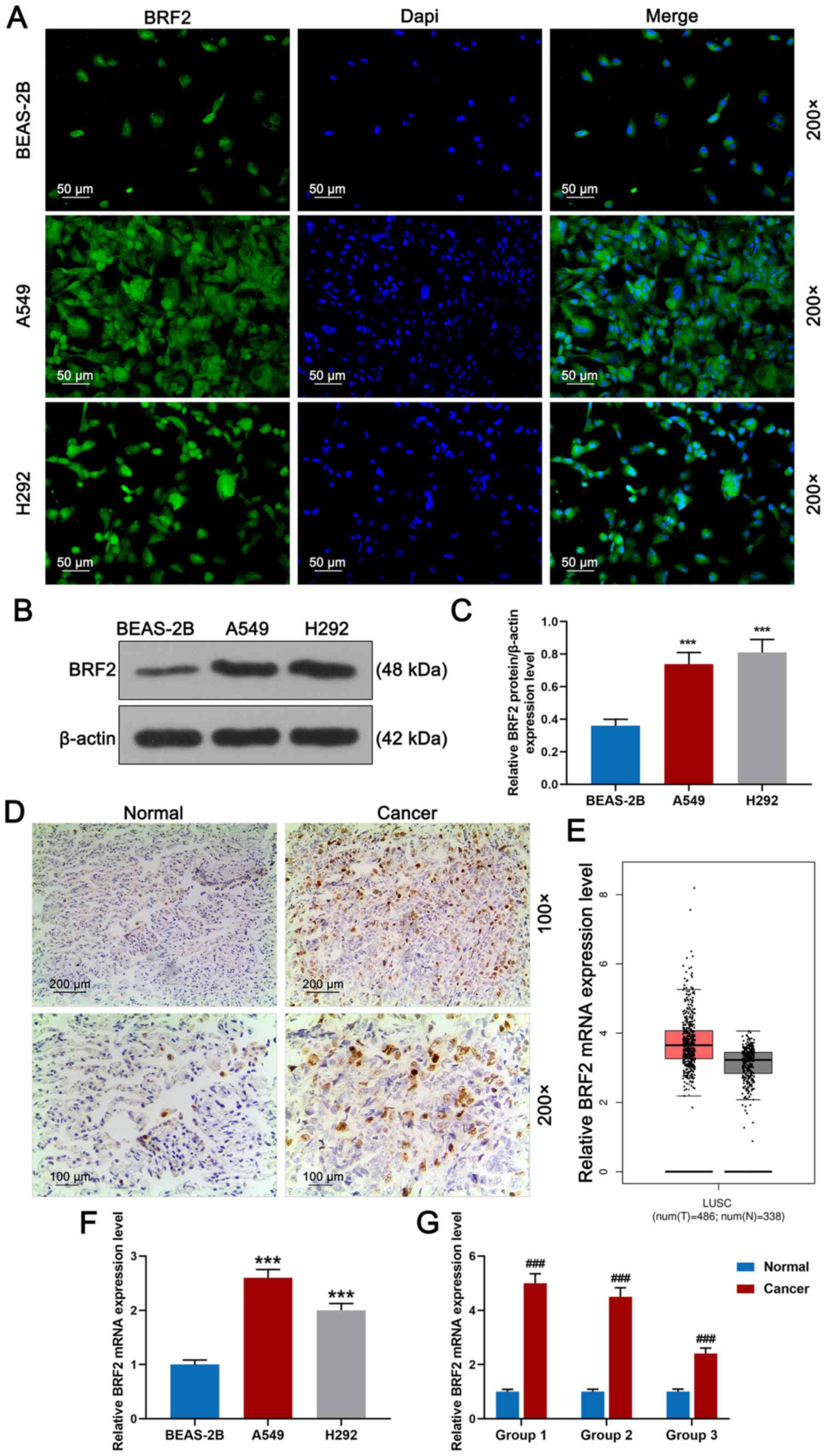 | Figure 1.BRF2 protein expression in lung
cancer tissue and cells. (A) BRF2 expression in normal human lung
epithelial BEAS-2B cells and lung cancer A549 and H292 cells was
detected by immunofluorescence staining. Magnification, ×200; scale
bar, 50 µm. (B and C) BRF2 expression was detected by western
blotting. (D) BRF2 expression in a normal adjacent tissue group
(normal, n=3) and cancer group (n=3) was detected by
immunohistochemistry. Magnification, ×100; scale bar, 200 µm. (E)
The BRF2 mRNA expression level in lung cancer tissues and normal
tissues was compared by bioinformatics website
(gepia.cancer-pku.cn). (F) RT-qPCR revealed that the A549 cell
group had the highest relative BRF2 mRNA expression level, while
the expression level in the BEAS-2B cell group was the lowest. (G)
Normal adjacent tissues and cancer tissues were divided into groups
1 (onset age, 35–50 years), 2 (onset age, 50–62 years) and 3 (onset
age, 62–72 years). RT-qPCR demonstrated that BRF2 protein had a
high expression in the cancer groups. The experiments were repeated
at least 3 times. ***P<0.001 vs. BEAS-2B;
###P<0.001 vs. normal group. BRF2, transcription
factor II B-related factor 2; RT-qPCR, reverse
transcription-quantitative PCR. |
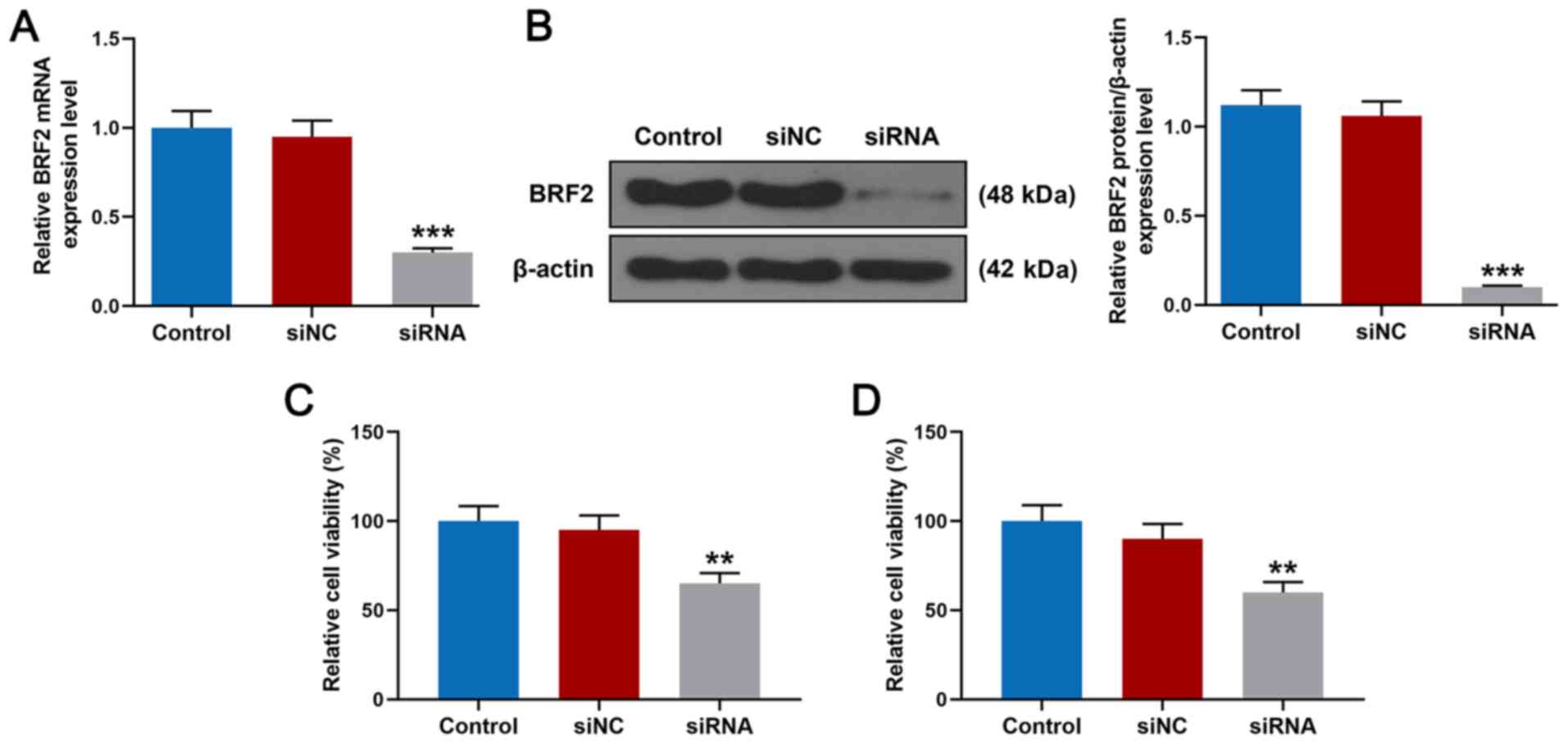
Knockdown of BRF2 inhibits the
proliferation of A549 cells
To explore the role of BRF2 in lung cancer cells,
A549 cells were transfected with siRNA for silencing BRF2. As shown
in Fig. 2A, 48 h after BRF2 siRNA
transfection, the result of RT-qPCR demonstrated that the relative
mRNA expression level of BRF2 in A549 cells was significantly lower
than that in the siNC group (P<0.001). The result of western
blotting (Fig. 2B) demonstrated
that the relative protein expression level of BRF2 was
significantly inhibited (P<0.001 vs. siNC). In addition, the
results of the CCK-8 and MTT assays demonstrated that after
transfection for 48 h the relative cell viability of A549 cells in
the siRNA group was significantly inhibited compared with the siNC
group (Fig. 2C and D,
**P<0.01).
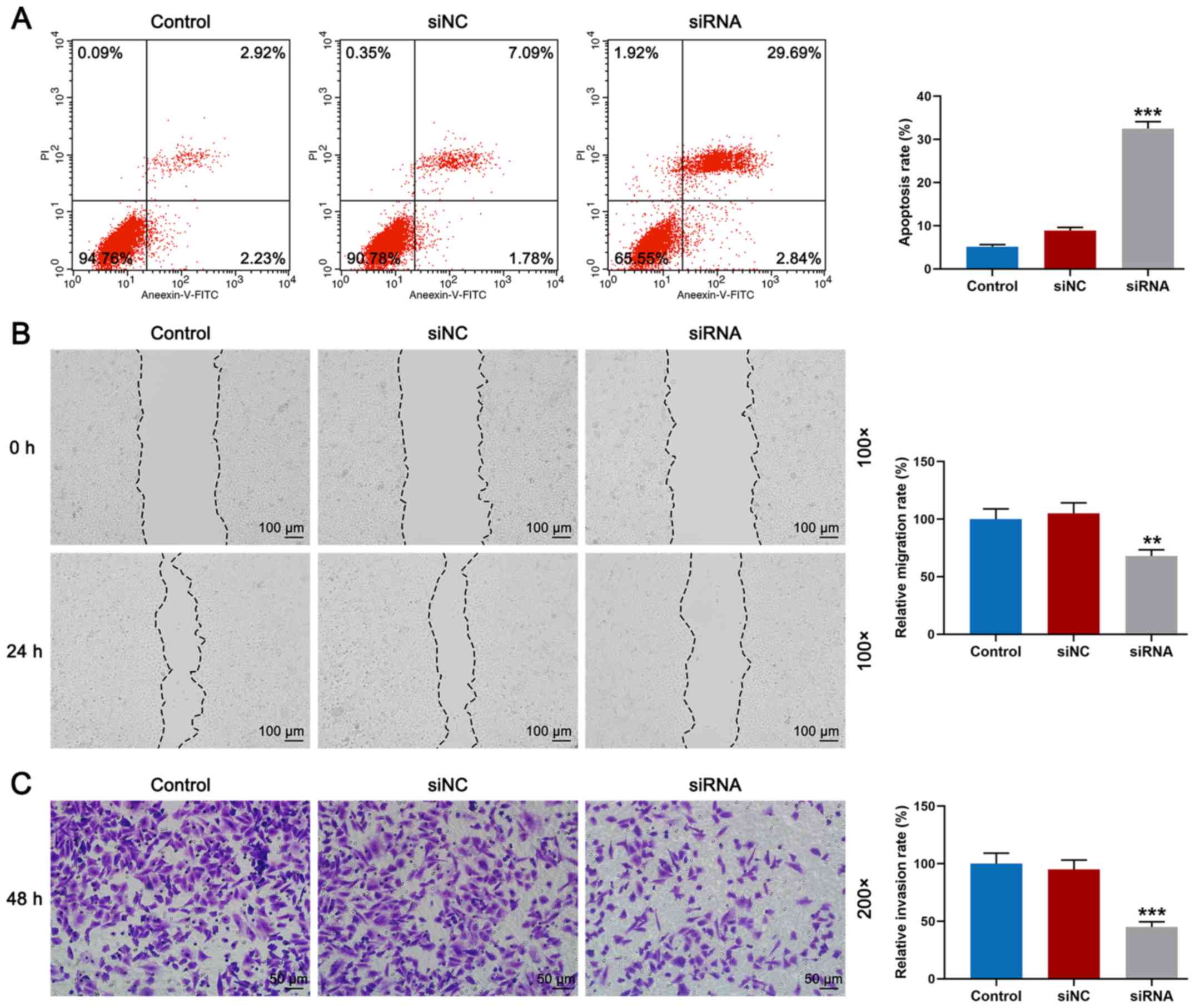
Knockdown of BRF2 promotes apoptosis
and inhibits cell migration and invasion of A549 cells
Following transfection, the effects of siRNA on cell
apoptosis and migration were assessed. As shown in Fig. 3A, flow cytometry demonstrated that
the rate of apoptosis of the siRNA group was significantly higher
compared with that noted in the siNC group (Fig. 3A, P<0.001). Meanwhile, cell
migration and invasion were detected by wound-healing and Transwell
assays. As shown in Fig. 3B, it
was observed that the relative migration rate of the siRNA group
was significantly lower than that noted in the siNC group (Fig. 3B, P<0.01). Furthermore, the
result of Fig. 3C revealed that
invasion of A549 cells transfected with BRF2 siRNA was
significantly reduced (Fig. 3C,
P<0.001).
Knockdown of BRF2 inhibits the protein
expression levels of Akt, p-Akt, Bcl-2, N-cadherin, Snail and EGFR
and increases those of Bax and E-cadherin
The western blot analysis (Fig. 4A and B) revealed that following
transfection of BRF2 siRNA, relative protein expression levels of
Akt (P<0.01 vs. siNC), p-Akt (P<0.001 vs. siNC) and Bcl-2
(P<0.01 vs. siNC) were suppressed, while relative protein
expression level of Bax was significantly promoted. Furthermore, as
demonstrated in Fig. 4C, relative
protein expression levels of N-cadherin (P<0.01 vs. siNC), Snail
(P<0.01 vs. siNC) and EGFR (P<0.001 vs. siNC) were
significantly inhibited. Notably, protein expression of E-cadherin
(P<0.01 vs. siNC) was significantly promoted following
transfection of BRF2 siRNA. In addition, the results further
confirmed that BRF2 siRNA suppressed phosphorylation of Akt
(Fig. 4D, P<0.001 vs.
siNC).
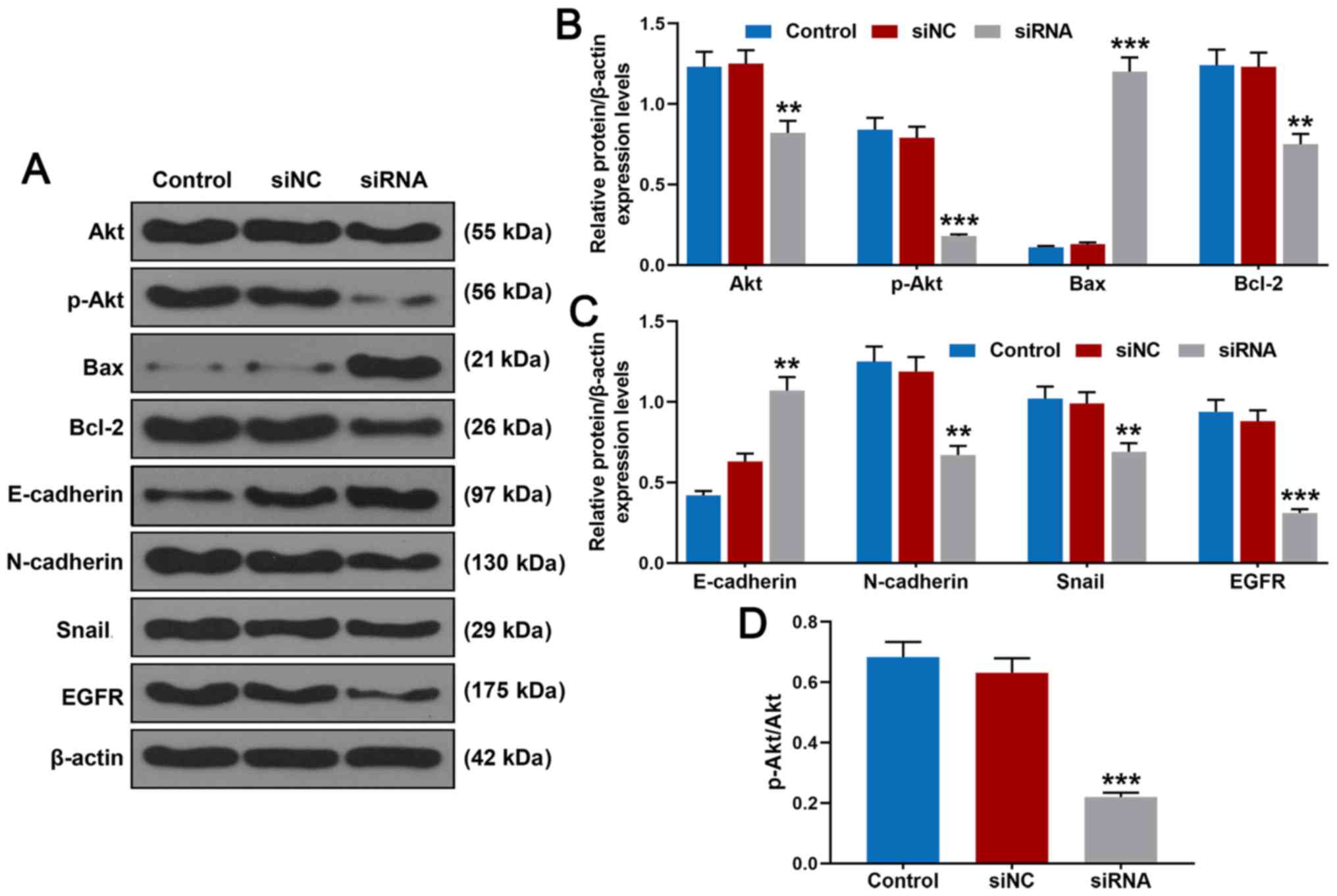 | Figure 4.Effects of BRF2 siRNA on Akt, p-Akt,
Bax, Bcl-2, E-cadherin, N-cadherin, Snail and EGFR in A549 cells.
(A) Akt, p-Akt, Bax, Bcl-2, E-cadherin, N-cadherin, Snail and EGFR
protein expression in A549 cells was detected by western blotting.
(B) BRF2 siRNA inhibited p-Akt, Akt and Bcl-2 protein expression
but increased Bax protein expression. (C) BRF2 siRNA reduced
N-cadherin, Snail and EGFR protein expressions but promoted
E-cadherin protein expression. (D) BRF2 siRNA suppressed
phosphorylation of Akt. The experiments were repeated at least 3
times. **P<0.01 and ***P<0.001 vs. siNC. BRF2, transcription
factor II B-related factor 2; siNC, negative control for siRNA;
siRNA, small interfering RNA for BRF2; p, phosphorylated. |
Discussion
Lung cancer accounts for a large proportion of
cancer-related deaths (24).
Although diagnostic and therapeutic techniques have been improved,
treatment results are far from satisfactory (25,26).
Therefore, it is crucial to improve current understanding on tumor
pathogenesis, gene expression profiles and tumor biology (27).
Melchor et al (28) observed increased transcription
factor II B (TFIIB)-related factor 2 (BRF2) gene expression in
breast cancer, and that BRF2 gene expression products are
significantly higher in breast cancer tissues compared normal
breast tissues. Furthermore, they identified a correlation between
BRF2 overexpression and clinical outcomes (29). It has been shown that ~40% of lung
squamous cell carcinoma is closely related to the local
amplification of chromosome 8p12 through comparative genomic
hybridization (29). During the
invasion stage of lung squamous cell carcinoma, the oncogene BRF is
frequently activated (30). These
findings indicate that BRF2 protein has an active expression in
various tumors and is closely related to invasion and migration of
various tumors. The present study found that BRF2 expression was
increased in lung cancer cells compared with that in normal
adjacent tissues.
miR-4299 is a key molecule of the cell survival
pathway PI3K/Akt; the study of Yang et al (31) found that miR-4299 suppresses the
progression of non-small lung cancer (NSCLC) by modulating the
activation of the PTEN/AKT/PI3K signaling pathway; thus, it may
serve as an independent candidate marker for prognosis of NSCLC
patients. Croton tiglium extract can elevate expression of
Bax genes and reduce expression of Bcl-2 genes to induce A549 cell
apoptosis (32). In addition, the
physiological function of EGFR is to regulate epithelial tissue
development and homeostasis (33).
In pathological settings, mostly in lung and breast cancers and
glioblastoma, EGFR is a driver of tumorigenesis (33). In the present study, following cell
transfection with BRF2 siRNA, the protein expression levels of Akt,
p-Akt, Bcl-2 and EGFR in A549 cells were inhibited, while Bax
protein expression was increased, suggesting that silencing of BRF2
inhibited the activation of cell survival pathway PI3K/Akt, and
promoted the activation of cell apoptosis.
Farmakovskaya et al (34) demonstrated that suppression of
E-cadherin expression increases cancer stem cells in human A549
lung adenocarcinoma and stimulates tumor growth. In addition,
N-cadherin induces cell survival, migration, and invasion by
modulating intracellular signaling molecules. As shown by
Quintanal-Villalonga et al (35), the FGFR4-388arg variant promotes
lung cancer progression through N-cadherin induction. Reducing
miR-22 expression promotes epithelial-mesenchymal transition (EMT)
and invasion of lung cancer cells by elevating Snail expression
(36). According to the results of
western blot analysis in the present study, transfection of BRF2
siRNA into A549 cells increased E-cadherin expression but reduced
N-cadherin expression.
Consistent with a previous study (37), in the present study silencing of
BRF2 protein expression reduced the migration and invasion of
NSCLC, suggesting that BRF2 expression plays an important role in
invasiveness of NSCLC cells, possibly through EMT, which involves
increased Snail expression and abnormal expression of E-cadherin
and N-cadherin. Furthermore, Gouge et al (38) found that BRF2 redox-dependent
regulation constitutes a cellular blockade, which is capable of
generating pro-apoptotic signals upon prolonged oxidative stress in
lung and breast cancers by limiting the availability of SeCys tRNA.
Wang et al (2) suggested
that miR-373 may function as a tumor suppressor in NSCLC by
attenuating the expression of BRF2 to inhibit proliferation. These
above findings are in line with the results of the current study,
in which the knockdown of BRF2 inhibited A549 cell migration and
invasion, suppressed A549 cell proliferation, and enhanced
apoptosis of A549 cells. It should be noted that the present study
did not assess the findings in vivo in animal experiments
and that the results should be verified in more cell lines. In
addition, the mechanism of BRF2 remains to be further
elucidated.
The findings of the present study revealed that BRF2
protein plays an important role in lung cancer cells and supports
the hypothesis that BRF2 protein could serve as a novel target for
lung cancer therapy. However, the role of BRF2 in the occurrence
and development of lung cancer should be further verified. Thus, in
future research, the effect of BRF2 on tumors in vivo should
be explored by establishing animal models of lung cancer.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The analyzed data sets generated during the study
are available from the corresponding author on reasonable
request.
Authors' contributions
YB made substantial contributions to conception and
design. QiuL, QiaL and RP were responsible for data acquisition,
data analysis and interpretation. YB drafted the article and
critically revising it for important intellectual content. All
authors agreed to be accountable for all aspects of the work in
ensuring that questions related to the accuracy or integrity of the
work are appropriately investigated and resolved. All authors
reviewed and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Liu H, Han L, Liu Z and Gao N: Long
noncoding RNA MNX1-AS1 contributes to lung cancer progression
through the miR-527/BRF2 pathway. J Cell Physiol. 234:13843–13850.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Wang L, Qu J, Zhou L, Liao F and Wang J:
MicroRNA-373 Inhibits Cell Proliferation and Invasion via Targeting
BRF2 in Human Non-small Cell Lung Cancer A549 Cell Line. Cancer Res
Treat. 50:936–949. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lu J and Han B: Liquid Biopsy Promotes
Non-Small Cell Lung Cancer Precision Therapy. Technol Cancer Res
Treat. 17:15330338188018092018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Collins LG, Haines C, Perkel R and Enck
RE: Lung cancer: Diagnosis and management. Am Fam Physician.
75:56–63. 2007.PubMed/NCBI
|
|
5
|
Tian Y, Wang C and Lu M: BRF2 as a
promising indicator for radical lymph-node dissection surgery in
patients with cN0 squamous cell carcinoma of the middle thoracic
esophagus. Surg Today. 49:158–169. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Verma N, Hurlburt AM, Wolfe A, Kim MK,
Kang YS, Kang JJ and Stumph WE: Bdp1 interacts with SNAPc bound to
a U6, but not U1, snRNA gene promoter element to establish a stable
protein-DNA complex. FEBS Lett. 592:2489–2498. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
McQueen C, Hughes GL and Pownall ME:
Skeletal muscle differentiation drives a dramatic downregulation of
RNA polymerase III activity and differential expression of Polr3g
isoforms. Dev Biol. 454:74–84. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Geiduschek EP and Kassavetis GA: The RNA
polymerase III transcription apparatus. J Mol Biol. 310:1–26. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Saxena A, Ma B, Schramm L and Hernandez N:
Structure-function analysis of the human TFIIB-related factor II
protein reveals an essential role for the C-terminal domain in RNA
polymerase III transcription. Mol Cell Biol. 25:9406–9418. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Song W, Filonov GS, Kim H, Hirsch M, Li X,
Moon JD and Jaffrey SR: Imaging RNA polymerase III transcription
using a photostable RNA-fluorophore complex. Nat Chem Biol.
13:1187–1194. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Scott PH, Cairns CA, Sutcliffe JE,
Alzuherri HM, McLees A, Winter AG and White RJ: Regulation of RNA
polymerase III transcription during cell cycle entry. J Biol Chem.
276:1005–1014. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lei J, Chen S and Zhong S: Abnormal
expression of TFIIIB subunits and RNA Pol III genes is associated
with hepatocellular carcinoma. Liver Res. 1:112–120. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
White RJ: RNA polymerase III transcription
- a battleground for tumour suppressors and oncogenes. Eur J
Cancer. 40:21–27. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Cabarcas S, Jacob J, Veras I and Schramm
L: Differential expression of the TFIIIB subunits Brf1 and Brf2 in
cancer cells. BMC Mol Biol. 9:742008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Johnson SA, Dubeau L, Kawalek M, Dervan A,
Schönthal AH, Dang CV and Johnson DL: Increased expression of
TATA-binding protein, the central transcription factor, can
contribute to oncogenesis. Mol Cell Biol. 23:3043–3051. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Srihari S, Kalimutho M, Lal S, Singla J,
Patel D, Simpson PT, Khanna KK and Ragan MA: Understanding the
functional impact of copy number alterations in breast cancer using
a network modeling approach. Mol Biosyst. 12:963–972. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ng CK, Martelotto LG, Gauthier A, Wen HC,
Piscuoglio S, Lim RS, Cowell CF, Wilkerson PM, Wai P, Rodrigues DN,
et al: Intra-tumor genetic heterogeneity and alternative driver
genetic alterations in breast cancers with heterogeneous HER2 gene
amplification. Genome Biol. 16:1072015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lockwood WW, Chari R, Coe BP, Thu KL,
Garnis C, Malloff CA, Campbell J, Williams AC, Hwang D, Zhu CQ, et
al: Integrative genomic analyses identify BRF2 as a novel
lineage-specific oncogene in lung squamous cell carcinoma. PLoS
Med. 7:e10003152010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Almanza A, Carlesso A, Chintha C,
Creedican S, Doultsinos D, Leuzzi B, Luís A, McCarthy N,
Montibeller L, More S, et al: Endoplasmic reticulum stress
signalling - from basic mechanisms to clinical applications. FEBS
J. 286:241–278. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Garcia MJ, Pole JC, Chin SF, Teschendorff
A, Naderi A, Ozdag H, Vias M, Kranjac T, Subkhankulova T, Paish C,
et al: A 1 Mb minimal amplicon at 8p11-12 in breast cancer
identifies new candidate oncogenes. Oncogene. 24:5235–5245. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Guo Q, Li H, Liu J, Xu L, Yang L, Sun Z
and Zhou B: Tunicamycin aggravates endoplasmic reticulum stress and
airway inflammation via PERK-ATF4-CHOP signaling in a murine model
of neutrophilic asthma. J Asthma. 54:125–133. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Jin J, Zhao L, Zou W, Shen W, Zhang H and
He Q: Activation of Cyclooxygenase-2 by ATF4 During Endoplasmic
Reticulum Stress Regulates Kidney Podocyte Autophagy Induced by
Lupus Nephritis. Cell Physiol Biochem. 48:753–764. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Thawani R, McLane M, Beig N, Ghose S,
Prasanna P, Velcheti V and Madabhushi A: Radiomics and
radiogenomics in lung cancer: A review for the clinician. Lung
Cancer. 115:34–41. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hirsch FR, Scagliotti GV, Mulshine JL,
Kwon R, Curran WJ Jr, Wu YL and Paz-Ares L: Lung cancer: Current
therapies and new targeted treatments. Lancet. 389:299–311. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Herbst RS, Heymach JV and Lippman SM: Lung
cancer. N Engl J Med. 359:1367–1380. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Hashemi ZS, Khalili S, Forouzandeh
Moghadam M and Sadroddiny E: Lung cancer and miRNAs: A possible
remedy for anti-metastatic, therapeutic and diagnostic
applications. Expert Rev Respir Med. 11:147–157. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Melchor L, Garcia MJ, Honrado E, Pole JC,
Alvarez S, Edwards PA, Caldas C, Brenton JD and Benítez J: Genomic
analysis of the 8p11-12 amplicon in familial breast cancer. Int J
Cancer. 120:714–717. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Cabarcas S and Schramm L: RNA polymerase
III transcription in cancer: The BRF2 connection. Mol Cancer.
10:472011. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Rosenwald IB: The role of translation in
neoplastic transformation from a pathologist's point of view.
Oncogene. 23:3230–3247. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yang WB, Zhang WP, Shi JL and Wang JW:
miR-4299 suppresses non-small cell lung cancer cell proliferation,
migration and invasion through modulating PTEN/AKT/PI3K pathway.
Eur Rev Med Pharmacol Sci. 22:3408–3414. 2018.PubMed/NCBI
|
|
32
|
Khodapasand E, Jafarzadeh N, Farrokhi F,
Kamalidehghan B and Houshmand M: Is Bax/Bcl-2 ratio considered as a
prognostic marker with age and tumor location in colorectal cancer?
Iran Biomed J. 19:69–75. 2015.PubMed/NCBI
|
|
33
|
Sigismund S, Avanzato D and Lanzetti L:
Emerging functions of the EGFR in cancer. Mol Oncol. 12:3–20. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Farmakovskaya M, Khromova N, Rybko V,
Dugina V, Kopnin B and Kopnin P: E-Cadherin repression increases
amount of cancer stem cells in human A549 lung adenocarcinoma and
stimulates tumor growth. Cell Cycle. 15:1084–1092. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Quintanal-Villalonga Á, Ojeda-Márquez L,
Marrugal Á, Yagüe P, Ponce-Aix S, Salinas A, Carnero A, Ferrer I,
Molina-Pinelo S and Paz-Ares L: The FGFR4-388arg Variant Promotes
Lung Cancer Progression by N-Cadherin Induction. Sci Rep.
8:23942018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhang K, Li XY, Wang ZM, Han ZF and Zhao
YH: miR-22 inhibits lung cancer cell EMT and invasion through
targeting Snail. Eur Rev Med Pharmacol Sci. 21:3598–3604.
2017.PubMed/NCBI
|
|
37
|
Tian Y, Lu M, Yue W, Li L, Li S, Gao C, Si
L, Qi L, Hu W and Tian H: TFIIB-related factor 2 is associated with
poor prognosis of nonsmall cell lung cancer patients through
promoting tumor epithelial-mesenchymal transition. BioMed Res Int.
2014:5307862014. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Gouge J, Satia K, Guthertz N, Widya M,
Thompson AJ, Cousin P, Dergai O, Hernandez N and Vannini A: Redox
Signaling by the RNA Polymerase III TFIIB-Related Factor Brf2.
Cell. 163:1375–1387. 2015. View Article : Google Scholar : PubMed/NCBI
|