Introduction
Gastric cancer (GC) is one of the most common
malignant tumors in the digestive system worldwide, accounting for
~720,000 GC-related deaths every year (1). According to Globocan 2018 data, the
incidence and mortality rates of GC among all malignant tumors
occupied fifth and third place respectively, and >70% of these
cases occurred in developing countries, with 50% in East Asia
(2). Early diagnosis and treatment
of GC can often produce an improved therapeutic effect and longer
overall survival time. However, most patients are diagnosed by
endoscopy only when they present with symptoms, at which point the
disease is in advanced stage, and the optimal opportunity for
surgical intervention has been missed. Even after complete R0
resection, one-third of patients experience recurrence (3). Most GC cases are adenocarcinomas,
representing a highly heterogeneous disease with differences in
epidemiology and histopathology. There has been no evident
breakthrough for the treatment of patients with advanced GC, and
surgery is still the primary therapy, with the majority of cancer
patients dying due to tumor recurrence and metastasis, and a median
overall survival (OS) <1 year (4).
GC can be categorized using different classification
systems, such as the Bormann, the Lauren, and the World Health
Organization classifications (5–7). In
addition, some scholars have attempted to classify GC using
molecular and genetic features, such as The Cancer Genome Atlas
(8) and Asian Cancer Research Group
classifications (9). Since the
Lauren classification was proposed in 1965, it has been widely
recognized by both clinicians and pathologists and continues to be
used. The Lauren classification primarily divides GC into
intestinal, diffuse and mixed types based on cell morphology and
histochemistry (6). Histologically,
intestinal-type GC cells are large, clear in boundary, variable in
morphology and closely arranged, exhibiting tubular and glandular
differentiation. In contrast, diffuse-type GC cells are typically
scattered and often appear as solitary cells or in small clusters
due to lack of adhesion. Thus, gland formation is hard to observe
in diffuse-type GC tumor tissue, and is easy to disseminate.
Mixed-type GC presents all of the aforementioned characteristics
(6). Epidemiologically, intestinal
GC is the most common type, has the highest five-year survival
rate, and is more common in men and the elderly. Diffuse-type GC is
more likely in women and younger patients and has a lower 5-year
survival rate (10–12). Mix-type has the highest degree of
malignancy due to variable biological behavior (13).
With the development of medicine and bioinformatics,
high-throughput sequencing has become a common tool for medical
research (14). For instance, data
from gene expression profiling studies can be uploaded to public
repositories, such as, the Gene Expression Omnibus (GEO) within the
National Center for Biotechnology Information. Reanalyzing and
reintegrating such datasets often provides some meaningful insights
for research. A number of microarray datasets of GC have been
developed in recent years (15–17)
and a large number of significant differentially expressed genes
(DEGs) have already been identified.
In the present study, the GSE62254 dataset was
downloaded from the GEO and screened for DEGs using the ‘limma’ and
‘survival’ R packages. Gene Ontology (GO) and the Kyoto
Encyclopedia of Genes and Genomes (KEGG) analysis was carried out
on selected DEGs, identifying key biological features and signaling
pathways. Moreover, protein-protein interaction (PPI) network of
DEGs associated with diffuse-type was constructed, identifying
three hub genes using the Cytoscape. Lastly, Kaplan-Meier analysis
was used to evaluate overall survival (OS) in patients with
different expression levels of these newly identified hub genes in
GSE62254 and GSE15459 datasets.
Materials and methods
Data collection
The GSE62254 (9,18)
microarray dataset was downloaded from the GEO database (http://www.ncbi.nlm.nih.gov/geo/). GSE62254 was
obtained using the GPL570 platform with an Affymetrix Human Genome
U133 Plus 2.0 Array (Thermo Fisher Scientific, Inc.) and contains
300 different Lauren subtypes GC samples (128 diffuse-type samples,
137 intestinal-type samples and 35 mixed-type samples).
Analysis of differentially expressed
genes
The Robust multi-array average (RMA) algorithm
(19) in the R environment (v3.6.1)
(20) was used to normalize and
transform the raw data to expression values. DEGs between diffuse-
and intestinal-type samples were screened using the ‘limma’ package
in R (v3.6.1) (21), using the
cut-off criteria of adjusted P<0.05 and
|log2FC|>0.585. Accordingly, DEGs with
log2FC>0.585 were considered associated with
diffuse-type GC, whereas DEGs with log2FC<-0.585 were
considered associated with intestinal-type GC. To identify genes
associated with OS, the ‘survival’ package (22) in R (v3.6.1) was used for Cox
regression analysis. The resuls are presented as hazard ratios
(HRs) and P-values for all genes in the GSE62254 dataset. Genes
with P<0.05 were identified as OS-related genes. Venny's (v2.1)
online software (http://bioinfogp.cnb.csic.es/tools/venny/index.html)
was used to draw Venn diagrams of OS-related genes associated with
diffuse or intestinal GC.
GO and KEGG enrichment analysis
The Database for Annotation, Visualization and
Integrated Discovery (DAVID, http://david.ncifcrf.gov/) provides a comprehensive
set of functional annotation tools for investigators to examine
biological meaning behind large lists of genes. DAVID (v6.8) was
used for GO functional annotation and KEGG pathway analysis of DEGs
associated with diffuse and intestinal GC. P<0.05 was the
cut-off for statistically significant terms.
PPI network construction and screening
of hub genes in diffuse-type GC
PPI data of diffuse-type GC DEGs were constructed
using the Search Tool for the Retrieval of Interacting Genes
(STRING) database (version 10.0; http://string-db.org) (23). Interactions with an interaction
score of >0.700 were used to construct the PPI network.
Cytoscape (https://cytoscape.org; v3.7.1) (24) was used to visualize the PPI network,
and the cytoHubba (v1.6) (25)
plug-in was used to identify hub genes of the PPI network via four
different algorithms. The algorithms used for analysis included
degree, Edge Percolated component, Closeness and EcCentricity.
Genes overlapping in the four groups were deemed hub genes.
Patients and tissues samples
A total of 40 GC patients who received a gastrectomy
in The Third Affiliated Hospital of Anhui Medical University
between December 2016 to July 2018 were recruited in this study.
This cohort included 27 male and 13 female patients aged 41–83
years (average, 56.35±9.60). The inclusion criteria were: i) The
postoperative pathological diagnosis of diffuse- or intestinal-type
GC was consistent by two pathologists; ii) patients were newly
diagnosed with GC; and iii) patients had not received any
radiotherapy, chemotherapy or biological therapy. The exclusion
criteria were: i) Patients were diagnosed with other types of
tumor; ii) postoperative recurrent patients; and iii) patients had
severe functional diseases, such as heart, liver, kidney and
immunological diseases; iv) patients were treated with other
therapies before surgery; and v) patients were pregnant.
All specimens were handled and made anonymous
according to ethical and legal standards. Tissue samples were
collected during the surgery for GC and were confirmed by tissue
pathology examination. The patients were divided into diffuse-type
and intestinal-type GC groups (n=20 in each group) according to the
postoperative pathology results. All tumor tissues specimens were
collected from formalin-fixed paraffin-embedded tissues of
resection surgical procedures.
Immunohistochemical analysis
Immunohistochemistry was performed to determine the
expressions of angiotensinogen (AGT), C-X-C motif chemokine ligand
12 (CXCL12) and adrenoceptor β2 (ADRB2) in diffuse-type
intestinal-type GC tissue samples. GC tissues were fixed in 10%
neutral buffered formalin at room temperature for 48 h and embedded
in paraffin at 62°C for 45 min. The sections were then cut into
4-µm thick sections and dried overnight at 56°C. Paraffin-embedded
tissue were passed through dimethylbenzene and gradient ethanol
solution to deparaffinize and rehydrate the sections. Antigen
retrieval was performed by heating the sections in a microwave oven
in 10 mM sodium citrate-hydrochloric acid buffer (pH 6.0) for ~15
min at 95°C. To block endogenous peroxidase activity, 0.3%
peroxidase quenching solution was used at 37°C for 10 min. After
blocking for 30 min with 10% skim milk at 37°C, each section was
incubated with a rabbit anti-human AGT antibody (1:50; cat. no.
ab108334; Abcam), rabbit anti-human CXCL12 antibody (1:200; cat.
no. ab9797; Abcam) or rabbit anti-human ADRB2 antibody (1:100; cat.
no. ab182136; Abcam) overnight at 4°C. The slides were then
incubated with HRP-secondary antibodies (Abcam; cat. no. ab6721;
1:1,000) at room temperature for 30 min. The sections then were
incubated with 3,3′-diaminobenzidine (DAB) solution at room
temperature for 5 min, and every slide were counterstained with
hematoxylin at room temperature for 1 min, dehydrated, and sealed
with cover slips. The stained sections were examined under an
optical microscope.
For quantitative analysis, images from each sample
were analyzed using Image-Pro Plus 6.0 software (Media
Cybernetics). For each slide, three randomly selected regions were
examined under the same exposure time and white balance setting.
The blank area of the images was selected for optical density
correction. Positive expression was defined as brown-yellow
granules in the cytoplasm. The expression intensity was calculated
as MD=IOD/area of target region, where MD is the mean density and
IOD is the accumulated value of integrated optical density. The MD
of the three fields of view was used to quantify the expression
levels of the hub genes.
RNA extraction and reverse
transcription-quantitative PCR (RT-qPCR)
RT-qPCR was used to examine the expression levels of
AGT, CXCL12 and ADRB2 between diffuse- and intestinal-type GC
tissue samples. Total RNA was extracted with the TRIzol®
reagent (Invitrogen; Thermo Fisher Scientific, Inc.) according to
the manufacturer's instructions. RNA purity was detected using a
microplate reader (Infinite M1000 PRO; Tecan Group, Ltd.). A
PrimeScript™ RT reagent kit (Takara Bio, Inc.) was used for cDNA
synthesis, according to the manufacturer's protocol, at 37°C for 15
min and 85°C for 5 sec. RT-qPCR was carried out using
SYBR® Premix ExTaq™ (Takara Bio, Inc.) on an ABI Prism
7500 Sequence Detection System (Applied Biosystems; Thermo Fisher
Scientific, Inc.). GAPDH was used as reference gene. The relative
mRNA expression levels were quantified using the 2−ΔΔCq
method (26). The following
thermocycling conditions were used: Initial denaturation at 95°C
for 7 min; followed by 40 cycles of 95°C for 15 sec and 60°C for 30
sec; and a final extension at 72°C for 30 sec. Primers were as
follows: i) AGT forward, 5′-CCCTGGCTTTCAACACCTAC-3′ and reverse,
5′-CTGTGGGCTCTCTCTCATCC-3′; ii) CXCL12 forward,
5′-GATTGTAGCCCGGCTGAAGA-3′ and reverse,
5′-TTCGGGTCAATGCACACTTGT-3′; iii) ADRB2 forward,
5′-AACTGGTTGGGCTATGTCAA-3′ and reverse, 5′-GTTAGTGTCCTGTCAGGGAG-3′;
and iv) GAPDH forward, 5′-TGTGGGCATCAATGGATTTGG-3′ and reverse
5′-ACACCATGTATTCCGGGTCAAT-3′.
Kaplan-Meier analysis of hub
genes
Kaplan-Meier Plotter (https://kmplot.com/analysis/) was used to examine the
effect of hub genes on OS of patients with diffuse-type GC. In
order to improve the reliability of the results, the GEO GSE62254
(9,18) and GSE15459 (27,28)
datasets were used as the research target. The three genes (AGT,
CXCL12 and ADRB2) were uploaded into the database to obtain the
Kaplan-Meier survival plots. Log rank P-value and hazard ratio (HR)
with 95% confidence intervals were calculated. Patients with
expression above the median (high expression) are indicated in the
red line, and patients with expressions below the median (low
expression) are indicated in the black line. P<0.05 was the
cut-off criterion.
Statistical analysis
The differences between the groups were compared
using an unpaired Student's t-test on Graph Prism 7.0 (GraphPad
Software, Inc.). P<0.05 was considered to indicate a
statistically significant difference.
Results
DEG identification
A flowchart of the bioinformatics analytical methods
is presented in Fig. 1. The
GSE62254 database includes 300 different Lauren-subtype GC samples.
Of these, 265 genes with definite Lauren subtypes and survival data
were singled out, including 128 diffuse-type and 137
intestinal-type GC samples. Detailed information regarding these
samples is presented in Table SI.
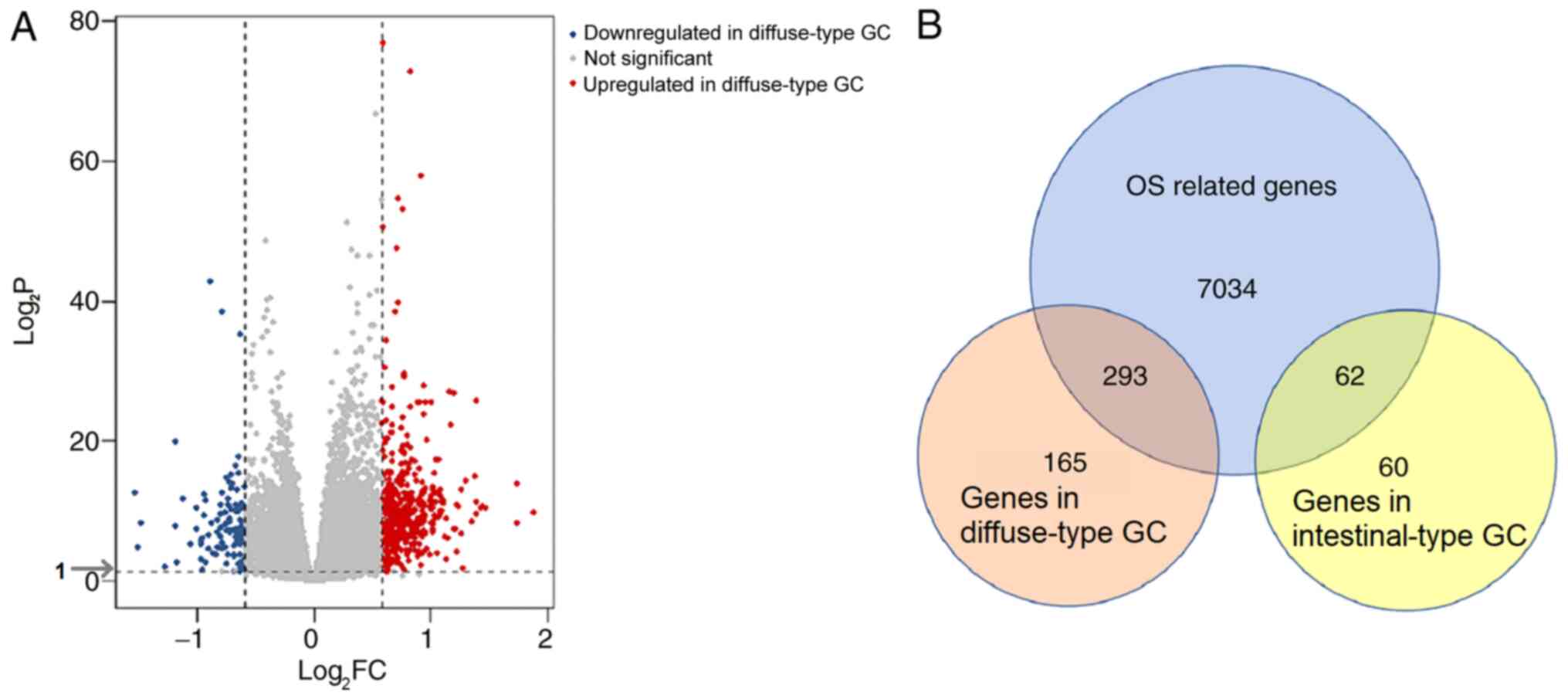
According to the screening criteria of |log2FC|>0.585
and adjusted P<0.05, 584 DEGs, including 458 genes associated
with diffuse-type and 122 genes associates with intestinal-type GC,
were identified (Fig. 2A). To
identify genes with prognostic value, Cox regression analysis was
carried out on the GSE62254 dataset. A total of 7,389 genes were
identified as significantly associated with OS. Of these OS-related
DEGs, 293 genes were identified in diffuse-type GC samples and 62
in intestinal-type GC (Fig.
2B).
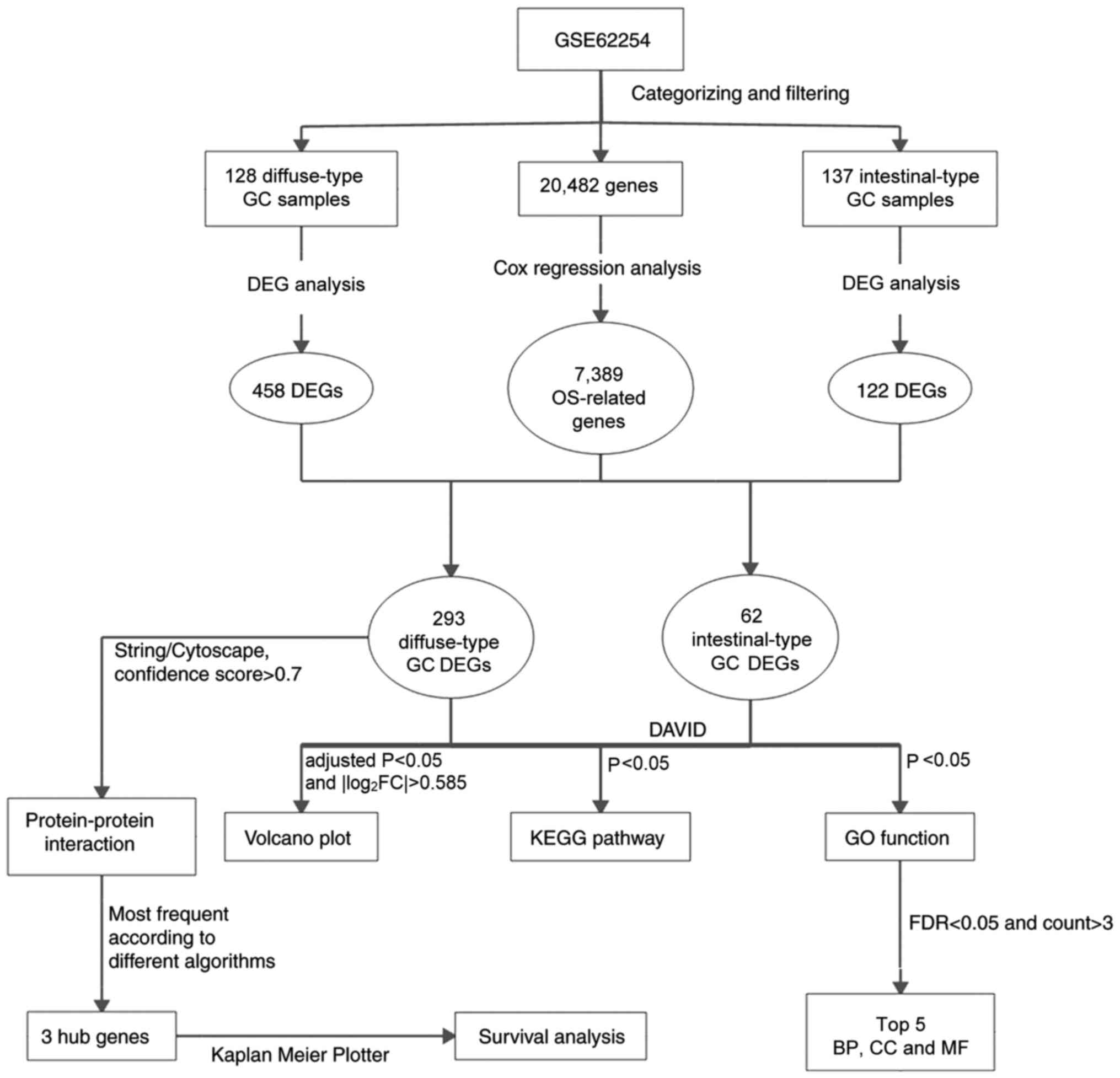 | Figure 1.Flowchart of the bioinformatics
analysis. GC, gastric cancer; OS, overall survival; DEG,
differentially expressed gene; GO, Gene Ontology; KEGG, Kyoto
Encyclopedia of Genes and Genomes; DAVID, Database for Annotation,
Visualization and Integrated Discovery; FDR, false discovery rate;
BP, biological process; CC, cellular component; MF, molecular
function. |
GO enrichment and KEGG pathway
analyses of DEGs
DEGs associated with diffuse-type and
intestinal-type GC were functionally annotated using GO and
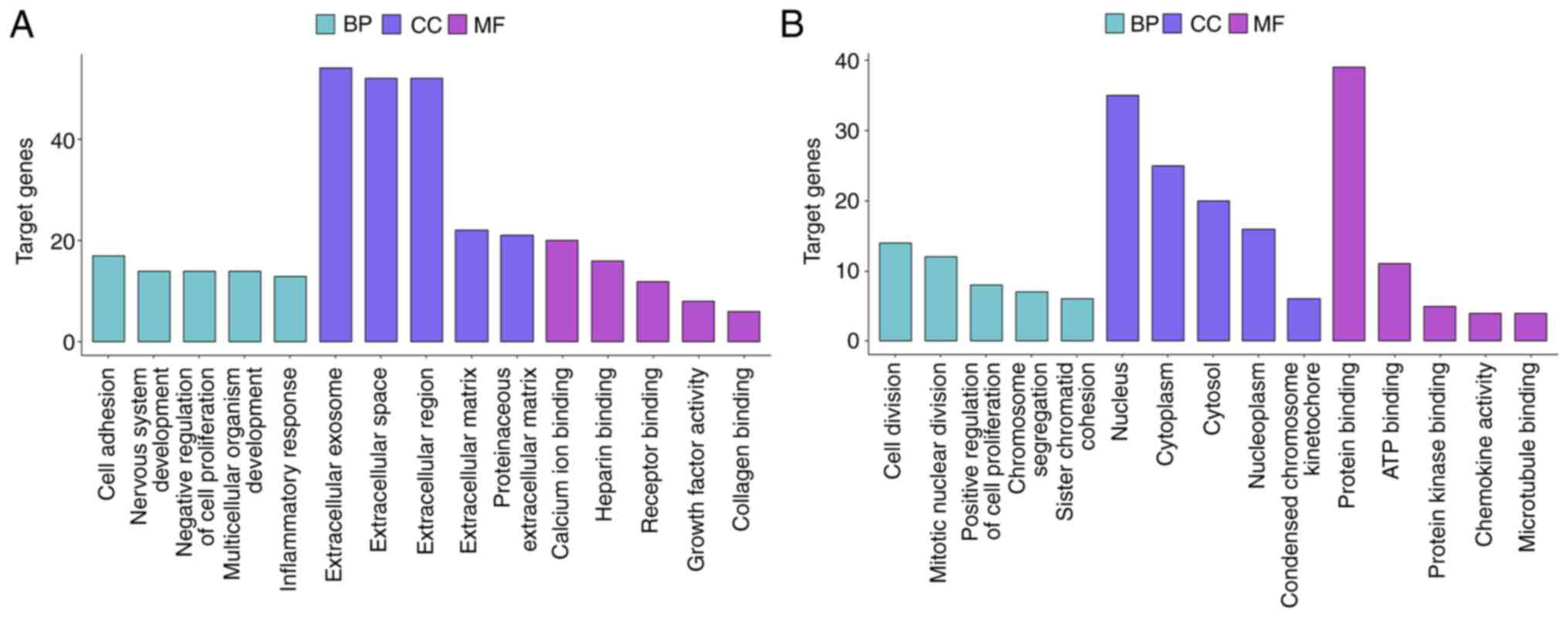
analyzed using KEGG pathway analysis. GO analysis suggested that
diffuse-type DEGs were primarily enriched in the ‘cell adhesion’
BP, the ‘extracellular exosome’ CC and the ‘calcium ion’ binding
MF. By contrast, intestinal-type DEGs were primarily enriched in
the ‘cell division’ BP, the ‘nucleus’ CC and the ‘protein binding’
MF (Figs. 3 and S1A and B). The top 5 BP, CC and MF
results from the GO enrichment analysis of the subtype-specific
DEGs are listed in Table I.
 | Table I.Top 5 enriched GO terms associated
with GC subtype-specific differentially expressed genes. |
Table I.
Top 5 enriched GO terms associated
with GC subtype-specific differentially expressed genes.
| A, Diffuse-type
GC |
|---|
|
|---|
| Category | Term | GO ID | Count | P-value |
|---|
| BP | Cell adhesion | GO:0007155 | 17 |
6×10−4 |
| BP | Nervous system
development | GO:0007399 | 14 |
1×10−4 |
| BP | Negative regulation
of cell proliferation | GO:0008285 | 14 |
3×10−3 |
| BP | Multicellular
organism development | GO:0007275 | 14 |
2×10−2 |
| BP | Inflammatory
response | GO:0006954 | 13 |
6×10−3 |
| CC | Extracellular
exosome | GO:0070062 | 54 |
1×10−2 |
| CC | Extracellular
space | GO:0005615 | 52 |
5×10−11 |
| CC | Extracellular
region | GO:0005576 | 52 |
2×10−8 |
| CC | Extracellular
matrix | GO:0031012 | 22 |
1×10−9 |
| CC | Proteinaceous
extracellular matrix | GO:0005578 | 21 |
1×10−9 |
| MF | Calcium ion
binding | GO:0005509 | 20 |
2×10−3 |
| MF | Heparin
binding | GO:0008201 | 16 |
2×10−9 |
| MF | Receptor
binding | GO:0005102 | 12 |
6×10−3 |
| MF | Growth factor
activity | GO:0008083 | 8 |
5×10−3 |
| MF | Collagen
binding | GO:0005518 | 6 |
1×10−3 |
|
| B,
Intestinal-type GC |
|
|
Category | Term | GO ID | Count | P-value |
|
| BP | Cell division | GO:0051301 | 14 |
7×10−11 |
| BP | Mitotic nuclear
division | GO:0007067 | 12 |
3×10−10 |
| BP | Positive regulation
of cell proliferation | GO:0008284 | 8 |
7×10−4 |
| BP | Chromosome
segregation | GO:0007059 | 7 |
8×10−8 |
| BP | Sister chromatid
cohesion | GO:0007062 | 6 |
2×10−5 |
| CC | Nucleus | GO:0005634 | 35 |
2×10−6 |
| CC | Cytoplasm | GO:0005737 | 25 |
2×10−2 |
| CC | Cytosol | GO:0005829 | 20 |
4×10−3 |
| CC | Nucleoplasm | GO:0005654 | 16 |
2×10−2 |
| CC | Condensed
chromosome kinetochore | GO:0000777 | 6 |
7×10−6 |
| MF | Protein
binding | GO:0005515 | 39 |
2×10−3 |
| MF | ATP binding | GO:0005524 | 11 |
1×10−2 |
| MF | Protein kinase
binding | GO:0019901 | 5 |
3×10−2 |
| MF | Chemokine
activity | GO:0008009 | 4 |
4×10−4 |
| MF | Microtubule
binding | GO:0008017 | 4 |
2×10−2 |
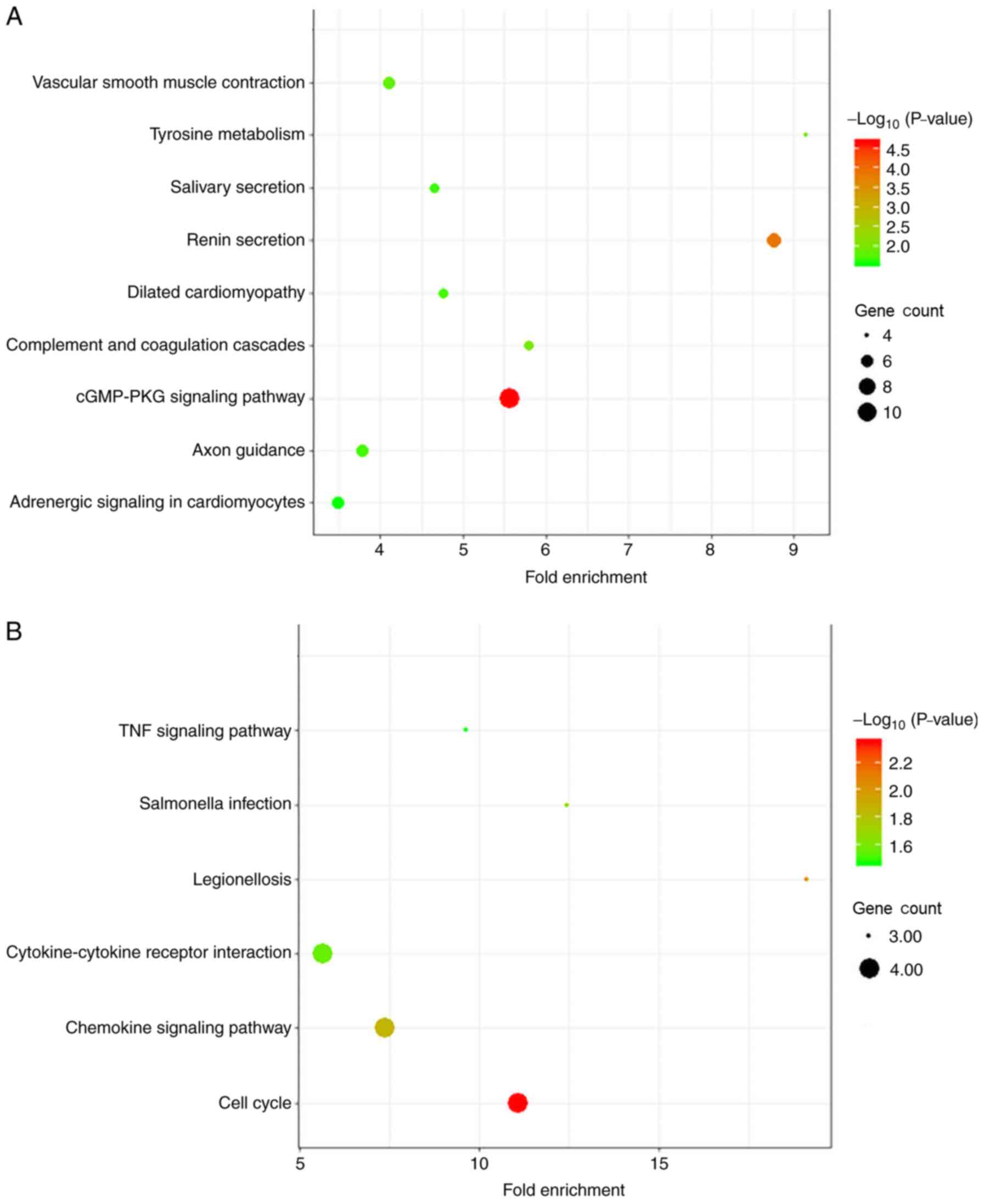
In the KEGG pathway analysis, diffuse-type GC was
primarily enriched in DEGs associated with ‘cGMP-PKG signaling
pathway’. Intestinal-type DEGs were primarily enriched in ‘cell
cycle’ (Fig. 4 and Table II).
 | Table II.KEGG pathway analysis of GC
subtype-specific differentially expressed genes. |
Table II.
KEGG pathway analysis of GC
subtype-specific differentially expressed genes.
| A, Diffuse-type
GC |
|---|
|
|---|
| Term | KEGG ID | Count | P-value |
|---|
| cGMP-PKG signaling
pathway | hsa04022 | 11 | 2.19E-05 |
| Renin
secretion | hsa04924 | 7 | 1.26E-04 |
| Tyrosine
metabolism | hsa00350 | 4 | 0.00896 |
| Complement and
coagulation cascades | hsa04610 | 5 | 0.01021 |
| Vascular smooth
muscle contraction | hsa04270 | 6 | 0.01456 |
| Dilated
cardiomyopathy | hsa05414 | 5 | 0.01983 |
| Axon guidance | hsa04360 | 6 | 0.02008 |
| Salivary
secretion | hsa04970 | 5 | 0.02142 |
| Adrenergic
signaling in cardiomyocytes |
hsa04261 | 6 | 0.02758 |
|
|
|
|
| B,
Intestinal-type GC |
|
| Term | KEGG ID | Count | P-value |
|
|
|
|
| Cell cycle | hsa04110 | 4 | 0.00449 |
| Legionellosis | hsa05134 | 3 | 0.0095 |
| Chemokine signaling
pathway | hsa04062 | 4 | 0.0137 |
| Salmonella
infection | hsa05132 | 3 | 0.02153 |
| Cytokine-cytokine
receptor interaction | hsa04060 | 4 | 0.02776 |
| TNF signaling
pathway | hsa04668 | 3 | 0.03451 |
Construction of the PPI network
To further examine subtype-specific genes in
diffuse-type GC, a PPI network of the 293 DEGs associated with
diffuse-type GC was constructed. A total of 112 nodes and 182
interactions were involved in the PPI network (Fig. S1C). The results were analyzed in
Cytoscape software, and the top 15 hub genes were ranked using four
different algorithms of the CytoHubba plugin according to predicted
scores. Only those genes that overlapped in the results of all four
ranking methods were used for further study, and three overlapping
hub genes were identified for further analysis, namely, AGT, CXCL12
and ADRB2 (Table III).
 | Table III.Hub genes for diffuse type DEGs
ranked in cytoHubba plugin of Cytoscape. |
Table III.
Hub genes for diffuse type DEGs
ranked in cytoHubba plugin of Cytoscape.
|
| Rank methods in
cytoHubba |
|---|
|
|
|
|---|
| Rank | Degree | EPC | Closeness | EcCentricity |
|---|
| 1 | AGTa | AGTa | AGTa | PRKAR2B |
| 2 | CXCL12a | CXCL12a | CXCL12a | ITGA1 |
| 3 | IGF1 | S1PR1 | PRKAR2B | GRP |
| 4 | EDNRB | EDNRB | EDNRB | ACTG2 |
| 5 | FBN1 | CXCL13 | IGF1 | MYH11 |
| 6 | IGFBP5 | ARHGEF25 | ADRB2a | AGTa |
| 7 | CCL19 | GRP | ITGA1 | CXCL12a |
| 8 | MYH11 | CX3CR1 | FGF2 | LMOD1 |
| 9 | TAC1 | TAC1 | CCL19 | ADRB2a |
| 10 | GRP | EDNRA | CXCL13 | ITGA8 |
| 11 | ADRB2a | CCL19 | CX3CR1 | CAV1 |
| 12 | ITGA1 | P2RY12 | S1PR1 | THBS4 |
| 13 | SCG2 | HTR2B | P2RY12 | EDNRA |
| 14 | EDNRA | ADRB2a | FGF13 | SLIT2 |
| 15 | CXCL13 | PRKAR2B | TAC1 | ROBO1 |
AGT, CXCL12 and ADRB2 expression is
increased in diffuse-type GC tissues
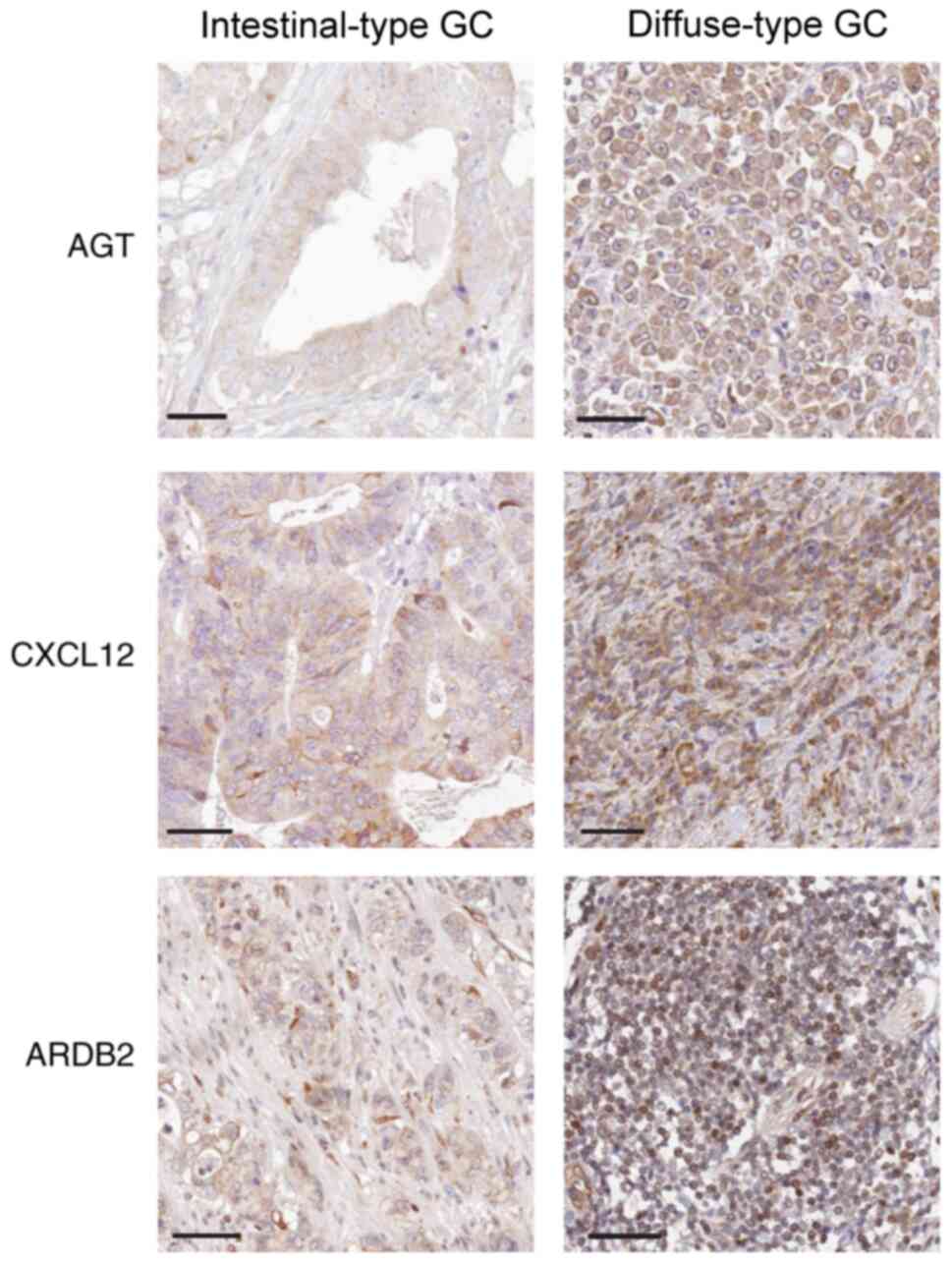
Immunohistochemical analysis indicated that the
staining intensity of AGT, CXCL12 and ADRB2 was associated with GC
Lauren subtypes. Indeed, MD values for AGT, CXCL12 and ADRB2
proteins in the diffuse-type GC samples were significantly
increased, compared with intestinal-type GC samples (P=0.023 for
AGT, P=0.011 for CXCL12 and P=0.007 for ADRB2, respectively;
Table IV). Thus, compared with
intestinal-type GC tissue samples, AGT, CXCL12 and ADRB2 stained
more strongly in diffuse-type GC tissues (Fig. 5).
 | Table IV.Immunohistochemical staining
intensity results for AGT, CXCL12 and ADRB2. |
Table IV.
Immunohistochemical staining
intensity results for AGT, CXCL12 and ADRB2.
| Gene | Intestinal-type
GC | Diffuse-type
GC | P-value |
|---|
| AGT | 2.36±1.29 |
6.87±4.37a | 0.023 |
| CXCL12 | 1.92±1.38 |
6.51±4.83a | 0.011 |
| ADRB2 | 2.32±0.95 |
7.68±5.13a | 0.007 |
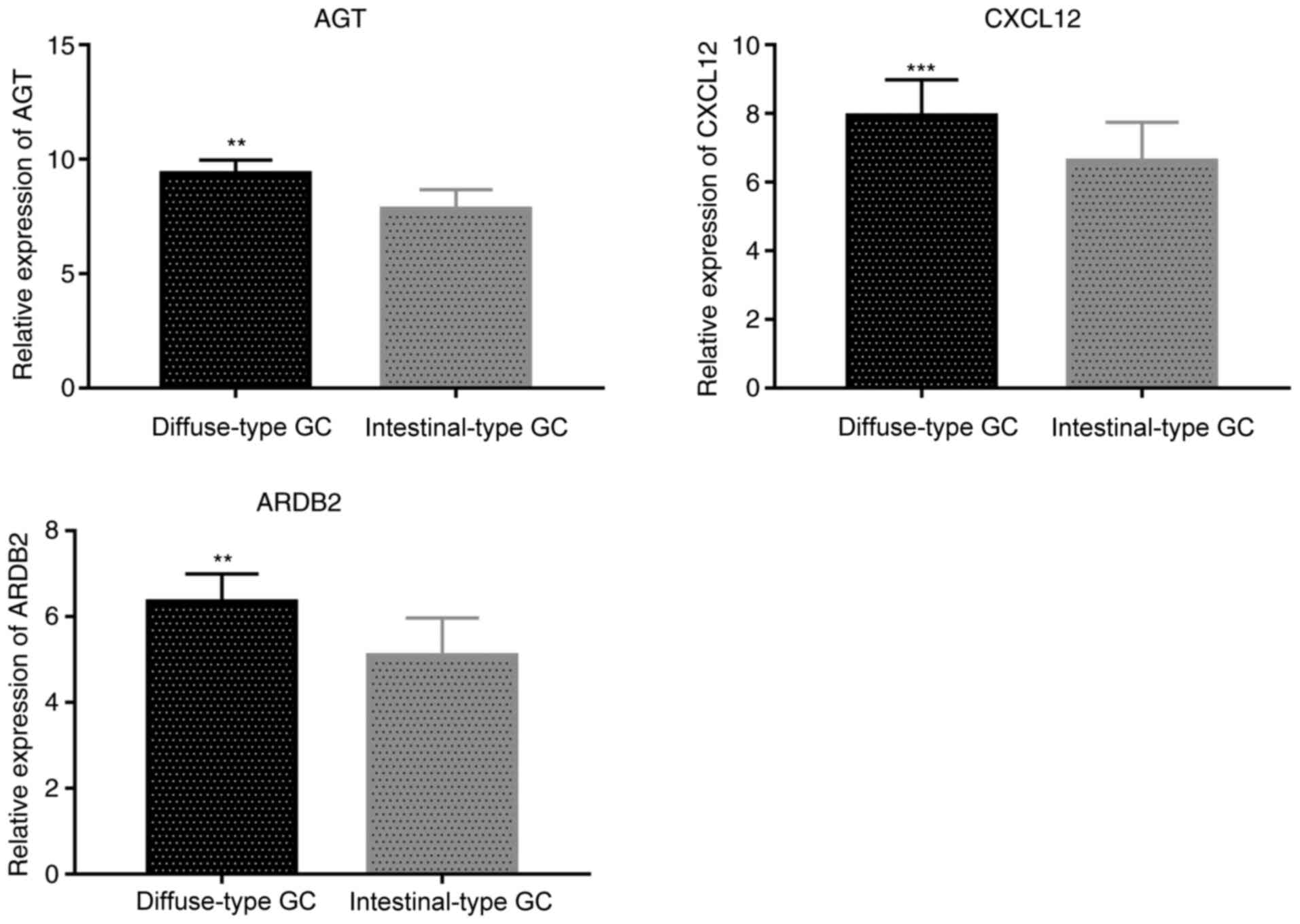
RT-qPCR results were consistent with
immunohistochemistry analysis, showing that expression levels of
AGT, CXCL12 and ADRB2 were significantly higher in diffuse-type GC
than in intestinal-type GC tissues (P=0.0067 for AGT, P=0.00018 for
CXCL12 and P=0.0043 for ADRB2, respectively; Fig. 6).
Association between hub genes and OS
in patients with diffuse-type GC
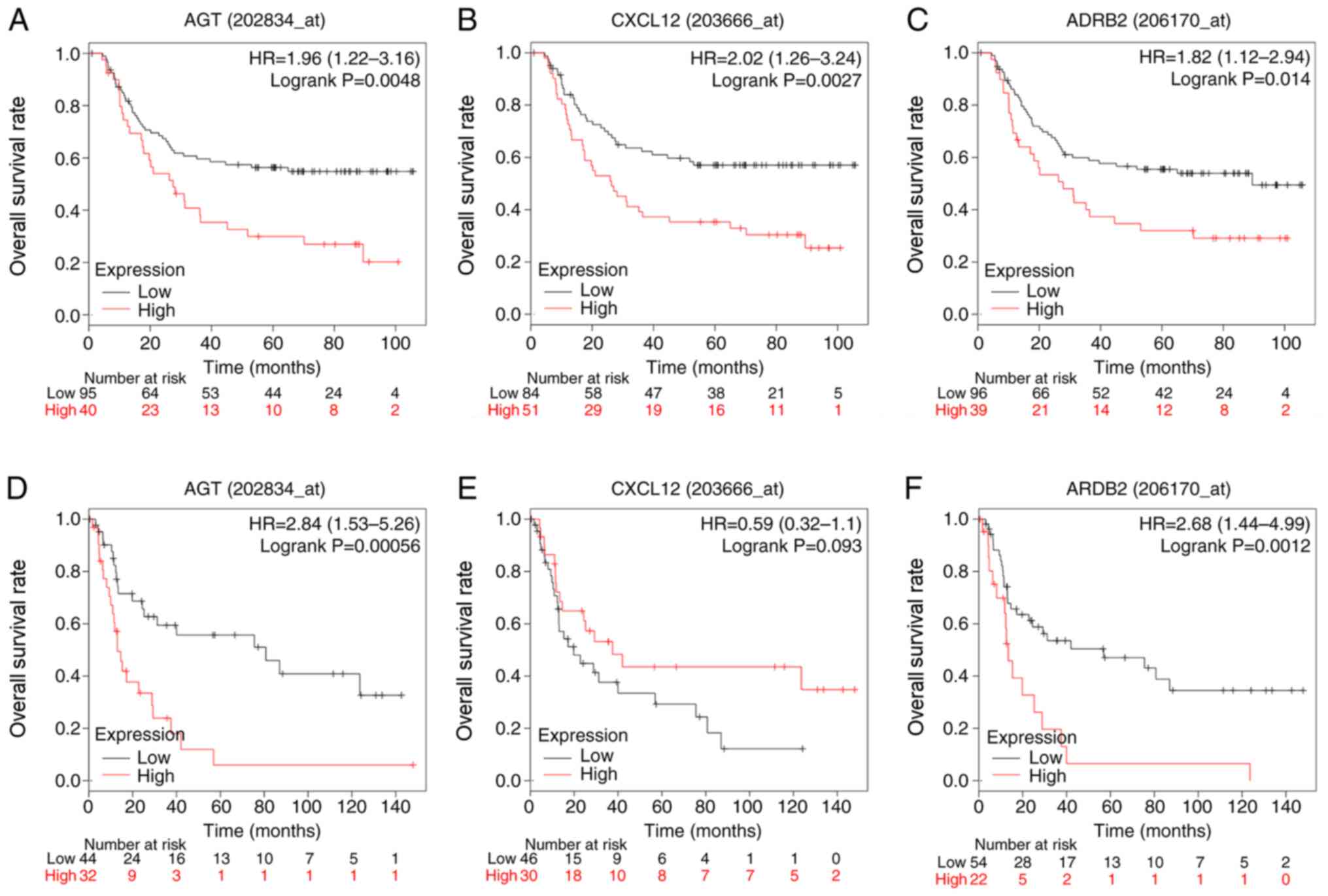
To evaluate the prognostic value of the three
identified hub genes in diffuse-type GC, Kaplan-Meier analysis was
carried out. To improve the reliability of results, two datasets,
GSE62254 and GSE15459, were used. In the GSE62254, high expression
of AGT (P=0.0048), CXCL12 (P=0.0027) and ADRB2 (P=0.014) was
associated with reduced overall survival rates in patients with
diffuse-type GC patients. In GSE15459, high expression of AGT
(P=0.00056) and ARDB2 (P=0.0012) presented similar results,
indicating a poor prognosis for diffuse-type GC patients. However,
expression of CXCL12 (P=0.093) was not associated with overall
survival in this dataset (Fig.
7).
Discussion
GC is a highly heterogeneous disease. Since the
Lauren classification was first proposed in 1965, it has been
widely recognized by clinicians and pathologists and is still
currently used. For many years, the value of histopathological
classification for evaluating the prognosis of GC has been very
limited, and Lauren classification is considered the most valuable
clinicopathological classification. There are significant
differences between different Lauren subtypes (29–31),
suggesting that some specific biomarkers might play an important
role during the pathogenesis and development of GC. Although
several studies have examined the mechanism of the occurrence and
development of GC, few have evaluated specific GC subtypes.
To identify genes specifically associated with
Lauren GC subtypes, the GSE62254 dataset was used, which contains
265 samples of diffuse or intestinal GC with survival data. A total
of 598 DEGs identified, including 293 diffuse-type DEGs and 62
intestinal-type DEGs. To predict the biological pathways and
functions involving these DEGs, GO and KEGG analyses were
performed. To identify key genes for diffuse-type GC progression
among the numerous DEGs, the top 15 hub genes in a PPI network were
identified and screened using four different algorithms. Genes
overlapping in the results of all four algorithms were then
identified as hub genes and selected for further study, namely,
AGT, CXCL12 and ADRB2. All three hub genes were associated with OS
in patients with diffuse-type GC. Thus, these hub genes may
represent novel diagnostic and prognostic biomarkers for
diffuse-type GC.
The AGT gene encodes pre-angiotensinogen, also known
as the angiotensinogen precursor protein, which is primarily
expressed in the liver and cleaved by the renin enzyme in response
to reduced blood pressure. The resulting product, angiotensin 1
(Ang I), is cleaved by the AGT-converting enzyme (ACE) to produce
angiotensin 2 (Ang II). Thus, the product of AGT constitutes a key
component of the renin-angiotensin system (RAS) (32). RAS has been implicated in arterial
hypertension, kidney disease, and other cardiovascular conditions
(33–35), as well as cancer growth and
dissemination (36). RAS components
are expressed in tumor microenvironments and directly or indirectly
affect cell proliferation, invasion, migration, metastasis,
apoptosis, angiogenesis, and cancer-associated inflammation and
immunomodulation (36,37). RAS can also promote tumor growth
indirectly, for instance, by regulating cancer-associated
fibroblasts (38) and promoting
VEGF-mediated angiogenesis (39,40) in
solid tumors. Based on the results of the present study, it was
hypothesized that AGT might be a potential indicator for the
diagnosis and prognosis of diffuse-type GC.
The CXCL12 gene, also known as stromal cell-derived
factor 1 (SDF1), encodes a chemokine of the intercrine family. The
CXCL12 chemokine binds primarily to the CXC receptor 4 (CXCR4)
receptor, playing an essential role in diverse cellular functions
(41–43). CXCR4 is widely expressed on
hematopoietic cells, embryonic pluripotent stem cells and several
types of tissue-committed stem cells (44), which have direct or indirect
proangiogenic properties. The CXCL12/CXCR4 axis is associated with
tumor progression, angiogenesis, metastasis, and survival. For
example, CXCL12 overexpression enhances the proliferation and
invasion of colon cancer cells through the MAPK/PI3K/AP-1 signaling
pathway (45). High CXCR4
expression may represent a biomarker indicating poor prognosis for
hepatocellular carcinoma patients (46). However, in the present study, the
survival curves from two datasets of patients with diffuse-type GC
divided according to CXCL12 expression exhibited opposite trends.
This discrepancy could stem from differential patient
characteristics, such as ethnicity, sex and Helicobacter
pylori infection. CXCL12 has been reported as a prognostic
marker in a variety of cancers according to the literature,
including GC (47). According to
the present results, the relationship between the expression of
CXCL12 and its prognosis in human diffuse type GC between GSE62254
and GSE15459 presented different results. To explain differences
between the two datasets, more comprehensive and precise analysis
is needed.
The ADRB2 gene encodes the β2-adrenergic receptor,
which belongs to the G protein-coupled receptor superfamily. The
ADRB2 protein increases cAMP, and downstream L-type calcium channel
interaction via adenylate cyclase stimulation through trimeric
G-proteins, thereby mediating physiological responses, such as
bronchodilation and smooth muscle relaxation (48). Previous studies have indicated that
ARDB2 also plays an important role in several cancer types. For
instance, ADRB2 signaling negatively regulates autophagy, leading
to hypoxia-inducible factor-1α stabilization and inducing sorafenib
resistance in hepatocellular carcinoma (HCC) (49). Moreover, ADRB2 expression is
associated with prognosis of patients with HCC (50). In prostate cancer, high ADRB2
expression levels activated an angiogenic switch, regulated
angiogenesis and affected the phenotype of prostate cells and
promoted their ability to migrate and invade (51). In GC, chronic stress caused by
stress hormone-induced activation of the ADRB2 signaling pathway
plays a crucial role in progression and metastasis (52). Additionally, ADRB2 signaling
regulates GC progression (53). In
the present study, ARDB2 was highly expressed in diffuse-type GC
and was associated with OS.
There are certain limitations to the present study.
Only using immunohistochemical and RT-qPCR methods to study the
qualitative and semiquantitative expression of AGT, CXCL12 and
ADRB2 in diffuse-type GC may yield results that are not
sufficiently rigorous. These data should be supplemented with
further experiments to validate expression levels of AGT, CXCL12
and ADRB2 in GC tissue. Furthermore, key signaling pathways related
to the identified hub genes should also be examined in in
vitro experiments.
In summary, the present study demonstrated that
expression of AGT, CXCL12 and ADRB2 was increased in diffuse-type
GC, relative to intestinal-type GC, and that expression levels of
these proteins negatively correlated with disease prognosis.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets generated and/or analyzed during the
current study are available in the GEO data repository, [https://www.ncbi.nlm.nih.gov/geo]. The datasets
used and/or analyzed during the current study are available from
the corresponding author on reasonable request.
Authors' contributions
GW conceived and designed the present study. SL and
CY collected, extracted and analyzed the data. SL and FD performed
the experiments and interpreted the data. SL wrote the manuscript.
YC, FD and GW contributed to the data collection and performed the
statistical analysis. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study was approved by The Ethics
Committee of The Third Affiliated Hospital of Anhui Medical
University. Written informed consent was obtained from all
participants.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Allemani C, Weir HK, Carreira H, Harewood
R, Spika D, Wang XS, Bannon F, Ahn JV, Johnson CJ, Bonaventure A,
et al: Global surveillance of cancer survival 1995–2009: Analysis
of individual data for 25,676,887 patients from 279
population-based registries in 67 countries (CONCORD-2). Lancet.
385:977–1010. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Spolverato G, Ejaz A, Kim Y, Squires MH,
Poultsides GA, Fields RC, Schmidt C, Weber SM, Votanopoulos K,
Maithel SK and Pawlik TM: Rates and patterns of recurrence after
curative intent resection for gastric cancer: A United States
multi-institutional analysis. J Am Coll Surg. 219:664–675. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
In H, Solsky I, Palis B, Langdon-Embry M,
Ajani J and Sano T: Validation of the 8th edition of the AJCC TNM
staging system for gastric cancer using the national cancer
database. Ann Surg Oncol. 24:3683–3691. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Borrmann R: Geschwulste des magens und des
duodenums. Handbuch spez pathol anat und histo. Henke F and
Lubarsch O: Springer Verlag; Berlin: pp. 812–1054. 1926, (In
German).
|
|
6
|
Lauren P: The two histological main types
of gastric carcinoma: Diffuse and so-called intestinal-type
carcinoma. An attempt at a histo-clinical classification. Acta
Pathol Microbiol Scand. 64:31–49. 1965. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Fléjou JF: WHO Classification of digestive
tumors: The fourth edition. Ann Pathol. 31:S27–S31. 2011.(In
French). View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Cancer Genome Atlas Research Network, .
Comprehensive molecular characterization of gastric adenocarcinoma.
Nature. 513:202–209. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cristescu R, Lee J, Nebozhyn M, Kim KM,
Ting JC, Wong SS, Liu J, Yue YG, Wang J, Yu K, et al: Molecular
analysis of gastric cancer identifies subtypes associated with
distinct clinical outcomes. Nat Med. 21:449–456. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zheng H, Takahashi H, Murai Y, Cui Z,
Nomoto K, Miwa S, Tsuneyama K and Takano Y: Pathobiological
characteristics of intestinal and diffuse-type gastric carcinoma in
Japan: An immunostaining study on the tissue microarray. J Clin
Pathol. 60:273–277. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Qiu MZ, Cai MY, Zhang DS, Wang ZQ, Wang
DS, Li YH and Xu RH: Clinicopathological characteristics and
prognostic analysis of Lauren classification in gastric
adenocarcinoma in China. J Transl Med. 11:582013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gong EJ, Lee JY, Bae SE, Park YS, Choi KD,
Song HJ, Lee GH, Jung HY, Jeong WJ, Cheon GJ, et al:
Characteristics of non-cardia gastric cancer with a high serum
anti-Helicobacter pylori IgG titer and its association with
diffuse-type histology. PLoS One. 13:e01952642018. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Tang CT, Zeng L, Yang J, Zeng C and Chen
Y: Analysis of the incidence and survival of gastric cancer based
on the lauren classification: A large population-based study using
SEER. Front Oncol. 10:12122020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Pan Q, Shai O, Lee LJ, Frey BJ and
Blencowe BJ: Deep surveying of alternative splicing complexity in
the human transcriptome by high-throughput sequencing. Nat Genet.
40:1413–1415. 2008. View
Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yang G, Zhang Y and Yang J: Identification
of potentially functional CircRNA-miRNA-mRNA regulatory network in
gastric carcinoma using bioinformatics analysis. Med Sci Monit.
25:8777–8796. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chen S, Pan S, Wu H, Zhou J, Huang Y, Wang
S and Liu A: ICAM1 regulates the development of gastric cancer and
may be a potential biomarker for the early diagnosis and prognosis
of gastric cancer. Cancer Manag Res. 12:1523–1534. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Li R, Jiang J, Shi H, Qian H, Zhang X and
Xu W: CircRNA: A rising star in gastric cancer. Cell Mol Life Sci.
77:1661–1680. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Oh SC, Sohn BH, Cheong JH, Kim SB, Lee JE,
Park KC, Lee SH, Park JL, Park YY, Lee HS, et al: Clinical and
genomic landscape of gastric cancer with a mesenchymal phenotype.
Nat Commun. 9:17772018. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wilson CL and Miller CJ: Simpleaffy: A
BioConductor package for Affymetrix quality control and data
analysis. Bioinformatics. 21:3683–3685. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Gautier L, Cope L, Bolstad BM and Irizarry
RA: affy-analysis of Affymetrix GeneChip data at the probe level.
Bioinformatics. 20:307–315. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW,
Shi W and Smyth GK: limma powers differential expression analyses
for RNA-sequencing and microarray studies. Nucleic Acids Res.
43:e472015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Therneau TM and Grambsch PM: The Cox
model. Modeling Survival Data: Extending the Cox Model. Springer;
Berlin: pp. 39–77. 2000
|
|
23
|
Szklarczyk D, Morris JH, Cook H, Kuhn M,
Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, et al:
The STRING database in 2017: Quality-controlled protein-protein
association networks, made broadly accessible. Nucleic Acids Res.
45(D1): D362–D368. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT and
Lin CY: cytoHubba: Identifying hub objects and sub-networks from
complex interactome. BMC Syst Biol. 8 (Suppl 4):S112014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ooi CH, Ivanova T, Wu J, Lee M, Tan IB,
Tao J, Ward L, Koo JH, Gopalakrishnan V, Zhu Y, et al: Oncogenic
pathway combinations predict clinical prognosis in gastric cancer.
PLoS Genet. 5:e10006762009. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Muratani M, Deng N, Ooi WF, Lin SJ, Xing
M, Xu C, Qamra A, Tay ST, Malik S, Wu J, et al: Nanoscale chromatin
profiling of gastric adenocarcinoma reveals cancer-associated
cryptic promoters and somatically acquired regulatory elements. Nat
Commun. 5:43612014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Qiu M, Zhou Y, Zhang X, Wang Z, Wang F,
Shao J, Lu J, Jin Y, Wei X, Zhang D, et al: Lauren classification
combined with HER2 status is a better prognostic factor in Chinese
gastric cancer patients. BMC Cancer. 14:8232014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Chen YC, Fang WL, Wang RF, Liu CA, Yang
MH, Lo SS, Wu CW, Li AF, Shyr YM and Huang KH: Clinicopathological
variation of lauren classification in gastric cancer. Pathol Oncol
Res. 22:197–202. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Janjigian YY, Werner D, Pauligk C,
Steinmetz K, Kelsen DP, Jäger E, Altmannsberger HM, Robinson E,
Tafe LJ, Tang LH, et al: Prognosis of metastatic gastric and
gastroesophageal junction cancer by HER2 status: A European and USA
international collaborative analysis. Ann Oncol. 23:2656–2662.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Lu H, Cassis LA, Kooi CW and Daugherty A:
Structure and functions of angiotensinogen. Hypertens Res.
39:492–500. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Kobori H, Nangaku M, Navar LG and
Nishiyama A: The intrarenal renin-angiotensin system: From
physiology to the pathobiology of hypertension and kidney disease.
Pharmacol Rev. 59:251–287. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Ferrario CM: Role of angiotensin II in
cardiovascular disease therapeutic implications of more than a
century of research. J Renin Angiotensin Aldosterone Syst. 7:3–14.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Paul M, Poyan Mehr A and Kreutz R:
Physiology of local renin-angiotensin systems. Physiol Rev.
86:747–803. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
George AJ, Thomas WG and Hannan RD: The
renin-angiotensin system and cancer: Old dog, new tricks. Nat Rev
Cancer. 10:745–759. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Ager EI, Neo J and Christophi C: The
renin-angiotensin system and malignancy. Carcinogenesis.
29:1675–1684. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Chauhan VP, Martin JD, Liu H, Lacorre DA,
Jain SR, Kozin SV, Stylianopoulos T, Mousa AS, Han X,
Adstamongkonkul P, et al: Angiotensin inhibition enhances drug
delivery and potentiates chemotherapy by decompressing tumour blood
vessels. Nat Commun. 4:25162013. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Ji Y, Wang Z, Li Z, Li K, Le X and Zhang
T: Angiotensin II induces angiogenic factors production partly via
AT1/JAK2/STAT3/SOCS3 signaling pathway in MHCC97H cells. Cell
Physiol Biochem. 29:863–874. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Anandanadesan R, Gong Q, Chipitsyna G,
Witkiewicz A, Yeo CJ and Arafat HA: Angiotensin II induces vascular
endothelial growth factor in pancreatic cancer cells through an
angiotensin II type 1 receptor and ERK1/2 signaling. J Gastrointest
Surg. 12:57–66. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Balkwill F: Cancer and the chemokine
network. Nat Rev Cancer. 4:540–550. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Murdoch C: CXCR4: Chemokine receptor
extraordinaire. Immunol Rev. 177:175–184. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Balabanian K, Lagane B, Infantino S, Chow
KY, Harriague J, Moepps B, Arenzana-Seisdedos F, Thelen M and
Bachelerie F: The chemokine SDF-1/CXCL12 binds to and signals
through the orphan receptor RDC1 in T lymphocytes. J Biol Chem.
280:35760–35766. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Ratajczak MZ, Zuba-Surma E, Kucia M, Reca
R, Wojakowski W and Ratajczak J: The pleiotropic effects of the
SDF-1-CXCR4 axis in organogenesis, regeneration and tumorigenesis.
Leukemia. 20:1915–1924. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Ma J, Su H, Yu B, Guo T, Gong Z, Qi J,
Zhao X and Du J: CXCL12 gene silencing down-regulates metastatic
potential via blockage of MAPK/PI3K/AP-1 signaling pathway in colon
cancer. Clin Transl Oncol. 20:1035–1045. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Liepelt A and Tacke F: Stromal
cell-derived factor-1 (SDF-1) as a target in liver diseases. Am J
Physiol Gastrointest Liver Physiol. 311:G203–G209. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Ishigami S, Natsugoe S, Okumura H,
Matsumoto M, Nakajo A, Uenosono Y, Arigami T, Uchikado Y, Setoyama
T, Arima H, et al: Clinical implication of CXCL12 expression in
gastric cancer. Ann Surg Oncol. 14:3154–3158. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Yang-Feng TL, Xue FY, Zhong WW, Cotecchia
S, Frielle T, Caron MG, Lefkowitz RJ and Francke U: Chromosomal
organization of adrenergic receptor genes. Proc Natl Acad Sci USA.
87:1516–1520. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Wu FQ, Fang T, Yu LX, Lv GS, Lv HW, Liang
D, Li T, Wang CZ, Tan YX, Ding J, et al: ADRB2 signaling promotes
HCC progression and sorafenib resistance by inhibiting autophagic
degradation of HIF1α. J Hepatol. 65:314–324. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Chen D, Xing W, Hong J, Wang M, Huang Y,
Zhu C, Yuan Y and Zeng W: The beta2-adrenergic receptor is a
potential prognostic biomarker for human hepatocellular carcinoma
after curative resection. Ann Surg Oncol. 19:3556–3565. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Zahalka AH, Arnal-Estapé A, Maryanovich M,
Nakahara F, Cruz CD, Finley LWS and Frenette PS: Adrenergic nerves
activate an angio-metabolic switch in prostate cancer. Science.
358:321–326. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Zhang X, Zhang Y, He Z, Yin K, Li B, Zhang
L and Xu Z: Chronic stress promotes gastric cancer progression and
metastasis: An essential role for ADRB2. Cell Death Dis.
10:7882019. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Zhi X, Li B, Li Z, Zhang J, Yu J, Zhang L
and Xu Z: Adrenergic modulation of AMPK-dependent autophagy by
chronic stress enhances cell proliferation and survival in gastric
cancer. Int J Oncol. 54:1625–1638. 2019.PubMed/NCBI
|