Introduction
Neuroblastoma is one of the most common solid tumors
in children and accounts for ~15% of pediatric cancer-related
deaths (1,2). Neuroblastoma cells originate from
neural crest cells that are committed to the development of the
sympathetic nervous system, and display extreme heterogeneity
ranging from spontaneous regression to malignant progression.
Although the prognosis for neuroblastoma patients has improved
considerably, <40% of high-risk patients can expect long-term
survival (3). The majority of
cancer-related deaths in neuroblastoma are due to tumor relapse
caused by minimal residual disease (MRD).
Tumor-initiating cells (TICs) account for only a
small population of tumor cells that generate the bulk of the tumor
through continuous self-renewal and differentiation, and are found
in a variety of cancers (4–6). TICs can persist as MRD locally in the
primary tumor or distantly in the blood, bone marrow and metastatic
organs (7,8). Neuroblastoma TICs were recently
identified as spheres grown in serum-free non-adherent culture and
were shown to express stem cell markers, exhibit self-renewal, and
form metastatic tumors in immunodeficient mice (9,10).
Sensitive detection of MRD is fundamental when
monitoring the disease status and evaluating the treatment response
(7,11). In acute lymphoblastic leukemia
(ALL), MRD detection with real-time PCR is directed against
tumor-specific junctions using genomic DNA as a template. This
sensitive MRD detection in ALL patients has revealed the most
important prognostic factors and is used to stratify treatments in
the clinic (12–14). In neuroblastoma, the rationale for
introducing the current MRD detection protocols to stratify
treatments in the clinic remains unclear (15,16).
Since no recurrent fusion genes have been found in neuroblastoma
cells, MRD detection in neuroblastoma patients requires
well-defined real-time RT-PCR markers that can be assessed using
cDNA as a template. Although ideal MRD markers should be
exclusively expressed in neuroblastoma cells and not in normal
cells, commonly used markers are selected by their ability to
experimentally define a cut-off value that distinguishes
neuroblastoma cells from normal cells.
Tyrosine hydroxylase (TH) was initially evaluated as
a single marker for neuroblastoma MRD detection, but was hampered
by its expression in normal cells (17–20).
Paired-like homeobox 2b (PHOX2B) was subsequently identified as a
more specific single marker, but was not highly expressed in all
neuroblastoma tumors (21). To
overcome tumor heterogeneity and achieve more sensitive detection
of MRD in neuroblastoma patients, several sets of multiple markers
have been reported at different centers. Based on gene expression
analyses in neuroblastoma tumors, Viprey et al(22) identified the following three
markers: doublecortin (DCX), PHOX2B and TH, while Cheung et
al(23) independently
identified the following 8 markers: cyclin D1 (CCND1), collapsin
response mediator protein 1 (CRMP1), dopa decarboxylase (DDC), GABA
A receptor β3 (GABRB3), ISL LIM homeobox 1 (ISL1), kinesin family
member 1A (KIF1A), PHOX2B, and transforming acidic coiled-coil
containing protein 2 (TACC2). Furthermore, Stutterheim et
al(24) selected the following
6 markers: cholinergic receptor, nicotinic, α3 (CHRNA3), dopamine
β-hydroxylase (DBH), DDC, growth-associated protein 43 (GAP43),
PHOX2B and TH. These MRD markers are currently being validated.
In the present study, we validated 14 commonly used
real-time RT-PCR markers based on their expression in neuroblastoma
TICs, and developed a novel MRD detection protocol. By using this
protocol, we prospectively monitored MRD in 73 bone marrow (BM), 12
peripheral blood stem cell (PBSC), and 8 peripheral blood (PB)
samples from 14 neuroblastoma patients treated at Kobe Children's
Hospital, Japan, between November, 2010 and August, 2012.
Materials and methods
Patients and samples
Normal PB samples were obtained from three healthy
volunteers. PB, PBSC and BM samples were obtained from 14
neuroblastoma patients who were treated at Kobe Children's Hospital
between November, 2010 and August, 2012. Written informed consent
was obtained from the patients, and the study was approved by Kobe
Children's Hospital Ethics Board. The samples were handled in
accordance with the Guidelines for the Clinical Research of Kobe
University Graduate School of Medicine. All PB, PBSC and BM samples
were separated using Mono-Poly Resolving Medium (DS Pharma
Biomedical, Osaka, Japan), and nucleated cells were collected
according to the manufacturer's instructions.
Neuroblastoma TICs
Human neuroblastoma BE(2)-C cells were obtained from
the American Type Culture Collection (Manassas, VA, USA), and
neuroblastoma TICs were isolated as spheres of BE(2)-C cells as
described previously (25).
Briefly, BE(2)-C cells were cultured at 37°C (5% CO2 and
95% air) in sphere medium consisting of Dulbecco's modified Eagle's
medium (DMEM)/Ham's F12 (3:1) (Wako Pure Chemical-Industries,
Osaka, Japan), 100 U/ml penicillin/streptomycin (PC/SM)
(Invitrogen), 2% B27 supplement (Life Technologies, Carlsbad, CA,
USA), 40 ng/ml basic fibroblast growth factor (bFGF) (R&D
Systems, Minneapolis, MN, USA) and 20 ng/ml epidermal growth factor
(EGF) (R&D Systems). The cells were subcultured with
non-enzymatic cell dissociation solution (Sigma, St. Louis, MO,
USA) and maintained for more than 4 weeks in Ultra-Low Attachment
culture dishes (Corning, Inc., Corning, NY, USA).
RNA extraction and cDNA synthesis
Total RNA from normal BM was obtained from Clonetech
(Palo Alto, CA, USA). Total RNA from neuroblastoma TICs as well as
PB, PBSC and BM samples was extracted with a TRIzol Plus RNA
Purification kit (Life Technologies) according to the
manufacturer's instructions. RNA integrity was evaluated by agarose
gel electrophoresis. cDNA was synthesized from 1 (or 0.5) μg of
total RNA using a Quantitect Reverse Transcription kit (Qiagen,
Valencia, CA, USA) and diluted to a total volume of 80 (or 40)
μl.
Real-time RT-PCR
Real-time RT-PCR was performed using an ABI 7500
Fast Real-time PCR system (Applied Biosystems, Foster City, CA,
USA) in a total volume of 15 μl consisting of 7.5 μl of 2X
FastStart Universal SYBR-Green Master (Roche, Mannheim, Germany),
1.5 μl each of 3 μM sense and antisense primers, and 1 μl of the
sample cDNA (corresponding to 12.5 ng of total RNA). Each cDNA was
amplified with a precycling hold at 95°C for 10 min, followed by 40
cycles at 95°C for 15 sec and 60°C for 60 sec, and one cycle at
95°C for 15 sec, 60°C for 60 sec, 95°C for 15 sec and 60°C for 15
sec. Each sample was analyzed in triplicate. To correct for
differences in the amount of total RNA and in the efficiency of
cDNA synthesis, the expression of each marker was calculated based
on the relative standard curve method using β2-microglobulin (B2M)
as an endogenous reference for normalization.
Primers
The following primers were used: GD2 synthase
(B4GALNT1, NM_001478) 5′-CCAACTCAACAGGCAACTACAA-3′ (sense) and
5′-ATGTCCCTCGGTGGAGAAC-3′ (antisense); CCND1 (NM_053056)
5′-GAAGATCGTCGCCACCTG-3′ (sense) and 5′-GACCTCCTCCTCGCACTTCT-3′
(antisense); CHRNA3 (NM_000743) 5′-TGAAATGGAACCCCTCTGAC-3′ (sense)
and 5′-GGAAATCCCCAACAGCATT-3′ (antisense); CRMP1 (NM_001014809)
5′-GAGTGCAGCCGACATCATC-3′ (sense) and 5′-GGGCTCTCCAAAAACTAGGG-3′
(antisense); DBH (NM_000787) 5′-GACCCCAAGGATTACCTCATT-3′ (sense)
and 5′-GTTGATGGCCTCCAGTGAC-3′ (antisense); DCX (NM_000555)
5′-CATCCCCAACACCTCAGAAG-3′ (sense) and 5′-GGAGGTTCCGTTTGCTGA-3′
(antisense); DDC (NM_000790) 5′-GGAGAAGGGGGAGGAGTG-3′ (sense) and
5′-CAGCCGATGGATCACTTTG-3′ (antisense); GABRB3 (NM_000814)
5′-GGGTGTCCTTCTGGATCAATTA-3′ (sense) and
5′-TTGTCAGCACAGTTGTGATCC-3′ (antisense); GAP43 (NM_001130064)
5′-GAGGATGCTGCTGCCAAG-3′ (sense) and 5′-GGCACTTTCCTTAGGTTTGGT-3′
(antisense); ISL1 (NM_002202) 5′-AAGGACAAGAAGCGAAGCAT-3′ (sense)
and 5′-TTCCTGTCATCCCCTGGATA-3′ (antisense); KIF1A (NM_004321)
5′-CTTGGCGACATCACTGACAT-3′ (sense) and 5′-GCTGGACAGGGCTGAGAG-3′
(antisense); PHOX2B (NM_003924) 5′-CTACCCCGACATCTACACTCG-3′ (sense)
and 5′-CCTGCTTGCGAAACTTGG-3′ (antisense); TACC2 (NM_006997)
5′-CCCCACTATTCGCTCAGAAA-3′ (sense) and 5′-GGGCTTCTATCCGCATGAT-3′
(antisense); TH (NM_199292) 5′-GCCAAGGACAAGCTCAGG-3′ (sense) and
5′-AGCGTGTACGGGTCGAACT-3′ (antisense); and β2-microglobulin (B2M,
NM_004048) 5′-TTCTGGCCTGGAGGCTATC-3′ (sense) and
5′-TCAGGAAATTTGACTTTCCATTC-3′ (antisense).
Standard curves
To mimic MRD-positive PB, PBSC and BM samples, total
RNA from neuroblastoma TICs was serially diluted into total RNA
from normal PB to yield a total amount of 1.0 μg. The resulting
six-step 5-fold dilutions were converted to cDNA yielding 100, 20,
4, 0.8, 0.16 and 0.032% dilutions. To obtain a standard curve, the
CT values for each marker were obtained from triplicate
real-time RT-PCR and plotted against the logarithm of the %
dilution/100. Due to slight expression of each marker in normal PB,
we generated standard curves with R2 >0.90 for each
marker by limiting the plotted points of the % dilution. The
quantitative range was defined as the minimum % dilution that
plotted to the standard curve or above.
Normal range
Based on the threshold defined by the European Study
Group on MRD detection in ALL (ESG-MRD-ALL), the mean CT
value of a normal sample −3 was set as the maximum CT
value for the normal sample (12).
The normal range was then defined as any value less than the %
dilution of MRD that corresponded to the maximum CT
value of the normal sample. If the mean CT value of the
normal sample was >40, the normal range was defined as less than
the % dilution of MRD that corresponded to the minimum % dilution
of MRD in the quantitative range.
Results
Expression profile of real-time RT-PCR
markers in neuroblastoma TICs
To achieve more sensitive detection of MRD, we first
analyzed the expression of commonly used real-time RT-PCR markers
in normal PB, normal BM and neuroblastoma TICs that were isolated
as spheres of BE(2)-C cells (25).
In addition to the 13 MRD markers that were collectively reported
by Viprey et al(22), Cheung
et al(23) and Stutterheim
et al(24), a well-examined
MRD marker, B4GALNT1, was also included in the present study
(26). When the expression of these
14 MRD markers were analyzed by real-time RT-PCR, CHRNA3, CRMP1,
DBH, DCX, DDC, GABRB3, GAP43, ISL1, KIF1A, PHOX2B and TH were
predominantly expressed in neuroblastoma TICs in which the mean
CT value was >9.0 lower when compared with that in
normal PB and BM (Table I).
Considering the variation of the mean CT value for
normal PB, BM and neuroblastoma TICs (12), B4GALNT1, CCND1 and TACC2 were
excluded from the present study.
 | Table IExpression profile of real-time
RT-PCR markers in neuroblastoma TICs. |
Table I
Expression profile of real-time
RT-PCR markers in neuroblastoma TICs.
| Gene name | Normal PB (mean
CT ± SD) | Normal BM (mean
CT ± SD) | Neuroblastoma TICs
(mean CT ± SD) |
|---|
| B4GALNT1 | >40 | 31.593±0.382 | 25.535±0.122 |
| CCND1 | 34.051±0.329 | 28.725±0.096 | 22.243±0.056 |
| CHRNA3 | >40 | 31.028±0.709 | 21.538±0.102 |
| CRMP1 | >40 | 34.863±0.957 | 21.779±0.103 |
| DBH | >40 | >40 | 23.133±0.059 |
| DCX | >40 | 31.683±0.727 | 22.037±0.544 |
| DDC | >40 | 36.150±0.329 | 21.538±0.102 |
| GABRB3 | 35.996±0.130 | 31.952±0.865 | 22.340±0.102 |
| GAP43 | 35.639±0.922 | 31.828±0.920 | 19.777±0.010 |
| ISL1 | >40 | >40 | 24.929±0.044 |
| KIF1A | >40 | 32.601±0.728 | 19.901±0.044 |
| PHOX2B | >40 | >40 | 20.351±0.140 |
| TACC2 | >40 | 28.007±1.272 | 34.782±0.001 |
| TH | >40 | 32.741±1.224 | 18.666±0.034 |
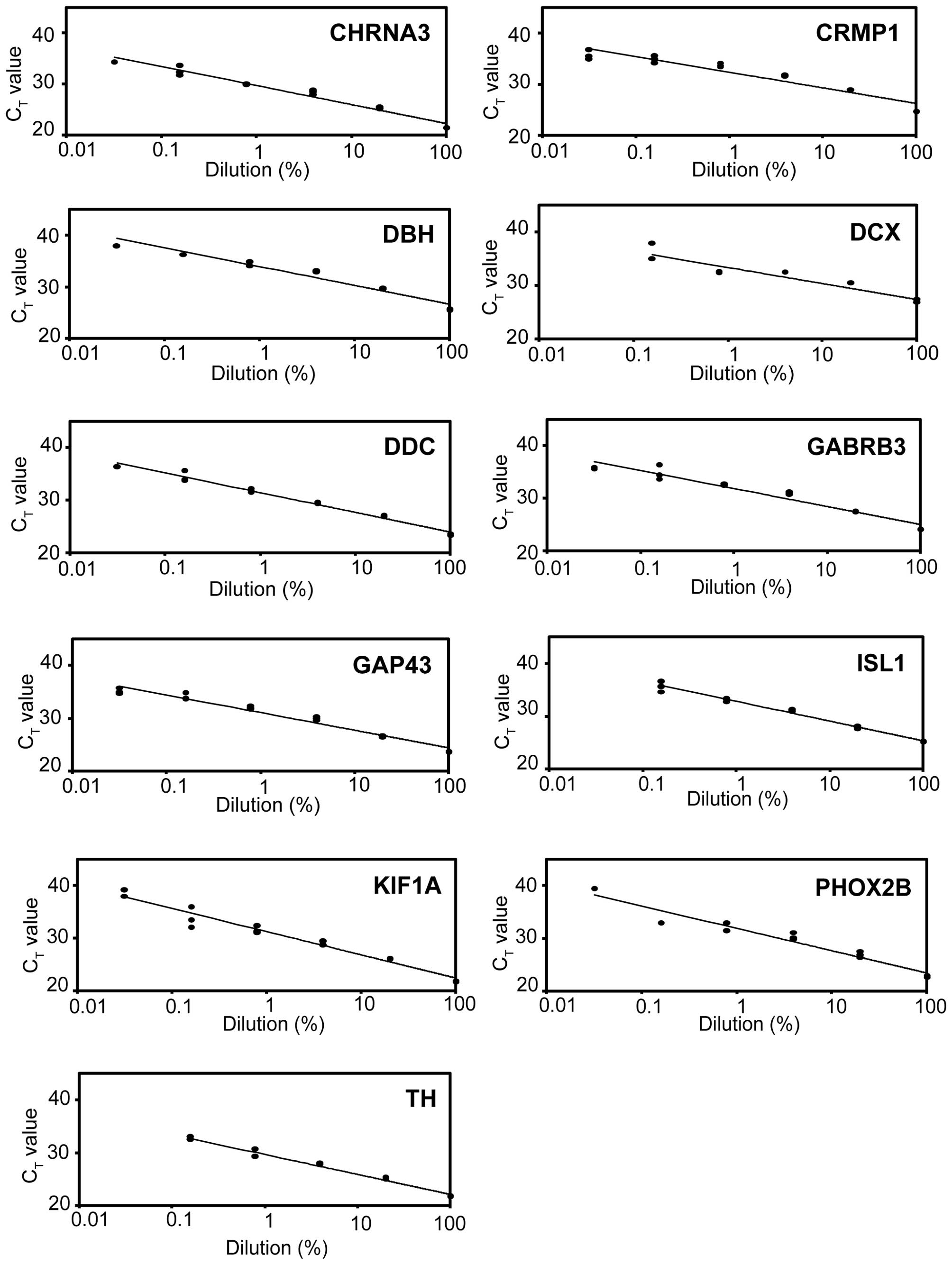
Standard curves for real-time RT-PCR
markers
To mimic the actual expression of neuroblastoma MRD
markers in clinical samples as precisely as possible, we generated
their standard curves by serially diluting neuroblastoma TICs into
normal PB. The 11 MRD markers (CHRNA3, CRMP1, DBH, DCX, DDC,
GABRB3, GAP43, ISL1, KIF1A, PHOX2B and TH) were amplified by
real-time RT-PCR, and triplicate CT values for each MRD
marker were plotted against the logarithm of the % dilution/100.
Although the standard curve for the real-time RT-PCR assay should
theoretically have R2>0.99, the actual R2
value is lower than expected due to slight expression in normal PB.
In the present study, we generated the standard curve with
R2>0.90 by limiting the plotted points of the %
dilution (Table II). Accordingly,
5 points (100, 20, 4, 0.8 and 0.16% dilution) were plotted against
triplicate CT values for DCX, ISL1 and TH (Fig. 1). For CHRNA3, CRMP1, DBH, DDC,
GABRB3, GAP43, KIF1A and PHOX2B, 6 points (100, 20, 4, 0.8, 0.16
and 0.032% dilution) were used (Fig.
1).
 | Table IINormal range of real-time RT-PCR
markers. |
Table II
Normal range of real-time RT-PCR
markers.
| Gene name | Standard curve | R2 | Quantitative range
(%) | Normal range in
PB/PBSC (%) | Normal range in BM
(%) |
|---|
| CHRNA3 | CT =
−3.69log (% dilution/100) + 22.28 | 0.969 | ≥0.032 | <0.032 | <0.413 |
| CRMP1 | CT =
−3.04log (% dilution/100) + 26.21 | 0.911 | ≥0.032 | <0.032 | <0.207 |
| DBH | CT =
−3.66log (% dilution/100) + 26.60 | 0.946 | ≥0.032 | <0.032 | <0.032 |
| DCX | CT =
−3.02log (% dilution/100) + 27.39 | 0.912 | ≥0.160 | <0.160 | <5.598 |
| DDC | CT =
−3.71log (% dilution/100) + 23.99 | 0.983 | ≥0.032 | <0.032 | <0.051 |
| GABRB3 | CT =
−3.43log (% dilution/100) + 25.00 | 0.948 | ≥0.032 | <0.032 | <1.059 |
| GAP43 | CT =
−3.34log (% dilution/100) + 24.37 | 0.966 | ≥0.032 | <0.032 | <0.694 |
| ISL1 | CT =
−3.71log (% dilution/100) + 25.36 | 0.985 | ≥0.160 | <0.160 | <0.160 |
| KIF1A | CT =
−4.40log (% dilution/100) + 22.43 | 0.962 | ≥0.032 | <0.032 | <0.351 |
| PHOX2B | CT =
−4.19log (% dilution/100) + 23.46 | 0.944 | ≥0.032 | <0.032 | <0.032 |
| TH | CT =
−3.78log (% dilution/100) + 22.13 | 0.987 | ≥0.160 | <0.160 | <0.160 |
Normal range of real-time RT-PCR
markers
Although the quantitative range frequently reflects
the portion of the standard curve in which the MRD levels can be
accurately quantitated, we here defined the quantitative range as
the minimum % dilution that was plotted to the standard curve or
above. While DCX, ISL1 and TH had a quantitative range of ≥0.160%
dilution, the quantitative range of CHRNA3, CRMP1, DBH, DDC,
GABRB3, GAP43, KIF1A and PHOX2B was ≥0.032% dilution (Table II). To account for tumor
heterogeneity, we set the maximum CT value for the
PB/PBSC and BM samples as the mean CT value of the
normal PB and BM cells −3, respectively, according to the threshold
defined by ESG-MRD-ALL (12). The
normal range for each MRD marker in the PB/PBSC and BM samples was
then defined as less than the % dilution that corresponded to the
maximum CT value for the PB/PBSC and BM samples,
respectively. If the mean CT value in normal PB and BM
cells was >40, the normal range was defined as less than the %
dilution that corresponded to the minimum % dilution in the
quantitative range. For the PB samples, the normal ranges of all
MRD markers did not exceed the minimum % dilution in the
quantitative range (Table II). For
the BM samples, the normal ranges of DBH, ISL1, PHOX2B and TH did
not exceed the minimum % dilution in the quantitative range,
whereas the normal ranges of CHRNA3, CRMP1, DCX, DDC, GABRB3, GAP43
and KIF1A were determined to be <0.413, <0.207, <5.598,
<0.051, <1.059, <0.694 and <0.351%, respectively
(Table II).
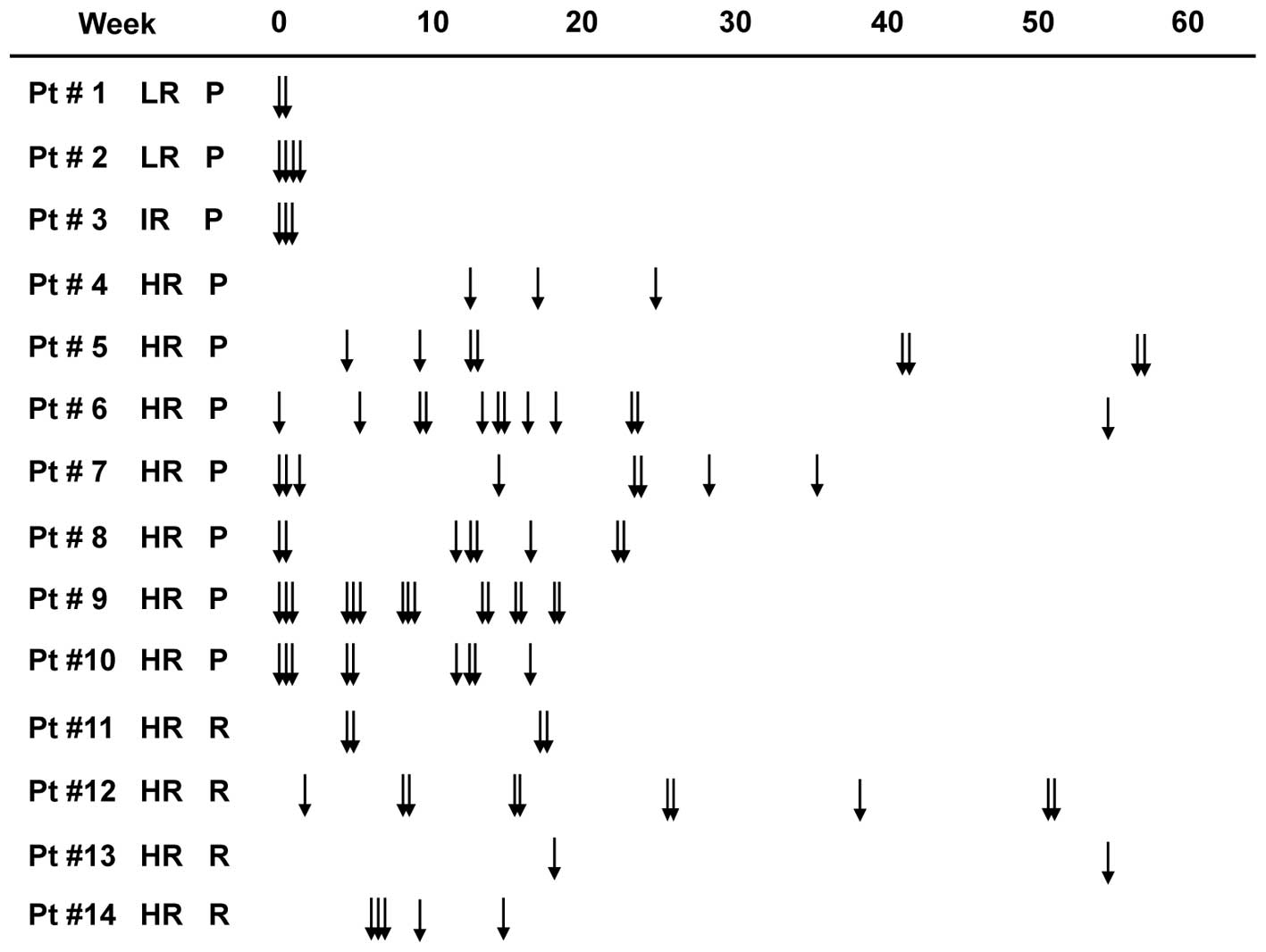
MRD sample characteristics
All 93 MRD samples were obtained from 14
neuroblastoma patients (#1–14) who were prospectively treated at
Kobe Children's Hospital between November, 2010 and August, 2012.
Patients #1–10 developed primary tumors and were treated according
to the Japan Neuroblastoma Study Group (JNBSG) protocol. Low-risk
patient #1 was subjected to surgical therapy followed by watchful
waiting. Low-risk patient #2 was treated with one course of the
LI-A regimen (1.5 mg/m2 VCR and 300 mg/m2
CPA). Intermediate-risk patient #3 was treated with one course of
the LI-D regimen (1.5 mg/m2 VCR, 900 mg/m2
CPA, 30 mg/m2 THP and 60 mg/m2 CDDP).
High-risk patient #4 was treated with five courses of the 05A3
regimen (2,400 mg/m2 CPA, 1.5 mg/m2 VCR, 40
mg/m2 THP and 100 mg/m2 CDDP) followed by
peripheral blood stem cell transplantation (PBSCT), radiation
therapy and surgical therapy. High-risk patient #5 was treated with
five courses of the 05A3 regimen followed by PBSCT and radiation
therapy. High-risk patient #6 was treated with one course of the
05A1 regimen (1,200 mg/m2 CPA, 1.5 mg/m2 VCR,
40 mg/m2 THP and 100 mg/m2 CDDP) and four
courses of the 05A3 regimen followed by PBSCT, radiation therapy
and surgical therapy. High-risk patient #7 was treated with one
course of the 05A1 regimen and four courses of the 05A3 regimen
followed by PBSCT and radiation therapy. High-risk patient #8 was
treated with one course of the 05A1 regimen and four courses of the
05A3 regimen. High-risk patients #9 and 10 were treated with one
course of the 05A1 regimen and three courses of the 05A3 regimen.
Patients #11–14 experienced tumor relapses and were subjected to
salvage chemotherapy. Patient #11 had one course of the IREC
regimen (300 mg/m2 CPT-11, 300 mg/m2 VP-16,
240 mg/m2 CBDCA) and five courses of the topotecan/CPA
regimen (3.75 mg/m2 topotecan, 1,250 mg/m2
CPA). Patient #12 had three courses of the ICE regimen (6,000
mg/m2 IFO, 800 mg/m2 CBDCA, 500
mg/m2 VP-16) and two courses of the CPT-11 regimen (125
mg/m2 CPT-11) followed by bone marrow transplantation
(BMT). Patient #13 had two courses of the CPT-11 regimen, six
courses of IREC regimen, five courses of the topotecan/CPA regimen,
and three courses of the topotecan/TMZ regimen (3.75
mg/m2 topotecan, 750 mg/m2 TMZ). Patient #14
had three courses of the ICE regimen and two courses of the IREC
regimen. All patients were subjected to MRD sampling as frequently
as possible during the course of treatment (Fig. 2).
Expression profile of real-time RT-PCR
markers in MRD samples
CHRNA3, CRMP1, DBH, DCX, DDC, GABRB3, GAP43, ISL1,
KIF1A, PHOX2B and TH expression was determined by real-time RT-PCR
in the 93 MRD (73 BM, 12 PBSC and 8 PB) samples. Each marker was
scored as positive when its expression exceeded the normal range.
The number of positive samples for each marker varied from 4
(GABRB3) to 33 (PHOX2B). PHOX2B and DBH seemed to be sensitive
markers in the present BM samples (Table III). Although only a limited
number of PBSC and PB samples were analyzed, the sensitivity of
each marker also varied between the BM, PBSC and PB samples as
described previously (24). Only
KIF1A, DCX, CRMP1 and GABRB3 were able to detect MRD in the present
PBSC and PB samples (Table
III).
 | Table IIIExpression profile of real-time
RT-PCR markers in MRD samples. |
Table III
Expression profile of real-time
RT-PCR markers in MRD samples.
| BM sample | PBSC sample | PB sample |
|---|
| CHRNA3 |
| Positive | 10 | 0 | 0 |
| Negative | 63 | 12 | 8 |
| CRMP1 |
| Positive | 14 | 0 | 1 |
| Negative | 59 | 12 | 7 |
| DBH |
| Positive | 22 | 0 | 0 |
| Negative | 51 | 12 | 8 |
| DCX |
| Positive | 9 | 0 | 2 |
| Negative | 64 | 12 | 6 |
| DDC |
| Positive | 14 | 0 | 0 |
| Negative | 59 | 12 | 8 |
| GABRB3 |
| Positive | 3 | 0 | 1 |
| Negative | 70 | 12 | 7 |
| GAP43 |
| Positive | 9 | 0 | 0 |
| Negative | 64 | 12 | 8 |
| ISL1 |
| Positive | 14 | 0 | 0 |
| Negative | 55 | 11 | 7 |
| KIF1A |
| Positive | 11 | 2 | 1 |
| Negative | 62 | 10 | 7 |
| PHOX2B |
| Positive | 33 | 0 | 0 |
| Negative | 40 | 12 | 8 |
| TH |
| Positive | 12 | 0 | 0 |
| Negative | 61 | 12 | 8 |
Comparison of the present MRD detection
protocol with clinical examinations
To validate the applicability of a set of these 11
commonly used real-time RT-PCR markers to monitor MRD, we compared
these results with BM cytology, urinary vanillylmandelic acid
(VMA), urinary homovanillic acid (HVA) and serum neuron-specific
enolase (NSE) data. BM cytology data were obtained from 73 MRD (73
BM) samples, whereas urinary VMA, urinary HVA, and serum NSE data
were obtained from 89 MRD (71 BM, 10 PBSC, and 8 PB) samples
(Table IV). MRD was scored as
positive when the expression of one of the 11 markers exceeded the
normal range. Thus, 100, 56, 56 and 57% of BM cytology-positive
(n=10), elevated VMA (>15 μg/mg Cr, n=45), elevated HVA (>30
μg/mg Cr, n=43), and elevated NSE (>20 ng/ml, n=49) samples were
MRD-positive, respectively (Table
IV). MRD was also positive in 48, 45, 46 and 43% of BM
cytology-negative (n=63), normal VMA (≤15 μg/mg Cr, n=44), normal
HVA (≤30 μg/mg Cr, n=46), and normal NSE (≤20 ng/ml, n=40) samples,
respectively (Table IV). As BM
samples were obtained from at least two sites (left and right iliac
crests), we also compared real-time RT-PCR results from the paired
BM samples. Of the 23 pairs of BM cytology-negative samples, 11
pairs were MRD-positive at only one site. The remaining 12 pairs
had identical results for both sites; 6 pairs were MRD-negative and
6 pairs MRD-positive.
 | Table IVComparison of the present MRD
detection protocol with clinical examinations. |
Table IV
Comparison of the present MRD
detection protocol with clinical examinations.
| MRD Positive | MRD Negative |
|---|
| BM cytology |
| Positive | 10 | 0 |
| Negative | 30 | 33 |
| Urinary VMA |
| >15 μg/mg
Cr | 25 | 20 |
| ≤15 μg/mg Cr | 20 | 24 |
| Urinary HVA |
| >30 μg/mg
Cr | 24 | 19 |
| ≤30 μg/mg Cr | 21 | 25 |
| Serum NSE |
| >20 ng/ml | 28 | 21 |
| ≤20 ng/ml | 17 | 23 |
Discussion
In the present study, we validated the ability of 14
commonly used real-time RT-PCR markers to detect neuroblastoma MRD
based on their expression in neuroblastoma TICs. We then developed
a novel MRD detection protocol and prospectively monitored MRD in
73 BM, 12 PBSC and 8 PB samples from 14 neuroblastoma patients
treated at Kobe Children's Hospital between November, 2010 and
August, 2012. Compared to currently used protocols, our protocol
had two advantages. First, it determined 11 real-time RT-PCR
markers based on their expression in neuroblastoma TICs. Second, it
calculated the expression of these markers using the relative
standard curve method and scored their expression levels as %
dilution.
Several sets of real-time RT-PCR markers have been
reported for MRD monitoring in neuroblastoma patients.
Neuroblastoma is characterized by extreme heterogeneity that is
observed in three levels: i) between different patients, ii)
between different tumors within a patient, and iii) between
different cells within a tumor. Regarding the heterogeneity between
different patients, highly variable expression of CHRNA3, DBH, DDC,
GAP43, PHOX2B and TH was detected in tumor samples from 56 patients
(24). In terms of the
heterogeneity between different tumors within a patient, one of the
5 MRD markers (CHRNA3, DDC, GAP43, PHOX2B and TH) showed a
>10-fold difference in relative expression in 6 out of 56 pairs
of BM samples obtained from two different sites (27). Regarding heterogeneity between
different cells within a tumor, Stutterheim et al(27) recently reported that GAP43 and TH
expression differed by >10 CT values across subclones
of the same neuroblastoma IMR-32 cells isolated by serial dilution.
Due to the differential gene expression between whole tumor cells
and TICs (9,25), we validated MRD markers based on
their expression in neuroblastoma TICs instead of whole
neuroblastoma cells used by previous studies. Accordingly,
B4GALNT1, CCND1 and TACC2 were determined to be inappropriate MRD
markers, even though they were selected by their ability to
experimentally define a cut-off value with which to distinguish
neuroblastoma cells from normal cells in previous studies (23,26).
Although most previous studies determined the
expression of MRD markers by real-time RT-PCR using the comparative
CT method, we hypothesized that standard curves
generated by serially diluting neuroblastoma TICs into normal PB
would reflect more accurate MRD marker expression in clinical
samples. Therefore, we calculated MRD marker expression by the
relative standard curve method and scored it as the % dilution.
While previous studies recommended a set of 3, 6 or 8 markers by
which to detect MRD in neuroblastoma patients (22–24),
we used a set of 11 MRD markers (CHRNA3, CRMP1, DBH, DCX, DDC,
GABRB3, GAP43, ISL1, KIF1A, PHOX2B and TH). PHOX2B was the most
sensitive MRD marker in the present BM samples. Among 40
PHOX2B-negative BM samples, only 7 samples were positive for CRMP1,
DBH, DDC, ISL1 and/or TH. Although CRMP1, DCX, GABRB3 and KIF1A
were not as sensitive as PHOX2B in the BM samples, they were able
to detect MRD in the present PB and PBSC samples. Consequently, a
set of these 11 markers was able to detect MRD in more than 40% of
the BM cytology-negative, normal urinary VMA, normal urinary HVA
and normal serum NSE samples analyzed in the present study.
In summary, the present MRD detection protocol based
on the expression of a set of 11 real-time RT-PCR markers (CHRNA3,
CRMP1, DBH, DCX, DDC, GABRB3, GAP43, ISL1, KIF1A, PHOX2B and TH) in
neuroblastoma TICs achieved sensitive MRD monitoring in
neuroblastoma patients who were prospectively treated at a single
center.
Acknowledgements
This study was supported in part by the
Grants-in-Aid for the Scientific Research from the Ministry of
Education, Culture, Sports, Science and Technology of Japan, and
grants from the Showa-hokokai, the Hyogo Prefecture Health
Promotion Association, the Foundation for Promotion of Cancer
Research, and the Hyogo Medical Association.
References
|
1
|
Brodeur GM: Neuroblastoma: biological
insights into a clinical enigma. Nat Rev Cancer. 3:203–216. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Maris JM, Hogarty MD, Bagatell R and Cohn
SL: Neuroblastoma. Lancet. 369:2106–2120. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Maris JM: Recent advances in
neuroblastoma. N Engl J Med. 362:2202–2211. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Jordan CT, Guzman ML and Noble M: Cancer
stem cells. N Engl J Med. 355:1253–1261. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Visvader JE and Lindeman GJ: Cancer stem
cells in solid tumours: accumulating evidence and unresolved
questions. Nat Rev Cancer. 8:755–768. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Vermeulen L, de Sousa e Melo F, Richel DJ
and Medema JP: The developing cancer stem-cell model: clinical
challenges and opportunities. Lancet Oncol. 13:e83–e89. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Müller V, Alix-Panabières C and Pantel K:
Insights into minimal residual disease in cancer patients:
implications for anti-cancer therapies. Eur J Cancer. 46:1189–1197.
2010.PubMed/NCBI
|
|
8
|
Mordant P, Loriot Y, Lahon B, et al:
Minimal residual disease in solid neoplasia: new frontier or
red-herring? Cancer Treat Rev. 38:101–110. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Hansford LM, McKee AE, Zhang L, et al:
Neuroblastoma cells isolated from bone marrow metastases contain a
naturally enriched tumor-initiating cell. Cancer Res.
67:11234–11243. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Coulon A, Flahaut M, Mühlethaler-Mottet A,
et al: Functional sphere profiling reveals the complexity of
neuroblastoma tumor-initiating cell model. Neoplasia. 13:991–1004.
2011.PubMed/NCBI
|
|
11
|
Lin H, Balic M, Zheng S, Datar R and Cote
RJ: Disseminated and circulating tumor cells: role in effective
cancer management. Crit Rev Oncol Hematol. 77:1–11. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
van der Velden VHJ, Cazzaniga G, Schrauder
A, et al: Analysis of minimal residual disease by Ig/TCR gene
rearrangements: guidelines for interpretation of real-time
quantitative PCR data. Leukemia. 21:604–611. 2007.PubMed/NCBI
|
|
13
|
Flohr T, Schrauder A, Cazzaniga G, et al:
Minimal residual disease-directed risk stratification using
real-time quantitative PCR analysis of immunoglobulin and T-cell
receptor gene rearrangements in the international multicenter trial
AIEOP-BFM ALL 2000 for childhood acute lymphoblastic leukemia.
Leukemia. 22:771–782. 2008. View Article : Google Scholar
|
|
14
|
Kågedal B: Detecting minimal residual
disease in neuroblastoma: still a ways to go. Clin Chem.
55:1268–1270. 2009.PubMed/NCBI
|
|
15
|
Stutterheim J, Zappeij-Kannegieter L,
Versteeg R, Caron HN, van der Schoot CE and Tytgat GAM: The
prognostic value of fast molecular response of marrow disease in
patients aged over 1 year with stage 4 neuroblastoma. Eur J Cancer.
47:1193–1202. 2011.PubMed/NCBI
|
|
16
|
Yáñez Y, Grau E, Oltra S, et al: Minimal
disease detection in peripheral blood and bone marrow from patients
with non-metastatic neuroblastoma. J Cancer Res Clin Oncol.
137:1263–1272. 2011.PubMed/NCBI
|
|
17
|
Burchill SA, Bradbury FM, Smith B, Lewis
IJ and Selby P: Neuroblastoma cell detection by reverse
transcriptase-polymerase chain reaction (RT-PCR) for tyrosine
hydroxylase mRNA. Int J Cancer. 57:671–675. 1994. View Article : Google Scholar
|
|
18
|
Miyajima Y, Kato K, Numata SI, Kudo K and
Horibe K: Detection of neuroblastoma cells in bone marrow and
peripheral blood at diagnosis by the reverse
transcriptase-polymerase chain reaction for tyrosine hydroxylase
mRNA. Cancer. 75:2757–2761. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kuçi Z, Seitz G, Kuçi S, et al: Pitfalls
in detection of contaminating neuroblastoma cells by tyrosine
hydroxylase RT-PCR due to catecholamine-producing hematopoietic
cells. Anticancer Res. 26:2075–2080. 2006.PubMed/NCBI
|
|
20
|
Viprey VF, Corrias MV, Kågedal B, et al:
Standardisation of operating procedures for the detection of
minimal disease by QRT-PCR in children with neuroblastoma: quality
assurance on behalf of SIOPEN-R-NET. Eur J Cancer. 43:341–350.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Stutterheim J, Gerritsen A,
Zappeij-Kannegieter L, et al: PHOX2B is a novel and specific marker
for minimal residual disease testing in neuroblastoma. J Clin
Oncol. 26:5443–5449. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Viprey VF, Lastowska MA, Corrias MV,
Swerts K, Jackson MS and Burchill SA: Minimal disease monitoring by
QRT-PCR: guidelines for identification and systematic validation of
molecular markers prior to evaluation in prospective clinical
trials. J Pathol. 216:245–252. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Cheung IY, Feng Y, Gerald W and Cheung NK:
Exploiting gene expression profiling to identify novel minimal
residual disease markers of neuroblastoma. Clin Cancer Res.
14:7020–7027. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Stutterheim J, Gerritsen A,
Zappeij-Kannegieter L, et al: Detecting minimal residual disease in
neuroblastoma: the superiority of a panel of real-time quantitative
PCR markers. Clin Chem. 55:1316–1326. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Nishimura N, Hartomo TB, Pham TVH, et al:
Epigallocatechin gallate inhibits sphere formation of neuroblastoma
BE(2)-C cells. Environ Health Prev Med. 17:246–251. 2012.
View Article : Google Scholar
|
|
26
|
Cheung IY and Cheung NK: Quantitation of
marrow disease in neuroblastoma by real-time reverse
transcription-PCR. Clin Cancer Res. 7:1698–1705. 2001.PubMed/NCBI
|
|
27
|
Stutterheim J, Zappeij-Kannegieter L, Ora
I, et al: Stability of PCR targets for monitoring minimal residual
disease in neuroblastoma. J Mol Diagn. 14:168–175. 2012. View Article : Google Scholar : PubMed/NCBI
|