Introduction
Glioblastoma is the most common primary central
nervous system malignancy with features of localization in the
brain, highly invasive behavior and extremely poor prognosis. The
effects of current conventional therapeutic means for glioblastoma
are not satisfactory (1). To
improve the life expectancy and the quality of life of the
patients, it is extremely critical to search for novel diagnostic
and therapeutic targets. New hope for the treatment of human
glioblastoma include microRNAs (miRNAs, miRs) that have recently
emerged as key regulators of cancers (2). miRNAs, a class of small, non-coding,
single-stranded RNAs, can post-transcriptionally regulate protein
expression by targeting the 3′-untranslated region (3′-UTR) of
target mRNAs, which cause either degradation or repression of
translation. miRNAs are involved in multiple biological activities
as well as progression of diseases including cancer (3). Many studies have shown that
dysregulation of miRNAs plays an important role in the initiation
and malignant progression of cancer (4–6).
miRNAs can act as either tumor suppressors or
oncogenes depending on the functions of their targets. It has been
reported that miR-223 functions as a tumor suppressor in certain
types of cancers, for example, in the inhibition of osteosarcoma by
targeting Hsp90B1 (7). miR-223 is
weakly expressed in colorectal cancer, cervical cancer and hepatoma
cells; and overexpression of miR-223 can suppress proliferation of
the three types of tumor cells by targeting FOXO1 (8). Notably, miR-223 is high expressed in
gastric cancer and promotes cellular proliferation and invasion by
targeting EPB41L3 (9) or
FBXW7/hCdc4 (10), functioning as
an oncogene. In addition, miR-223 can target Stathmin1 which plays
an oncogenic role in gastric cancer (11). Thus, the role of miR-223 may be
opposing in different types of cancer or tumor cell lines via the
targeting of different functional genes. Current studies have
revealed that the growth and metastasis of glioblastoma are
associated with various miRNAs (12–17).
However, whether miR-223 serves as a tumor suppressor or an
oncogene in glioblastoma remains unclear at present.
The paired box 6 (PAX6) encodes a transcription
factor that is involved in the development of the brain, where its
expression persists. It can suppress cell growth and invasion of
glioblastoma cells, and has currently been demonstrated to function
as a tumor suppressor in glioblastoma (18,19).
Yet, reports related to miRNAs of PAX6 are rare (20). Particularly, whether or not miR-223
targets PAX6 in glioblastoma requires elucidation.
Therefore, in the present study, we determined the
expression levels of miR-223 and PAX6 in glioblastoma cells or
tissues. Then, the regulation of miR-223 on PAX6 expression and
their effects on proliferation, cell cycle and invasion of
glioblastoma U251 and U373 cells were investigated. It was
confirmed that PAX6 is a target of miR-223 in glioblastoma. Our
results may support miR-223 and/or PAX6 as novel diagnostic or
therapeutic targets for glioblastoma.
Materials and methods
Cell culture
Glioblastoma A172, U251, U373 and U138 cell lines,
and the human fetal glial HFGC cell line were purchased from the
China Center for Type Culture Collection (CCTCC, Wuhan, China) and
maintained in RPMI-1640 medium (HyClone) containing 10% fetal
bovine serum (Hangzhou Sijiqing) in a humidified atmosphere of 5%
CO2 at 37°C.
Recombinant plasmid construction
To generate the PAX6-overexpression plasmid, PAX6
CDS (NM_001258462) was amplified by PCR using the following
primers: sense, 5′-GGATCCATGCAGAACAGTCACAGC-3′ and antisense,
5′-GAATTCTTACTGTAATCTTGGCCAGT-3′, then the sequence was cloned into
the BamHI/EcoRI sites of the pcDNA3.1. The
recombinant plasmid was termed pcDNA3.1-PAX6.
Transfection
For the miR-223 functional analysis, cells were
transfected with the miR-223 scrambled miRNA control (negative
control, NC), miR-223 mimics, or the miR-223 inhibitor (Ambion). To
generate gain-of-function models of PAX6, cells were transfected
with PAX6-pcDNA3.1 or its negative control (pcDNA3.1 vector),
respectively. Transfection was performed using Lipofectamine 2000
(Invitrogen) according to the manufacturer’s recommendations.
Lentivirus package and infection
The lentivirus system (Neuron, Shanghai, China)
consisted of the pLKD-CMV-G&NR-U6-shRNA vector together with
the plasmids containing the imperative elements for virus
packaging. Short hairpin RNA (shRNA) targeting PAX6
(5′-ATGGGCGGAGTTATGATACCTAC-3′) and the scrambled control
(5′-AATTCTCCGAACGTGTCACGT-3′) were coloned into the vector,
respectively, which were then cotransfected with packaging plasmids
into 293T cells to generate individual lentiviruses. After 48 h,
the PAX6-RNAi-lentivirus and control lentivirus were harvested and
purified. The lentivirus expressing PAX6 shRNA and the control
lentivirus were termed Lv-PAX6-shRNA and Lv-Ctr, respectively. U251
and U373 cells were plated in 6-well plates to reach 40–50%
confluency. Appropriately titered lentivirus was added at a MOI of
20 and incubated at 37°C.
Real-time RT-PCR
Total RNA was extracted from the cells with TRIzol
reagent (Invitrogen) following the manufacturer’s instructions. The
relative expression level of miR-124 was determined by quantitative
real-time RT-PCR using mirVana™ qRT-PCR microRNA detection kit
(Ambion) following the manufacturer’s instructions. Specific primer
sets for miR-223 and U6 (used as an internal reference) were
obtained from Ambion. Expression of PAX6 mRNA was detected by
real-time RT-PCR using the standard SYBR-Green RT-PCR kit (Takara,
Otsu, Japan) following the manufacturer’s instructions. The
specific primer pairs were as follows: PAX6 (157 bp) sense,
5′-AGACACAGCCCTCACAAAC-3′ and antisense, 5′-ATCATAACTCCGCCCATTC-3′;
β-actin (202 bp) as an internal control, sense,
5′-AGGGGCCGGACTCGTCATACT-3′ and antisense,
5′-GGCGGCACCACCATGTACCCT-3′. The relative expression of PAX6 mRNA
or miR-223 was quantified using the GraphPad Prism 4.0 software
(GraphPad Software, San Diego, CA, USA) and the 2−ΔΔCt
method (21).
Immunohistochemical analysis
A human brain gliocytoma and normal tissue
microarray (Auragene Bioscience Co., Changsha, China) containing 5
cases of normal tissue, 5 cases of T1N0M0 glioblastoma, 4 cases of
T2N0M0 glioblastoma and other subtypes of brain gliocytoma (data
not shown), duplicate cores per case, was processed for
immunohistochemical analysis using anti-PAX6 antibody (Millipore)
as previously described (22).
Images were analyzed using Leica Q550 image analysis system (Leica,
Wetzlar, Germany). The mean optical density value (D) and area (A)
of brown particles in three visual fields of each section were
calculated by the system. The expression levels of PAX6 in tissues
were evaluated using the formula: integral density = D × A.
Dual luciferase reporter assay
The 3′-UTR of PAX6 (NM_000280) containing the
miRNA-223 binding sites and its corresponding mutated sequence were
cloned into the psi-CHECK2 luciferase reporter vector (Promega)
downstream of Renilla luciferase, named PAX6
3′-UTR-psi-CHECK2 and PAX6 Mut 3′-UTR-psi-CHECK2, respectively.
Using Lipofectamine 2000 (Invitrogen), U251 and U373 cells were
co-transfected with the reporter constructs and miR-223 mimics, the
miR-223 inhibitor, negative control (NC) or the negative control
inhibitor. Luciferase activity was determined after 48 h using the
Dual-Glo substrate system (Promega) and a Beckman Coulter LD400
luminometer. Data are presented as the ratio of experimental
(Renilla) luciferase to control (Firefly) luciferase.
Cell proliferation assay
For analysis of the functions of miR-223, U251 and
U373 cells were transfected with miR-223 mimics or its inhibitor,
respectively. For analysis of PAX6 functions, U251 and U373 cells
were either transfected with pcDNA3.1-PAX6 or infected with
Lv-PAX6-shRNA. Cells in exponential growth were plated at a final
concentration of 2×103 cells/well in 96-well plates. The
viability of the cells was evaluated by MTT assay after 24, 48, 72
and 96 h of seeding. The optical density at 570 nm (OD570) of each
well was measured with an ELISA reader (ELX-800 type, Bio-Tek).
Flow cytometric (FCM) analysis
The cells were digested and collected after 48 h
post-transfection or 96 h post-infection, and washed with PBS
twice. The cells were resuspended in PBS and then fixed in 70%
ethanol at 4°C for 18 h. The cells were washed with PBS and
resuspended in staining solution (50 μg/ml of propidium iodide, 1
mg/ml of RNase A, 0.1% Triton X-100 in PBS). The stained cells
(1×105) were then analyzed with a flow cytometer
(Beckman Coulter).
Cell invasion assay
The cell invasion assay was performed using a Cell
Invasion Assay kit (Chemicon International, Temecula, CA, USA)
according to the manufacturer’s guidelines. Briefly, U251 or U373
cells were placed in the upper compartment of the chambers, and
RPMI-1640 containing 10% fetal bovine serum was added to the lower
chambers. After 24 h of incubation at 37°C, cells on the upper face
of the membrane were scraped using a cotton swab and cells on the
lower face were fixed, stained and observed under a microscope. The
dye on the membrane was then dissolved with 10% acetic acid,
dispensed into 96-well plates (150 μl/well), and the optical
density at 570 nm (OD570) of each well was measured with an ELISA
reader (ELX-800 type; Bio-Tek).
Western blotting
Cells were lysed in cell lysate, and then
centrifuged at 12,000 × g for 20 min at 4°C. The supernatant was
collected and denatured. Proteins were separated using 10% SDS-PAGE
and blotted onto a polyvinylidene difluoride membrane (PVDF). The
PVDF membrane was treated with TBST containing 50 g/l skimmed milk
at room temperature for 4 h, followed by incubation with the
primary antibodies anti-PAX6, anti-MMP-2, anti-MMP-9, anti-VEGFA
(Abcam) and anti-β-actin (Abzoom, Dallas, TX, USA), respectively,
at 37°C for 1 h. Membranes were rinsed and incubated for 1 h with
the corresponding peroxidase-conjugated secondary antibodies.
Chemiluminent detection was performed with the ECL kit (Pierce
Chemical, Rockford, IL, USA). The amount of the protein of
interest, expressed as arbitrary densitometric units, was
normalized to the densitometric units of β-actin.
Statistical analysis
Data are expressed as the means ± SD from at least
three separate experiments. Statistical analysis was carried out
using SPSS 15.0 software. The difference between two groups was
analyzed by the Student’s t-test. A value of P<0.05 was
considered to indicate a statistically significant result.
Results
Expression of miR-223 and PAX6 in
glioblastoma
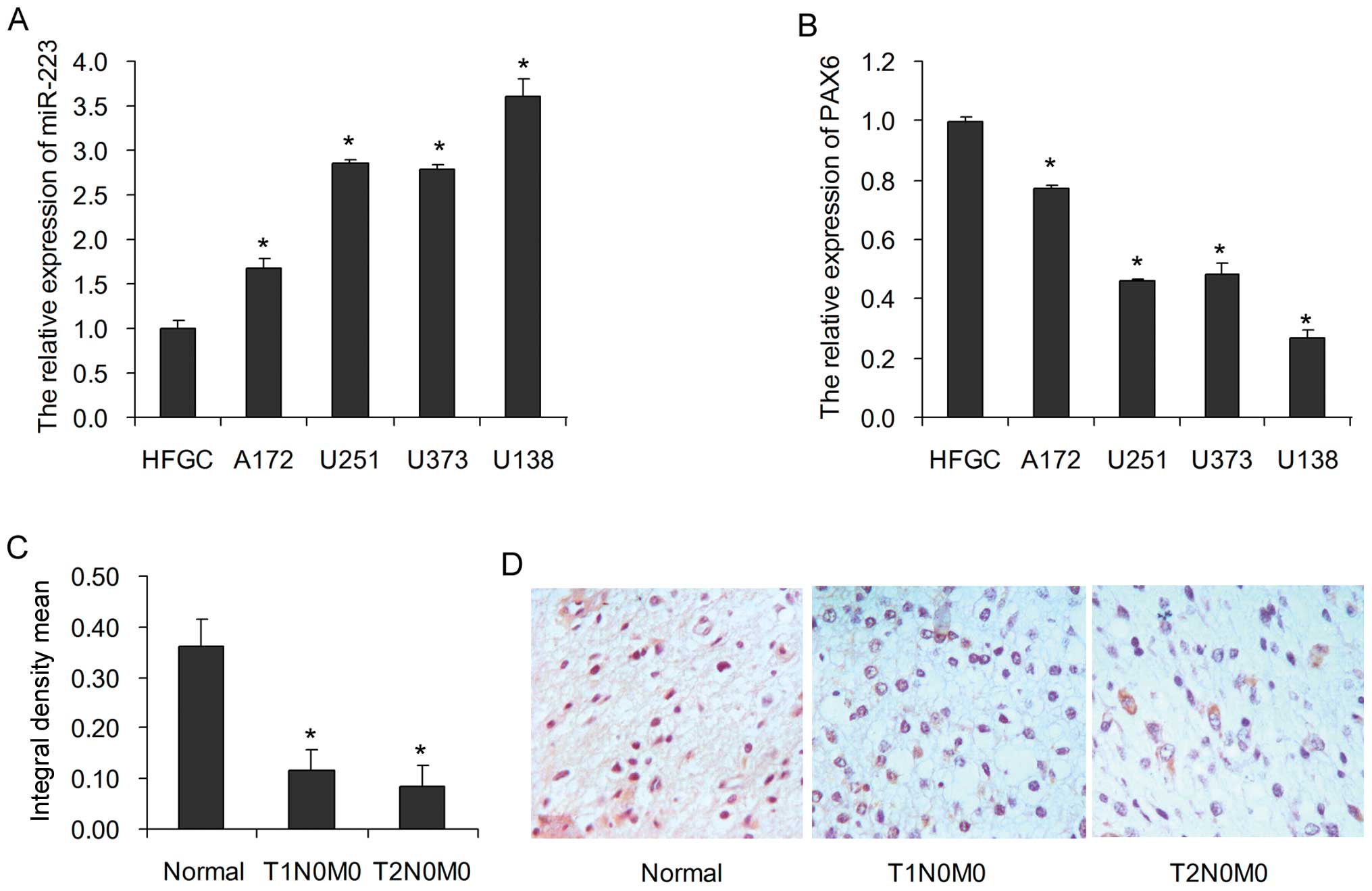
The expression levels of miR-223 and PAX6 in human
glioblastoma A172, U251, U373 and U138 cells were determined using
quantitative RT-PCR (qRT-PCR). Compared to the human fetal glial
HFGC cell line, the miR-223 expression level was apparently
increased in the glioblastoma cells (Fig. 1A); however, the PAX6 expression
level was markedly decreased (Fig.
1B). Moreover, we found that the PAX6 protein was weakly
expressed in the glioblastoma tissues (Fig. 1C and D), using the glioblastoma
tissue microarrays. These results suggest that miR-223 and PAX6 are
potential regulatory molecules in the progression of glioblastoma,
and PAX6 is a potential diagnostic biomarker for glioblastoma.
PAX6 is a direct target of miR-223
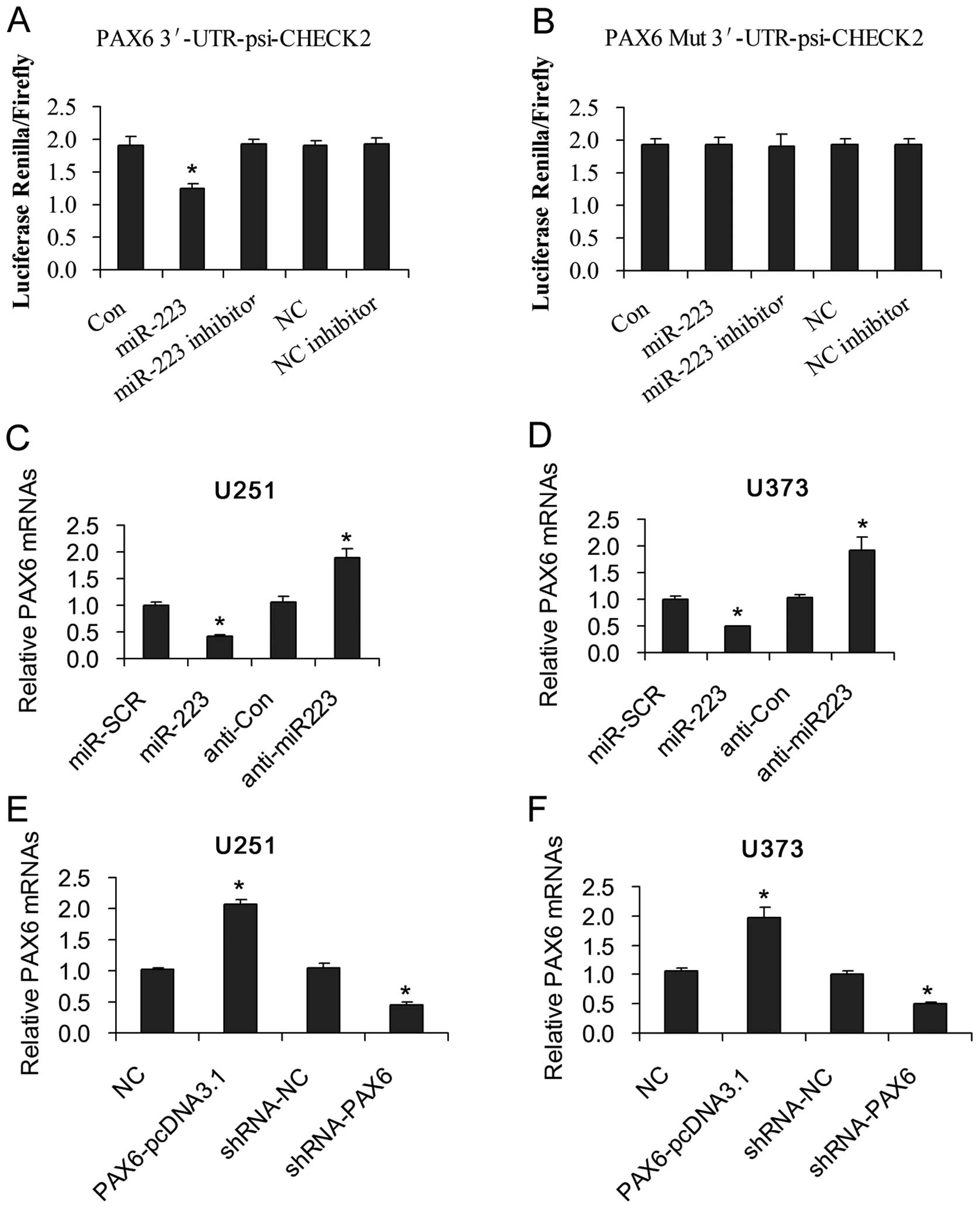
To confirm whether PAX6 is a direct target of
miR-223, we subcloned the PAX6 3′-UTR fragment containing the
miR-223 binding site and mutated targeting sequence, and cloned
them into psi-CHECK2 dual luciferase reporter vectors. The miR-223
mimics significantly inhibited the luciferase activity in the
HEK293T cells transfected with the PAX6 3′-UTR-psi-CHECK2 reporter
vector (Fig. 2A). However, miR-223
mimics did not suppress the luciferase activity levels in the
HEK293T cells transfected with the PAX6 Mut 3′-UTR-psi-CHECK2
reporter vector (Fig. 2B). Thus,
PAX6 was confirmed as a direct target of miR-223.
To further investigate the regulation of PAX6 by
miR-223 in glioblastoma cells, the expression levels of PAX6 mRNA
in the U251 and U373 cells transfected with miR-223 mimics or the
miR-223 inhibitor (anti-miR-223) were detected. The results showed
that the introduction of miR-223 mimics to glioblastoma U251 and
U373 cells decreased PAX6 mRNA expression. Moreover, the miR-223
inhibitor increased the PAX6 mRNA expression levels (Fig. 2C and D). This indicates that miR-223
functions as a suppressor of PAX6 by targeting PAX6 in glioblastoma
cells.
PAX6 inhibits the growth and invasion of
glioblastoma cells
To investigate the roles of PAX6 in the glioblastoma
cells, PAX6 was overexpressed or knocked down in the U251 and U373
cells by transfection of the PAX6-pcDNA3.1 construct or infection
of Lv-shRNA-PAX6, respectively (Fig. 2E
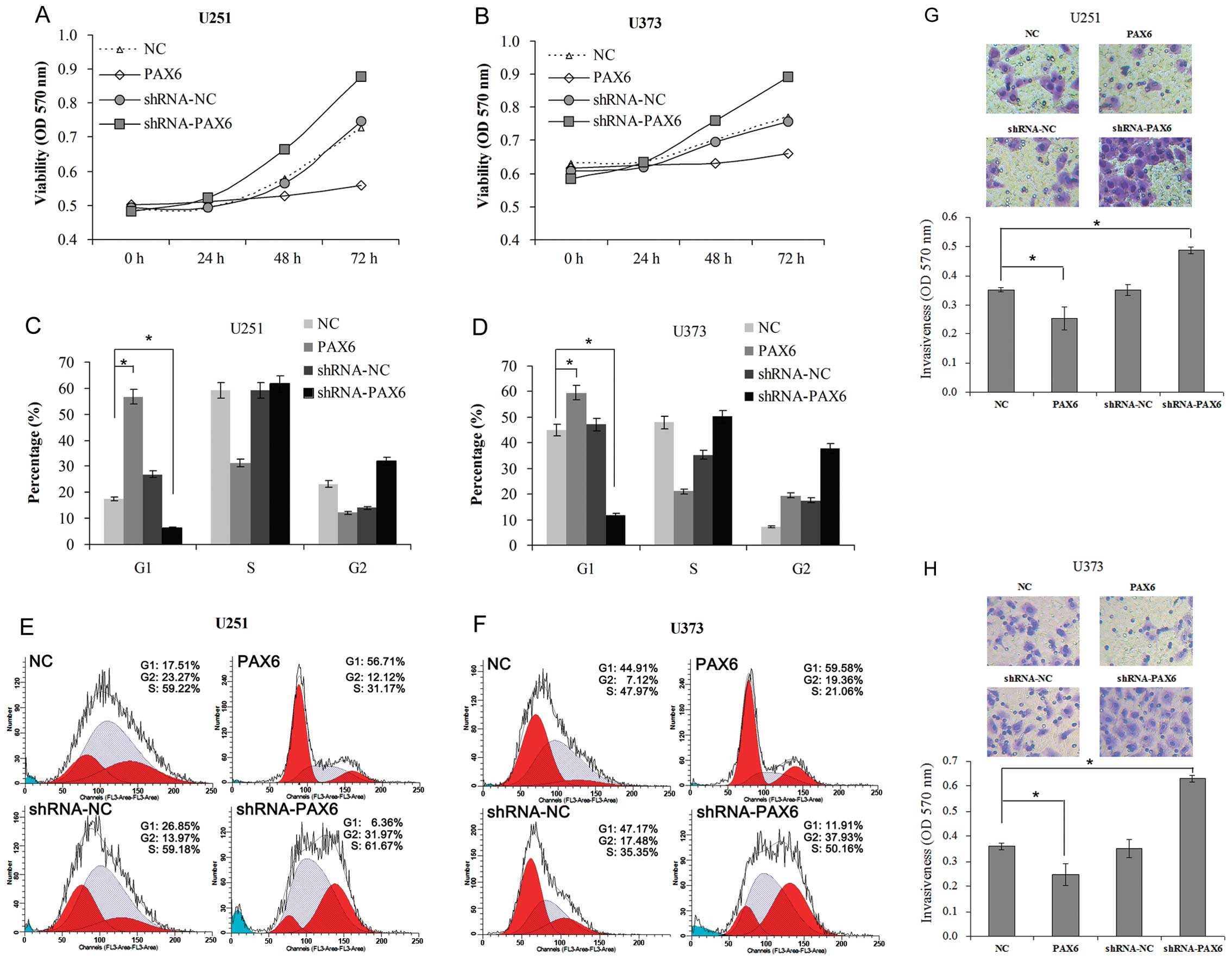
and F). The data showed that overexpression of PAX6 decreased
the cell viability (Fig. 3A and B),
the percentage of cells in the G1 phase (Fig. 3C–F) and cell invasiveness (Fig. 3G and H) of the U251 and U373 cells;
and knockdown of PAX6 caused a reverse result. This suggests that
PAX6, which serves as a tumor suppressor, plays an important role
in the control of growth and metastasis of glioblastoma.
miR-223 promotes the growth and invasion
of glioblastoma cells
Based on the above data, we hypothesized that
miR-223 is also able to affect the growth and invasion of
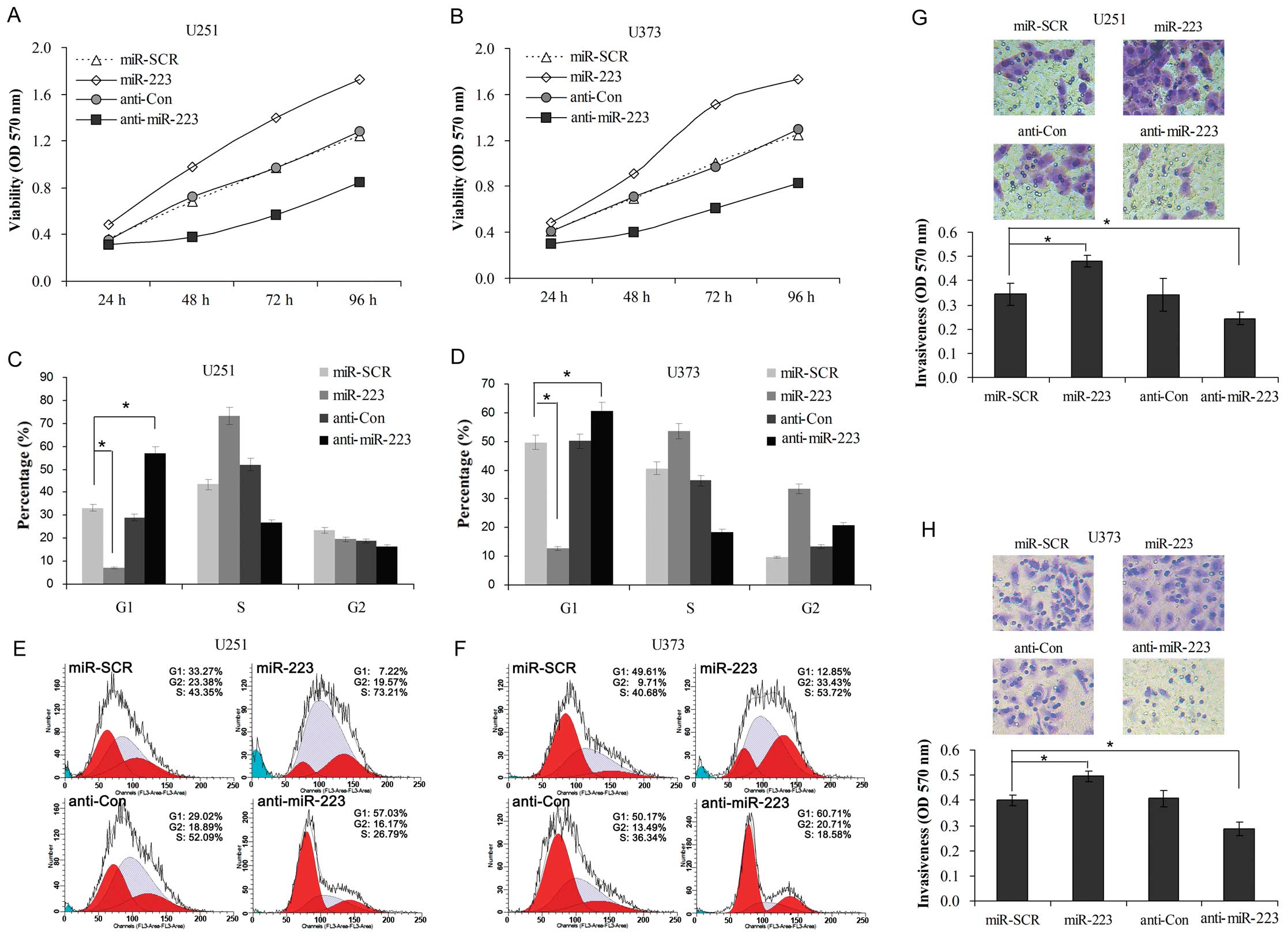
glioblastoma cells by regulation of PAX6 expression. Our results
showed that introduction of miR-223 mimics to U251 and U373 cells
increased their cell viability (Fig. 4A
and B), the percentage of cells in the G1 phase (Fig. 4C–F) and cell invasiveness (Fig. 4G and H); however, introduction of
the miR-223 inhibitor (anti-miR-223) to both cell lines decreased
their cell viability, the percentage of cells in the G1 phase and
cell invasiveness. This suggests that miR-223, which functions as
an oncogene, plays a role in the regulation of growth and
metastasis of glioblastoma; and its underlying mechanisms may
include its targeted regulation of PAX6 expression.
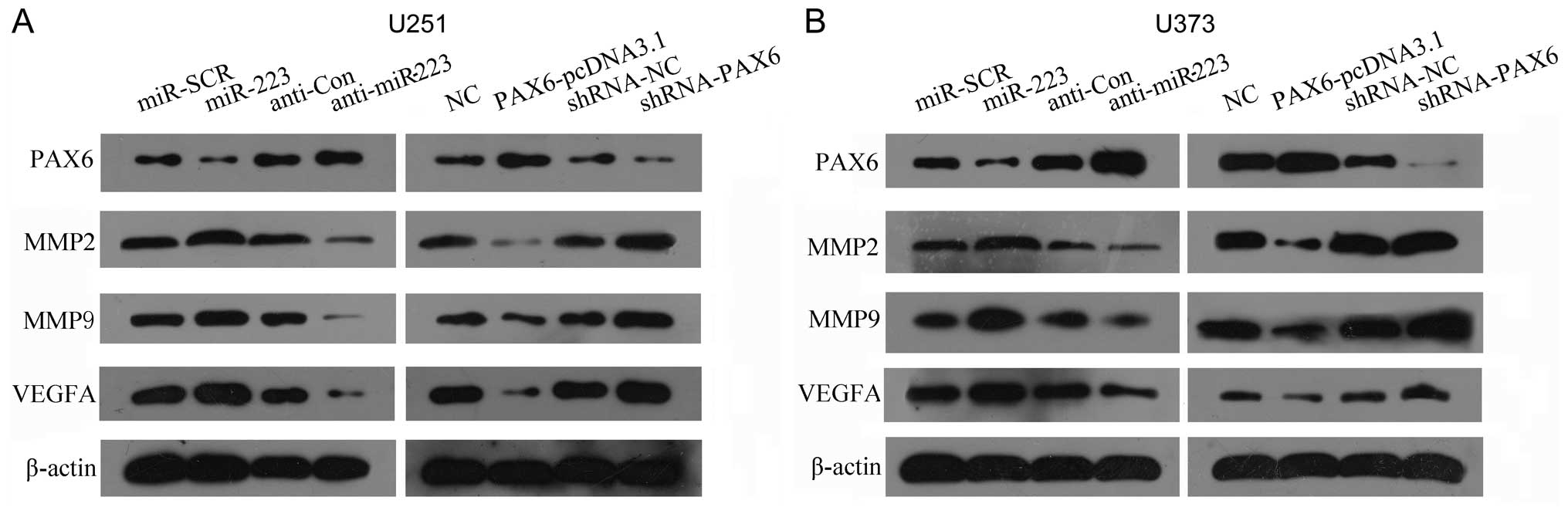
Effects of PAX6 and miR-223 on MMP2, MMP9
and VEGFA expression
Western blot analysis showed that overexpression of
PAX6 resulted in the downregulation of MMP2, MMP9 and VEGFA, and
knockdown of PAX6 caused upregulation of MMP2, MMP9 and VEGFA in
glioblastoma U251 (Fig. 5A) and
U373 (Fig. 5B) cells.
Correspondingly, introduction of miR-223 mimics to glioblastoma
U251 and U373 cells led to effects similar to those following
knockdown of PAX6 in both cell lines: enhanced MMP2, MMP9 and VEGFA
expression. Introduction of the miR-223 inhibitor resulted in
effects similar to those following overexpression of PAX6 in both
cell lines: decreased MMP2, MMP9 and VEGFA expression. This
demonstrates that the miR-223-mediated process of the promotion of
growth and metastasis of glioblastoma possibly occur through
targeting PAX6 and regulating MMP2, MMP9 and VEGFA expression.
Discussion
Cancer initiation and progression are associated
with abnormal expression of various genes or miRNAs. Identification
of dysregulated miRNAs and their target genes is important for
elucidating molecular mechanisms of cancer pathology and
discovering novel diagnostic and therapeutic targets. Studies
indicate that miR-223 functions as an oncogene or a tumor
suppressor in cancer related to the cancer type (7–11). Our
results showed that the miR-223 expression level in the human
glioblastoma A172, U251, U373 and U138 cells was higher than that
in human fetal glial HFGC cells, indicating that miR-223 is
increased in glioblastoma. It has been reported that PAX6 regulates
growth, invasion and angiogenesis in multiple types of cancer
including glioblastoma (15,18,19,23).
We found that the PAX6 expression level was decreased in the
glioblastoma A172, U251, U373 and U138 cells and tumor tissues from
patients with glioblastoma. This demonstrated that increased
miR-223 expression and reduced PAX6 expression coexist in
glioblastoma, implying a potential correlation between them.
We next confirmed that miR-223 directly targets PAX6
through binding to its 3′-UTR using a dual luciferase reporter
assay. In addition, post-transcriptional regulation of PAX6
expression by miR-223 was observed in glioblastoma U251 and U373
cells. In the glioblastoma cells, upregulation of miR-223 decreased
PAX6 mRNA and protein expression; however, inhibition of miR-223
increased PAX6 mRNA and protein expression. This suggests that PAX6
is a direct target of miR-223 in glioblastoma. To investigate their
functional correlations, we analyzed the effects of PAX6 and
miR-223 on the growth, invasion and the related signaling molecules
of glioblastoma cells. In glioblastoma U251 and U373 cells,
overexpression of miR-223 led to effects similar to those following
knockdown of PAX6: increased cell viability, percentage of cells in
the G1 phase and cell invasiveness parallel to increased MMP2, MMP9
and VEGFA expression. Moreover, inhibition of miR-223 resulted in
effects similar to those following overexpression of PAX6:
decreased cell viability, percentage of cells in the G1 phase and
cell invasiveness parallel to reduced MMP2, MMP9 and VEGFA
expression. These results indicate that miR-223 functions as an
oncogene in glioblastoma and the underlying mechanisms include its
targeting of PAX6, which serves as a tumor suppressor in
glioblastoma.
It is worth noting that PAX6 is also considered an
oncogene promoting proliferation and tumorigenesis in some types of
cancers such as human retinoblastoma (24) and in breast cancer cells (25). However, in the present study, we
demonstrated that PAX6 functions as a tumor suppressor in
glioblastoma cells, and our conclusion is in accordance with the
results obtained by Mayes et al(18) and Chang et al(26). PAX6, as a transcription factor,
suppresses cell growth by inhibition of G1/S transition (19) and decreases invasiveness by
inhibiting the expression of MMP2, MMP9 and VEGFA possibly through
its direct binding to their promoters (18,23,27).
Cell migration and invasion are initial steps in cancer metastasis.
PAX6 downregulation facilitates enhancement of MMP2 and MMP9
expression, promoting glioblastoma cell enzymatic degradation of
the extracellular matrix and increase in its invasive phenotype
(28). Vascular endothelial growth
factors (VEGF) play important roles in angiogenesis. VEGFA is
frequently overexpressed in cancers and its overexpression promotes
cancer vasculogenesis, growth and metastasis (29,30).
Therefore, PAX6 inhibits the growth and invasion of glioblastoma by
suppression of its cell cycle transition and the expression of
MMP2, MMP9 and VEGFA. Correspondingly, miR-223 promotes
proliferation, cell cycle transition and invasion of glioblastoma
cells possibly by downregulation of PAX6 expression.
Although our data confirm that miR-223 functions as
an oncogenic miRNA in glioblastoma cells by its targeting of PAX6,
whether PAX6 is regulated by other miRNAs [for example, miR-328
(20) or miR-7a (31)] or whether miR-223 also targets other
downstream genes [for example, EPB41L3 (9) or FBXW7/hCdc4 (10)] in glioblastoma warrants further
investigation.
In brief, PAX6 is weakly expressed in glioblastoma
parallel to increased miR-223 expression, and is a target of
miR-223. miR-223 can promote growth and invasion of glioblastoma
U251 and U373 cells by targeting PAX6, which serves as a tumor
suppressor in glioblastoma exerting the functions of inhibition of
cell cycle progression, and the expression of MMP2, MMP9 and VEGFA.
Collectively, the present study suggests that miR-223 and PAX6 are
novel therapeutic targets for glioblastoma.
Acknowledgements
The present study was supported by the National
Natural Science Foundation of China (30770838).
References
|
1
|
Preusser M, de Ribaupierre S, Wohrer A, et
al: Current concepts and management of glioblastoma. Ann Neurol.
70:9–21. 2011. View Article : Google Scholar
|
|
2
|
Qu S, Yao Y, Shang C, et al: MicroRNA-330
is an oncogenic factor in glioblastoma cells by regulating SH3GL2
gene. PloS One. 7:e460102012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lages E, Ipas H, Guttin A, Nesr H, Berger
F and Issartel JP: MicroRNAs: molecular features and role in
cancer. Front Biosci. 17:2508–2540. 2012. View Article : Google Scholar
|
|
4
|
Chang HT, Li SC, Ho MR, et al:
Comprehensive analysis of microRNAs in breast cancer. BMC Genomics.
13(Suppl 7): S182012.PubMed/NCBI
|
|
5
|
Chen PS, Su JL and Hung MC: Dysregulation
of microRNAs in cancer. J Biomed Sci. 19:902012. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Liu H: MicroRNAs in breast cancer
initiation and progression. Cell Mol Life Sci. 69:3587–3599. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Li G, Cai M, Fu D, et al: Heat shock
protein 90B1 plays an oncogenic role and is a target of
microRNA-223 in human osteosarcoma. Cell Physiol Biochem.
30:1481–1490. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wu L, Li H, Jia CY, et al: MicroRNA-223
regulates FOXO1 expression and cell proliferation. FEBS Lett.
586:1038–1043. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Li X, Zhang Y, Zhang H, et al: miRNA-223
promotes gastric cancer invasion and metastasis by targeting tumor
suppressor EPB41L3. Mol Cancer Res. 9:824–833. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li J, Guo Y, Liang X, et al: MicroRNA-223
functions as an oncogene in human gastric cancer by targeting
FBXW7/hCdc4. J Cancer Res Clin Oncol. 138:763–774. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kang W, Tong JH, Chan AW, et al: Stathmin1
plays oncogenic role and is a target of microRNA-223 in gastric
cancer. PloS One. 7:e339192012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhao S, Deng Y, Liu Y, et al: MicroRNA-153
is tumor suppressive in glioblastoma stem cells. Mol Biol Rep.
40:2789–2798. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhang QQ, Xu H, Huang MB, et al:
MicroRNA-195 plays a tumor-suppressor role in human glioblastoma
cells by targeting signaling pathways involved in cellular
proliferation and invasion. Neuro Oncol. 14:278–287. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yue X, Wang P, Xu J, et al: MicroRNA-205
functions as a tumor suppressor in human glioblastoma cells by
targeting VEGF-A. Oncol Rep. 27:1200–1206. 2012.PubMed/NCBI
|
|
15
|
Lin J, Teo S, Lam DH, Jeyaseelan K and
Wang S: MicroRNA-10b pleiotropically regulates invasion,
angiogenicity and apoptosis of tumor cells resembling mesenchymal
subtype of glioblastoma multiforme. Cell Death Dis. 3:e3982012.
View Article : Google Scholar
|
|
16
|
Fang X, Yoon JG, Li L, et al: The SOX2
response program in glioblastoma multiforme: an integrated
ChIP-seq, expression microarray, and microRNA analysis. BMC
Genomics. 12:112011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Li Y, Guessous F, Zhang Y, et al:
MicroRNA-34a inhibits glioblastoma growth by targeting multiple
oncogenes. Cancer Res. 69:7569–7576. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Mayes DA, Hu Y, Teng Y, et al: PAX6
suppresses the invasiveness of glioblastoma cells and the
expression of the matrix metalloproteinase-2 gene. Cancer Res.
66:9809–9817. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhou YH, Wu X, Tan F, et al: PAX6
suppresses growth of human glioblastoma cells. J Neurooncol.
71:223–229. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen KC, Hsi E, Hu CY, Chou WW, Liang CL
and Juo SH: MicroRNA-328 may influence myopia development by
mediating the PAX6 gene. Invest Ophthalmol Vis Sci. 53:2732–2739.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Arocho A, Chen B, Ladanyi M and Pan Q:
Validation of the 2-DeltaDeltaCt calculation as an alternate method
of data analysis for quantitative PCR of BCR-ABL P210 transcripts.
Diagn Mol Pathol. 15:56–61. 2006.PubMed/NCBI
|
|
22
|
Jiang X, Yue J, Lu H, et al: Inhibition of
filamin-A reduces cancer metastatic potential. Int J Biol Sci.
9:67–77. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhou YH, Hu Y, Mayes D, et al: PAX6
suppression of glioma angiogenesis and the expression of vascular
endothelial growth factor A. J Neurooncol. 96:191–200. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Li L, Li B, Zhang H, et al: Lentiviral
vector-mediated PAX6 overexpression promotes growth and inhibits
apoptosis of human retinoblastoma cells. Invest Ophthalmol Vis Sci.
52:8393–8400. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zong X, Yang H, Yu Y, et al: Possible role
of Pax-6 in promoting breast cancer cell proliferation and
tumorigenesis. BMB Rep. 44:595–600. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Chang JY, Hu Y, Siegel E, Stanley L and
Zhou YH: PAX6 increases glioma cell susceptibility to detachment
and oxidative stress. J Neurooncol. 84:9–19. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Sivak JM, West-Mays JA, Yee A, Williams T
and Fini ME: Transcription factors Pax6 and AP-2α interact to
coordinate corneal epithelial repair by controlling expression of
matrix metalloproteinase gelatinase B. Mol Cell Biol. 24:245–257.
2004.
|
|
28
|
Zhang W, Fulci G, Buhrman JS, et al:
Bevacizumab with angiostatin-armed oHSV increases antiangiogenesis
and decreases bevacizumab-induced invasion in U87 glioma. Mol Ther.
20:37–45. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Qiu JF, Zhang ZQ, Wang Y and You J:
Lentivirus-mediated RNAi knockdown of VEGFA in RKO colorectal
cancer cells decreases tumor formation and growth in vitro and in
vivo. Int J Clin Exp Pathol. 5:290–298. 2012.PubMed/NCBI
|
|
30
|
Chen CH, Lai JM, Chou TY, et al: VEGFA
upregulates FLJ10540 and modulates migration and invasion of lung
cancer via PI3K/AKT pathway. PloS One. 4:e50522009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
de Chevigny A, Core N, Follert P, et al:
miR-7a regulation of Pax6 controls spatial origin of forebrain
dopaminergic neurons. Nat Neurosci. 15:1120–1126. 2012.PubMed/NCBI
|