Introduction
Bladder cancer is the most common malignancy of the
urinary system and the sixth most common cancer in China. The
age-standardized incidence rate is 11.4 per 100,000 for men, and
3.51 per 100,000 for women (1). The
incidence of bladder cancer is even higher in Western countries
(2). Approximately 75% of bladder
cancers are noninvasive cancers, which have a high rate of
recurrence and progression. The remaining 25% of bladder cancers
are muscle-invasive cancers and are associated with extremely poor
prognosis despite systemic therapy (3). Although great advances have been
achieved in the understanding of this disease, the molecular
mechanisms involved in the development and progression of bladder
cancer are still largely unknown.
Increasing evidence has demonstrated the existence
of cancer stem cells (CSCs) in most cancers. The CSC hypothesis
suggests that a minor population of tumor cells, termed cancer stem
cells or tumor-initiating cells, share various properties with
normal stem cells and possess tumorigenic potential, as well as
self-renewal, differentiation, and proliferation capacities
(4). CSCs seem to be chemoresistant
and radioresistant, leading to local recurrence (5,6).
Isolation of rare CSCs is an important step in CSC
research. The most common method of CSC isolation is the use of
specific surface markers, such as CD44, CD133, CD24, ESA and
aldehyde dehydrogenase1 (ALDH1). Notably, the expression of CSC
surface markers is specific to each tissue type, but some tumors do
not have specific surface markers available for isolation. Side
population (SP) cell analysis and sorting have been successfully
used for the enrichment of stem cells in a variety of tissues
(5,7). SP cells isolated from diverse cancer
cell lines harbor stem cell-like properties (7–9).
B-cell-specific Moloney murine leukemia virus
insertion site 1 (Bmi1) is a member of the Polycomb group (PcG)
gene family, and was initially found to induce lymphoma upon
cooperation with c-Myc by inhibiting c-Myc-induced apoptosis via
p16INK4a/p14ARF (10,11).
Bmi1 plays a critical role in the self-renewal and differentiation
of a variety of normal stem cells and CSCs (10,12).
Recent studies suggest that Bmi1 is involved in cancer initiation,
and targeting Bmi1 using gene therapy abolishes the chemoresistance
of tumor cells (10,13). However, we have limited
understanding of the underlying mechanisms of Bmi1 function.
In the present study, we detected SP cells from the
human bladder cancer cell line T24. These cells exhibited stem
cell-like properties, providing the foundation for further study on
the molecular mechanism of bladder cancer. The objectives were then
to examine the functional significance of Bmi1 in the maintenance
of the stem cell-like properties of the SP subpopulation in T24
cells, and investigate whether Bmi1 is involved in the
tumorigenicity of SP T24 cells in vivo. Results from our
study may reveal the potential strategy for targeting Bmi1 for
bladder cancer therapy.
Materials and methods
Cell culture
The human bladder cancer cell line T24 [American
Type Culture Collection (ATCC), Manassas, VA, USA] was cultured in
RMPI-1640 medium (Gibco, Invitrogen, Carlsbad, CA, USA),
supplemented with 10% fetal bovine serum (FBS) (Gibco), 100 U/ml
penicillin G and 100 μg/ml streptomycin. All cell lines were
maintained in a humidified 5% CO2 incubator at 37°C.
Sorting of SP cells by flow cytometry
using Hoechst 33342
T24 cells were resuspended in pre-warmed complete
RPMI-1640 medium at a concentration of 1×106 cells/ml.
Cells were cultured either in the absence or in the presence of
verapamil (50 μmol/l; Sigma, St. Louis, MO, USA) for 30 min at
37°C, before adding the DNA-binding dye, Hoechst 33342 (Sigma), at
a final concentration of 5 μg/ml and incubating for 90 min in the
dark with gentle mixing. Cells were then washed twice with PBS and
resuspended in ice-cold PBS/2% FBS to a final concentration of
1×106 cells/ml. Propidium iodide (PI) (2 μg/ml; Sigma)
was added to gate viable cells, and the cells were maintained at
4°C in the dark before flow cytometry (Beckman Coulter, Brea, CA,
USA) sorting. Hoechst 33342 was excited at 355 nm and a fluorescent
profile was generated for dual-wavelength analysis (450/50 and
675/20 nm). We collected both SP and non-SP (NSP) cells to evaluate
the sorting purity and to conduct further experiments.
Immunofluorescence analysis
Freshly isolated SP and NSP cells from the T24 cells
were cultured on glass slides overnight. Cells were then fixed in
4% paraformaldehyde for 20 min, incubated with 0.5% Triton-X in PBS
for 5 min, and blocked in 10% normal goat serum for 30 min. After
incubation, cells were stained with a primary antibody against Bmi1
(Santa Cruz Biotechnology, Santa Cruz, CA, USA), at a dilution of
1:100 for 12 h at 4°C. Cells were then washed and incubated with
FITC-conjugated goat anti-mouse IgG (Proteintech Group Inc.,
Chicago, IL, USA) at a dilution of 1:200 for 1 h at room
temperature. After washing in PBS, cells were coverslipped with a
mounting medium containing 4′,6-diamidino-2-phenylindole (DAPI).
Stained cells were examined under a BX51 laser scanning confocal
microscope (Olympus, Tokyo, Japan).
Small-interfering RNA transfection
Specific siRNA targeting human Bmi1 cDNA (GenBank
no. NM_005180.8) (5′-CCU CCA CCU CUU CUU GUU UTT-3′) named
Bmi1-siRNA and a scramble-siRNA (5′-UUC UCC GAA CGU GUC ACG UTT-3′)
were synthesized by GenePharma (Shanghai, China). siRNAs (100 nM)
were transfected into T24 cells or SP cells sorted from T24 cells
using Lipofectamine 2000 (Invitrogen), according to the
manufacturer’s instructions. Transfection efficiency was then
assessed using quantitative real-time RT-PCR and western blotting
for Bmi1. Transfected cells were cultured for further
experiments.
Real-time RT-PCR analysis
Total RNA was extracted from cells using TRIzol
(Invitrogen). RNA purity was determined using absorbance at 260 and
280 nm (A260/280), and the integrity of the RNA was verified by
electrophoresis on formaldehyde gels. Each sample (1 μg) was
reverse transcribed using a RT-PCR kit (Takara Bio, Otsu, Japan).
Real-time PCR analysis was performed using a standard SYBR Green
PCR kit (Roche Applied Science, Penzberg, Germany) on a LightCycler
480 real-time PCR system (Roche Diagnostics, Basel, Switzerland),
according to the manufacturer’s instructions. RNA levels of the
GAPDH gene were monitored as an internal control. Primer sequences
were: GAPDH, forward 5′-AAG GTG AAG GTC GGA GTC AAC-3′ and reverse
5′-GGG GTC ATT GAT GGC AAC AAT A-3′; Bmi1, forward 5′-CGT GTA TTG
TTC GTT ACC TGG A-3′ and reverse 5′-TTC AGT AGT GGT CTG GTC TTG
T-3′; p14ARF, forward 5′-ATG GAG CCT TCG GCT GAC T-3′ and reverse
5′-GTA ACT ATT CGG TGC GTT GGG-3′; p16INK4a, forward 5′-GGG TTT TCG
TGG TTC ACA TCC-3′ and reverse 5′-CTA GAC GCT GGC TCC TCA GTA-3′;
Oct4, forward 5′-CTT GAA TCC CGA ATG GAA AGG G-3′ and reverse
5′-GTG TAT ATC CCA GGG TGA TCC TC-3′; ABCG2, forward 5′-ACG AAC GGA
TTA ACA GGG TCA-3′ and reverse 5′-CTC CAG ACA CAC CAC GGA T-3′. The
relative expression was calculated using the 2−ΔΔCt
method.
Western blotting
Equal amounts of protein lysates (20 μg) were
subjected to SDS-PAGE and transferred to a PVDF membrane by
electroblotting. Antibodies against Bmi1 (1:2,000 dilution; Santa
Cruz Biotechnology), P14ARF (1:2,000 dilution; Abcam,
USA) and P16INK4a (1:1,000 dilution, Abcam) were used as
the primary antibodies. An anti-β-actin antibody at a 1:10,000
dilution was used as a loading control. The protein bands were
visualized using an imaging system (Bio-Rad, Hercules, CA,
USA).
Cell proliferation assays
Cells were plated at a density of 1×103
cells/well in complete RPMI-1640, in triplicate, in a 96-well plate
(100 μl/well) and incubated over 7 days. The cell titer
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)
was added to each well according to the manufacturer’s
instructions. After 4 h in culture, the MTT medium was removed, and
then DMSO was added into each well to dissolve the formazan. Cell
viability was determined by measuring the absorbance at 490 nm
using a Model 550 plate reader (Bio-Rad).
Cells were plated at 1,000 cells/well in 96-well
plates. Cisplatin (Sigma) was added the following day at a
concentration gradient (0.5, 2 and 4 μg/ml), in triplicate. After
48 h, the cell survival rate (SR) (%) was determined using the MTT
method. SR was calculated using the formula: SR (%) = (mean
absorbance of the test well/mean absorbance of the control) × 100.
Inhibition rate (IR) was calculated as IR (%) = 100% − SR (%).
Cell migration assays
For assessment of cell migration in vitro,
SP, NSP, SP-Bmi1-siRNA cells and SP-scramble-siRNA cells
(1×105 in 100 μl of serum-free RPMI-1640 medium) were
placed in the upper chamber of Transwell migration chambers with an
8-μm pore size (BD Biosciences, Franklin Lakes, NJ, USA). The lower
chamber contained 0.6 ml of 10% FBS medium as a chemoattractant.
Cells were incubated for 12 h. Non-migrated cells were removed from
the upper surface of the Transwell membrane with a cotton swab, and
the migrated cells on the lower side of the membrane were fixed
with 4% paraformaldehyde and stained with 0.25% crystal violet.
After staining, cell migration was quantified by counting cells in
5 random fields (x200) under a microscope.
Self-renewal as assessed by tumor sphere
formation
Cells were plated in ultra-low adhesion 6-well
plates at a density of 1×104 cells/well and grown in a
serum-free DMEM/F12 medium (Gibco) containing 20 ng/ml of epidermal
growth factor (EGF; R&D Systems, Minneapolis, MN, USA), 5 μg/ml
of insulin, 0.4% bovine serum albumin (Gibco), and 2% B27
(Invitrogen), as previously described (14). After 7 days in culture, the number
of spheres was quantified by counting spheres in 5 random fields
(x100) under a phase contrast microscope (Olympus).
Cell cycle analysis
For cell cycle analysis, SP cells transfected with
Bmi1-siRNA or scramble-siRNA were washed twice with PBS and fixed
overnight with 70% ethanol at −20°C. Cells were then washed with
PBS and incubated in PBS containing 50 μg/ml PI, 0.2% Triton X-100,
and 200 mg/ml RNase A for 30 min in the dark. Cell cycle
distribution was analyzed using FACScan System (BD Diagnostics,
Sparks, MD, USA). Data were analyzed with FlowJo software (Tree
Star Inc., Ashland, OR, USA).
Mouse xenograft models
NOD/SCID mice (n=4, male, 6–10 weeks of age) were
purchased from the Laboratory Animal Center of the Sun Yat-Sen
University (Guangzhou, China). All animal experiments were approved
by the Animal Ethics and Welfare Committee of the Sun Yat-Sen
University. SP cells (1×105) transfected with Bmi1-siRNA
or scramble-siRNA were suspended in RMPI-1640 and Matrigel (BD
Biosciences, 1:1) and subcutaneously injected into the left and
right sides of the back of recipient mice, respectively. Tumor
formation was observed weekly for 6 weeks.
Statistical analysis
Data analysis was performed using SPSS 19.0 (SPSS
Inc., Chicago, IL, USA). Data are expressed as means ± standard
deviation (SD). Differences between groups were compared using the
Student’s t-test or one-way analysis of variance (ANOVA) with
Fisher’s least significant difference (LSD) test for post hoc
analysis. Differences were considered to be statistically
significant at P<0.05.
Results
Identification of SP cells from human
bladder cancer T24 cells
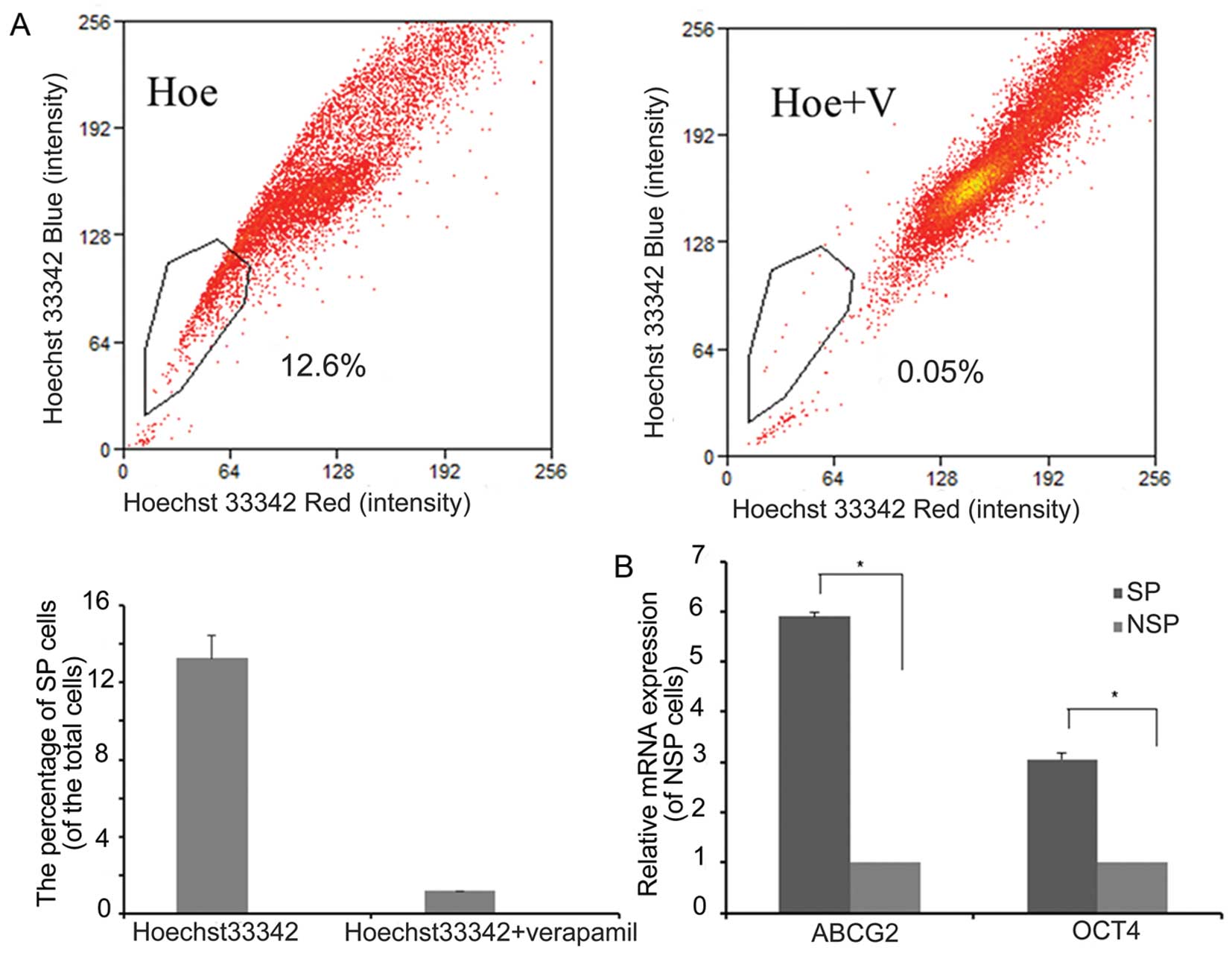
To explore the effect of Bmi1 in the maintenance of
stem-like SP cells, the human bladder cancer cell line T24 was
sorted by flow cytometry after incubation with Hoechst 33342 for 90
min. SP cells represented 13.2±1.2% of the total cells. When
preincubated with verapamil for 30 min, the proportion of SP cells
dropped to 0.05±0.01% of the total cells (Fig. 1A), which indicated that Hoechst
33342 exclusion is verapamil-sensitive.
To determine whether SP cells had a stem cell
phenotype, we compared the expression levels of cancer stem cell
markers, such as Oct4 and ABCG2, between SP and NSP cells by
real-time RT-PCR. The mRNA expression of ABCG2 and OCT4 was
significantly higher in SP cells than in NSP (both P<0.05)
(Fig. 1B).
Bmi1 is highly expressed in SP cells when
compared with the expression level in NSP cells
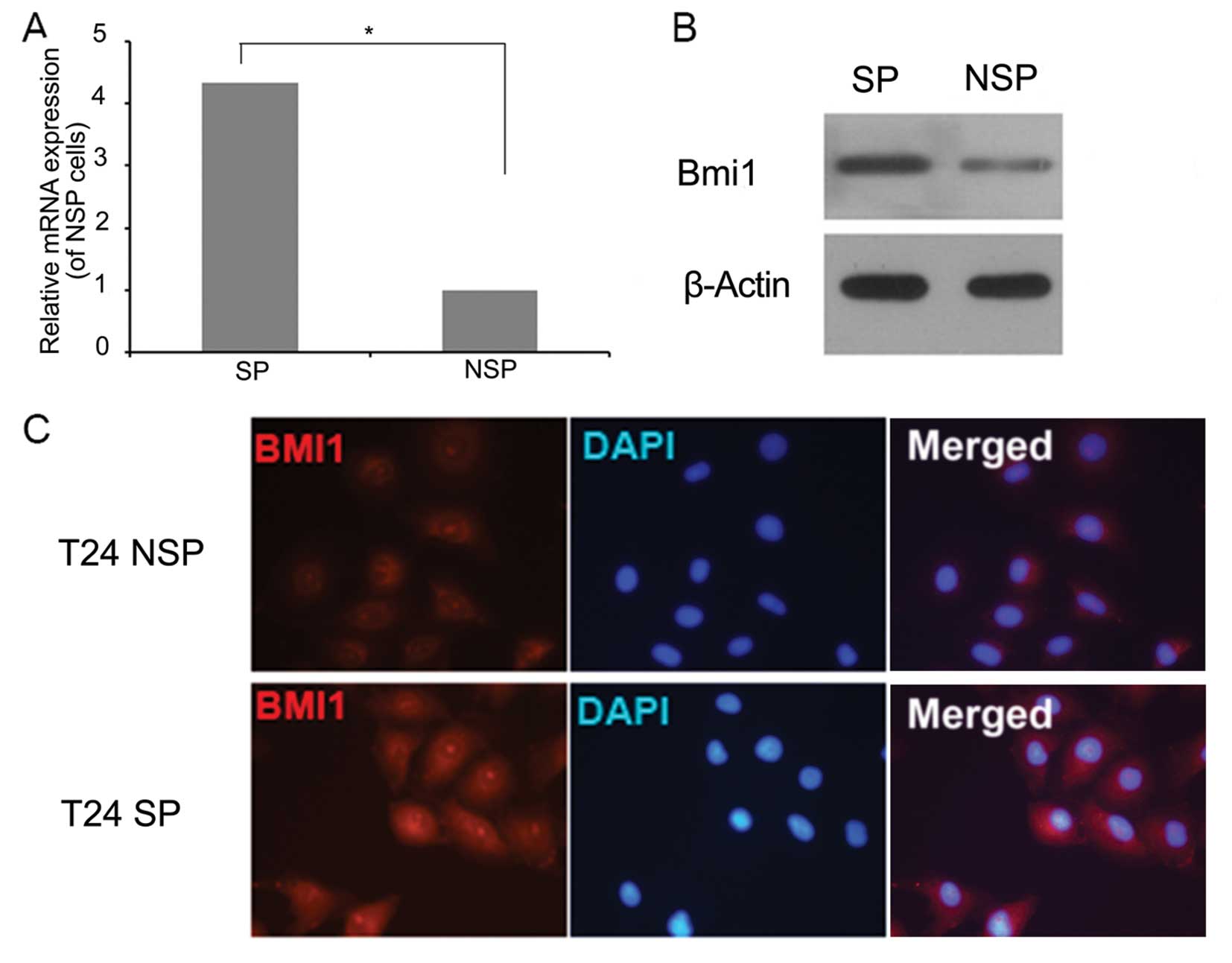
We observed that the mRNA expression of Bmi1 was
upregulated 4.34 times in SP cells when compared with the
expression level in NSP cells (Fig.
2A). Western blot and immunofluorescence analyses confirmed
that Bmi1 was highly expressed in the nuclei and cytoplasm of SP
cells when compared with the NSP cells (Fig. 2B and C). These results suggest that
Bmi1 may play a regulatory role in the stem-like SP phenotype.
Effects of Bmi1 knockdown on the SP T24
cell phenotype, cell proliferation and migration in vitro
Our previous study confirmed that the purified SP
cells can generate non-stem-like NSP cells through asymmetric cell
division in vitro (17).
Firstly, we transfected SP cells with Bmi1 siRNA and scramble siRNA
for 48 h, and the efficient knockdown of Bmi1 was confirmed by
real-time RT-PCR and western blotting (Fig. 6A and B). Secondly, we transfected
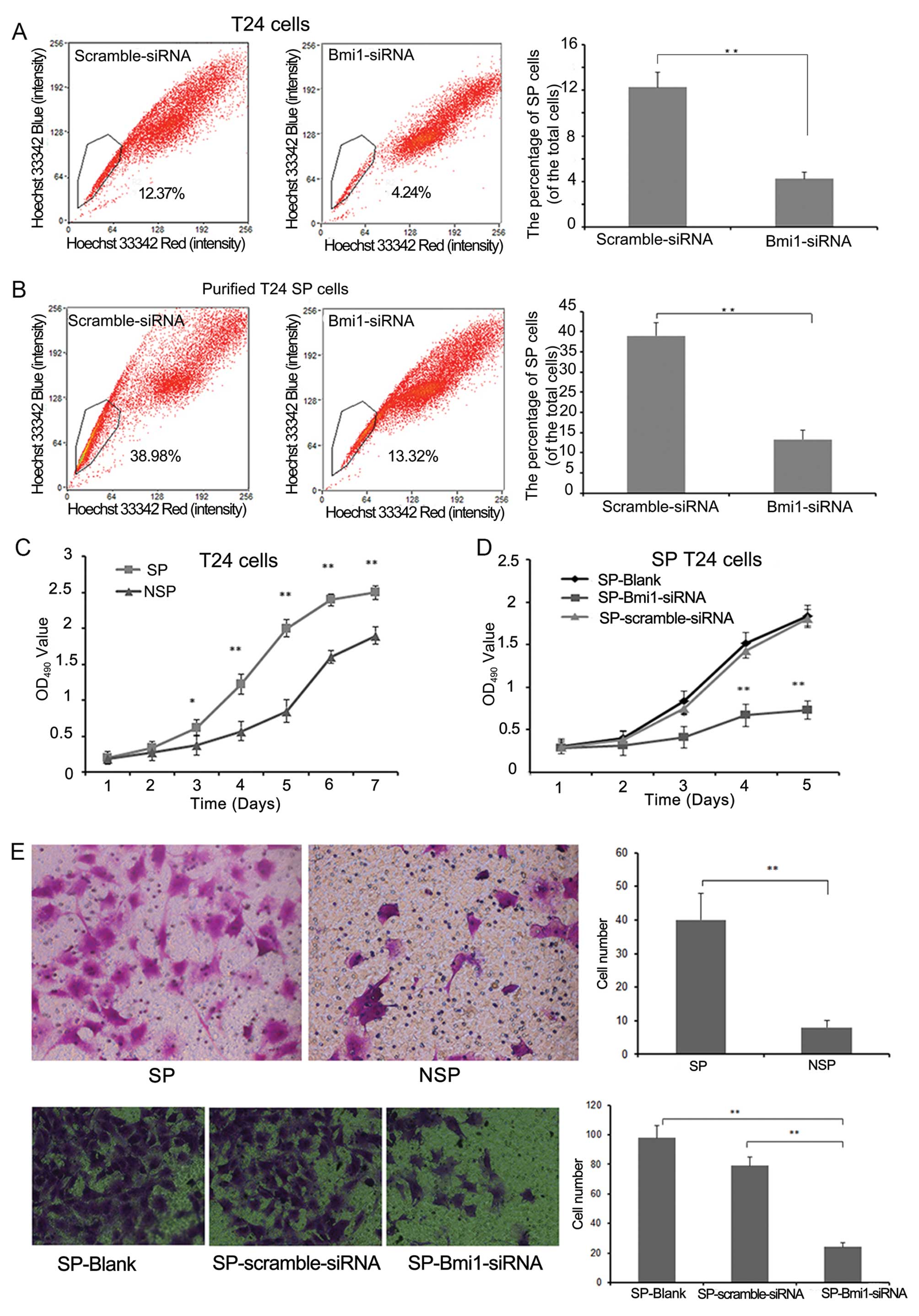
Bmi1 siRNA and scramble siRNA into the T24 cells (Fig. 3A), and the result showed that
knockdown of Bmi1 reduced the proportion of SP cells from 12.26±1.3
to 4.24±0.6% (P<0.01) in the T24 cells. Thirdly, we transfected
Bmi1 siRNA and scramble siRNA into purified SP cells and cultured
them for 10 days to examine the role of Bmi1 in this process. The
SP subpopulation in the Bmi1 siRNA group was decreased when
compared with the negative control cells (13.32±2.2 vs. 38.98±3.2%,
P<0.01; Fig. 3B). These results
demonstrate that Bmi1 plays an important role in maintaining SP
cell self-renewal and differentiation abilities, and knockdown of
Bmi1 promoted SP cell differentiation toward non-stem-like NSP
cells.
We then detected the proliferation and migration
abilities of SP and NSP cells using MTT and Transwell assays. The
cell growth curve indicated that the SP cells proliferated faster
than the NSP cells after day 2 (Fig.
3C). In the Transwell assay, more SP cells penetrated through
the gel membrane compared with NSP cells (P<0.01) (Fig. 3E). After transfection with siRNA
targeting Bmi1, the proliferation and migration of SP-Bmi1-siRNA
cells were sharply decreased compared with the SP-scramble-siRNA
cells (Fig. 3D and E).
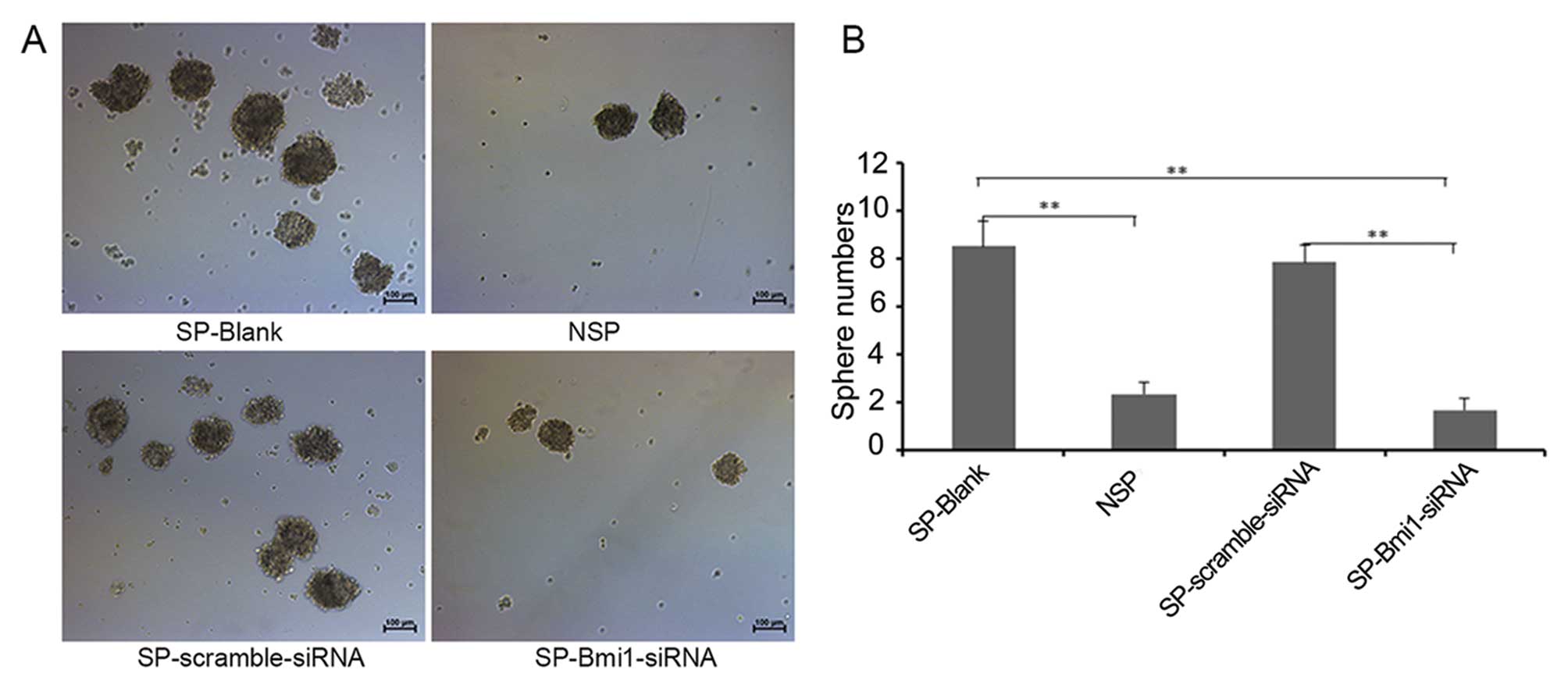
Bmi1 regulates the self-renewal capacity
of SP cells
Tumor sphere generation in vitro is
considered to be indicative of self-renewal potential, one of the
characteristics of CSCs (15). We
assessed the self-renewal properties of SP cells by detecting their
ability to form tumor spheres in serum-free media and non-adherent
conditions. Results showed that the number of tumor spheres formed
by SP cells was nearly four times higher than that for NSP cells.
In order to ascertain whether Bmi1 regulates the self-renewal
capacity of SP cells, we silenced the Bmi1 expression in SP cells
using siRNA, and the results of the sphere formation assay showed
that this ability of SP cells was significantly inhibited compared
with that of SP-scramble- siRNA cells (P<0.01) (Fig. 4). These data revealed that Bmi1
plays an important role in regulating the self-renewal ability of
SP cells.
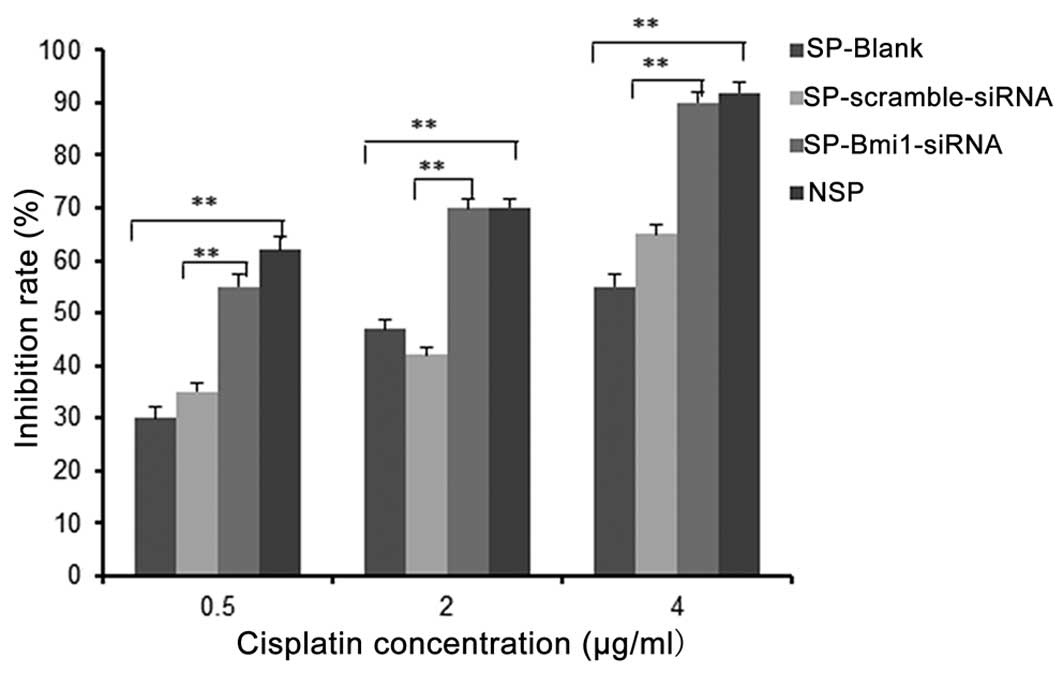
Bmi1 knockdown enhances SP cell sensitive
to cisplatin in vitro
Chemoresistance is considered to be one of the
characteristics of CSCs (16). We
previously reported that the SP cells sorted from the T24 cells
were significantly more resistant to chemotherapy when compared
with the resistance of NSP cells (17). In order to ascertain whether Bmi1
knockdown influences cisplatin sensitivity of SP cells, we treated
SP, NSP, SP-Bmi1-siRNA and SP-scramble-siRNA cells with various
concentrations of cisplatin for 48 h (Fig. 5). We then observed the inhibition
rate (IR) of each cell group by MTT assays. Results revealed that
SP cells showed a stronger resistance to cisplatin compared with
the NSP cells at all concentrations (all P<0.01). After
knockdown of Bmi1, the IR of the SP-Bmi1-siRNA cells was obviously
higher when compared with the IR of the SP-scramble-siRNA cells
(all P<0.01). These results demonstrated SP chemoresistance and
that Bmi1 knockdown could sensitize SP cells to cisplatin.
Bmi1 knockdown induces cell cycle arrest
in the G0/G1 phase through derepression of the
p16INK4a/p14ARF locus in SP T24 cells
Bmi1 has been shown to be important for the
self-renewal of neural stem cells by inhibition of the
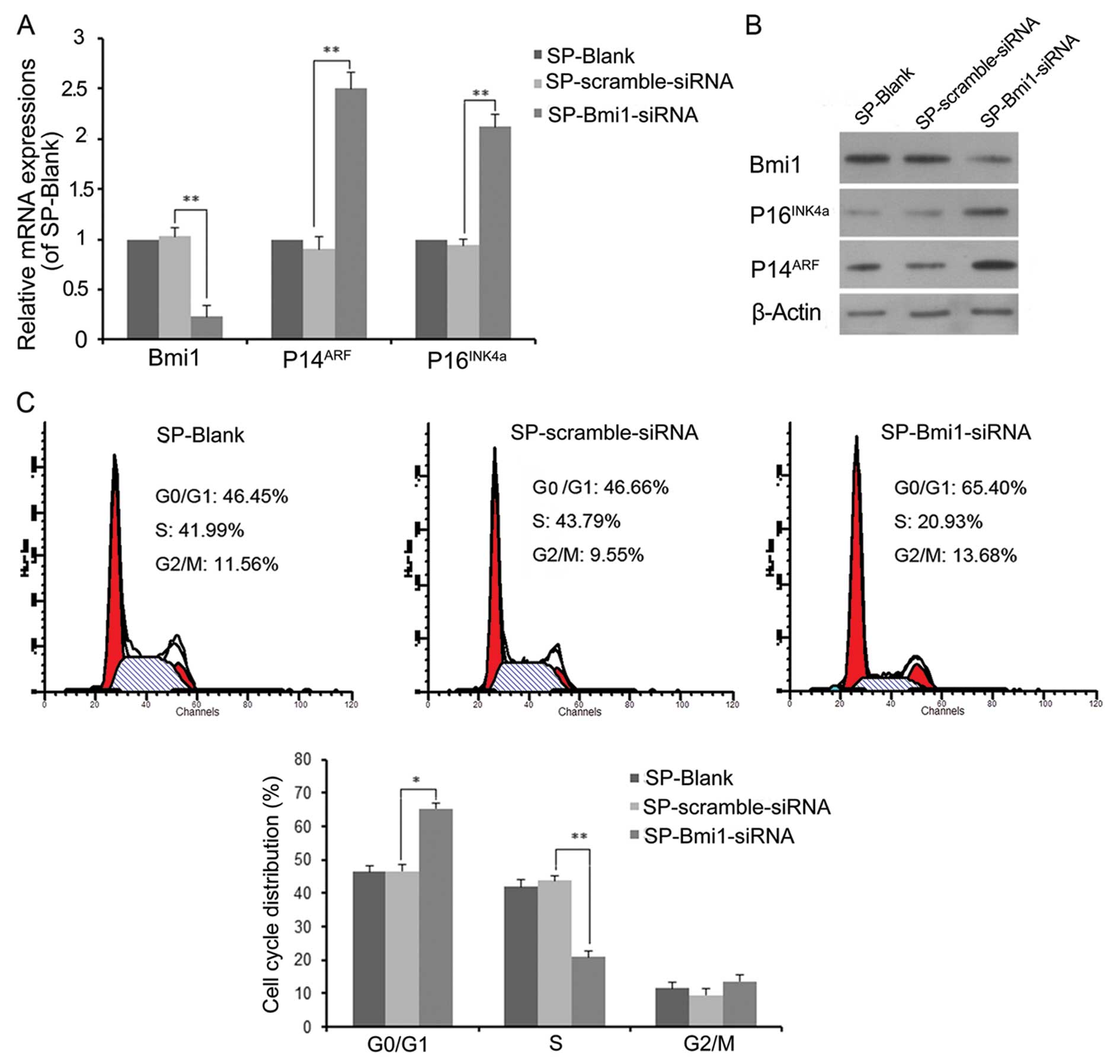
p16INK4a/p14ARF locus (18). To ascertain whether knockdown of
Bmi1 results in the upregulation of
p16INK4a/p14ARF, we transfected SP cells with
Bmi1 siRNA and scramble siRNA for 48 h. The expression of
p16INK4a and p14ARF was upregulated at the
mRNA and protein levels (Fig. 6A and
B). We also assessed the relationship between Bmi1 knockdown
and cell cycle progression by flow cytometric analysis (Fig. 6C). Compared with the negative
control SP cells, Bmi1 knockdown caused a significant decrease in
the proportion of S-phase cells (from 43.8±1.5 to 20.9±1.9%,
P<0.01) and a significant increase in the proportion of
G0/G1-phase cells (from 46.7±2.0 to 65.4±1.6%, P<0.05). These
results suggest that Bmi1 promotes stem-like SP cell proliferation
by regulating cell cycle progression through suppression of the
p16INK4a/p14ARF locus.
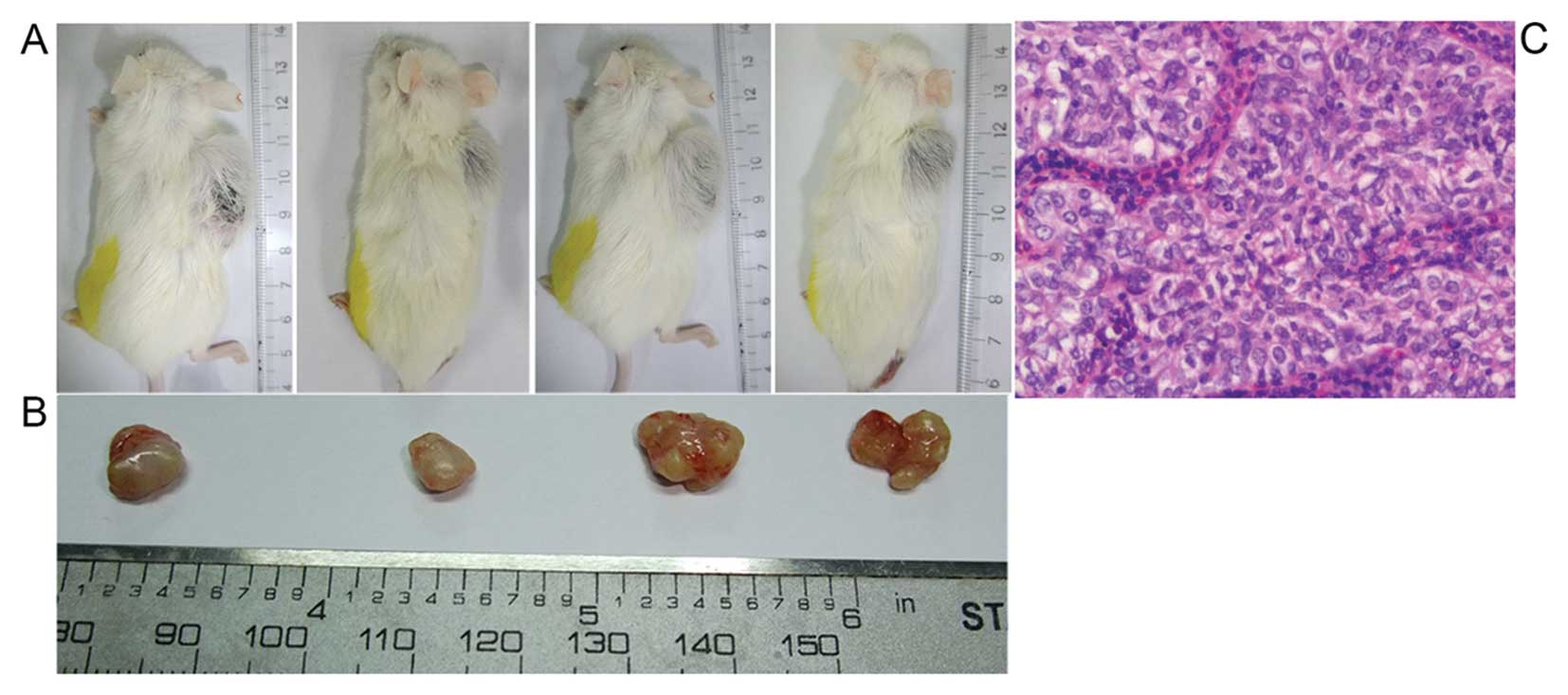
Bmi1 knockdown suppresses the
tumorigenicity of SP T24 cells in vivo
To further explore the role of Bmi1 in the
tumorigenesis of bladder cancer CSCs in vivo, equal numbers
of SP-Bmi1-siRNA and SP-scramble-siRNA cells were subcutaneously
injected into the left and right sides of the back of recipient
mice, respectively. The development of tumors was detected
throughout the 6 weeks after injection. The SP-scramble-siRNA cells
generated varying sizes of tumors in all four mice (Fig. 7B). In contrast, SP-Bmi1-siRNA cells
failed to form subcutaneous tumors in any of the recipient mice
(Fig. 7A). Results of hematoxylin
and eosin (H&E) staining (Fig.
7C) showed that the tumors formed from the subcutaneously
injected SP-scramble-siRNA cells was a typical urothelial
carcinoma. This in vivo result further supports that Bmi1 is
essential for the tumorigenic capacity of SP T24 cells.
Discussion
Increasing evidence supports the existence of CSCs,
which are considered to be responsible for tumor initiation, tumor
recurrence and therapy resistance (4,5).
Therefore, a better understanding of the biology and regulatory
mechanisms of CSCs is of great significance for developing new
strategies targeting CSCs for cancer treatment (11). In a previous study, we enriched
bladder cancer stem cells from the T24 cell line and characterized
their CSC properties (17).
Furthermore, SP-enriched cells possessing stem cell characteristics
have been identified in a number of cancers (4,8,9).
We used SP cells as a bladder cancer stem cell model
to elucidate their self-renewal and tumor initiation mechanisms. In
the present study, we observed that SP cells from the T24 cell line
possessed CSC properties and that Bmi1 was highly expressed in the
SP population when compared with that in the NSP cells. Our results
also demonstrated that Bmi1 plays an important role in regulating
proliferation, self-renewal and tumorigenic properties of SP cells
by the inhibition of the p16INK4a/p14ARF
locus. Knockdown of Bmi1 expression by siRNA sensitized SP cells to
cisplatin and suppressed their tumorigenicity in vitro and
in vivo.
CSCs are characterized by self-renewal,
chemoresistance and tumorigenesis (19). To identify CSCs among tumor cells,
two different methods are usually proposed. The first is to use
surface markers selectively expressed on CSCs. However, in many
tumor types, no markers are known to prospectively identify CSCs;
in such cases, the ability of stem cells to extrude dyes such as
Hoechst 33342 can be used to identify them (20). Thus, in a number of tumors and
cancer cell lines, SP cells have been isolated and identified to be
potential candidates for CSCs (9,20–22).
The ATP-binding cassette transporter ABCG2/BCRP1 has been revealed
to be the main mediator of the SP phenotype (23,24).
In contrast, some studies have demonstrated that in some normal
tissues or certain tumors, SP cells are not enriched in stem-like
cells (25–27). Triel et al (27) demonstrated that SP cells found
within human epidermis lack stem cell characteristics, and Broadley
et al (28) revealed that SP
cells from glioblastoma multiforme cell lines had no stem-like
activity in vitro and in vivo when compared with
these characteristics in non-SP and parental cells. These
conflicting data indicate that SP cells are not necessarily stem
cells and that further characterization is necessary to prove their
nature. In one of our previous studies (17) and in the present study, results
showed that SP cells exhibited a stronger ability for
proliferation, self-renewal and chemoresistance than NSP cells in
the bladder cancer cell line T24. Furthermore, SP cells had high
expression of ABCG2 and stem cell marker OCT4 when compared with
that in the NSP cells. Our results suggest that SP cells can serve
as a model for the study of bladder CSCs.
Bmi1, a component of PRC1, is thought to be
essential for maintaining self-renewal in several normal stem cell
systems and CSCs, including haematopoietic stem cells (12), neural stem cells (18) and leukemic stem cells (29). Recent studies have demonstrated that
Bmi1 is highly expressed and regulates the stem-like properties of
SP cells in hepatocellular carcinoma, breast cancer and pancreatic
adenocarcinoma (30–32). In our previous studies, Bmi1 was
found to be highly expressed in bladder cancer specimens and to be
correlated to clinicopathological characteristics and patient
prognosis (33,34). Therefore, we hypothesized that Bmi1
may also play an important regulatory role in bladder CSCs. In the
present study, we functionally validated the importance of Bmi1
expression in stem-like SP cells from the bladder cancer cell line
T24. As expected, our results showed that both the mRNA and protein
levels of Bmi1 were higher in SP cells. Knockdown of Bmi1 by siRNA
significantly decreased the proportion of SP cells among the T24
cells and purified SP cells after long-term differentiation.
Furthermore, analysis of the growth, migration and tumor sphere
formation of purified SP cells transfected with siRNA revealed that
loss of Bmi1 caused a considerable inhibition in the proliferation
and self-renewal of SP cells. These observations were further
confirmed by xenograft transplantation in NOD/SCID mice, where Bmi1
knockdown in SP cells resulted in the failure to develop
tumors.
Bmi1 has been reported to be associated with the
protection of cancer cells from apoptosis induced by chemotherapy.
Yin et al (35) observed
that Bmi1 promoted the chemoresistance of pancreatic cancer cells
to gemcitabine. Recently, Wang et al (36) reported that Bmi1 enhanced the
chemoresistance of ovarian cancer cells, and that silencing Bmi1
sensitized ovarian cancer cells to cisplatin. A previous study also
showed that CTCs extracted from T24 cells were resistant to
cisplatin, and that this resistance was related to aldehyde
dydrogenase (37); however, the
relationship between Bmi1 and aldehyde dydrogenase is unknown.
Another study showed that the cisplatin resistance of T24 SP cells
was correlated with Bmi1, and that Bmi1 was also correlated with
Nanog expression (38). Consistent
with these reports, our results revealed that SP cells from bladder
cancer cells showed a stronger resistance to cisplatin, and that
Bmi1 knockdown significantly increased the IR by cisplatin in SP
cells. These results strongly suggest that Bmi1 is essential for
maintaining tumorigenesis and chemoresistance in stem-like SP
cells.
However, the molecular mechanisms of Bmi1 function
in CSCs are not completely understood. Some studies revealed that
Bmi1 preserves the self-renewal and proliferation characteristics
of stem cells through the repression of the
p16INK4a/p14ARF locus, which encodes two
tumor-suppressor proteins (P16Ink4a and P14Arf) (18,39,40).
Conversely, some studies revealed an
p16INK4a/p14ARF-independent contribution of
Bmi1 to self-renewal in neural stem cells and tumorigenesis in a
hepatocellular carcinoma mouse model (41,42).
Douglas et al (43) reported
that Bmi1 knockdown significantly inhibited cell proliferation in
both wild-type and p16INK4a-null Ewing sarcoma.
Nevertheless, the molecular mechanisms by which Bmi1 affects CSCs
in human bladder cancer have not been previously identified. In the
present study, we observed that Bmi1 knockdown using siRNA could
promote p16INK4a/p14ARF mRNA and protein
expression in stem-like SP cells from the human bladder cancer cell
line T24. Bmi1 knockdown also caused S-phase cell cycle arrest in
bladder cancer T24 SP cells. These results demonstrated that Bmi1
is involved in stem-like SP cell self-renewal and tumorigenesis
through the repression of the p16INK4a/p14ARF
locus in the human bladder cancer cell line T24.
In conclusion, our results demonstrated that the SP
phenotype can be used to isolate CSCs in the bladder cancer cell
line T24, and that Bmi-1 modulates various cancer stem-like
characteristics of bladder cancer SP cells, such as self-renewal,
chemoresistance and tumorigenesis. In addition, Bmi1 performed
these functions to a great extent through the suppression of the
p16INK4a/p14ARF locus. Based on our findings,
Bmi1 may be a target by which to decrease the CSC population of
tumors and to improve treatment outcomes.
Acknowledgements
This study was supported by grants from the National
Natural Science Foundation of China (no. 81072102 to W. Xie) and
the Collaborative Funds from Panyu Hospital of Chinese
Medicine.
References
|
1
|
Han S, Zhang S and Chen W: Analysis of the
status and trends of bladder cancer incidence in China. Oncol
Progress. 11:89–95. 2013.
|
|
2
|
Ploeg M, Aben KK and Kiemeney LA: The
present and future burden of urinary bladder cancer in the world.
World J Urol. 27:289–293. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Babjuk M, Burger M, Zigeuner R, et al: EAU
guidelines on non-muscle-invasive urothelial carcinoma of the
bladder: update 2013. Eur Urol. 64:639–653. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Enderling H, Hlatky L and Hahnfeldt P:
Cancer stem cells: a minor cancer subpopulation that redefines
global cancer features. Front Oncol. 3:762013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Reya T, Morrison SJ, Clarke MF and
Weissman IL: Stem cells, cancer, and cancer stem cells. Nature.
414:105–111. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
6
|
Siddique HR and Saleem M: Role of BMI1, a
stem cell factor, in cancer recurrence and chemoresistance:
preclinical and clinical evidences. Stem Cells. 30:372–378. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Sugihara E and Saya H: Complexity of
cancer stem cells. Int J Cancer. 132:1249–1259. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kondo T: Stem cell-like cancer cells in
cancer cell lines. Cancer Biomark. 3:245–250. 2007.PubMed/NCBI
|
|
9
|
Wu C and Alman BA: Side population cells
in human cancers. Cancer Lett. 268:1–9. 2008. View Article : Google Scholar
|
|
10
|
Park IK, Morrison SJ and Clarke MF: Bmi1,
stem cells, and senescence regulation. J Clin Invest. 113:175–179.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Jacobs JJ, Kieboom K, Marino S, DePinho RA
and van Lohuizen M: The oncogene and Polycomb-group gene bmi-1
regulates cell proliferation and senescence through the ink4a
locus. Nature. 397:164–168. 1999. View
Article : Google Scholar : PubMed/NCBI
|
|
12
|
Park IK, Qian D, Kiel M, et al: Bmi-1 is
required for maintenance of adult self-renewing haematopoietic stem
cells. Nature. 423:302–305. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Siddique HR, Parray A, Tarapore RS, et al:
BMI1 polycomb group protein acts as a master switch for growth and
death of tumor cells: regulates TCF4-transcriptional factor-induced
BCL2 signaling. PLoS One. 8:e606642013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Fan X, Chen X, Deng W, Zhong G, Cai Q and
Lin T: Up-regulated microRNA-143 in cancer stem cells
differentiation promotes prostate cancer cells metastasis by
modulating FNDC3B expression. BMC Cancer. 13:612013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ponti D, Costa A, Zaffaroni N, et al:
Isolation and in vitro propagation of tumorigenic breast cancer
cells with stem/progenitor cell properties. Cancer Res.
65:5506–5511. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Tang C, Ang BT and Pervaiz S: Cancer stem
cell: target for anti-cancer therapy. FASEB J. 21:3777–3785. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ning ZF, Huang YJ, Lin TX, et al:
Subpopulations of stem-like cells in side population cells from the
human bladder transitional cell cancer cell line T24. J Int Med
Res. 37:621–630. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Molofsky AV, He S, Bydon M, Morrison SJ
and Pardal R: Bmi-1 promotes neural stem cell self-renewal and
neural development but not mouse growth and survival by repressing
the p16Ink4a and p19Arf senescence pathways.
Genes Dev. 19:1432–1437. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Cetin I and Topcul M: Cancer stem cells in
oncology. J BUON. 17:644–648. 2012.PubMed/NCBI
|
|
20
|
Moserle L, Ghisi M, Amadori A and
Indraccolo S: Side population and cancer stem cells: therapeutic
implications. Cancer Lett. 288:1–9. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ho MM, Ng AV, Lam S and Hung JY: Side
population in human lung cancer cell lines and tumors is enriched
with stem-like cancer cells. Cancer Res. 67:4827–4833. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Oates JE, Grey BR, Addla SK, et al:
Hoechst 33342 side population identification is a conserved and
unified mechanism in urological cancers. Stem Cells Dev.
18:1515–1522. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhou S, Schuetz JD, Bunting KD, et al: The
ABC transporter Bcrp1/ABCG2 is expressed in a wide variety of stem
cells and is a molecular determinant of the side-population
phenotype. Nat Med. 7:1028–1034. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Hepburn AC, Veeratterapillay R, Williamson
SC, et al: Side population in human non-muscle invasive bladder
cancer enriches for cancer stem cells that are maintained by MAPK
signalling. PLoS One. 7:e506902012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Stingl J, Eirew P, Ricketson I, et al:
Purification and unique properties of mammary epithelial stem
cells. Nature. 439:993–997. 2006.PubMed/NCBI
|
|
26
|
Mitsutake N, Iwao A, Nagai K, et al:
Characterization of side population in thyroid cancer cell lines:
cancer stem-like cells are enriched partly but not exclusively.
Endocrinology. 148:1797–1803. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Triel C, Vestergaard ME, Bolund L, Jensen
TG and Jensen UB: Side population cells in human and mouse
epidermis lack stem cell characteristics. Exp Cell Res. 295:79–90.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Broadley KW, Hunn MK, Farrand KJ, et al:
Side population is not necessary or sufficient for a cancer stem
cell phenotype in glioblastoma multiforme. Stem Cells. 29:452–461.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lessard J and Sauvageau G: Bmi-1
determines the proliferative capacity of normal and leukaemic stem
cells. Nature. 423:255–260. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Chiba T, Miyagi S, Saraya A, et al: The
polycomb gene product BMI1 contributes to the maintenance of
tumor-initiating side population cells in hepatocellular carcinoma.
Cancer Res. 68:7742–7749. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Liu S, Dontu G, Mantle ID, et al: Hedgehog
signaling and Bmi-1 regulate self-renewal of normal and malignant
human mammary stem cells. Cancer Res. 66:6063–6071. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Proctor E, Waghray M, Lee CJ, et al: Bmi1
enhances tumorigenicity and cancer stem cell function in pancreatic
adenocarcinoma. PLoS One. 8:e558202013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Qin ZK, Yang JA, Ye YL, et al: Expression
of Bmi-1 is a prognostic marker in bladder cancer. BMC Cancer.
9:612009. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Liang W, Zhu D, Cui X, et al: Knockdown
BMI1 expression inhibits proliferation and invasion in human
bladder cancer T24 cells. Mol Cell Biochem. 382:283–291. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yin T, Wei H, Leng Z, et al: Bmi-1
promotes the chemoresistance, invasion and tumorigenesis of
pancreatic cancer cells. Chemotherapy. 57:488–496. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wang E, Bhattacharyya S, Szabolcs A, et
al: Enhancing chemotherapy response with Bmi-1 silencing in ovarian
cancer. PLoS One. 6:e179182011. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Falso MJ, Buchholz BA and White RW:
Stem-like cells in bladder cancer cell lines with differential
sensitivity to cisplatin. Anticancer Res. 32:733–738.
2012.PubMed/NCBI
|
|
38
|
Zhang Y, Wang Z, Yu J, et al: Cancer
stem-like cells contribute to cisplatin resistance and progression
in bladder cancer. Cancer Lett. 322:70–77. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Molofsky AV, Pardal R, Iwashita T, Park
IK, Clarke MF and Morrison SJ: Bmi-1 dependence distinguishes
neural stem cell self-renewal from progenitor proliferation.
Nature. 425:962–967. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Biehs B, Hu JK, Strauli NB, et al: BMI1
represses Ink4a/Arf and Hox genes to regulate stem cells in the
rodent incisor. Nat Cell Biol. 15:846–852. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Fasano CA, Dimos JT, Ivanova NB, Lowry N,
Lemischka IR and Temple S: shRNA knockdown of Bmi-1 reveals a
critical role for p21-Rb pathway in NSC self-renewal during
development. Cell Stem Cell. 1:87–99. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Xu CR, Lee S, Ho C, et al: Bmi1 functions
as an oncogene independent of Ink4A/Arf repression in hepatic
carcinogenesis. Mol Cancer Res. 7:1937–1945. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Douglas D, Hsu JH, Hung L, et al: BMI-1
promotes Ewing sarcoma tumorigenicity independent of CDKN2A
repression. Cancer Res. 68:6507–6515. 2008. View Article : Google Scholar : PubMed/NCBI
|