Introduction
Colon cancer is one of the most frequently diagnosed
cancer and the third most fatal malignancy worldwide (1). Growing urbanization and
industrialization have led to the steady rise in colon cancer
incidence in China during the last 20 years (2). In certain high incidence areas, colon
cancer has become the second leading cause of cancer-related
mortality. Surgical resection is widely considered as the most
useful therapy for colon cancer. Tumor metastasis is the major
cause for the recurrence of colon cancer and leads to the failure
of colon cancer treatment following radical operation. At present,
clinicopathologic staging is the main risk assessment for colon
cancer metastasis. Clinical outcomes, however, are quite variable,
even among patients at the same stage. Therefore, research focusing
on the molecular basis of colon tumor metastasis with the aim of
implementing individualized therapeutic regimens for patients with
colon cancer is urgently required.
The complicated process of tumor metastasis involves
a number of molecular biological changes, including genetic and
epigenetic mutations (3). Recently,
the contribution of long non-coding RNAs in cancer progression has
attracted increased attention. Long non-coding RNAs (lncRNAs)
(typically >200 nt), a type of regulator non-coding RNAs, are
increasingly reported to play key roles in numerous biological
processes such as development, differentiation, diseases via
integrity of nuclear structure, chromatin remodeling and
post-transcription regulation (4).
Gupta et al (5) first
discovered that HOTAIR (Hox transcript antisense intergenic
RNA) is one of the cancer metastasis-associated lncRNAs, and is a
powerful biomarker of breast cancer metastasis and poor prognosis.
Many other groups successively reported that HOTAIR
overexpression is linked to the malignant features of many other
cancer types (6–13). Moreover, Rinn et al (14) and Tsai et al (15) suggested that HOTAIR lncRNA
serves as a scaffold for at least two histone modification
complexes in order to induce transcriptional silencing of hundreds
of genes including tumor-suppressor genes. Nevertheless, the
mechanism of how HOTAIR is involved in colon cancer
progression remains unknown.
In the present study, we evaluated the expression of
HOTAIR in colon cancer tissue and paired normal mucosa and
metastasized lymph node, and then determined the correlation
between HOTAIR expression and clinicopathologic
characteristics. Using RNA interference experiments, we ascertained
whether HOTAIR has effects on colon cancer cell biological
properties. Furthermore, we revealed how HOTAIR initiates
epithelial-mesenchymal transition of colon cancer, the main
mechanistic step resulting in metastasis.
Materials and methods
Tissue samples and cell lines
Samples from 120 patients undergoing colectomy
between October 2007 and December 2010 by the same surgical team at
the Shanghai Jiao Tong University Affiliated First People’s
Hospital were collected and archived for the study. Fresh colon
cancer samples and matched adjacent non-tumor tissues were obtained
from 52 males and 68 females with ages ranging from 35 to 88 years.
The samples were stored at −80°C immediately following removal.
None of the patients had received any preoperative therapy, and
patients with stage II, III and IV disease underwent standard
adjuvant chemotherapy according to The National Comprehensive
Cancer Network Practice Guidelines for Colon Cancer (1). Follow-up was carried out completely
(median follow-up time, 55.5 months; range, 10–72 months). The
study was approved by the Ethics Committee of Shanghai Jiaotong
University Affiliated First People’s Hospital. Informed written
consent was obtained from each patient enrolled in our study
according to the Declaration of Helsinki. Colon cancer cell lines
were obtained from the American Type Culture Collection and
cultured in Dulbecco’s modified Eagle’s medium (DMEM), supplemented
with 10% fetal bovine serum (FBS) in a humidified atmosphere
containing 5% CO2 at 37°C.
RNA isolation and quantitative RT-PCR for
assessment of mRNA expression
Total RNAs from all samples were isolated with
AllPrep DNA/RNA Mini kit (Qiagen, Hilden, Germany). After the RNA
integrity and purity were checked, the first single-stranded cDNAs
were synthesized from 3 μg of RNA, according to the manufacturer’s
instructions (Fermentas, Shenzhen, China). One microliter of the
cDNAs from each sample was used as a template for qRT-PCR analysis
with the ABsolute qPCR SYBR-Green Mix (Fermentas), employing the
ABI Prism 7900 system (Applied Biosystems Inc., Foster City, CA,
USA).
The following primers were used for qRT-PCR:
HOTAIR sense 5′-GGTAGAAAAAGCAACCACGAAGC-3′ and antisense
5′-ACATAAACCTCTGTCTGTGAGTGCC-3′; GAPDH sense
5′-GGAGCGAGATCCCTCCAAAAT-3′ and antisense
5′-GGCTGTTGTCATACTTCTCATGG-3′.
The expression level of HOTAIR was normalized
to the transcription level of GAPDH. The quantitative PCR reaction
for each sample was repeated in triplicate. The relative
HOTAIR expression was calculated using the 2−ΔΔCt
comparative method.
RNA interference
Two colon cancer cell lines were transfected with 50
nM positive siRNAs (si-HOTAIR 5′-GAACGGGAG UACAGAGAGAUU-3′)
or negative control siRNA (Mock 5′-CUACAACAGCCACAACGUCdTdT-3′)
employing Lipofectamine RNAiMAX (Life Technologies, Carlsbad, CA,
USA) following the manufacturer’s instructions. Total RNA isolation
was performed 72 h later for real-time PCR analysis, while other
functional assays were carried out 48 h post-transfection. All RNA
interference assays were carried out in triplicates
independently.
Cell proliferation assay
For cell proliferation assay, cells were seeded in
96-well plates (2×103 cells/well) in triplicate. At the
appropriate time (24, 48, 72, 96 and 120 h), the cells were
incubated with MTT (5 mg/ml; Sigma, St. Louis, MO, USA) for 4 h at
37°C. MTT solution was then carefully removed and replaced with
dimethyl sulfoxide (DMSO; Sigma). Absorbance was measured at a
wavelength of 570 nm.
Cell migration and invasion assays
Cell migration and invasion assays were carried out
using Transwell chambers (micropore size, 8 μm, 24-well; BD
Biosciences, Franklin Lakes, NJ, USA) without Matrigel (for
migration assay) or with Matrigel (for invasion assay). Both were
performed according to the manufacturer’s protocol. Briefly, the
treated cells were plated in the upper chamber at a concentration
of 5×104 in 500 μl FBS-free media. The bottom chambers
maintained suitable media with 10% FBS for 48 h. Then, the bottom
of the chamber insert was fixed and stained with Giemsa solution.
The stained cells were counted under a microscope at ×200
magnification; meanwhile, images were captured and saved. The
experiments were carried out in triplicate.
Western blot analysis of protein
expression
The total protein was extracted from cells using
Thermo Scientific Pierce RIPA buffer combined with Thermo
Scientific Halt proteinase inhibitor cocktail and then quantified
using Thermo Scientific BCA protein assay kit (Thermo Fisher
Scientific, Waltham, MA, USA). The equivalent amounts of protein
(20 μg) for each sample were electrophoresed on 10% sodium dodecyl
sulfate-polyacrylamide gels and transferred onto PVDF membranes
(Millipore, Billerica, MA, USA). Membranes were blocked using 5%
non-fat milk solution for 1 h, and then probed with primary
polyclonal antibodies at 4°C overnight, followed by secondary
antibodies (1:5,000; Santa Cruz Biotechnology, Santa Cruz, CA, USA)
for 1 h at room temperature. β-actin was used as a loading control.
After washing with TBST (Tris-buffered saline with 0.01% Tween-20),
the proteins were detected by enhanced electrochemiluminescence
(Millipore) and radiography. The primary antibodies used above were
anti-E-cadherin (1:1,000; Abcam, Cambridge, UK) and anti-Vimentin
(1:1,000; Abcam), anti-MMP9 (1:500; Cell Signaling Technology,
Boston, MA, USA) and anti-β-actin antibody (1:1,000; Cell Signaling
Technology).
Statistical analysis
All statistical analyses were set with a
significance level of P≤0.05 and carried out using SPSS 19.0
statistical software (SPSS Inc., Chicago, IL, USA). The differences
between 2 groups were analyzed with the Student’s t-test and
χ2 test, as appropriate. The relationship between
HOTAIR expression and clinicopathological characteristics
was estimated with the Mann-Whitney U test and the Kruskal-Wallis
test appropriately. Survival curves were calculated by Kaplan-Meier
method, with the log-rank test employed for the comparison of
differences. Variables estimated with significant means by
univariate analysis were subjected to the multivariate Cox
proportional hazards regression model.
Results
Overexpression of HOTAIR in colon cancer
tissues
qPCR was applied to examine the expression level of
HOTAIR in 120 pairs of cancer and matched normal tissues,
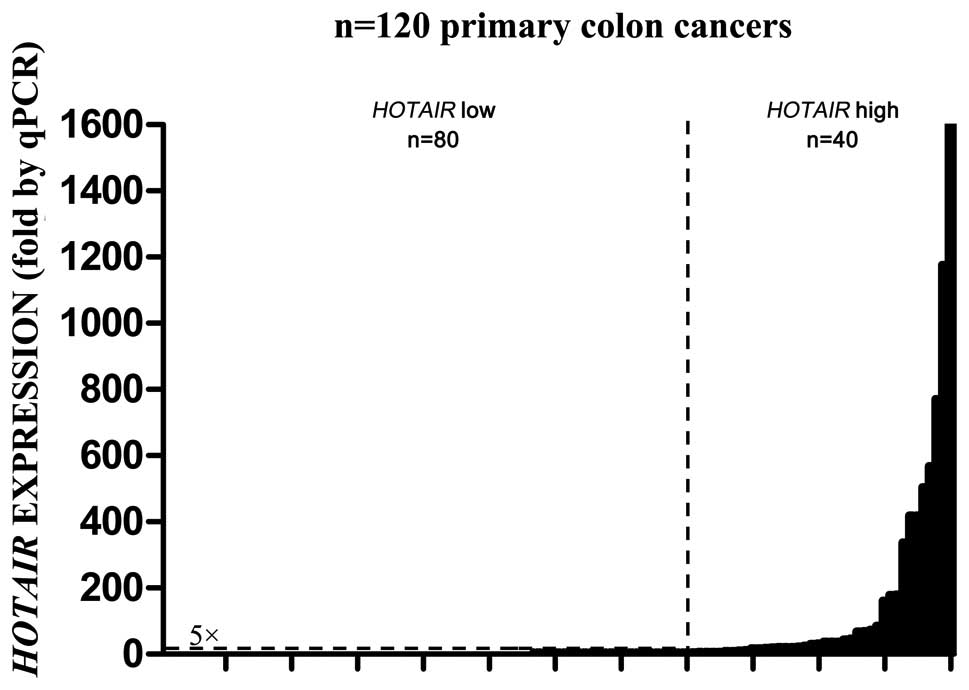
and 32 metastasized lymph node tissues. As shown in Fig. 1, HOTAIR was overexpressed
from 2- to nearly 1,600-fold in the cancer tissues. According to a
HOTAIR expression level of >5-fold compared to normal
tissues, 120 patients were divided into a high expression group
(n=40) and a low expression group (n=80). In addition, 27 of 32
(84.4%) lymph node metastasis tissues showed high expression of
HOTAIR, with significant means between primary cancer
tissues and metastatic lymph node tissues (Table I).
 | Table IExpression of HOTAIR in colon
cancer tissues and lymph node metastatic tissues. |
Table I
Expression of HOTAIR in colon
cancer tissues and lymph node metastatic tissues.
| Tissue sample | n | Expression of
HOTAIR | P-value |
|---|
|
|---|
Low
n (%) | High
n (%) |
|---|
| Cancer tissue | 120 | 80 (66.7) | 40 (33.3) | <0.001a |
| LNM tissue | 32 | 5 (15.6) | 27 (84.4) | |
Correlation between HOTAIR expression and
clinicopathological characteristics
The correlation between HOTAIR expression and
clinicopathological characteristics is summarized in Tables II and III. Increased HOTAIR expression
was significantly associated with AJCC stage (P<0.001), lymph
node metastasis (pN stage, P=0.002), organ metastasis (pM stage,
P<0.001), vascular invasion (P=0.019) and histologic
differentiation (P<0.001). No significant correlations were
found between HOTAIR expression and age, gender or tumor
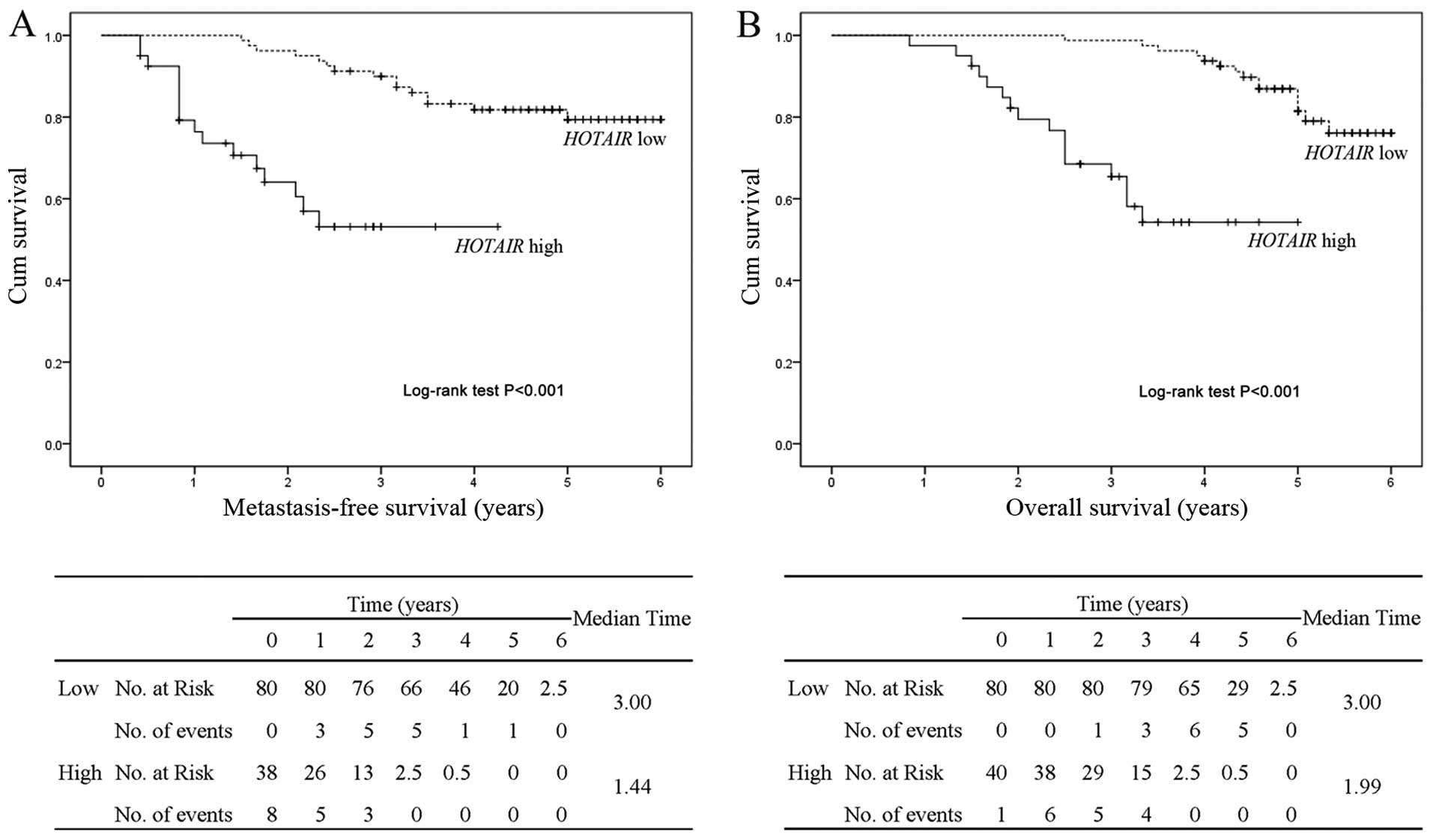
location. The Kaplan-Meier curves (Fig.
2) revealed that patients in the high expression group had a
significant poorer clinical prognosis than those in the low
expression group, both for metastasis-free survival and overall
survival (log-rank test P<0.001). With regard to Cox univariate
proportional hazards regression model, the level of HOTAIR
expression, lymph node metastasis, organ metastasis, AJCC stage,
degree of differentiation and vascular invasion were prognostic
factors. Additionally, multivariate analysis, including the
variables above with P<0.05, indicated that HOTAIR
expression was an independent clinical risk indicator for both
metastasis-free survival (HR, 3.878; 95% CI, 1.369–10.981; P=0.011)
and overall survival (HR, 3.915; 95% CI, 1.226–12.499;
P=0.021).
 | Table IICorrelation between HOTAIR
expression and clinicopathological characteristics in the colon
cancer cases. |
Table II
Correlation between HOTAIR
expression and clinicopathological characteristics in the colon
cancer cases.
| HOTAIR
expression | |
|---|
|
| |
|---|
| Low (n=80) | High (n=40) | P-value |
|---|
| Age (years) |
| <65 | 28 | 17 | 0.424 |
| ≥65 | 52 | 23 | |
| Gender |
| Male | 40 | 24 | 0.301 |
| Female | 40 | 16 | |
| Location |
| Right | 20 | 11 | 0.768 |
| Others | 60 | 29 | |
| AJCC stage |
| I | 12 | 1 | <0.001a |
| II | 37 | 7 | |
| III | 26 | 23 | |
| IV | 5 | 9 | |
| pT stage |
| T1 | 3 | 0 | 0.226 |
| T2 | 17 | 4 | |
| T3 | 41 | 26 | |
| T4 | 19 | 10 | |
| pN stage |
| N0 | 48 | 12 | 0.002a |
| N1 | 25 | 17 | |
| N2 | 7 | 11 | |
| pM stage |
| M0 | 78 | 28 | <0.001a |
| M1 | 2 | 12 | |
|
Differentiation |
| Well | 11 | 3 | 0.001a |
| Moderate | 61 | 21 | |
| Poor | 8 | 16 | |
| Vascular
invasion |
| Yes | 6 | 9 | 0.019a |
| No | 74 | 31 | |
 | Table IIIAssociation between clinical
characteristics and metastasis-free and overall survival by Cox
regression model analysis. |
Table III
Association between clinical
characteristics and metastasis-free and overall survival by Cox
regression model analysis.
| Metastasis-free
survival | Overall
survival |
|---|
|
|
|
|---|
| Univariate | Multivariate | Univariate | Multivariate |
|---|
|
|
|
|
|
|---|
| HR (95% CI) | P-value | HR (95% CI) | P-value | HR (95% CI) | P-value | HR (95% CI) | P-value |
|---|
| HOTAIR
expression |
| Low | - | | - | | - | | | |
| High | 6.330
(2.610–15.348) |
<0.001a | 3.878
(1.369–10.981) |
0.011a | 8.147
(3.466–9.152) |
<0.001a | 3.915
(1.226–12.499) |
0.021a |
| Age (years) |
| <65 | - | | | | - | | | |
| ≥65 | 0.803
(0.385–1.675) | 0.559 | | | 0.977
(0.487–1.961) | 0.948 | | |
| Gender |
| Male | - | | | | - | | | |
| Female | 1.492
(0.693–3.213) | 0.307 | | | 1.048
(0.514–2.135) | 0.898 | | |
| Location |
| Right | - | | | | - | | | |
| Other | 0.596
(0.280–1.266) | 0.178 | | | 0.696
(0.338–1.432) | 0.325 | | |
| pT stage |
| T1 | - | | | | - | | | |
| T2 | 1.146
(0.101–12.970) | 0.912 | | | 1.169
(0.104–13.150) | 0.900 | | |
| T3 | 1.137
(0.319–9.333) | 0.905 | | | 1.085
(0.132–8.939) | 0.940 | | |
| T4 | 4.167
(0.560–30.992) | 0.163 | | | 4.383
(0.588–32.663) | 0.149 | | |
| pN stage |
| N0 | - | | - | | - | | - | |
| N1 | 3.172
(1.288–7.812) |
0.012a | 2.885
(1.147–7.255) |
0.024a | 3.270
(1.315–8.130) |
0.011a | 2.626
(0.995–6.931) | 0.051 |
| N2 | 10.485
(4.046–27.170) |
<0.001a | 7.878
(2.903–21.384) |
<0.001a | 11.135
(4.227–29.331) |
<0.001a | 6.982
(2.365–20.614) |
<0.001a |
| pM stage |
| N0 | - | | - | | - | | - | |
| M1 | 8.386
(12.739–25.670) |
<0.001a | 3.580
(1.121–11.437) |
0.031a | 6.935
(2.365–10.864) |
<0.001a | 2.856
(0.745–10.945) | 0.126 |
| AJCC |
| N0 | - | | - | | - | | - | |
| III-IV | 4.644
(2.060–10.466) |
<0.001a | 5.331
(0.959–29.636) | 0.056 | 4.782
(2.105–10.864) |
<0.001a | 7.571
(1.330–43.103) |
0.023a |
|
Differentiation |
| N0 | - | | - | | - | | - | |
| Moderate | 1.930
(0.814–4.576) | 0.135 | 1.570
(0.667–3.699) | 0.302 | 1.681
(0.709–3.985) | 0.239 | 1.489
(0.591–3.751) | 0.399 |
| Poor | 4.094
(1.565–10.715) |
0.004a | 1.889
(0.677–5.273) | 0.225 | 4.106
(1.568–10.751) |
0.004a | 2.234
(0.761–6.556) | 0.143 |
| Vascular
invasion |
| No | - | | | | - | | | |
| Yes | 2.282
(0.531–9.809) | 0.267 | | | 2.124
(0.501–9.010) | 0.307 | | |
HOTAIR promotes migration and invasion of
colon cancer cells
As reported by other research groups, HOTAIR
may play a pro-oncogenic role in certain types of cancers such as
breast (5), hepatocellular
(6), gastrointestinal (8), pancreatic (9) and lung cancer (11). To examine the effect of
HOTAIR on colon cancer, a series of functional experiments
concerning cell proliferation and cell metastasis was performed. We
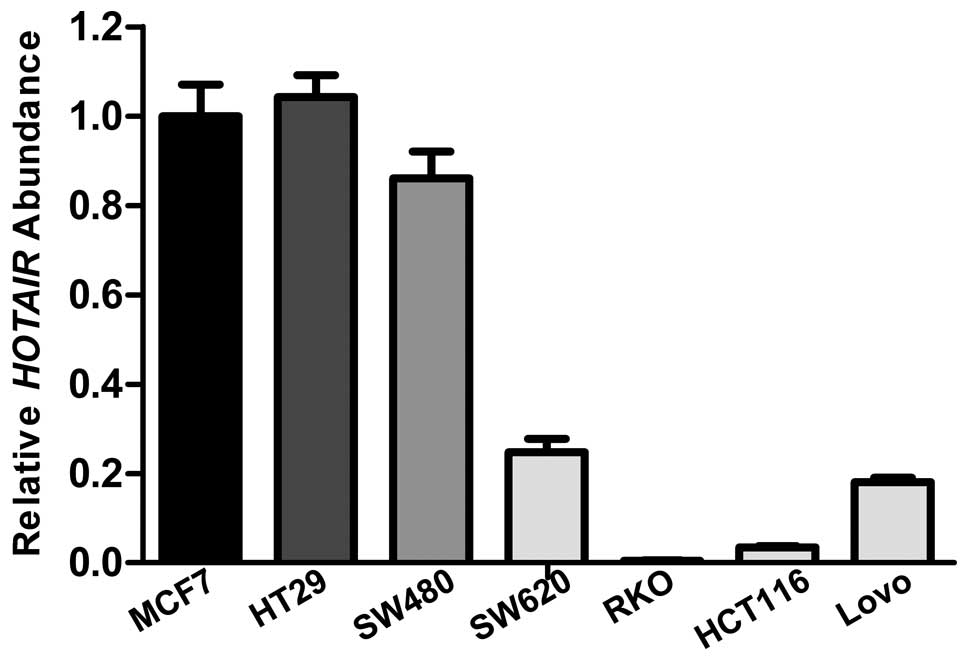
quantified the expression levels of HOTAIR in the colon
cancer cell lines by qRT-PCR, and compared the levels to the level
in MCF-7, a breast cancer cell line that overexpresses endogenous
HOTAIR (Fig. 3).
HOTAIR expression levels in colon cancer cell lines SW480
and HT29 were higher than that in the other 4 cell lines.
HOTAIR knockdown was carried out by using small interfering
RNA transfection in the cell lines. The efficiency of
siRNA-mediated knockdown in the SW480 or HT29 cells was >60%
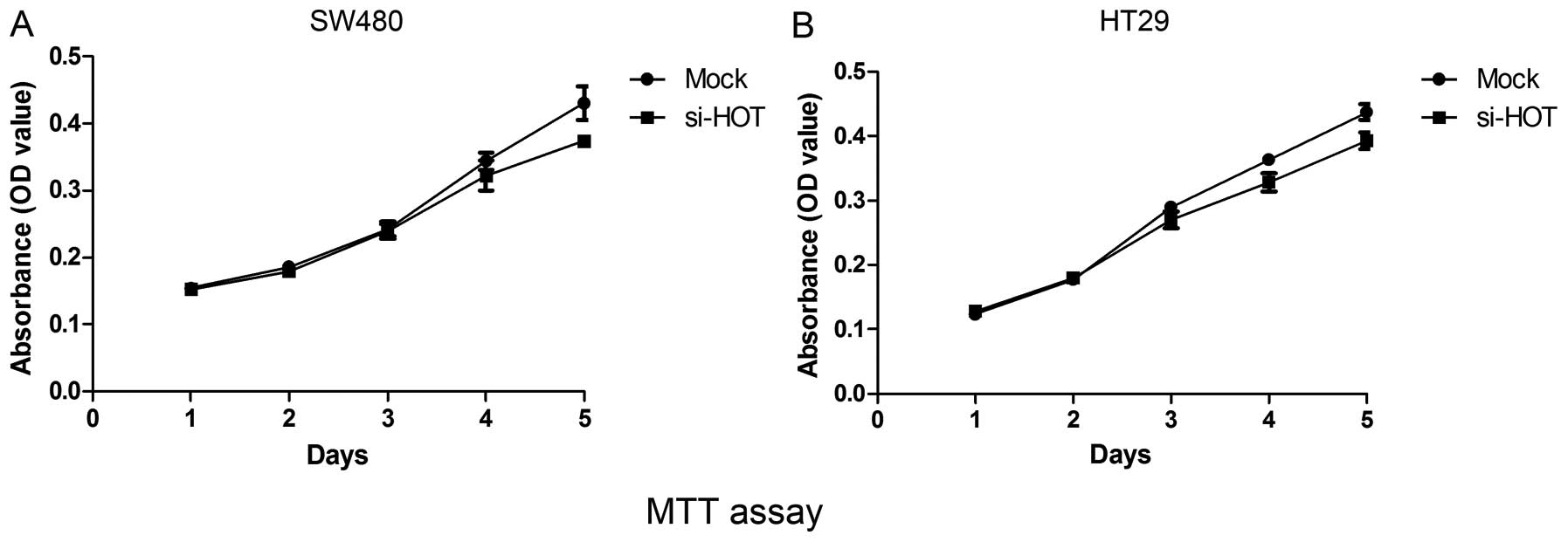
(data not shown). As illustrated by Fig. 4, results from the MTT assay showed
that the proliferation rate of cells depleted of HOTAIR was
slightly slower than the rate of cells treated with mock siRNA
throughout 5 continuous days.
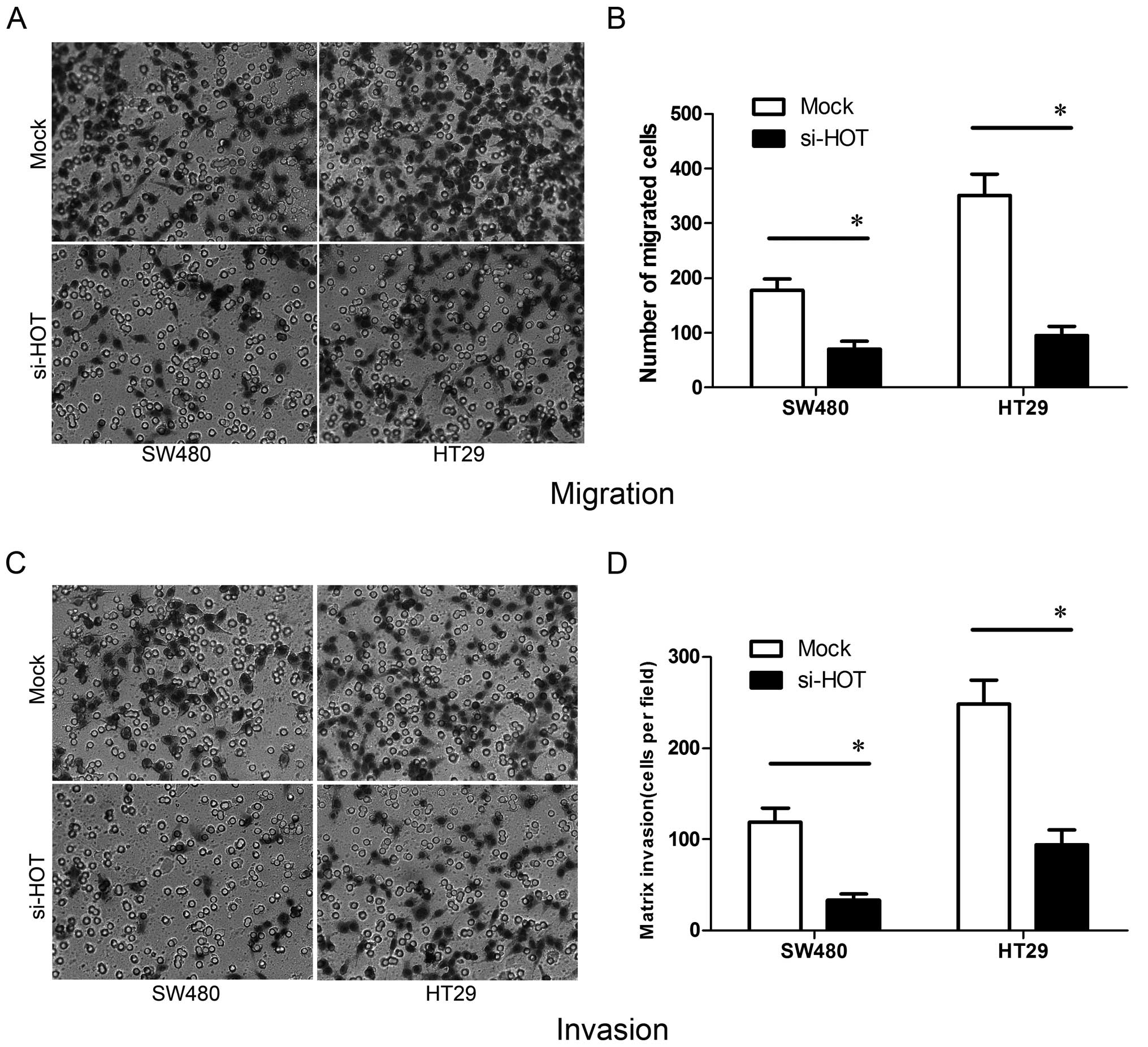
We next ascertained whether manipulation of the
HOTAIR level affects cell metastasis in vitro.
Migration and invasiveness of tumor cells are key aspects of
metastasis. In the Transwell migration assay, fewer SW480 and HT29
colon cancer cells treated by si-HOTAIR migrated to the 8-μm
pores of the Transwell chambers when compared to the cells treated
with Mock (Fig. 5A and B).
Furthermore, similar results were obtained in the invasion assay
using a Matrigel environment study (Fig. 5C and D). All of these studies were
repeated three times and the differences were statistically
significant (P<0.05). Based on these outcomes, it was suggested
that HOTAIR is involved in the progression of colon cancer
mainly by promoting cancer cell migration and invasiveness.
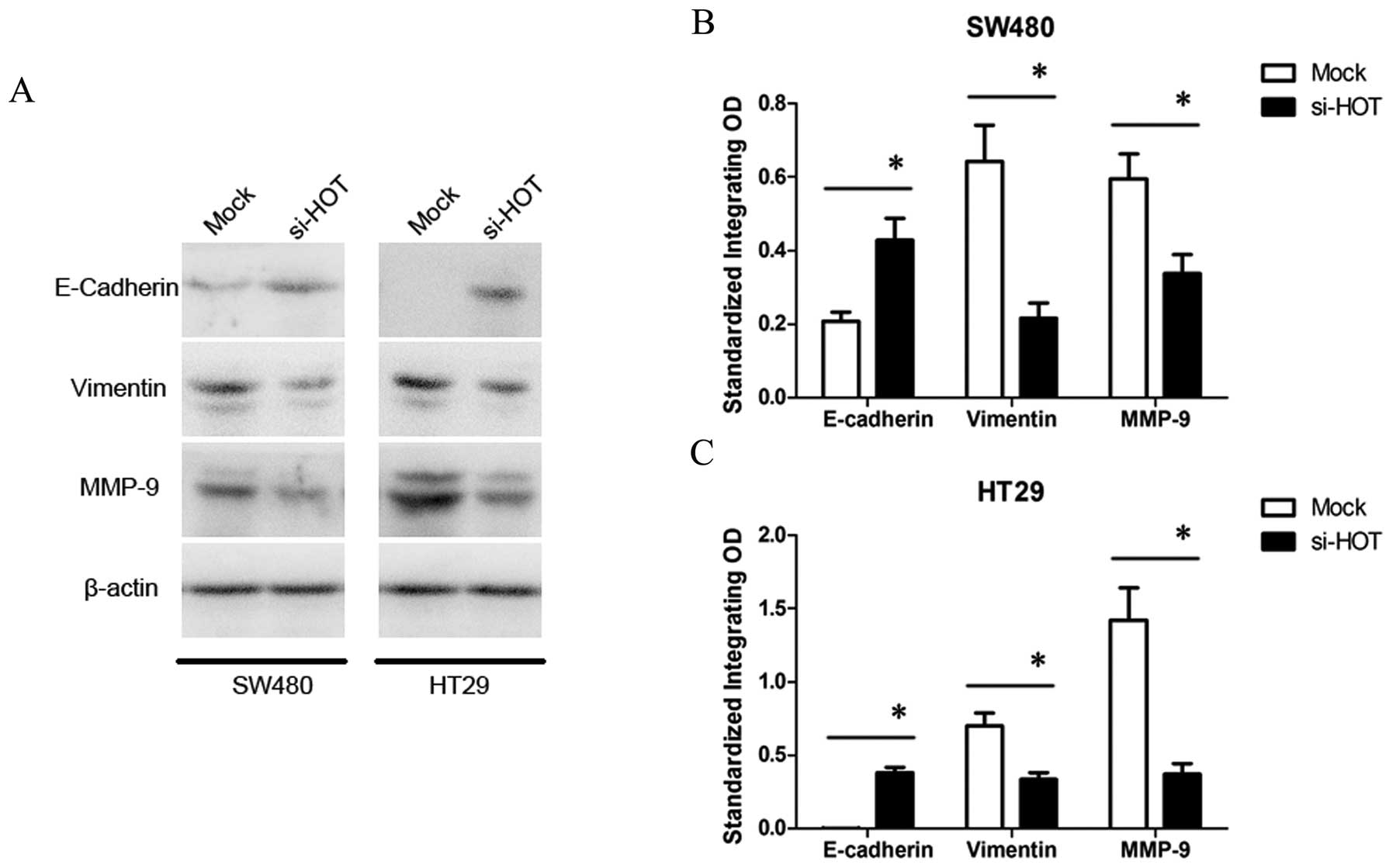
HOTAIR regulates expression of
E-cadherin, vimentin and MMP9
Epithelial-mesenchymal transition (EMT) with
alterations in gene expression profiles of cells initiates the
invasion-metastasis cascade. Therefore, we ascertained whether
HOTAIR promotes EMT to regulate tumor progression. Several
key genes involved in EMT of colon cancer were selected for further
western blot analysis (Fig. 6).
When SW480 and HT29 cells were treated by HOTAIR knockdown,
expression of E-cadherin, a hallmark of epithelial cell was
increased, while expression of vimentin, a mesenchymal cell marker,
was decreased. Additionally, in the si-HOTAIR-treated cells,
expression of matrix metalloproteinase 9 (MMP9), which functions as
an important proteinase to degrade the extracellular matrix for
facilitating cell motility, was inhibited as well. These data
indicate that the regulation of EMT is partly involved in the
mechanism of HOTAIR-induced metastasis.
Discussion
HOTAIR lncRNA, with a 2158-nucleotide
transcript, is transcribed from the HOXC locus in an antisense
manner, which is also the reason why it is termed HOXC
antisense intergenic RNA (14).
HOTAIR has very high nucleotide conservation in vertebrates
and an anatomic conservative expression pattern from embryo to
adulthood. Moreover, the lncRNA is selectively required to
physically interact with polycomb repressive complex 2 (PRC2) to
regulate the chromatin methylation state. PRC2, comprising H3K27
histone methyl transferase EZH2, SUZ2 and EED, is involved in stem
cell pluripotency and cancer progression (16–18).
In 2010, HOTAIR was reported as a powerful predictor of
breast cancer metastasis and poor prognosis. Gupta and his
coworkers (5)found a positive
association among high expression pattern of HOTAIR, breast
cancer malignant biologic characteristics and reduced survival. The
present study showed that the level of HOTAIR lncRNA
expression in tumor tissues was distinctly higher than the level in
paired normal mucosa. Additionally, HOTAIR was more highly
expressed in invaded lymph nodes than in primary cancer tissues.
Increased HOTAIR expression was significantly correlated
with the depth of tumor invasion, lymph node metastasis, organ
metastasis, histological differentiation, vascular invasion and
advanced tumor stage. Patients with high HOTAIR expression
had higher recurrence rates and poorer metastasis-free and overall
survival rates than patients with low HOTAIR expression.
Multivariate analyses revealed that HOTAIR expression was an
independent factor for colon cancer prognosis after surgery. These
results indicated that HOTAIR also could be a meaningful
predictor for colon cancer management.
The 5′ domain of HOTAIR binds to PRC2 and is
required for histone H3 lysine 27 trimethylation in order to
silence HOXD and other target genes (5,14). On
the other hand, the 3′ domain of HOTAIR maintains
LSD1-binding activity (15). LSD1,
the first identified demethylase that specifically catalyzes the
demethylation of histone H3 lysine 4, is believed to be linked to
the transcriptional silencing of tumor-suppressor genes (19–22).
Tsai et al (15) suggested
that HOTAIR, as a modular scaffold, is required to target
both PRC2 and LSD1 to many promoter elements of genes in order to
coordinate chromatin modification for gene repression. Loss of
HOTAIR notably weakens the activities of PRC2 and LSD1 in
modifying the histone state. Therefore, it is reasonable to expect
that HOTAIR plays a crucial role in cancer development other
than a merely novel cancer biomarker. At present, the study results
revealed that HOTAIR had a limited effect on cell
proliferation. As HOTAIR knockdown using siRNA significantly
reduced the ability of cancer cells to invade the Transwell
membranes, this lncRNA may promote colon cancer cell migration and
invasion, which is mainly consistent with previous studies in other
cancer types.
We further addressed how HOTAIR-dependent
gene regulation in colon tumors is involved in the enhancement of
an aggressive biological behavior. The early step leading to
metastasis is migration and invasion of cancer cells from original
tissues into the surrounding stroma. In order to acquire such
abilities, carcinoma cells must inherit a drastic phenotypic
alteration - the epithelial-mesenchymal transition (EMT). EMT is a
biological process that allows polarized epithelial cells to
experience multiple biochemical changes that enable them to assume
a mesenchymal cell phenotype (23).
Upon the phenotype change, cancer cells lose intercellular
junction, breach the basement membrane and acquire enhanced
motility. The complexity of EMT means that only small numbers of
centrally pleiotropic regulators orchestrate the complement, such
as Twist, Slug and Snail and the miR200 family. The present study
demonstrated that depletion of HOTAIR increased expression
of E-cadherin while concomitantly decreasing expression of vimentin
and MMP9. Together with previous studies using global gene
expression analysis, the emerging evidence indicates that increased
expression of HOTAIR in cancer cells appears to reprogram
the chromatin profile of epithelial cells to that of mesenchymal
cells, cooperating with PRC2 and LSD1 at least. Hence,
HOTAIR may be another pleiotropic modulator participating in
EMT.
Collectively, our data indicate that HOTAIR
is a valuable factor for colon cancer prognosis. Moreover,
HOTAIR can promote colon cancer cell migration and
invasiveness and may participate in epithelial-mesenchymal
transition. Further studies are warranted to advance our
understanding of the involvement of HOTAIR in cancer
development, since this lncRNA is a potential target for cancer
prevention and treatment.
Acknowledgements
The present study was supported by funds from the
National Natural Science Foundation of China (81172328), the
Important International Cooperation grants from the National
Natural Science Foundation of China (81220108021), the Medical
Guidance Project of Shanghai Science and Technology Commission
(124119a1700), the Science and Technology Innovation Plan of
Shanghai Science and Technology Commission (11431921000), the
Frontier Technology Union Research Project of Shanghai Municipal
Hospitals (SHDC12012105), and the Medical Climbing Project from
Songjiang Health Bureau of Shanghai (2011PD03).
References
|
1
|
NCCN Clinical Practice Guidelines in
Oncology for Colon Cancer (Version 3). 2013, [EB/OL] Available at
www.nccn.org/professionals/physician_gls/f_guidelines.asp.
|
|
2
|
Li M and Gu J: Changing patterns of
colorectal cancer in China over a period of 20 years. World J
Gastroenterol. 11:4685–4688. 2005.PubMed/NCBI
|
|
3
|
Fearon ER: Molecular genetics of
colorectal cancer. Annu Rev Pathol. 6:479–507. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ulitsky I and Bartel DP: lincRNAs:
genomics, evolution, and mechanisms. Cell. 154:26–46. 2013.
View Article : Google Scholar
|
|
5
|
Gupta RA, Shah N, Wang KC, et al: Long
non-coding RNA HOTAIR reprograms chromatin state to promote
cancer metastasis. Nature. 464:1071–1076. 2010.PubMed/NCBI
|
|
6
|
Yang Z, Zhou L, Wu LM, et al:
Overexpression of long non-coding RNA HOTAIR predicts tumor
recurrence in hepatocellular carcinoma patients following liver
transplantation. Ann Surg Oncol. 18:1243–1250. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lu L, Zhu G, Zhang C, et al: Association
of large noncoding RNA HOTAIR expression and its downstream
intergenic CpG island methylation with survival in breast cancer.
Breast Cancer Res Treat. 136:875–883. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Niinuma T, Suzuki H, Nojima M, et al:
Upregulation of miR-196a and HOTAIR drive malignant
character in gastrointestinal stromal tumors. Cancer Res.
72:1126–1136. 2012.PubMed/NCBI
|
|
9
|
Kim K, Jutooru I, Chadalapaka G, et al:
HOTAIR is a negative prognostic factor and exhibits pro-oncogenic
activity in pancreatic cancer. Oncogene. 32:1616–1625. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li D, Feng J, Wu T, et al: Long intergenic
noncoding RNA HOTAIR is overexpressed and regulates PTEN
methylation in laryngeal squamous cell carcinoma. Am J Pathol.
182:64–70. 2013. View Article : Google Scholar
|
|
11
|
Nakagawa T, Endo H, Yokoyama M, et al:
Large noncoding RNA HOTAIR enhances aggressive biological behavior
and is associated with short disease-free survival in human
non-small cell lung cancer. Biochem Biophys Res Commun.
436:319–324. 2013. View Article : Google Scholar
|
|
12
|
Zhuang Y, Wang X, Nguyen HT, et al:
Induction of long intergenic non-coding RNA HOTAIR in lung cancer
cells by type I collagen. J Hematol Oncol. 6:352013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kogo R, Shimamura T, Mimori K, et al: Long
noncoding RNA HOTAIR regulates polycomb-dependent chromatin
modification and is associated with poor prognosis in colorectal
cancers. Cancer Res. 71:6320–6326. 2011.PubMed/NCBI
|
|
14
|
Rinn JL, Kertesz M, Wang JK, et al:
Functional demarcation of active and silent chromatin domains in
human HOX loci by noncoding RNAs. Cell. 129:1311–1323. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Tsai MC, Manor O, Wan Y, et al: Long
noncoding RNA as modular scaffold of histone modification
complexes. Science. 329:689–693. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sparmann A and van Lohuizen M: Polycomb
silencers control cell fate, development and cancer. Nat Rev
Cancer. 6:846–856. 2006. View
Article : Google Scholar : PubMed/NCBI
|
|
17
|
Richly H, Aloia L and Di Croce L: Roles of
the Polycomb group proteins in stem cells and cancer. Cell Death
Dis. 2:e2042011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Crea F, Fornaro L, Bocci G, et al: EZH2
inhibition: targeting the crossroad of tumor invasion and
angiogenesis. Cancer Metastasis Rev. 31:753–761. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ding J, Zhang ZM, Xia Y, et al:
LSD1-mediated epigenetic modification contributes to proliferation
and metastasis of colon cancer. Br J Cancer. 109:994–1003. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Huang Z, Li S, Song W, et al:
Lysine-specific demethylase 1 (LSD1/KDM1A) contributes to
colorectal tumorigenesis via activation of the Wnt/β-catenin
pathway by down-regulating Dickkopf-1 (DKK1). PLoS One.
8:e700772013.PubMed/NCBI
|
|
21
|
Tang M, Shen H, Jin Y, et al: The
malignant brain tumor (MBT) domain protein SFMBT1 is an integral
histone reader subunit of the LSD1 demethylase complex for
chromatin association and epithelial-to-mesenchymal transition. J
Biol Chem. 288:27680–27691. 2013. View Article : Google Scholar
|
|
22
|
Abdel-Wahab O and Dey A: The ASXL-BAP1
axis: new factors in myelopoiesis, cancer and epigenetics.
Leukemia. 27:10–15. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kalluri R and Weinberg RA: The basics of
epithelial-mesenchymal transition. J Clin Invest. 119:1420–1428.
2009. View
Article : Google Scholar : PubMed/NCBI
|