Introduction
Esophageal cancer is the eighth most common cancer
worldwide and the sixth most common cause of cancer-related
mortality (1,2). Although Barrett’s adenocarcinoma is
the most rapidly increasing cancer in Western countries, esophageal
squamous cell carcinoma (ESCC) remains dominant in East Asia
(3). Often diagnosed at later
stages, the prognosis of affected patients is unsatisfactory
despite the development of therapeutic options such as surgery,
chemotherapy and radiotherapy (4).
Consequently, there is a great need for ESCC biomarkers to allow a
tailored multimodal approach with increased efficacy. Nevertheless,
efforts to identify molecular markers in association with the
pathogenesis of ESCC have, to date, been unsuccessful (5).
Mature microRNAs (miRNAs) are a class of small,
well-conserved, non-coding RNA molecules that silence gene
expression usually by interfering with mRNA stability or protein
translation (6,7). The function is performed by
identifying 3′-UTRs (untranslated regions) of target mRNAs with
conserved complementarities to the seed (nucleotides 2–7) of the
miRNA. Up to 30% of human protein-coding genes may be regulated by
miRNAs (8,9). miRNAs are involved in biological and
pathological processes including cell differentiation,
proliferation, apoptosis and metabolism, and they are emerging as
highly tissue-specific biomarkers with potential clinical
application for defining cancer types and origins. miRNAs can
either function as oncogenes or tumor suppressors (10,11).
It has been reported that miR-126 is located on
chromosome 9q34.3 within the host gene encoding for epidermal
growth factor like-7 (EGFL-7), an endothelial cell-derived,
secreted inhibitor of smooth muscle cell migration and a regulator
of blood vessel formation (12,13).
Vasculature is required for the expansion of tumor masses, as
inhibition of new vessel formation prevents tumor growth. Studies
have shown that the expression of miR-126 is downregulated in
cancers such as non-small cell lung cancer and colon cancer
(14,15), suggesting that the downregulation of
miR-126 is significantly related to the occurrence and development
of cancer. However, studies have also reported that aberrant
overexpression of miR-126 contributes to carcinogenesis, suggesting
that miR-126 acts as an oncogene in gastric cancer and oral
squamous cell carcinoma (16,17).
Therefore, miR-126 may have diverse roles in tumor carcinogenesis.
To date, some genes have been identified as miR-126 target genes,
including sex determining region Y-box 2 (SOX2), Crk,
vascular endothelial growth factor (VEGF), matrix
metalloproteinase-9 (MMP-9), insulin receptor substrate-1
(IRS-1), SLC7A5, TOM1, and
CD97/G-protein-coupled receptor (GPCR) (16,18–21).
Bioinformatics has also shown that the 3′-UTR of genes such as
IRS-1, FBXO33, PTPN9, PLK2, and Golgi phosphoprotein 3
(GOLPH3) contains a putative binding site for miR-126.
However, the functional role and mechanism of
miR-126 in regulating tumorigenesis of ESCC is still not fully
understood. To identify miRNAs which can be specifically expressed
and exert distinct biological actions in ESCC, we investigated
miR-126 expression and its functional role in ESCC. miR-126
expression was measured in ESCC tissue specimens and paired
paracancer tissues, and the relationship between the expression
level of miR-126 and the clinicopathological characteristics were
analyzed. Further functional studies of miR-126 showed that it
could inhibit ESCC cell growth, migration and invasion. IRS-1 and
GOLPH3 are potential downstream targets of miR-126 in ESCC.
Materials and methods
Tissue specimens and cancer cell
lines
Specimens of 102 ESCC tissues and paired paracancer
tissues were collected from Henan Cancer Hospital between July 2012
and July 2013. Clinicopathological data of patients were collected.
None of the patients received preoperative antitumor therapy. All
specimens were collected using two methods: one was snap frozen in
liquid nitrogen immediately after surgery and stored at −80°C for
further miRNA detection, the other was formalin-fixed for
histopathological assay. The clinicopathological information of the
ESCC patients is shown in Table I.
All patients provided informed consent under a protocol reviewed
and approved by the Institutional Review Board, Henan Cancer
Hospital.
 | Table IExpression of miR-126, IRS-1 protein
and GOLPH3 protein in ESCC patients. |
Table I
Expression of miR-126, IRS-1 protein
and GOLPH3 protein in ESCC patients.
| Variable | Total | miR-126 high | P-value | IRS-1-positive | P-value |
GOLPH3-positive | P-value |
|---|
| Age (years) | | | 0.298 | | 0.68 | | 0.833 |
| <60 | 45 | 13 | | 32 | | 14 | |
| ≥60 | 57 | 23 | | 34 | | 20 | |
| Gender | | | 0.513 | | 0.275 | | 0.408 |
| Male | 69 | 26 | | 43 | | 22 | |
| Female | 33 | 10 | | 23 | | 12 | |
| Tumor location | | | 0.413 | | 0.788 | | 0.582 |
| Upper | 15 | 4 | | 11 | | 4 | |
| Middle | 68 | 23 | | 45 | | 25 | |
| Lower | 19 | 9 | | 10 | | 5 | |
|
Differentiation | | | 0.017 | | 0.006 | | 0.462 |
| Well | 29 | 10 | | 19 | | 7 | |
| Moderate | 24 | 14 | | 10 | | 9 | |
| Poor | 49 | 12 | | 37 | | 18 | |
| Smoking | | | 0.539 | | 0.539 | | 0.294 |
| Yes | 44 | 14 | | 30 | | 12 | |
| No | 58 | 22 | | 36 | | 22 | |
| Alcoholic | | | 0.477 | | 0.477 | | 0.814 |
| Yes | 26 | 11 | | 15 | | 8 | |
| No | 76 | 25 | | 51 | | 26 | |
| Lymph node
metastasis | | | 0.019 | | 0.289 | | 0.01 |
| Yes | 39 | 8 | | 31 | | 7 | |
| No | 63 | 28 | | 35 | | 27 | |
| Invasion depth | | | 0.009 | | 0.515 | | <0.01 |
| T1 | 12 | 9 | | 3 | | 11 | |
| T2 | 34 | 11 | | 23 | | 12 | |
| T3,4 | 56 | 16 | | 40 | | 11 | |
| TNM stage | | | 0.03 | | 0.406 | | <0.01 |
| I | 10 | 7 | | 3 | | 9 | |
| II | 59 | 21 | | 38 | | 18 | |
| III | 33 | 8 | | 25 | | 7 | |
The human ESCC cell line Eca9706 was purchased from
the cell bank of the tumor hospital of the Chinese Academy of
Medical Sciences/Biological Detection Center (Beijing, China). The
cells were cultured in RPMI-1640 (Thermo, Waltham MA, USA)
supplemented with 10% fetal bovine serum (FBS; Thermo) and
penicillin/streptomycin in a 5% CO2 humidified incubator
at 37°C (normal conditions for cell culture).
RNA isolation and real-time reverse
transcription polymerase chain reaction (qRT-PCR)
Total-RNA in cancer tissues or cell lines was
isolated using QIAzol lysis reagent and miRNeasy mini kit (Qiagen,
Düsseldorf, Germany), according to the manufacturer’s instructions.
Total-RNA was isolated 48 h after cell transfection. Quantitation
of total-RNA was carried out using spectrophotometry. All samples
had an excellent 260/280 ratio. RNAs were reverse-transcripted into
cDNAs using miScript Reverse Transcription kit (Qiagen) and stored
at −20°C for immediate or further use. The qRT-PCR of miRNA and
mRNA expression was detected using miScript SYBR-Green PCR kit
(Qiagen) on an ABI 7500 detector (Applied Biosystems). RUN6 was
used as internal control according to the protocol. miR-126
miScript Primer assay (no. MS00003430), RNU6 miScript Primer assay
(no. MS00033740), IRS-1 QuantiTect Primer assay (QT00074144),
GOLPH3 QuantiTect Primer assay (no. QT00061320), and β-actin
QuantiTect Primer assay (no. QT00095431) for miRNA or mRNA
detection were commercially provided by Qiagen. Relative gene
expression determinations were made using the comparative ΔΔCt
method (2−ΔΔCt). Patients were divided into two groups
(high or low miR-126 expression) for further analysis. The high
miR-126 expression group was defined as an expression level of
miR-126 in tumor tissue higher than the paired paracancer
tissue.
Online prediction of miRNA targets
To predict potential targets of miR-126, we searched
PicTar (http://www.pictar.org), miRGenTargets
(http://microrna.gr) and TargetScanHuman (http://www.targetscan.org/vert_61/). The 3′-UTR
of IRS-1 mRNA (RefSeq 005544) and GOLPH3 mRNA (RefSeq 022130) may
have putative miRNA-126 binding sites.
Immunohistochemistry
To explore expression of IRS-1 and GOLPH3 proteins
in ESCC, immunohistochemistry experiments were carried out on the
collected 3–5 μm tissue slices. After deparaffinization and antigen
retrieval, slices were incubated at 4°C overnight with rabbit
polyclonal GOLPH3 antibody (1:100; Abcam, Hong Kong), or rabbit
polyclonal IRS-1 antibody (1:50; Santa Cruz Biotechnology, Inc.,
Santa Cruz, CA, USA; sc-559). After washing with phosphate-buffered
saline (PBS) three times, the slides were incubated with
biotinylated secondary antibody (1:80; goat anti-rabbit IgG,
ready-to-use; Dako, Denmark) for 1 h at 37°C, washing with PBS
three times and chromogenic use of 3,3′-diaminobenzidine (DAB) for
5 min. Counterstaining was carried out with hematoxylin. The
immunohistochemically stained tissue sections were reviewed and
scored separately by two pathologists blinded to the clinical
information. Any intensity of cell membrane, endosome or
cytoplasmic staining was considered a positive stain for GOLPH3,
whereas cytoplasmic or nuclear staining was considered positive for
IRS-1. Increased expression of proteins was considered if the
percentage of stained cells was ≥10%.
Transfection of miR-126 mimics or
inhibitor into ESCC cells
Eca9706 cells were seeded into 24-well plates and
transfected when the cells reached 60% confluence.
Syn-hsa-miR-126-3p miScript miRNA Mimic (no. MSY0000445),
anti-hsa-miR-126-3p miScript miRNA Inhibitor (no. MIN0000445), and
miScript miRNA Inhibitor Negative Control (mock sequence, no.
1027271), commercially provided by Qiagen, were transfected into
Eca9706 cells. siRNAs were allowed to form transfection complexes
with HiPerFect Transfection Reagent (Qiagen) in serum-free
Opti-MEM, according to the manufacturer’s protocol. All groups were
performed in triplicate. Cells were collected for RNA or protein
assay at different time points after transfection.
For the invasion and apoptosis assays, cells were
seeded in 6-well plates and the transfected cells were suspended
using tyrosine; a hemocytometer was used for cell counting.
Western blot analysis
Transfected cells were lysed in RIPA buffer (Sigma,
St. Louis, MO, USA) with protease inhibitors. Proteins were
resolved in 10% SDS-PAGE gel and electro transferred onto a PVDF
membrane, blocked with 5% milk, and then the membrane was incubated
with a primary antibody at 4°C overnight (rabbit polyclonal GOLPH3
antibody, 1:500; Abcam, or rabbit polyclonal IRS-1 antibody; 1:500;
Santa Cruz). After washing with Tris-buffered saline 3 times,
corresponding horseradish peroxidase-conjugated secondary antibody
(1:2,000; goat anti-rabbit IgG; Dako) was used to incubate the
membrane at room temperature for 2 h. The membrane was then washed
3 times and the blots were visualized using Western Lightning Ultra
(PerkinElmer, Waltham, MA, USA). Bands were quantified using
FluorChem FC3 Software® (ProteinSimple, San Francisco,
CA, USA) for band intensity and β-actin was used as internal
control.
Cell proliferation assay
The cells transfected with 50 nM miR-126 mimics,
inhibitors or mock sequence were initially seeded at a density of
5.0×103 cells/well in 96-well plates with siRNAs. After
6–8 h, non-adherent cells were removed by washing with PBS. The
remaining adherent cells were cultured under normal conditions, and
this time point was set as the start point. Cell counting kit-8
(CCK-8) analyses were performed at 5, 24, 48, 72 or 96 h for cell
proliferation observation. Absorbance was detected at 490 nm in an
enzyme microplate reader (Wellscan MK-3; Labsystems Dragon,
Finland).
Cell apoptosis assay
Cells were suspended in 0.25% trypsin without EDTA
and harvested at 48 h after transfection. After washing twice with
PBS, the cell concentration was adjusted to 1×106 using
Annexin V-FITC binding buffer. Then, 100 μl of the suspended cells
were stained with 5 μl of Annexin V for 30 min and 10 μl of
propidium iodide (4Abio, Beijing, China) for 15 min in the dark at
room temperature. After washing with PBS, 400 μl of PBS was added
to the samples and analysis was performed using a flow cytometer
(BD Biosciences, San Jose, CA, USA). The percentage of Annexin
V-positive cells was recorded for analysis.
Cell migration and invasion assay
The Transwell invasion system (8.0 μm polycarbonate
membrane, 6.5 mm insert in a 6-well plate; Corning, Lowell, MA,
USA) with or without Matrigel was used for the invasion or
migration assay. After transfection for 24 h, cells were harvested
using 0.25% trypsin-EDTA (Thermo). For the migration assay,
2×106 cells were added into the upper compartment of the
Transwell chamber. For the invasion assay, 1×105 cells
were plated on chambers preloaded with Matrigel. In both assays,
all cells were suspended in 100 μl of RPMI-1640 without serum,
1,000 μl RPMI-1640 supplemented with 10% FBS was added in the
bottom compartment of the chamber. After 12 h of incubation,
Matrigel, dead cells and non-migrated or non-invaded cells on the
upper surface of the membrane were removed with a cotton swab. The
membranes were fixed with 4% paraformaldehyde and stained with 1%
crystal purple. The invasive cells that were stuck to the lower
surface of the membrane were then counted.
Statistical analysis
Statistical analysis was performed using SPSS 17.0
software (SPSS, Inc., Chicago, IL, USA). Data obtained from qRT-PCR
were log2 transformed. Differences in miR-126
expressions between ESCC and paired paracancer tissues were
analyzed using non-parametric test. The two-sided Fisher’s exact
test was used to determine the relationship between miR-126, IRS-1
and GOLPH3 expression and clinicopathological variables.
Quantitative data expressed as means ± standard deviation and
differences between groups were calculated with the t-test. All
results were considered statistically significant when
P<0.05.
Results
Downregulation of miR-126 expression and
association with lymph node metastasis, tumor in-depth or cell
differentiation in ESCC
To investigate the expression of miR-126 in ESCC, we
first evaluated the status of miR-126 in samples of ESCC by
RT-qPCR. Total-RNA was isolated from 102 pairs of ESCC and
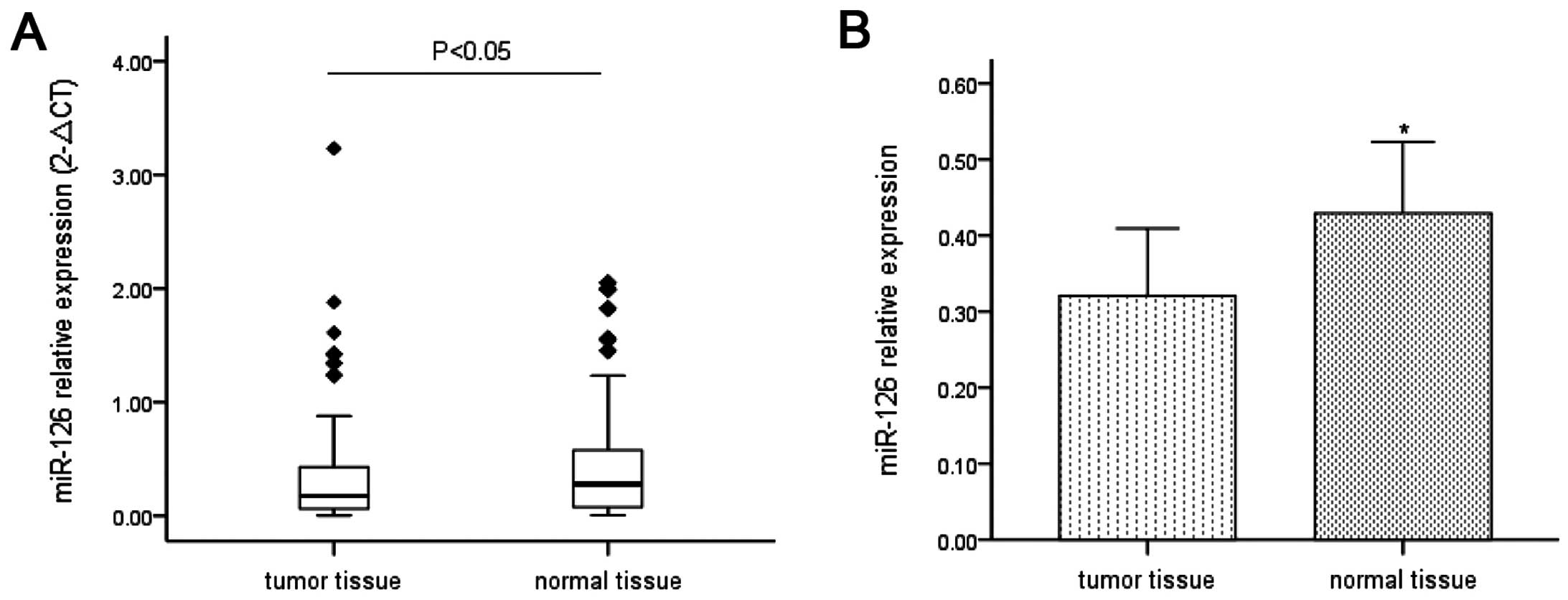
paracancer tissues. Lower expression of miR-126 was found in 66
(64.7%) patients. The expression of miR-126 in ESCC samples was on
average decreased ~68.36% of that in paracancer samples (P<0.05;
Fig. 1A and B).
To further investigate the relationship between the
expression level of miR-126 and clinicopathological
characteristics, all patients were divided into two groups. Lymph
node metastasis, tumor in-depth, cell differentiation and TNM stage
were all associated with miR-126 expression. Greater possibility of
lymph node metastasis was observed in the miR-126 low expression
group. There were no statistically significant associations between
miR-126 expression and other clinicopathological parameters, such
as gender, age, tumor location, alcoholism or smoking (P>0.05;
Table I).
IRS-1 and GOLPH3 are potential targets of
miR-126 predicted online
Since miR-126 was closely related to
clinicopathological characteristics in ESCC, we further analyzed
its function and molecular mechanism in the carcinogenesis of ESCC.
We predicted targets of miR-126 in humans using the online tools
miRGenTargets, TargetScanHuman and PicTar. Following computational
prediction, the proteins IRS-1, FBXO33, PLK2, PTPN9, and GOLPH3
were analyzed. We reviewed published reports that focused on
individual miRNAs of the proteins. Among them, IRS-1 has been
reported as a target of miR-126 in colon cancer, whereas GOLPH3
shows overexpression in ESCC and is associated with poor prognosis.
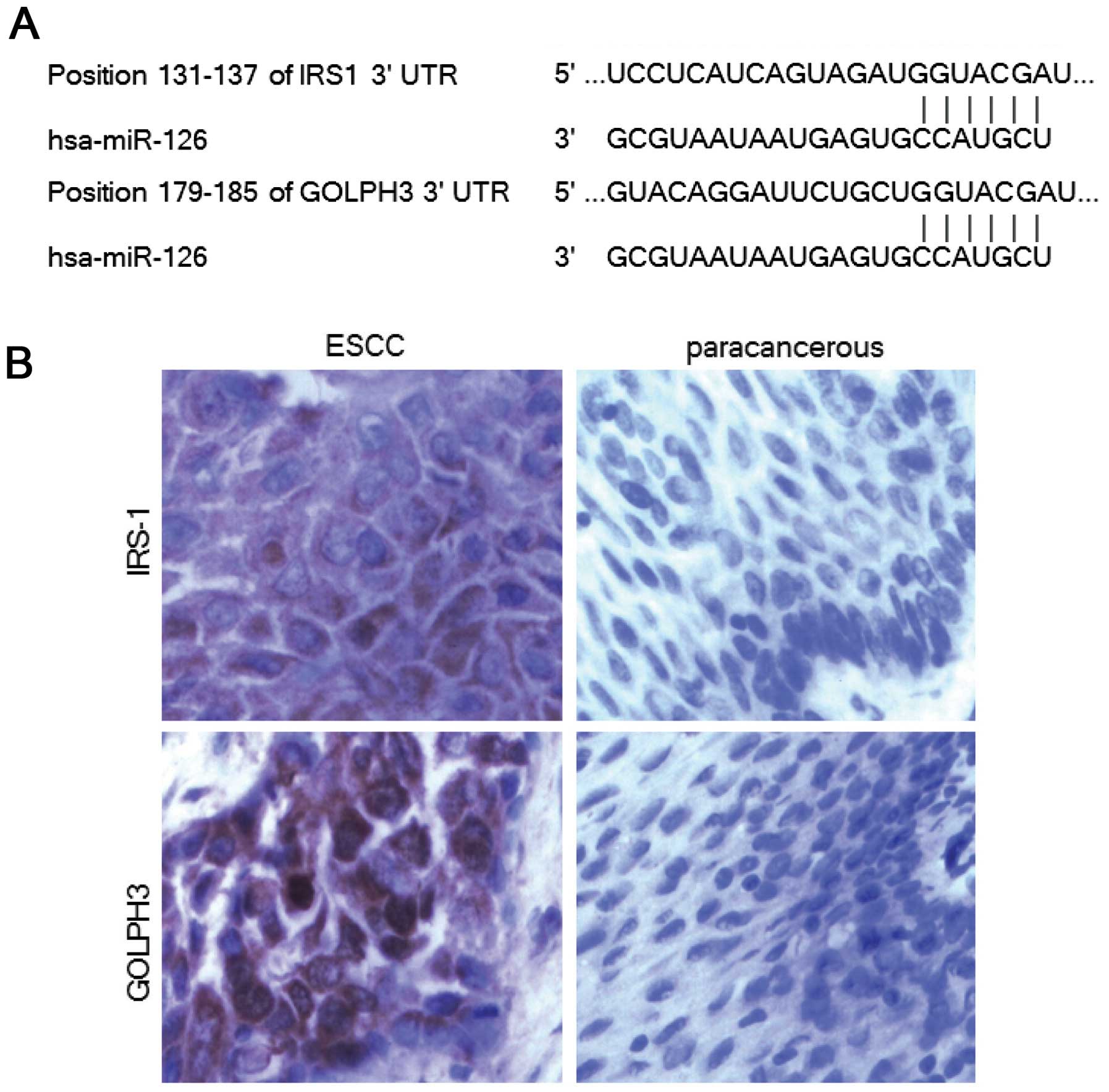
Therefore IRS-1 and GOLPH3 were selected as targets of our research
(Fig. 2A).
IRS-1 and GOLPH3 are overexpressed in
human ESCC
To verify our prediction, we first studied the
relationship between the two proteins in ESCC. IRS-1 and GOLPH3
were determined using immunohistochemistry in ESCC tissues and the
paracancer tissues respectively. The expression of IRS-1 protein
was significantly higher in ESCC samples compared with paracancer
tissues, and the same results were found for GOLPH3 expression
levels (P<0.01; Fig. 2B).
We also analyzed the relationship between IRS-1 or
GOLPH3 protein and clinical characteristics. Positive expression of
IRS-1 was associated with cell differentiation but no obvious
association with any other clinical characteristics. Meanwhile,
GOLPH3 was closely related to lymph node metastasis, tumor invasion
in-depth and TNM stage (Table
I).
IRS-1 and GOLPH3 protein expression after
up/down regulation of miR-126
To further explore the impact of aberrant expression
of miR-126 on IRS-1 and GOLPH3, we transfected miR-126 mimics,
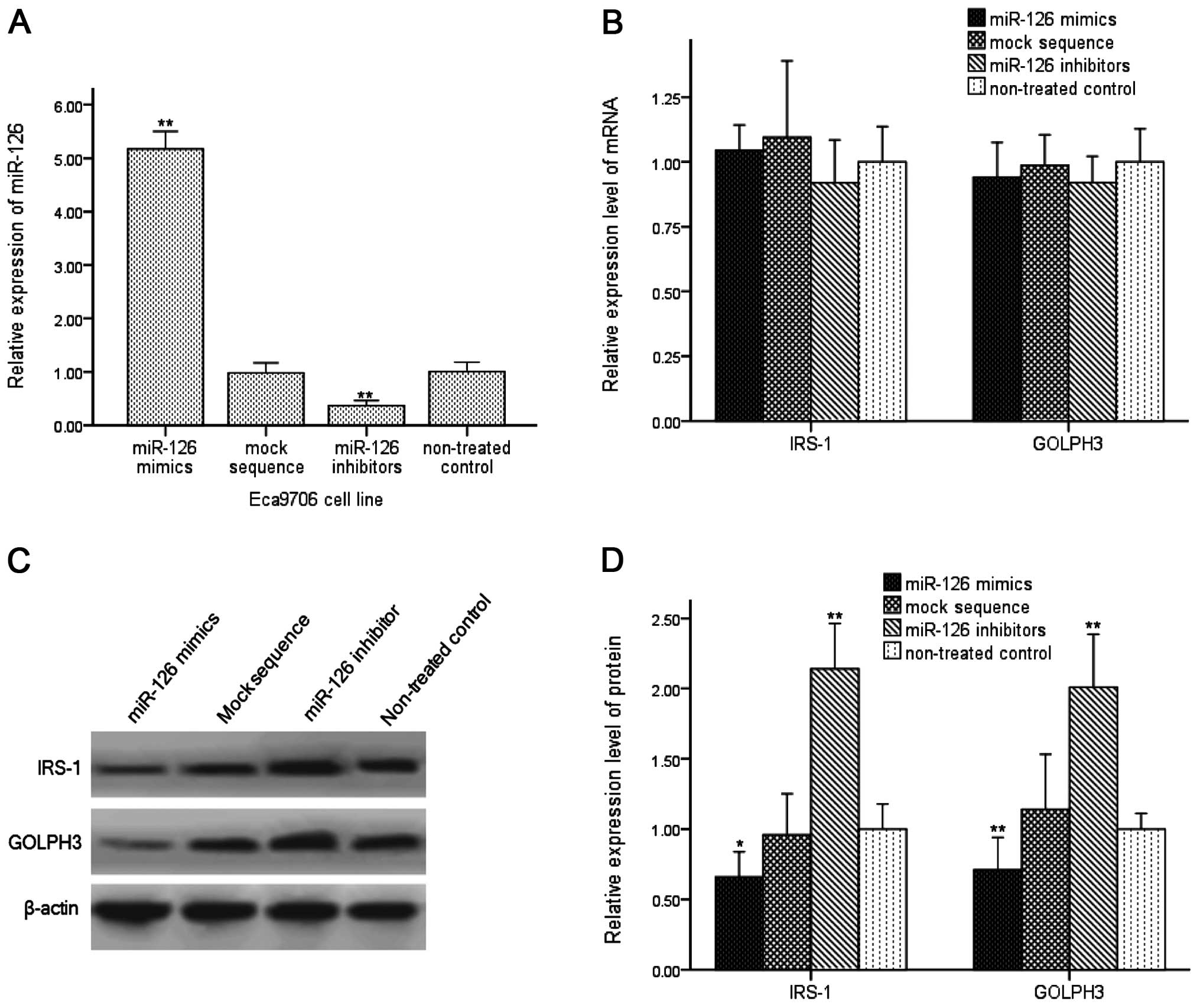
inhibitors or mock sequence into Eca9706 cells. miR-126 levels
increased on an average 5.17-fold (P<0.01) after transfection
with miR-126 mimics and decreased 0.37-fold (P<0.01) when
transfected with miR-126 inhibitors. There was no obvious change in
cells transfected with the mock sequence (Fig. 3A).
Initially, relative mRNA levels of IRS-1 and GOLPH3
in Eca9706 cells were detected. Then, 50 nM of miR-126 mimics,
inhibitors or mock sequence were transfected into Eca9706 cells.
After transfection for 48 h, we detected the mRNA levels of IRS-1
and GOLPH3 in these cells. In cells transfected with miR-126
mimics, altered relative mRNA levels of IRS-1 (1.05-fold) and
GOLPH3 (0.94-fold) had no statistical difference compared with
non-treated controls. Downregulation of miR-126 in Eca9706 cells
showed no significant difference in IRS-1 (0.92-fold) and GOLPH3
(0.92-fold) mRNA levels, and there were also no differences in the
mock sequence group (P>0.05; Fig.
3B).
We then analyzed the relationship between miR-126
and IRS-1 or GOLPH3 protein. The expression level of IRS-1 protein
decreased 0.71-fold (P<0.05) compared with the non-treated
control, and GOLPH3 protein decreased 0.66-fold (P<0.01) when
transfected with miR-126 mimics. In addition, when transfected with
miR-126 inhibitors, IRS-1 protein expression increased 2.01-fold
(P<0.01) and GOLPH3 protein increased 2.14-fold (P<0.01)
compared with the non-treated controls, and there were no
differences between the mock sequence group and the non-treated
controls (Fig. 3C and D).
miR-126 suppresses cell proliferation but
has no effect on apoptosis in ESCC cells
To further characterize the functional role of
miR-126 in ESCC, we observed the effect of miR-126 on the
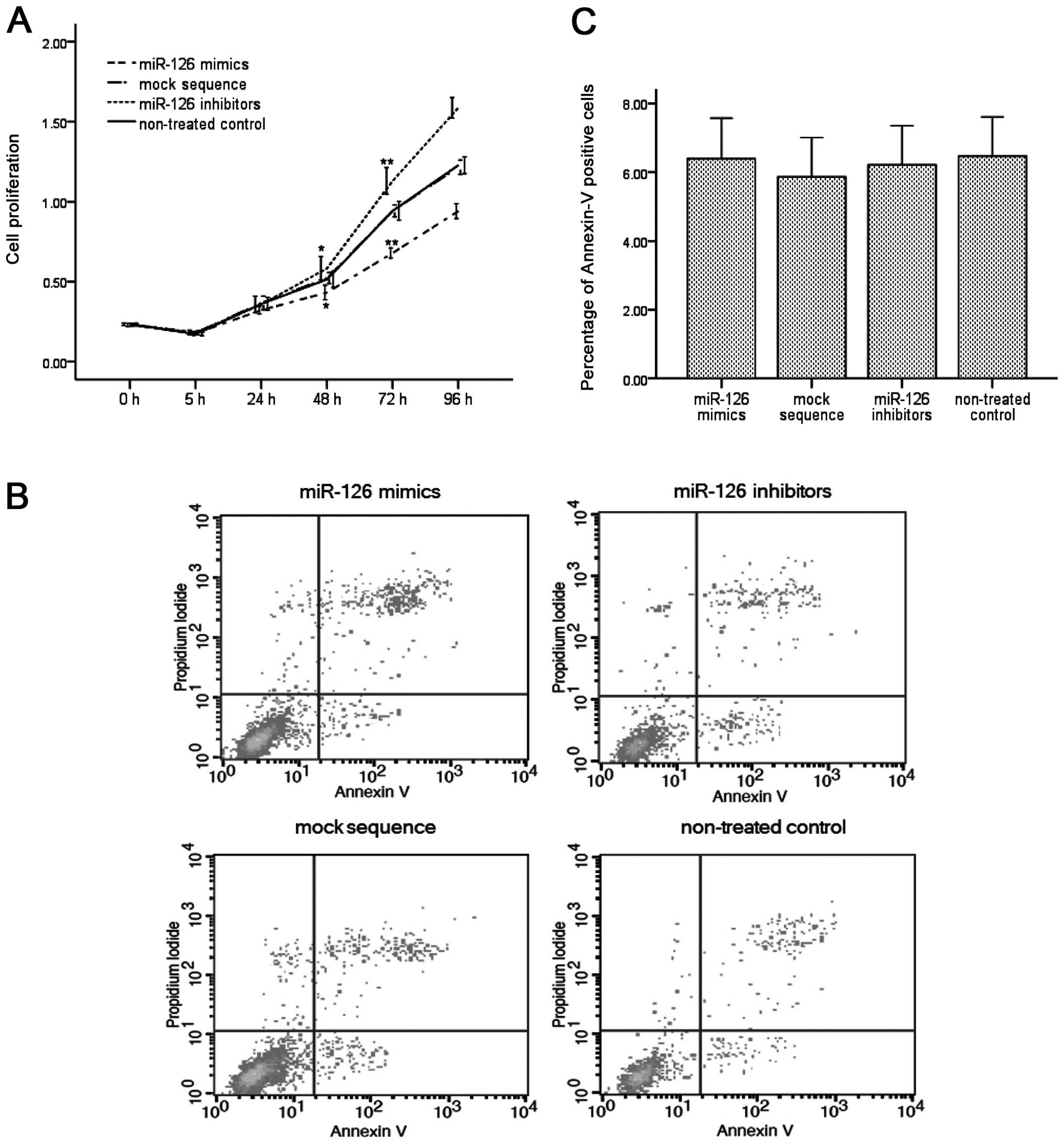
proliferation and apoptosis of ESCC cells. After transfection with
miR-126 mimics, cell proliferation was suppressed after 48 h
(0.433±0.054 vs. 0.514±0.061, P<0.05) and 72 h (0.680±0.038 vs.
0.943±0.070, P<0.01). In addition, transfection with miR-126
inhibitors showed an improvement in cell proliferation at 48 h
(0.584±0.088 vs. 0.514±0.061, P<0.05) and 72 h (1.130±0.098 vs.
0.943±0.070, P<0.01); no alteration of cell proliferation
occurred in the mock sequence group (P>0.05; Fig. 4A) compared with the non-treated
group. However, cell apoptosis, which was measured using flow
cytometry at 48 h after transfection, showed no significant
differences between all three groups compared to that in the
non-treated controls (P>0.05, Fig.
4B and C).
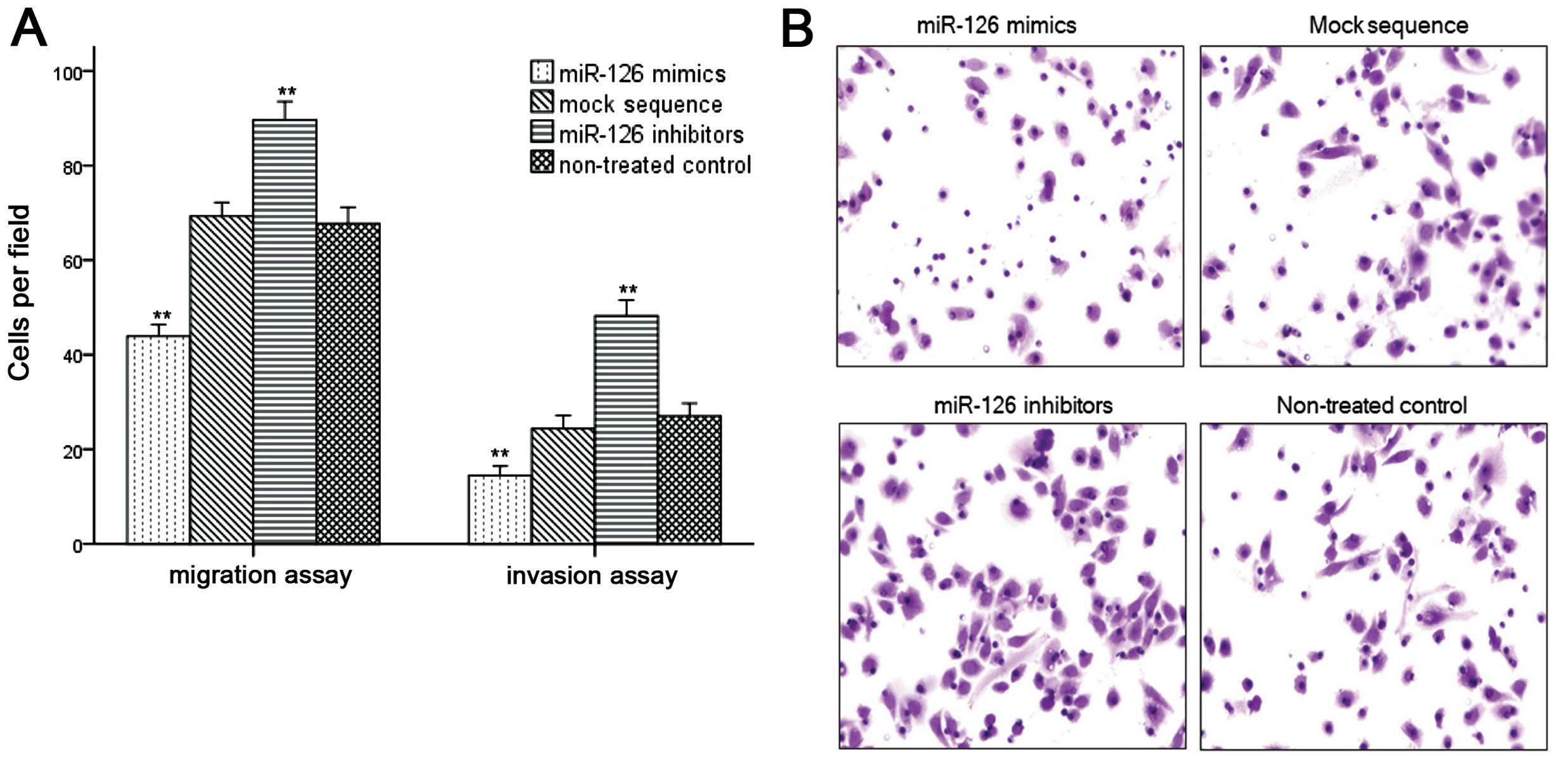
miR-126 regulates cell migration and
invasion
To explore the correlation between miR-126
expression and migration or invasion potential in human ESCC cell
lines, we carried out the Transwell assay. To reduce the bias of
proliferation suppression of miR-126, the incubation time was
controlled for 24 h. In the migration assay, the number of migrated
cells was significantly decreased in the group transfected with
miR-126 mimics compared to the non-treated controls (43.87±7.59 vs.
67.77±10.57, P<0.01), whereas the number of cells increased in
the miR-126 inhibitor group (89.59±12.16 vs. 67.77±10.57,
P<0.01, Fig. 5A and B). In the
invasion assay, cell invasiveness increased significantly when
transfected with the miR-126 inhibitor (48.21±10.21 vs. 27.03±8.36,
P<0.01), whereas overexpression of miR-126 significantly
suppressed Eca9706 cell invasion (14.44±6.14 vs. 27.03±8.36,
P<0.01); results in the mock sequence group were similar to the
non-treated controls (P>0.05, Fig.
5A).
Discussion
miRNAs have important roles in tumorigenesis by
various mechanisms. miRNAs can act as tumor suppressors depending
on the targeting of specific suppressor genes or oncogenes, or they
can alter tumorous epithelial-mesenchymal transitions to promote
tumor invasion and metastasis, and they can also regulate the
expression of metastasis-associated genes in tumors (22,23).
Thus, the aberrant expression of tumor-suppressive miRNAs may
contribute to human carcinogenesis. For example, downregulation of
miR-7 can promote metastasis of gastric cancer by targeting the
insulin-like growth factor-1 receptor, and the low level of miR-210
is associated with the carcinogenesis of ESCC (24,25).
The expression of miR-126 is downregulated in many cancers, such as
lung, breast and liver cancer (14,26),
suggesting that the suppression of miR-126 was significantly
related to the occurrence and development of cancer. The results of
the present study confirmed that miRNA-126 was downregulated in
ESCC compared with paracancer tissues, indicating that miR-126 may
be associated with the carcinogenesis of ESCC. We therefore
analyzed the relationship between downregulation of miR-126 and the
clinicopathological characteristics of ESCC patients. We found that
downregulation of miR-126 was associated with tumor cell
differentiation, lymph nodes metastasis and tumor invasion in-depth
in ESCC patients, which indicated that miR-126 acts as an oncogene
in ESCC and participates in tumor invasiveness and metastasis. This
result is similar to that found in colon and lung cancer (18,20).
To further explore the molecular mechanism of
miR-126 in suppressing ESCC development and invasion, we predicted
IRS-1 and GOLPH3 as novel targets of miR-126. The insulin receptor
substrates are cytoplasmic signaling adaptor proteins that function
as intermediates of the insulin receptor and insulin-like growth
factor receptor (IGF-IR), which are involved in cell growth and
survival. As a member of the IRS family, IRS-1 has been a hotspot
and is widely expressed. Tyrosine-phosphorylated IRS-1 bind
proteins containing Src homology 2 (SH2) domains, which activate
mitogen-activated protein kinase (MAPK) by effector cascades such
as the PI3K pathway (27). Porter
et al (28) reported that
IRS-1 was highly expressed in localized breast tumors and that high
IRS-1 levels can be a good indicator of the effectiveness of
specific types of chemotherapy in breast cancer, but were not
related to tumor aggressiveness. Several studies have also shown
that IRS-1 forced expression of VEGF expression via transcriptional
activation and promoted cell proliferation in human colon cancer
(29,30). The results of the present study
showed that IRS-1 was overexpressed in ESCC tissues and was
associated with cell differentiation. Forced expression of miR-126
downregulated the expression level of IRS-1 protein in Eca9706
cells. Therefore, we suggest that IRS-1 is a direct target of
miR-126 in human ESCC.
The endocytic protein, GOLPH3, is a highly conserved
34 kDa protein which was initially identified by proteomic
characterization of the Golgi apparatus. It was suggested an
enhanced activation of mTOR signaling represents a molecular basis
of the oncogenic activity of GOLPH3 (31). Several studies have reported that
the increased expression of GOLPH3 is closely associated with the
clinicopathological characteristics of cancers, such as prostate,
lung, ovary and gastric cancer (31–33).
Wang et al (34) showed that
GOLPH3 was overexpressed in ESCC and was positively associated with
clinical stage, TNM classification, histological differentiation
and vital status. The present study indicates that GOLPH3 is highly
expressed in ESCC and is related to tumor invasion in-depth, lymph
nodes metastasis and TNM stage, which is the same as the above
study. However, in our study, expression of GOLPH3 was not related
to histological differentiation. Although there are few
differences, results from the present study and previous studies
suggest that GOLPH3, as well as IRS-1, are targets of miR-126 in
ESCC, indicating that miR-126 can have different targets and roles
across malignancies. In the present study, the mRNA levels of IRS-1
and GOLPH3 did not change significantly with miR-126 levels (when
up/downregulated) in ESCC cell lines. This indicated that IRS-1 and
GOLPH3 were negatively regulated by miR-126 at the
post-transcriptional level in ESCC.
Crawford et al (18) indicated that miR-126 inhibited the
proliferation of the human lung cancer cell line A549 both in
vitro and in vivo. Li et al (35) found that miR-126 suppresses the
proliferation and invasion of the human colon cancer cell lines
HCT116, SW620 and HT-29. However, the functions of miR-126 in
esophageal cancer progression remain unresolved. In the present
study, the overexpression of miR-126 suppressed cell proliferation
in the ESCC cell line Eca9706. Moreover, the Transwell experiment
also showed that miR-126 can inhibit ESCC cell migration and
invasion in vitro. Collectively, these results suggest that
miR-126 may function as a tumor suppressor by inhibiting cell
proliferation, suppressing migration and invasion, which, in turn,
affect multiple clinicopathological characteristics of cancer such
as lymph node metastasis and depth of invasion. At the same time,
this is the first report indicating the suppressor action of
miR-126 in ESCC.
In conclusion, as a tumor suppressor, miR-126
expression decreased in ESCC and might have a tumor suppression
role via the regulation of IRS-1 and GOLPH3. It can suppress cell
proliferation, migration and invasion. miR-126 correlates with
lymph node metastasis, tumor invasion in-depth and TNM stage of
ESCC, and could serve as a potential marker and therapeutic target
in ESCC.
Acknowledgements
This study was supported by the Research Fund for
the National Key Clinical Speciality of China, and the Innovation
Foundation of Excellent Intellectuals of Henan Province (no.
20070214).
References
|
1
|
Ferlay J, Shin HR, Bray F, Forman D,
Mathers C and Parkin DM: Estimates of worldwide burden of cancer in
2008: GLOBOCAN 2008. Int J Cancer. 127:2893–2917. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Jemal A, Siegel R, Xu J and Ward E: Cancer
statistics, 2010. CA Cancer J Clin. 60:277–300. 2010. View Article : Google Scholar
|
|
3
|
Mathé EA, Nguyen GH, Bowman ED, et al:
MicroRNA expression in squamous cell carcinoma and adenocarcinoma
of the esophagus: associations with survival. Clin Cancer Res.
15:6192–6200. 2009.PubMed/NCBI
|
|
4
|
Song QK, Li J, Jiang HD, He YM, Zhou XQ
and Huang CY: Esophageal cancer mortality during 2004–2009 in
Yanting County, China. Asian Pac J Cancer Prev. 13:5003–5006.
2012.
|
|
5
|
Fareed KR, Kaye P, Soomro IN, et al:
Biomarkers of response to therapy in oesophago-gastric cancer. Gut.
58:127–143. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kloosterman WP and Plasterk RH: The
diverse functions of microRNAs in animal development and disease.
Dev Cell. 11:441–450. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Pillai RS: MicroRNA function: multiple
mechanisms for a tiny RNA? RNA. 11:1753–1761. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lewis BP, Burge CB and Bartel DP:
Conserved seed pairing, often flanked by adenosines, indicates that
thousands of human genes are microRNA targets. Cell. 120:15–20.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Croce CM and Calin GA: miRNAs, cancer, and
stem cell division. Cell. 122:6–7. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Rosenfeld N, Aharonov R, Meiri E, et al:
MicroRNAs accurately identify cancer tissue origin. Nat Biotechnol.
26:462–469. 2008. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Croce CM: Causes and consequences of
microRNA dysregulation in cancer. Nat Rev Genet. 10:704–714. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Schmidt M, De Maziere A, Smyczek T, et al:
The role of Egfl7 in vascular morphogenesis. Novartis Found Symp.
283:18–28; discussion 28–36, 238–241. 2007. View Article : Google Scholar
|
|
13
|
Schmidt M, Paes K, De Maziere A, et al:
EGFL7 regulates the collective migration of endothelial cells by
restricting their spatial distribution. Development. 134:2913–2923.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Li XM, Wang AM, Zhang J and Yi H:
Downregulation of miR-126 expression in colorectal cancer and its
clinical significance. Med Oncol. 28:1054–1057. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yang J, Lan H, Huang X, Liu B and Tong Y:
MicroRNA-126 inhibits tumor cell growth and its expression level
correlates with poor survival in non-small cell lung cancer
patients. PLoS One. 7:e429782012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Otsubo T, Akiyama Y, Hashimoto Y, Shimada
S, Goto K and Yuasa Y: MicroRNA-126 inhibits SOX2 expression and
contributes to gastric carcinogenesis. PLoS One. 6:e166172011.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Sasahira T, Kurihara M, Bhawal UK, et al:
Downregulation of miR-126 induces angiogenesis and
lymphangiogenesis by activation of VEGF-A in oral cancer. Br J
Cancer. 107:700–706. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Crawford M, Brawner E, Batte K, et al:
MicroRNA-126 inhibits invasion in non-small cell lung carcinoma
cell lines. Biochem Biophys Res Commun. 373:607–612. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ye P, Liu J, He F, Xu W and Yao K:
Hypoxia-induced deregulation of miR-126 and its regulative effect
on VEGF and MMP-9 expression. Int J Med Sci. 11:17–23. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Li N, Li X, Huang S, Shen S and Wang X:
miR-126 inhibits colon cancer proliferation and invasion through
targeting IRS1, SLC7A5 and TOM1 gene. Zhong Nan Da Xue Xue Bao Yi
Xue Ban. 38:809–817. 2013.(In Chinese).
|
|
21
|
Lu YY, Sweredoski MJ, Huss D, Lansford R,
Hess S and Tirrell DA: Prometastatic GPCR CD97 is a direct target
of tumor suppressor microRNA-126. ACS Chem Biol. 9:334–338. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kalluri R and Weinberg RA: The basics of
epithelial-mesenchymal transition. J Clin Invest. 119:1420–1428.
2009. View
Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhang Y, Wang Z, Chen M, et al:
MicroRNA-143 targets MACC1 to inhibit cell invasion and migration
in colorectal cancer. Mol Cancer. 11:232012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhao X, Dou W, He L, et al: MicroRNA-7
functions as an anti-metastatic microRNA in gastric cancer by
targeting insulin-like growth factor-1 receptor. Oncogene.
32:1363–1372. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Tsuchiya S: The role of microRNA-210 in
esophageal squamous cell carcinoma. Yakugaku Zasshi. 132:1069–1073.
2012.(In Japanese).
|
|
26
|
Yang R, Dick M, Marme F, et al: Genetic
variants within miR-126 and miR-335 are not associated with breast
cancer risk. Breast Cancer Res Treat. 127:549–554. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
White MF: IRS proteins and the common path
to diabetes. Am J Physiol Endocrinol Metab. 283:E413–E422. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Porter HA, Perry A, Kingsley C, Tran NL
and Keegan AD: IRS1 is highly expressed in localized breast tumors
and regulates the sensitivity of breast cancer cells to
chemotherapy, while IRS2 is highly expressed in invasive breast
tumors. Cancer Lett. 338:239–248. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Shi B, Sepp-Lorenzino L, Prisco M, Linsley
P, deAngelis T and Baserga R: Micro RNA 145 targets the insulin
receptor substrate-1 and inhibits the growth of colon cancer cells.
J Biol Chem. 282:32582–32590. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Yin Y, Yan ZP, Lu NN, et al:
Downregulation of miR-145 associated with cancer progression and
VEGF transcriptional activation by targeting N-RAS and IRS1.
Biochim Biophys Acta. 1829:239–247. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Scott KL, Kabbarah O, Liang MC, et al:
GOLPH3 modulates mTOR signalling and rapamycin sensitivity in
cancer. Nature. 459:1085–1090. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Hua X, Yu L, Pan W, et al: Increased
expression of Golgi phosphoprotein-3 is associated with tumor
aggressiveness and poor prognosis of prostate cancer. Diagn Pathol.
7:1272012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Hu BS, Hu H, Zhu CY, Gu YL and Li JP:
Overexpression of GOLPH3 is associated with poor clinical outcome
in gastric cancer. Tumour Biol. 34:515–520. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang JH, Chen XT, Wen ZS, et al: High
expression of GOLPH3 in esophageal squamous cell carcinoma
correlates with poor prognosis. PLoS One. 7:e456222012. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Li N, Tang A, Huang S, et al: MiR-126
suppresses colon cancer cell proliferation and invasion via
inhibiting RhoA/ROCK signaling pathway. Mol Cell Biochem.
380:107–119. 2013. View Article : Google Scholar : PubMed/NCBI
|