Introduction
Ribavirin, a synthetic nucleoside analogue
(1-β-D-ribofuranosyl-1,2,4-triazole-3-carboxamide) is a widely used
drug whose antiviral activity was described almost 40 years
(1). Currently, ribavirin is almost
solely used for hepatitis C infection (HCV) in combination with
interferon. This combination significantly increases sustained
virological response rates even in patients with interferon
resistance arising from HCV-related factors (2). The knowledge gained on the molecular
pathogenesis of cancer has led to considerable interest in
repurposing non-cancer drugs against several malignancies (3). It was previously demonstrated that
ribavirin and mycophenolic acid inhibit inosine 5′-phosphate
dehydrogenase (IMPDH), the first enzyme specific for the de
novo synthesis of guanosine monophosphate (GMP) (4). IMPDH encodes the rate-limiting
enzyme in de novo guanine nucleotide biosynthesis, which is
necessary for DNA and RNA synthesis. IMPDH has been linked to cell
growth, differentiation and malignant transformation (5,6). Two
isoforms of IMPDH have been described: Type I is constitutively
expressed in normal cells, whereas type II activity has been shown
to be increased in proliferating and particularly malignant cells
(7,8). Thus, IMPDH has been considered an
attractive target for cancer therapy (9).
On the other hand, Kentsis et al (10), demonstrated the direct binding of
ribavirin to eIF4E in vitro at micromolar concentrations and
an efficient competition with eIF4E:m7G mRNA cap binding in
vitro and in vivo, which disrupts eIF4E:m7G functions in
the transport and translation of eIF4E-regulated genes. These
effects lead to downregulation of the oncogenic proteins, cell
cycle arrest and suppression of the eIF4E-mediated transformation
in vitro and in vivo (10). eIF4E is overexpressed in many human
malignancies where it is typically a marker of poor prognosis. The
function of this factor is controlled by interactions with protein
cofactors together with many signaling pathways, including Ras,
Mnk, Erk, MAPK, PI3K, mTOR and Akt. Borden and Culjkovic-Kraljacic
suggest that ~30% of cancer types have elevated eIF4E (11). In a recent proof-of-principle study
on relapsed or refractory acute myeloid patients with subtypes M4
and M5 or other subtypes with high eIF4E levels, administered oral
ribavirin at daily doses ranging from 1,000 to 1,800 mg led to
plasma ribavirin between 5 and 36 μM, produced 1 complete, 2
partial remissions, 2 blast response, and four stable diseases out
of 11 evaluable patients. Best responses were observed at ~28 days
(12). These results clearly
support investigations on ribavirin as a cancer treatment.
Beyond DNA methylation and histone deacetylase
inhibitors, drugs able to block histone methylation are currently
pursued as epigenetic anticancer drugs (13). Enhancer of zeste homolog 2 (EZH2), a
core component of polycomb repressive complex 2 (PRC2), plays a
role in transcriptional repression through H3K27 trimethylation,
and is involved in various biological processes (14). It is well known that
3-deazaneplanocin A (DZNep), an inhibitor of
S-adenosylmethionine-dependent methyltransferase targets the
degradation of EZH2, leading to apoptosis in various malignancies,
suggesting that EZH2 may be a new target for epigenetic treatment
(15). The results of the present
study showed that ribavirin is also an EZH2 inhibitor and that its
antitumor effects stem from the inhibition of the IMPDH, eIF4E and
EZH2 targets.
Materials and methods
Cell culture and ribavirin treatment
The breast (MCF-7, MDA-231 and MDA-436), brain
(D54), cervical (HeLa), colon (SW-480) and prostate (PC3 and
DU-145) cancer cell lines were obtained from the ATCC (Manassas,
VA, USA) and cultured in Dulbecco’s modified Eagle’s medium
(DMEM)/F12 medium supplemented with 10% fetal bovine serum (FBS)
(both from Invitrogen Life Technologies, Carlsbad, CA, USA), and
penicillin-streptomycin at 37°C in a humidified 5% CO2
atmosphere. Ribavirin was obtained from Sigma (St. Louis, MO, USA),
and dissolved in dimethyl sulfoxide (DMSO), stored at −20°C and
thawed prior to use. Stock solutions were thawed/frozen ≤3 times.
Cell lines employed for the proliferation assay were treated with
10, 15, 20, 25 and 50 μM for 96 h. For the mRNA expression,
and protein immunodetection, ribavirin treatment with 25 μM
was maintained for 72 h.
Cell proliferation assay
Cells were seeded in 96-well microplates (Corning)
at a density of 2×5103 cells/well in 0.1 ml of complete
medium for 24 h, and then treated for 96 h with ribavirin at
concentrations of 10, 15, 20, 25 and 50 μM with changes of
medium containing ribavirin on alternate days. After 96 h, the
medium was aspirated and replaced for 10 min with 50 μl of
0.75% crystal violet in 50% ethanol, 0.25% NaCl and 1.75%
formaldehyde solution. The cells were then washed with water,
air-dried, and the dye was eluted with PBS + 1% sodium dodecyl
sulfate (SDS) solution. The absorbance of crystal violet is
proportional to the cell number and was assessed by dye absorbance
measured at 570 nm on an automated microplate reader. All assays
were performed in triplicate. The cytotoxic effect of each
treatment was expressed as a percentage of cell proliferation
relative to the untreated control cells (percentage of control) and
was defined as (A570 nm treated cells/A570 nm non-treated cells) ×
100.
Computational search of approved drugs
with a high structural similarity to DZNep
To identify in a systematic manner approved drugs
with similar activity as DZNep, we carried out a rapid
computational search of DrugBank, a database listing approved drugs
(16). Using the well-established
principles of similarity searching by two-dimensional fingerprints
(17), we used the chemical
structure of DZNep as a query and computed the similarity of each
of 1,491 compounds listed in DrugBank using the Molecular Access
System (MACCS) keys and the Tanimoto coefficientREF as
implemented in Molecular Operating Environment (MOE), version
2011.10 (18,19). In general, compounds in DrugBank had
an extremely low molecular similarity with DZNep (median and mean
similarity values of 0.35 with a standard deviation of 0.12).
Flexible alignment in silico
The chemical structures of ribavirin and DZNep were
constructed in MOE, version 2011.10 (18). The alignment of the two compounds
was performed following the flexible alignment protocol implemented
in MOE for small molecules. In this tool, the alignment is based on
the internal strain and overlap of molecular features (20). For the flexible alignment we used
default parameters including an iteration limit of 200, failure
limit of 20, and an energy cut-off of 15. Prior to the search, the
charges were computed using the MMFF94x force field. The search was
performed with the stochastic conformation search enabled. The
top-ranked solution was used for analysis.
RNA isolation and quantitative expression
assay
Total RNA was isolated using the PureLink RNA kit
(Ambion-Life Technologies) according to the manufacturer’s
instructions. RNA purity and integrity were assessed by
spectrophotometric analysis (NanoDrop 2000c; Thermo Scientific) and
denaturing agarose gel. Total RNA (1 μg) was used for cDNA
synthesis with the SuperScript III First-Strand Synthesis SuperMix
according to the manufacturer’s instructions (Invitrogen Life
Technologies). Quantitative PCR for amplification and detection of
DNA was performed on the CFX96 thermocycler (Bio-Rad, Hercules, CA,
USA). PCR was carried out in triplicate in a 10 μl reaction
volume with the SYBR-Green qPCR master mix (Bio-Rad), 0.35 nM
forward primer, and 0.35 nM reverse primer. Data were analyzed
using the 2−ΔΔCT method and reported as the fold-change
in gene expression normalized to the endogenous control gene
(GUSB) and relative to cells without treatment. The primers
used were: GUSB, forward: 5′-CCTGTGACCTTTGTGAGCAA-3′ and reverse:
5′-AAACCCTGCAATCGTTTCTG-3′; EZH2, forward:
5′-CAACACCAAGCAGTGCCCGTGC-3′ and reverse:
5′-CCTGCCACGTCAGATGGTGCC-3′; G9a, forward:
5′-CTCCGCTGATTTTCGAGTGTAA-3′ and reverse:
5′-GTCGAAGAGGTAAGAATCATCC-3′; LSD1, forward:
5′-GTGTCTCGTTGGCGTGCT-3′ and reverse: 5′-CCCGCAAAGAAGAGTCGTG-3′;
HDAC1, forward: 5′-GTTACACCATTCGTAACGTTGC-3′ and reverse:
5′-CAGTCGCTGTTTGATCTTCTC-3; eif4E, forward:
5′-GCTTTGGTTAAAAATGGCTCA-3′ and reverse:
5′-GAGACTGCCTTGCAATAAGAAG-3; Dnmt1, forward:
5′-TACCTGGACGACCCTGACCTC-3′ and reverse:
5′-CGTTGGCATCAAAGATGGACA-3′; Dnmt3a, forward: 5′-TATT
GATGAGCGCACAAGAGAGC-3′ and reverse: 5′-GGGTGTTCCAGGGTAACATTGAG-3′;
Dnmt3b, forward: 5′-GGCAAGTTCTCCGAGGTCTCTG-3′ and reverse:
5′-TGGTACATGGCTTTTCGATAGGA-3′.
Protein extraction and immunoblot
analysis
Whole-cell extraction was performed with lysis
buffer containing 50 mM Tris-HCl pH 7.4, 150 mM NaCl, 0.5% Nonidet
P-40, 1 mM EDTA, 1 mM phenylmethylsulfonyl fluoride (PMSF), 0.2 mM
Na3VO4 and a protease inhibitor cocktail
(Sigma). Histones proteins were obtained by sulfuric acid
extraction. Briefly, the nuclear pellet was resuspended in 0.4 M
H2SO4 for 4 h, and acid-soluble proteins in a
postcentrifugation supernatant (10,000 × g for 20 min) were
recovered by overnight precipitation with 20% trichloroacetic acid
and centrifugation (16,000 × g for 30 min). The pellets containing
the histone proteins were washed once in ice-cold acetone
containing 1% HCl and then washed once with ice-cold acetone. The
pellet was dried under vacuum and stored at −80°C. The protein
concentration was determined by the Bradford method, and the
integrity of the histone proteins in the acid soluble extract was
evaluated with Coomassie staining. Proteins were separated in an
SDS-PAGE gel 10–18% according to the molecular weight of the
studied protein, and transferred to a PVDF membrane (Bio-Rad). The
membrane was incubated for 1 h with blocking solution
(TBS/Tween-20, 5% of non-fat milk), and incubated overnight with
the specific primary antibody in blocking solution (dilution
1:1,000). The membranes were washed with TBS-T, and incubated 1 h
with a peroxidase secondary antibody at a dilution of 1:1,000 and
developed with chemiluminescence substrate cat. no. WBLUR0100
(Millipore, Billerica, MA, USA). Antibodies for EZH2 (cat. no.
36–6300) Invitrogen, G9a (cat. 07-551), histone 3 (cat. no.
06-755), H3K4me2 (cat. no. 07-030), H3K9me2 (cat. no. 07-212) and
H3K27me3 (cat. no. 07-449) were obtained from Millipore.
Inhibition of the histone
methyltranferase activity
Histone methyltransferase activity/inhibition assay
kit (Epigentek, New York, NY, USA) was employed to determine the
inhibitory activity of ribavirin with HMT that specifically target
histone H3 at lysine 27. Briefly, the nuclear extract was prepared
from the MCF-7 cell line using the EpiQuik Nuclear Extraction kit
(Epigentek), and incubated with ribavirin at 5, 10 and 15
μM. The percentage of ribavirin inhibition was calculated
according to the level of H3K27 obtained in the control assay in
the presence of 10 μg of nuclear protein plus DMSO.
siRNA transfection assay
MCF-7 cells were seeded in 24-well microplate
(Corning) at a density of 1.5×105 cells/well into 0.2 ml
of DMEM/F12 medium supplemented with 10% FBS. After 24 h, cells
were transfected with the Lipofectamine RNAimax transfection
reagent (Invitrogen-Life Technologies) and siRNA directed to EZH2
(4392420), eiF4E (4390771), IMPDH1 (4390771) and IMPDH2 (4390824)
(Applied Biosystems-Life Technologies). Negative control was
included as siRNA scramble (4390844) (Applied Biosystems-Life
Technologies). Transfected cells were employed for the cytotoxicity
assay as described above, and analyzed after 72 h with ribavirin
treatment.
Results
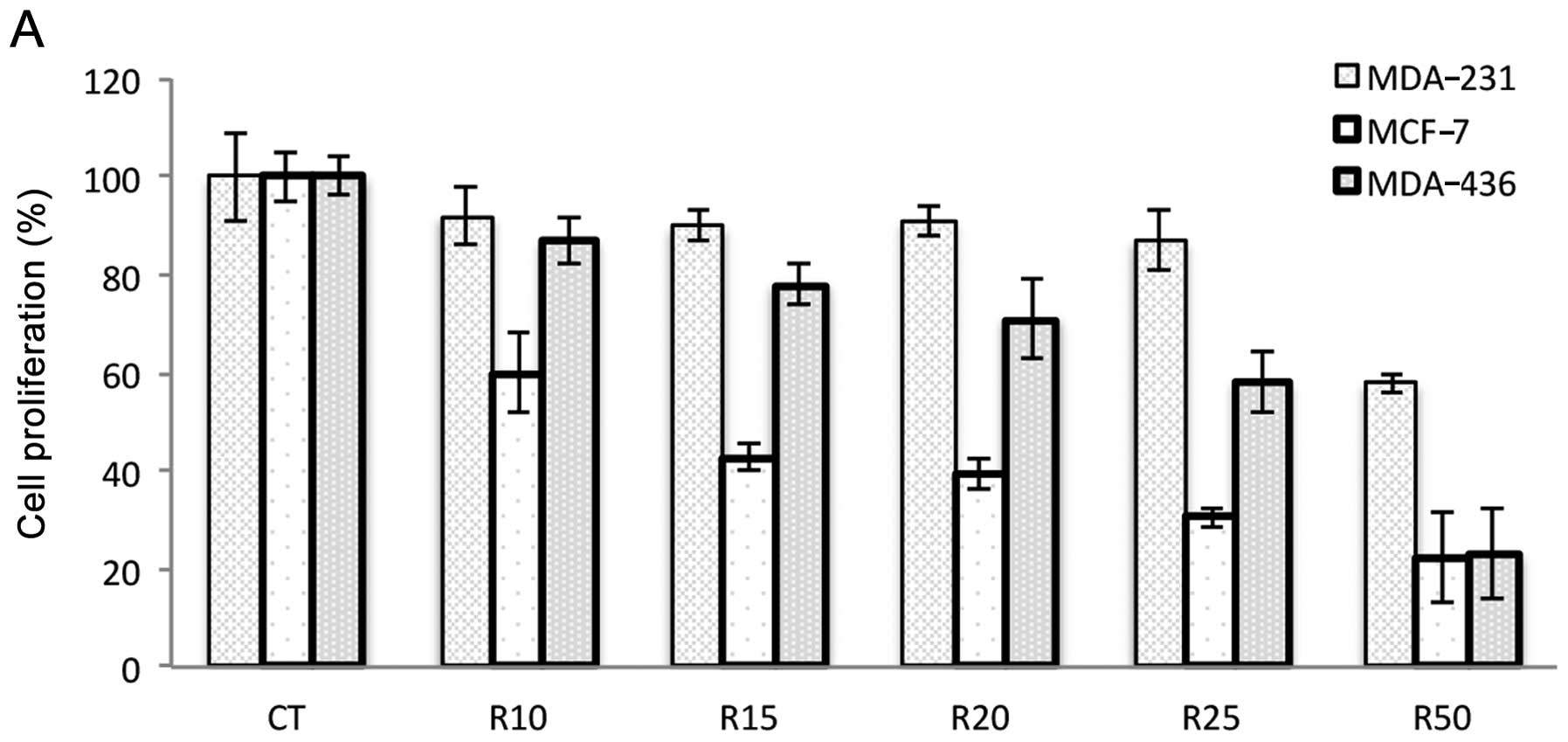
Ribavirin inhibits cell growth
To investigate the growth inhibitory effects of
ribavirin, the cell lines were exposed to different concentrations
of ribavirin (10–50 μM) and cell viability was analyzed at
72 h. The results showed that the cell lines were inhibited in a
dose-dependent manner although the extent of inhibition was cell
line-dependent. The MCF-7 and MDA-436 breast cancer, DU145 prostate
carcinoma and D54 glioma cell lines showed increased inhibition
whereas the SW480 colon cancer, prostate PC3 and breast MDA-231
cells were less inhibited. HeLa cells showed minimal inhibition
even at the highest dose of ribavirin (Fig. 1).
Ribavirin has a high structural
similarity to DZNep
The search for similarity using DZNep as a query and
after visual inspection of the 32 top-ranked compounds with a
significantly increased similarity value compared to the observed
mean value [i.e., mean plus 2 standard deviations (0.59)],
ribavirin (similarity value of 0.70) was selected for additional
three-dimensional comparisons using flexible alignment. Following
the ‘principle of molecular similarity’ (21) which states that, similar compounds
have similar properties (with the exception of the so-called
activity cliffs) (22) we performed
a comparison in silico of the chemical structures of
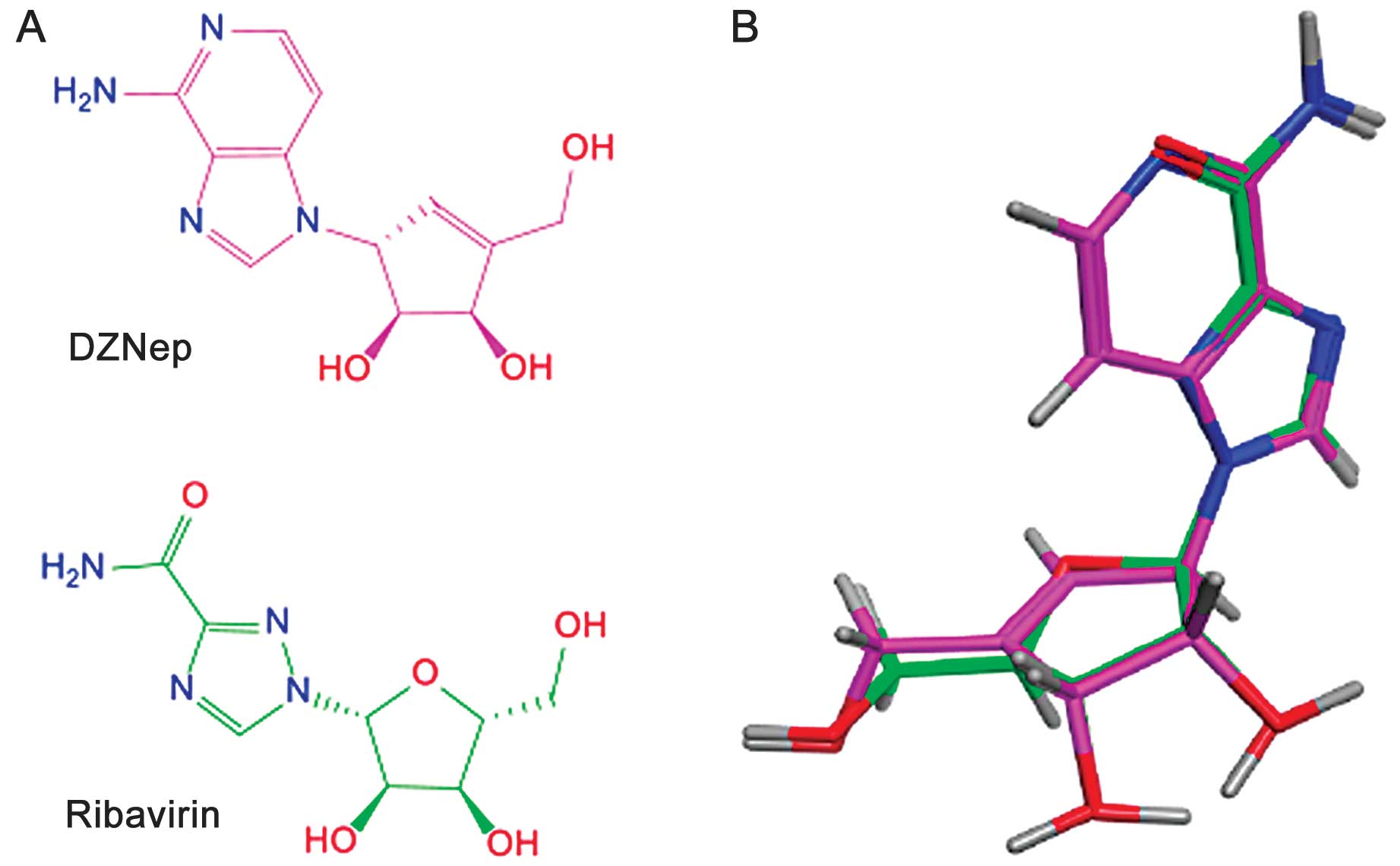
ribavirin and DZNep. Fig. 2A shows
the two-dimensional chemical structures of the two molecules and
the result of the top-ranked solution of the flexible alignment of
both compounds obtained with Molecular Operating Environment
software. Fig. 2B shows the almost
perfect overlay of the pentose ring of ribavirin with the
cyclopenten ring of DZNep. Furthermore, there was an excellent
overlay in three dimensions of the 1,2,4-triazole carboxamide
moiety of ribavirin with the 4-aminoimidazo[4,5-c]pyridine ring of
DZNep leading to a perfect match of several nitrogen atoms,
including the amino groups of the two molecules. The comparison of
the two compounds clearly demonstrated that ribavirin has a high
structural similarity to DZNep in the two- and three-dimensions.
This analysis in silico further supported the hypothesis
that the antiviral compound ribavirin has similar biological
properties as DZNep.
Effects of ribavirin on epigenetic
enzymes
To determine whether ribavirin exerts similar
epigenetic effects compared to DZNep, quantitative RT-PCR was
performed on MCF-7, PC3 and D54 cell lines to evaluate the effect
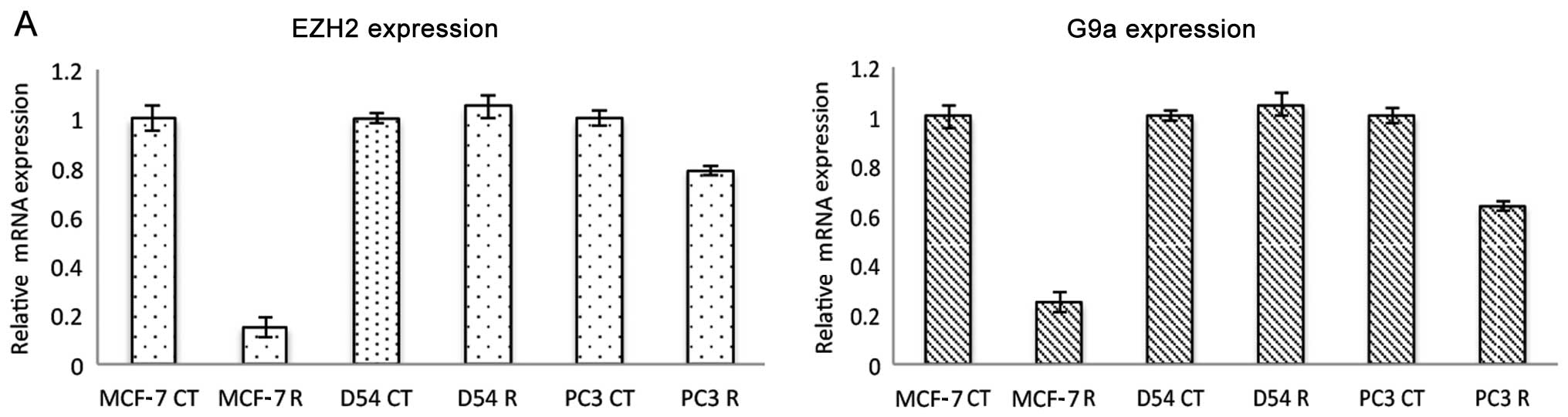
of ribavirin upon several epigenetic enzymes. In MCF-7 cells
ribavirin at 25 μM led to a profound decrease in the level
of EZH2 expression, while the effect was minor in PC3 cells and
absent in D54 cells (Fig. 3A).
Notably, a similar effect was observed for G9A histone
methyltransferase expression. This downregulation was observed at
the protein level as shown in Fig.
3B in MCF-7 cells treated with ribavirin at 25 μM for 72
h and the effect appeared as early as 24 h (Fig. 3C). These data suggested that
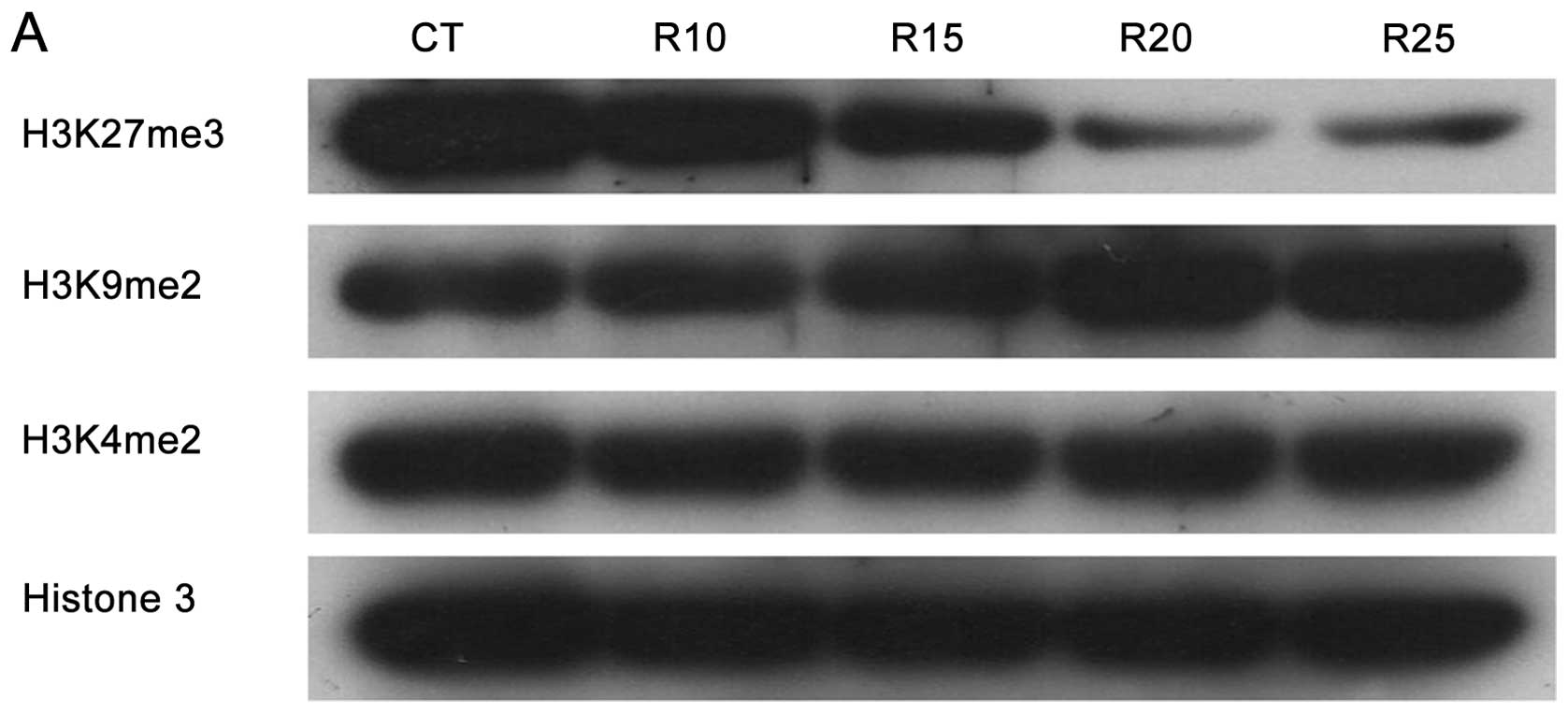
ribavirin inhibits histone methylation targets. Thus, the
methylation levels of H3K27, H3K9 and H3K4 were analyzed by western
blotting. Results in Fig. 4A shows
that while ribavirin clearly decreased trimethylation of H3K27, no
inhibition of methylation was observed for H3K9 and H3K4. To
confirm these findings, an in vitro histone
methyltransferase assay was performed in MCF-7 cells treated with
ribavirin. As shown in Fig. 4B,
ribavirin at concentrations starting at 10 μM showed a clear
inhibition of the histone methyltransferase activity following
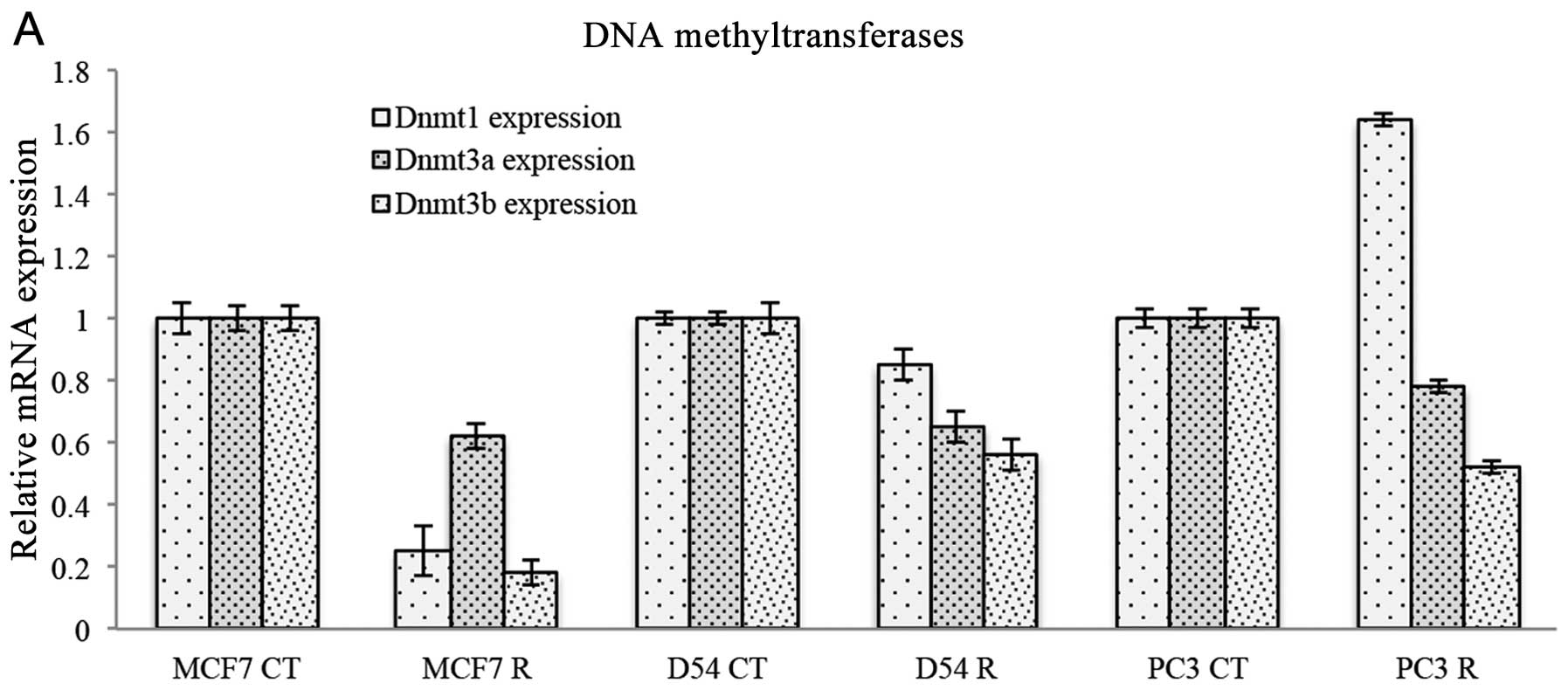
H3K27. The expression of DNA methyltransferases was also evaluated
following treatment with ribavirin. Fig. 5A shows that DNMT1 was inhibited only
in MCF-7 cells, had no effect on D54 cells but was increased in PC3
cells. By contrast, DNMT3a was decreased to almost the same extent
in the three cell lines, while DNMT3b was also decreased in three
cell lines, although an increase was observed in MCF-7 cells. To
explore other epigenetic enzymes that could be modified by
ribavirin, we analyzed HDAC1, which was decreased in MCF-7 and D54
but not in PC3. Ribavirin did not induce changes in the expression
of histone lysine demethylase 1 (LSD1) in these cell lines
(Fig. 5B).
Target inhibition by ribavirin
To determine the extent that each ribavirin target
contributed to the growth inhibitory effects of this drug on the
cancer cell lines investigated, siRNA assays were used to
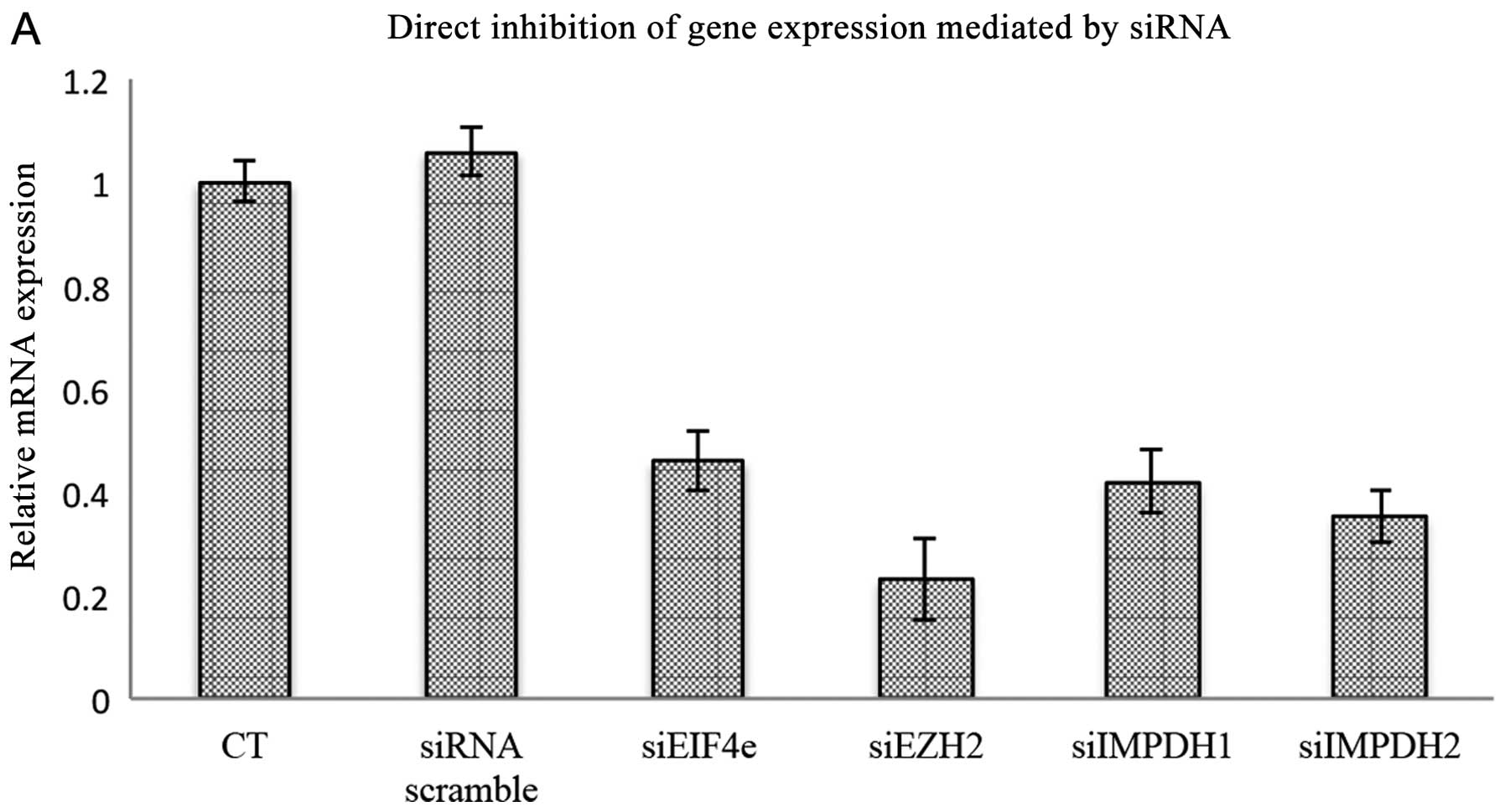
downregulate eIF4E, EZH2 and both IMPDH1 and IMPDH2. As shown in
Fig. 6A, each of the target genes
were downregulated by >50%. Downregulation of EZH2 and eIF4E
reduced cell viability almost to the same extent, with an ~40%
reduction as compared to the control. The extent of growth
inhibition was slightly higher, reaching almost 50% reduction for
IMPDH1 and IMPDH2. No attempt was made to knock the targets down
simultaneously, however, each individual target downregulation led
to higher growth inhibition as compared to ribavirin at 25
μM. Notably, no significant further reduction in viability
was observed when siRNA-transfected cells were treated with
ribavirin.
Discussion
The results of the present study demonstrate that
ribavirin at clinically achievable concentrations had growth
inhibitory effects in vitro on a number of malignant cancer
cell lines. Besides inhibiting eIF4E and IMPDH (4,10)
ribavirin led to downregulation of EZH2 at RNA and protein levels
and inhibition of EZH2 enzyme activity and reduction of H3K27
methylation. These data support the repositioning of ribavirin as
an antitumor drug, specifically in neoplasias whose growth is
associated with alterations in these targets.
The realization that genetic alterations in histone
methyltransferases occur in different types of tumor (23–25)
has led to the development of epigenetic drugs beyond DNA
methyltransferase (DNMTi) and histone deacetylase (HDACi)
inhibitors. Concerning EZH2 histone methyltransferase,
gain-of-function heterozygous point mutations in the SET-coding
domain of the gene have shown to lead transcriptional repression
via trimethylation of H3K27. In addition, the elevated levels of
EZH2 have been associated with silencing of EZH2 target genes and
poor prognosis in solid tumors such as breast, kidney and lung
carcinomas (26–28). A number of EZH2 small-molecule
inhibitors are in preclinical development, with E7438 already being
in a phase I-II trial (unpublished data). Due to the proven
tolerability and extensive use of ribavirin, the results from the
present study suggest its rapid introduction into clinical trials
for neoplasias with alterations at this oncogene.
DZNep is a potent inhibitor of
S-adenosylhomocysteine (AdoHcy) hydrolase, which results in the
accumulation of AdoHyc, which in turn causes by-product inhibition
of S-adenosyl-L-methionine-dependent MTases (29). Initial studies have demonstrated a
selective depletion of methylation targets on H3K27 and H3K20
(15). However, subsequent studies
have shown that according to its proposed mechanism of action as a
global inhibitor of SAM-dependent MTAses, DZNep inhibits histone
methylation in a non-selective manner (30). Nevertheless, our results, although
limited as only two additional methylation targets were analyzed,
suggest that ribavirin is a potential selective histone methylation
inhibitor. In particular, ribavirin induced a decrease in the
trimethylation of H3K27. However, ribavirin did not induce changes
in the methylation targets H3K9 and H3K20. A comprehensive analysis
of histone methylation targets is needed to confirm whether
ribavirin may be selected with H3K27 target.
As previously reported for DZNep, ribavirin also
leads to the depletion of EZH2 RNA in MCF-7 although this effect is
cell line-dependent as it was not observed in D54 glioma cells and
prostate cancer PC3 cells. Of note, ribavirin also depletes H3K9
methyltransferase G9A in MCF-7 cells but not or only slightly in
D54 and PC3, respectively. In this sense, DZNep has been shown to
downregulate another H3K9 methyltransferase, SETDB1, in lung cancer
cells (31). To demonstrate that
ribavirin inhibits the methylation reaction at H3K27, nuclear
extracts of MCF-7 cells treated with or without ribavirin between
10 and 15 μM were assayed with a kit containing the
substrate, the enzyme and adomet. The results in show an almost 50%
reduction in activity. As in mammals, EZH2 is the only enzyme that
catalyzes di- and trimethylation at H3K27 (32). Our results suggest that ribavirin
inhibits EZH2 in a similar manner to DZNep. Ribavirin treatment
also affected the expression of some elements of epigenetic
machinery in a variable manner depending on the cell line. In
particular, the majority of changes were identified in MCF-7 cells,
such as the reduction of DNMT1, DNMT3a, DNMT3b and HDAC1 but not
LSD1. Notably, DNMT3b was consistently decreased in the three cell
lines whereas LSD1 was not affected. To the best of our knowledge,
no information exists on the effect of any EZH2 inhibitor on the
expression of these enzymes. Since DZNep depleted cell proteins of
the core PRC2 complex, as well as several other interacting
proteins of this complex, such as Jarid2, HDAC1/2, Aebp2, Phf1,
MTF2 and Phf19, it is suggested that significant interactions
between polycomb proteins and other chromatin regulators may exist
(33). In this sense, if DNMTs and
HDACs also interact with the complex, then it is expected that they
may be downregulated by ribavirin as a consequence of its depleting
effect on EZH2; however, this remains to be demonstrated. Even for
highly selective drugs, off-target effects are common. Thus,
alternatively, the inhibition of IMPDH and eIF4E by ribavirin
affects the expression of epigenetic machinery components by
unknown mechanisms.
The present study on the antitumor effects of
ribavirin as a tri-targeted therapy has some limitations. It is
clear that growth inhibition while dose-dependent is also dependent
on the cell line. Of the eight lines tested, MCF-7, MDA-436, D54
and DU-145 were the most sensitive. However, we have no information
on the expression level of each of these putative targets on the
model system. In a model of K562 leukemia cells, ribavirin induces
differentiation and apoptosis while modulating the expression of
~60 out of 85 genes having roles in cell proliferation, purine
biosynthesis, translation initiation, oncogenic signaling and cell
survival (34). These effects are
mediated through the modulation of key molecular and metabolic
pathways but most likely result from the composed action of
ribavirin on these three targets. In a study of 6 breast cancer
cell lines investigating the antitumor effects of ribavirin as an
eIF4E antagonist, it was shown that while ribavirin induces growth
inhibition in all the cell lines, the variability in growth
response observed did not correlate with eIF4E expression level
(35). Our results on the
individual downregulation of EZH2, eIF4E and IMPDH1 and IMPDH2 by
siRNAs showed growth inhibition that varied between 36 and
55%, being lower for eIF4E and higher for IMPDH2, which suggests
that in this cell line viability is dependent on IMPDH2. Notably,
in all the cases, the depletion of each of these genes led to
higher growth inhibition as compared to ribavirin alone at 25
μM. However, ribavirin did not significantly increase the
growth inhibition already achieved by downregulating the mRNA of
the gene in any of the cases, suggesting that ribavirin is a
tri-targeted agent. Other potential antitumor effects of ribavirin
were not investigated in the present study. Recently, Kosaka et
al, using an early transposon Oct4 and Sox2 enhancer (EOS)
system to select human prostate cancer cells expressing high levels
of OCT4, a transcription factor that induces pluripotency in
somatic cells, found that ribavirin reverses docetaxel resistance
in prostatic cancer cells most likely by reprogramming cancer stem
cells by downregulating OCT4 expression (36).
Our results are of relevance in the field of drug
repositioning for cancer therapy as these three targets are
commonly altered in cancers, supporting the current shift of drug
identification from single- to multi-target drug development
(37). It is necessary however, to
investigate which and what proportion of tumors share alterations
in these targets and the manner in which their status correlates
with sensitivity to ribavirin. However, a preclinical study to
investigate whether non-Hodgkin lymphoma models with
gain-of-function mutations in EZH2 are as sensitive to ribavirin as
they are to the highly selective EZH2 inhibitor E7438 should be
conducted. If these results are yielded a follow-up clinical study
should be performed taking into account the known safety of
ribavirin.
In conclusion, highly potent and selective
small-molecule inhibitors of EZH2 are currently being investigated
under the vision of ‘rational drug design’. However, ‘promiscuous’
drugs that act on more than one relevant target for cancer biology
such as ribavirin, should be studied to increase therapeutic
armamentarium for cancer patients.
Acknowledgments
This study was supported by Fonsec CONACyT grant
161915, and UNAM PAPIIT grant IT206611.
References
|
1
|
Sidwell RW, Robins RK and Hillyard IW:
Ribavirin: an antiviral agent. Pharmacol Ther. 6:123–146. 1979.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Shepherd J, Jones J, Hartwell D, Davidson
P, Price A and Waugh N: Interferon alpha (pegylated and
non-pegylated) and ribavirin for the treatment of mild chronic
hepatitis C: a systematic review and economic evaluation. Health
Technol Assess. 11:1–205. iii2007. View
Article : Google Scholar : PubMed/NCBI
|
|
3
|
Dueñas-González A, García-López P, Herrera
LA, Medina-Franco JL, González-Fierro A and Candelaria M: The
prince and the pauper. A tale of anticancer targeted agents. Mol
Cancer. 7:822008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Scheidel LM, Durbin RK and Stollar V:
Sindbis virus mutants resistant to mycophenolic acid and ribavirin.
Virology. 158:1–7. 1987. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Itoh O, Kuroiwa S, Atsumi S, Umezawa K,
Takeuchi T and Hori M: Induction by the guanosine analogue
oxanosine of reversion toward the normal phenotype of
K-ras-transformed rat kidney cells. Cancer Res. 49:996–1000.
1989.PubMed/NCBI
|
|
6
|
Jackson RC, Weber G and Morris HP: IMP
dehydrogenase, an enzyme linked with proliferation and malignancy.
Nature. 256:331–333. 1975. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zimmermann A, Gu JJ, Spychala J and
Mitchell BS: Inosine monophosphate dehydrogenase expression:
transcriptional regulation of the type I and type II genes. Adv
Enzyme Regul. 36:75–84. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Nagai M, Natsumeda Y, Konno Y, Hoffman R,
Irino S and Weber G: Selective up-regulation of type II inosine
5′-monophosphate dehydrogenase messenger RNA expression in human
leukemias. Cancer Res. 51:3886–3890. 1991.PubMed/NCBI
|
|
9
|
Chen L and Pankiewicz KW: Recent
development of IMP dehydrogenase inhibitors for the treatment of
cancer. Curr Opin Drug Discov Devel. 10:403–412. 2007.PubMed/NCBI
|
|
10
|
Kentsis A, Topisirovic I, Culjkovic B,
Shao L and Borden KL: Ribavirin suppresses eIF4E-mediated oncogenic
transformation by physical mimicry of the 7-methyl guanosine mRNA
cap. Proc Natl Acad Sci USA. 101:18105–18110. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Borden KL and Culjkovic-Kraljacic B:
Ribavirin as an anticancer therapy: acute myeloid leukemia and
beyond? Leuk Lymphoma. 51:1805–1815. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Assouline S, Culjkovic B, Cocolakis E,
Rousseau C, Beslu N, Amri A, Caplan S, Leber B, Roy DC, Miller WH
Jr and Borden KL: Molecular targeting of the oncogene eIF4E in
acute myeloid leukemia (AML): a proof-of-principle clinical trial
with ribavirin. Blood. 114:257–260. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zagni C, Chiacchio U and Rescifina A:
Histone methyltransferase inhibitors: novel epigenetic agents for
cancer treatment. Curr Med Chem. 20:167–185. 2013. View Article : Google Scholar
|
|
14
|
Simon JA and Lange CA: Roles of the EZH2
histone methyltransferase in cancer epigenetics. Mutat Res.
647:21–29. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Tan J, Yang X, Zhuang L, Jiang X, Chen W,
Lee PL, Karuturi RK, Tan PB, Liu ET and Yu Q: Pharmacologic
disruption of Polycomb-repressive complex 2-mediated gene
repression selectively induces apoptosis in cancer cells. Genes
Dev. 21:1050–1063. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wishart DS, Knox C, Guo AC, Cheng D,
Shrivastava S, Tzur D, Gautam B and Hassanali M: DrugBank: a
knowledgebase for drugs, drug actions and drug targets. Nucleic
Acids Res. 36:D901–D906. 2008. View Article : Google Scholar :
|
|
17
|
Willett P: Similarity-based virtual
screening using 2D fingerprints. Drug Discov Today. 11:1046–1053.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Chemical Computing Group Inc., Montreal,
Quebec, Canada: Molecular Operating Environment (MOE), version
2011.10. http://www.chemcomp.com.
|
|
19
|
Willett P, Barnard JM and Downs GM:
Chemical similarity searching. J Chem Inf Comput Sci. 38:983–996.
1998. View Article : Google Scholar
|
|
20
|
Chan SL and Labute P: Training a scoring
function for the alignment of small molecules. J Chem Inf Model.
50:1724–1735. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bender A and Glen RC: Molecular
similarity: a key technique in molecular informatics. Org Biomol
Chem. 2:3204–3218. 2004. View
Article : Google Scholar : PubMed/NCBI
|
|
22
|
Medina-Franco JL: Scanning
structure-activity relationships with structure-activity similarity
and related maps: from consensus activity cliffs to selectivity
switches. J Chem Inf Model. 52:2485–2493. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Robinson G, Parker M, Kranenburg TA, Lu C,
Chen X, Ding L, Phoenix TN, Hedlund E, Wei L, Zhu X, Chalhoub N,
Baker SJ, Huether R, Kriwacki R, Curley N, Thiruvenkatam R, Wang J,
Wu G, Rusch M, Hong X, Becksfort J, Gupta P, Ma J, Easton J,
Vadodaria B, Onar-Thomas A, Lin T, Li S, Pounds S, Paugh S, Zhao D,
Kawauchi D, Roussel MF, Finkelstein D, Ellison DW, Lau CC, Bouffet
E, Hassall T, Gururangan S, Cohn R, Fulton RS, Fulton LL, Dooling
DJ, Ochoa K, Gajjar A, Mardis ER, Wilson RK, Downing JR, Zhang J
and Gilbertson RJ: Novel mutations target distinct subgroups of
medulloblastoma. Nature. 488:43–48. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ryan RJ and Bernstein BE: Molecular
biology. Genetic events that shape the cancer epigenome. Science.
336:1513–1514. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Shih AH, Abdel-Wahab O, Patel JP and
Levine RL: The role of mutations in epigenetic regulators in
myeloid malignancies. Nature Rev Cancer. 12:599–612. 2012.
View Article : Google Scholar
|
|
26
|
Kleer CG, Cao Q, Varambally S, Shen R, Ota
I, Tomlins SA, Ghosh D, Sewalt RG, Otte AP, Hayes DF, Sabel MS,
Livant D, Weiss SJ, Rubin MA and Chinnaiyan AM: EZH2 is a marker of
aggressive breast cancer and promotes neoplastic transformation of
breast epithelial cells. Proc Natl Acad Sci USA. 100:11606–11611.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wagener N, Macher-Goeppinger S, Pritsch M,
Hüsing J, Hoppe-Seyler K, Schirmacher P, Pfitzenmaier J, Haferkamp
A, Hoppe-Seyler F and Hohenfellner M: Enhancer of zeste homolog 2
(EZH2) expression is an independent prognostic factor in renal cell
carcinoma. BMC Cancer. 10:5242010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Takawa M, Masuda K, Kunizaki M, Daigo Y,
Takagi K, Iwai Y, Cho HS, Toyokawa G, Yamane Y, Maejima K, Field
HI, Kobayashi T, Akasu T, Sugiyama M, Tsuchiya E, Atomi Y, Ponder
BA, Nakamura Y and Hamamoto R: Validation of the histone
methyltransferase EZH2 as a therapeutic target for various types of
human cancer and as a prognostic marker. Cancer Sci. 102:1298–1305.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Chiang PK and Cantoni GL: Perturbation of
biochemical transmethylations by 3-deazaadenosine in vivo. Biochem
Pharmacol. 28:1897–1902. 1979. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Miranda TB, Cortez CC, Yoo CB, Liang G,
Abe M, Kelly TK, Marquez VE and Jones PA: DZNep is a global histone
methylation antagonist inhibitor that reactivates developmental
genes not silenced by DNA methylation. Mol Cancer Ther.
8:1579–1588. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Lee JK and Kim KC: DZNep, inhibitor of
S-adenosylhomocysteine hydrolase, down-regulates expression of
SETDB1 H3K9me3 HMTase in human lung cancer cells. Biochem Biophys
Res Commun. 438:647–652. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Margueron R, Li G, Sarma K, Blais A,
Zavadil J, Woodcock CL, Dynlacht BD and Reinberg D: Ezh1 and Ezh2
maintain repressive chromatin through different mechanisms. Mol
Cell. 32:503–518. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Simon JA and Kingston RE: Occupying
chromatin: Polycomb mechanisms for getting to genomic targets,
stopping transcriptional traffic, and staying put. Mol Cell.
49:808–824. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Kökény S, Papp J, Weber G, Vaszkó T,
Carmona-Saez P and Oláh E: Ribavirin acts via multiple pathways in
inhibition of leukemic cell proliferation. Anticancer Res.
29:1971–1980. 2009.PubMed/NCBI
|
|
35
|
Pettersson F, Yau C, Dobocan MC,
Culjkovic-Kraljacic B, Retrouvey H, Puckett R, Flores LM, Krop IE,
Rousseau C, Cocolakis E, Borden KL, Benz CC and Miller WH Jr:
Ribavirin treatment effects on breast cancers overexpressing eIF4E,
a biomarker with prognostic specificity for luminal B-type breast
cancer. Clin Cancer Res. 17:2874–2884. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Kosaka T, Nagamatsu G, Saito S, Oya M,
Suda T and Horimoto K: Identification of drug candidate against
prostate cancer from the aspect of somatic cell reprogramming.
Cancer Sci. 104:1017–1026. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Medina-Franco JL, Giulianotti MA, Welmaker
GS and Houghten RA: Shifting from the single to the multitarget
paradigm in drug discovery. Drug Discov Today. 18:495–501. 2013.
View Article : Google Scholar : PubMed/NCBI
|