Introduction
Cancer is a leading cause of mortality that
contributes the highest number of deaths from diseases. There were
14.1 million new cases and 8.2 million deaths worldwide in 2012
based on the GLOBOCAN series of the International Agency for
Research on Cancer (1). At the time
of diagnosis, most of the patients have developed invasive cancer.
At the same time, a large number of trials have been performed to
find an effective treatment strategy, but the overwhelming number
of various cancers still carry high risks of morbidity and
mortality (2,3). Therefore, there is a great need and
urgency in the search for early diagnosis biomarkers and novel
treatment measures to improve the survival and the cure rate of
cancer.
Genome-wide association analyses have shown that in
humans, over half of all miRNAs are located at chromosomal fragile
sites, which are associated closely with malignant tumors (4,5). This
suggests that miRNAs play an important role in the development of
malignant tumors. For example, the expression of microRNA-7 (miR-7)
is downregulated in tumor tissues or cancerous cells found in
breast cancer (6), tongue cancer
(7), gastric carcinoma (8), lung neoplasm (9), hepatocellular carcinoma (HCC)
(10), schwannoma (11) and cervical cancer (12). It also plays a significant role in
invasion and metastasis and other tumor biological processes. In
March 2013, two publications reported that circular RNAs (circRNAs)
function as miRNA ‘sponges’ that act to naturally sequester and
competitively suppress the activity of miRNAs (13,14).
These findings suggest that circRNAs may play a role in cancer.
CircRNAs are a large class of non-coding RNAs that
exist ubiquitously in the cytoplasm of eukaryotic cells (15). At present, they are the subject of
intense research in the field of RNAs. Recently, some circRNAs have
been identified, including circRNA sponge for miR-7, known as
ciRS-7. ciRS-7 is also named cerebellar degeneration-related
protein 1 antisense RNA (CDR1as) or CDR1NAT (13,16).
It is ~1,500 nucleotides in length and is found predominantly in
human and mouse brain (16), where
it naturally occurs as an antisense RNA strand and takes on the
structure of a closed loop. As a super sponge or competing
endogenous RNA (ceRNA) of miR-7, ciRS-7 competitively sequesters
and quenches the activity of miR-7 (13,14,17).
This has significant implications for the biology of cancer, as
miR-7 has tremendous effects on the development of a variety of
cancer, such as breast cancer, hepatocellular carcinoma, and
cervical cancer (12). To address
the question whether ciRS-7 is also implicated in cancer, we will
review the origin and functions of circRNAs and in particular,
ciRS-7, and discuss the connection between ciRS-7 and its molecular
targets that are involved in cancer. It is our hope that further
research in this area may provide a new direction toward the early
diagnosis and targeted therapy of cancer.
2. Origin of circRNAs
CircRNAs are ubiquitous in the cytoplasm of
eukaryotic cells, and have returned to the forefront of research
with the discovery of their abundance thanks to technological
breakthroughs in high-throughput deep sequencing. In 1979,
scientists observed for the first time the presence of small RNA
structural variants that were circular in shape in the cytoplasm of
eukaryotic cells by electron microscope (18). These were mistaken as rare existing
transcriptional ‘noise’ (19).
However, with the emergence of high-throughput sequencing and
RNA-seq, several studies have confirmed the widespread existence of
circRNAs in eukaryotic organisms and their role in carrying out
important regulatory functions. The question is then, how are
circRNAs generated?
CircRNAs may be generated from exons (exonic
circRNA) or introns (intronic circRNA). Jeck et al put forth
two models for the origination of circRNAs, namely lariat-driven
circularization and intron-pairing-driven circularization (20,21).
The main difference is in the first step of circRNAs generation.
The first model requires the covalent binding of the splice donor
with the splice acceptor in the exon, while the second model
involves complementary pairing within two introns to form a
circular structure. The subsequent steps of circRNA formation are
largely similar. Namely, the spliceosome excises residual introns
and forms the circRNAs.
In addition, recent studies have shown that there is
an alternative circularization pathway for the production of
circRNAs. This alternative circularization of RNAs relies on the
human genome containing a large number of complementary sequences
(such as Alu sequences) in the intronic regions. The selective
pairing and dynamic regulation of these complementary sequences
allows the same gene to produce multiple circRNAs (22,23).
However, not all of the complementary sequences can produce
circularization. The production of circRNAs is facilitated by short
intronic repeat sequences, while it is suppressed by increasing
hairpin stability between sequences (24). Endogenous conditions of circRNAs
formation are very complicated. For example, the number and
distance of complementary sequences across the circularization
exons can influence matching and competition between complementary
sequences. Alternative circularization is closely related to normal
physiological function, while aberrant alternative circularization
leads to a variety of important human diseases (22), such as cancer.
3. Functions of ciRS-7
miRNA sponges
There are thousands of circRNAs expressed at
different positions in the genome of various species, from archaea
to humans (25–28). Nearly 2,000 types of circRNAs have
been found in humans, 900 types have been identified in mice, and
~700 types exist in nematodes based on sequencing data (14,29).
Among these, ciRS-7 expression shows complex tissue specificity,
cell type specificity, and developmental stage specificity. It has
been reported that (14): i) ciRS-7
binds densely to the miR-7 effector; ii) ciRS-7 contains 74 binding
sites for miR-7, 63 of which exist in at least one other organism;
iii) ciRS-7 and miR-7 possess the same specific expression region
in the embryonic brain tissue of mice; iv) ciRS-7 is circularized
and is not detectable as a linear RNAs; v) ciRS-7 injection shows
similar phenotypes as miR-7 knock-out in zebrafish brains. The
above-described data show that ciRS-7 acts as a ceRNA of miR-7 and
functions as a natural inhibitor of miR-7 and competes miR-7 with
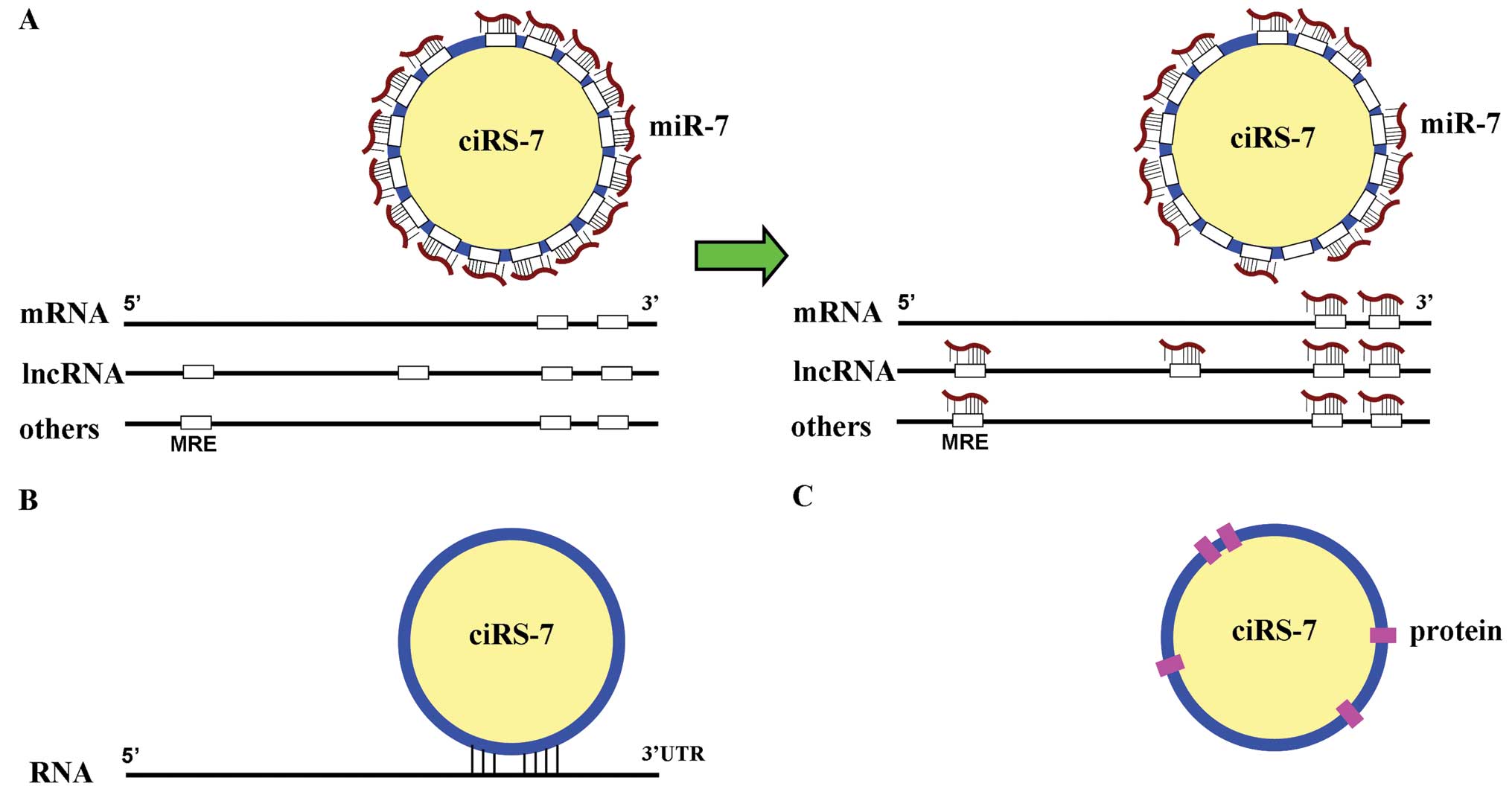
other RNAs, such as mRNAs, and lncRNAs (Fig. 1A).
Transcriptional and posttranscriptional
regulations
Artificial circRNAs contain an internal ribosome
entry site that can be translated in the cellular extract (30), but endogenous circRNAs have not yet
been found to be associated with ribosomes. Hence, linear RNAs and
circRNAs must be closely regulated, as their presence can radically
change the functional output of a gene (20,27).
Owing to its ability to sequester miR-7, ciRS-7 plays an important
role in fine-tuning of the expression of target genes by binding in
trans to the 3′UTRs of a target mRNA (Fig. 1B). The combination of miR-7 with
ciRS-7 may even silence the trans-acting activity. Stable
expression of ciRS-7 in HeLa cells results in significantly lower
expression of α-synuclein (SNCA), epidermal growth factor receptor
(EGFR), and IRS2 mRNA (13). This
result suggests that ciRS-7 presumably regulates the expression of
SNCA, EGFR, and IRS2 by a posttranscriptional mechanism in HeLa
cells. ciRS-7 may be associated with the assembly of large RNAs or
protein complexes, perhaps similar to other molecules with low
complexity (31). In addition,
ciRS-7 acts as an ‘mRNA trap’ by forming closed translation
initiation sites (21), and leaves
the non-coding linear transcript, thereby reducing the expression
levels of protein.
Interaction with binding protein
CiRS-7 may act as ‘scaffolding’ for RNA binding
protein (RBP) to combine multiple protein subunits (21,32)
and to facilitate the interaction by potentially increasing the
stability of the circRNAs transcripts. The capturing mechanism of
miRNA sponges is equally important to RBP. Similarly, ciRS-7 can
store, classify or find RBP. Besides, ciRS-7 also interacts widely
with the argonaute (AGO) protein (14) (Fig.
1C).
To summarize, ciRS-7 can regulate the activities of
miRNAs, mRNAs, and RBP by competing miRNAs with other RNAs
molecules, by binding in trans with the 3′UTRs of target
gene mRNAs, and by interacting with proteins, respectively
(Fig. 1). These are the mechanisms
by which ciRS-7 exerts its specific biological effects.
4. ciRS-7 and cancer
miR-7 and cancer
miRNAs are a group of single-stranded small
non-coding RNAs consisting of 19–24 nucleotides, which regulate
gene expression by interfering with the posttranscriptional process
(33,34). In the past 10 years, growing
evidence suggests that abnormal expression of miRNAs is a sign of
malignant tumors (35,36), as miRNAs can affect cell
proliferation, migration, apoptosis, and much other cancer-related
biological process. miRNAs involved in cancer can be divided into
oncomiRs and tumor suppressor miRNAs. As a tumor suppressor miRNA,
the absence/low expression or dysfunction of miR-7 participates in
the initiation and evolution of various malignancies in humans,
including breast cancer, hepatocellular carcinoma, and cervical
cancer. Liu et al found that (12) miR-7 can inhibit the growth of cancer
cells and promote apoptosis, and can, therefore potentially be a
treatment target in cancer.
miRNAs can prevent or promote the translation and
degradation of target gene by binding to the 3′UTR of target gene
mRNAs, thereby regulating the expression of target genes (37). However, miRNA-mediated upregulation
of target genes is usually specific and depends on distinct target
mRNAs (38). It has been proven
that target genes of miR-7 mainly including EGFR, X-linked
inhibitor of apoptosis protein (XIAP), Kruppel-like factor 4
(KLF4), SNCA, IRS2, IRS1, 4EBP1, ACK1, AKT, FAK, HNF4, IGF1R, mTOR,
NOTCH1, RAF1, p70S6K, PA28c, PAK1, and PIK3CD, all of which play
indispensable roles as oncogenes or tumor suppressor genes in a
variety of cancers.
In cervical cancer, as an example, EGFR is
constitutively highly expressed in pancreatic cancer, oral cancer,
cervical cancer and other cancers, and participates in the
pathogenesis and progression of these cancers (39–41).
The variation in EGFR regulates the risk of developing in
situ and invasive cervical cancer (42); while inhibition of EGFR results in
antitumor effects and can strengthen the curative effect of
chemoradiotherapy based on cisplatin (43). As an oncogene, XIAP expression is
increased in cervical cancer cells, and regulated positively by
TGF-β in a TGF-β/Smad signaling pathway-dependent manner (44). It is associated closely with
survival of cervical cancer patients and promotes oncogenesis.
Also, it regulates the sensitivity of human cervical cancer cells
to cisplatin, doxorubicin, and taxol (45). As a transcription factor of zinc
finger-like structure, KLF4 can inhibit the growth of cancer cells
and tumor formation (46,47). Thus, it functions as a tumor
suppressor in cervical cancer. The above research suggests that
both oncogenes (such as EGFR and XIAP) and tumor suppressor genes
(such as KLF4) can be treated as target molecules of miRNAs, and
play a vital role in the initiation, development, and prognosis of
cancer (such as cervical cancer).
ciRS-7 and cancer
It has been reported that a circular isoform of the
non-coding RNA ANRIL is associated with the risk for the
development of atherosclerosis (15,48).
This finding indicates that circRNAs have biochemical and
phenotypic consequences. Recently, the expression of ciRS-7 was
found to be decreased in the hippocampus of Alzheimer’s patients,
suggesting that there is a deficiency in the ‘sponging’ effects of
ciRS-7 when the level of miR-7 is increasing and the expression of
miR-7-sensitive mRNA targets is decreasing (49). It has been implied that ciRS-7
likely serves as a crucial factor that is significantly involved in
various diseases. In addition, a recent review reported that
expression analyses of various tumor cell lines showing wide spread
expression of ciRS-7 in neuroblastomas and frequent expression in
astrocytoma, renal cell, and lung carcinomas (50). The stable expression of ciRS-7 in
HeLa cells (13,15) indicates that ciRS-7 may be
associated with cervical cancer. There is a large degree of
subcellular co-localization between ciRS-7 and miR-7 in HeLa cells
when analyzed by RNA-fluorescence in situ hybridization and
IF-fluorescence in situ hybridization (13). Moreover, ciRS-7 can be used as an
ideal ceRNA of miR-7, as it sequesters and potently quenches the
normal activities of miR-7 (14).
miR-7 is involved in multiple biological processes of various
cancers; for example, it promotes the growth of cervical cancer
cells. Furthermore, ciRS-7 is sensitive to miR-671. miR-671 can
induce the degradation and endonucleolytic cleavage of ciRS-7, so
ciRS-7 may be responsible for bringing miR-7 to a subcellular
location where miR-671 promotes the release of miR-7 by ciRS-7
(14,51). Thus suggesting that miR-671 may
inhibit the expression of miR-7. From the interaction of ciRS-7
with cancer associated miR-7, we surmise that they have a potential
role in the regulation of cancer (such as cervical cancer).
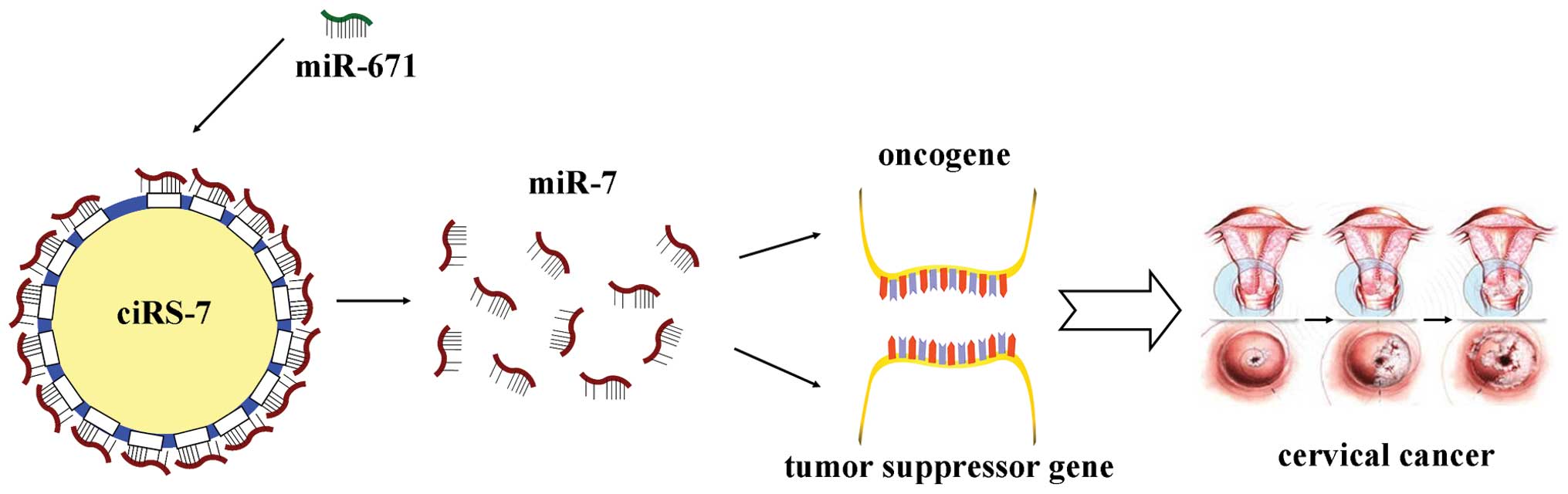
In summary, ciRS-7 may act as a ceRNA of miR-7,
competitively inhibiting the activity of miR-7, promotes the
expression of oncogenes (such as EGFR and XIAP), while inhibiting
tumor suppression genes (such as KLF4), therefore promoting the
initiation and development of cancer (take cervical cancer for
example) (Fig. 2).
5. Potential values of circRNAs in diagnosis
and treatment of cancer
As diagnostic and prognostic
biomarkers
According to the current studies, the main
characteristics of circRNAs are as the following. i) Universality:
a report in 2012 confirmed for the first time that circRNAs are the
most common molecules after linear RNAs in human cells (15) and that they are far more ubiquitous
than previously imagined. ii) Conservatism: the signal behind
circularization seems to be evolutionarily conserved in different
species (52); for example, some
circRNAs are detected in both humans and mice. iii) Cytoplasmic
location. iv) Definite specificity (53). v) Stability: the half-life of
circRNAs in most species is longer than 48 h, while the average
half-life of mRNAs is 10 h (21).
This may be attributable to the resistance of circRNAs to
ribonuclease (RNaseR). Nucleases usually recognize the end of
linear RNA molecules, and the closed structure of circRNAs likely
grants them their high tolerance to nucleases. vi) Highly abundant
expression: the gene product level of some exonic circRNAs has been
shown to be higher than that of linear RNAs as measured by the
sequencing method and quantitative PCR (21,54).
Sometimes, the abundance of circRNAs is over 10 times that of the
corresponding linear mRNAs. This abundance may be attributed to the
stability of circRNAs. Therefore, circRNAs possess distinct
advantages and have the potential to be biomarkers of diagnosis and
prognosis in cancer.
Contribution to targeted therapy
Circularization may be the future target of
treatment, either to reduce the circularization of functional
transcripts, or to sequester by an ‘mRNA trap’, exons contributing
to dysfunctional transcripts (21).
Compared to the miRNA sponges that only contain a string of single
miRNA response elements (MREs), ciRS-7 contains plenty of MREs that
bind miR-7 (51) (Fig. 1A). As such, it can instantaneously
bind or release a large number of miR-7 molecules, thereby
effectively regulating the network of diseases. In comparison with
single miRNA-targeted technology, targeting inhibition of the
expression of ciRS-7 can amplify the protective effect of the tumor
suppressor miR-7. Compared with single RNAi technology targeting
oncogenes, inhibiting the expression of ciRS-7 could affect the
activities of multiple oncogenes. In short, ciRS-7 contains a large
number of MREs, can regulate the expression of several targeted
molecules, and it possesses more advantages in comparison with a
single miRNA or single gene.
In conclusion, as a ceRNA of miR-7, ciRS-7 can
naturally sequester and inhibit the activity of miR-7 and promote
the expression of oncogenic EGFR and XIAP gene as well as suppress
the expression of tumor-suppressed KLF4, thus promoting the
initiation and development of cancer (cervical cancer was is shown
as an example) (Fig. 2). However,
the biological functions and its mechanism still have to be
validated and further investigated. Although studies on circRNAs
are just beginning, the significance in diseases/cancers has
gradually grown. With unique characteristics as biomarkers owing to
its stability and specificity, ciRS-7 can provide a new direction
to improve the early screening and diagnosis in cancer. Also,
ciRS-7 possesses unique characteristics as an ideal ceRNA, which
could provide a novel avenue for the development of treatments
against cancer.
Acknowledgments
This study was supported by National Key Basic
Research Program (973 Program) funded project (2010CB833605),
People’s Republic of China.
References
|
1
|
Ferlay J, Soerjomataram I, Dikshit R, Eser
S, Mathers C, Rebelo M, Parkin DM, Forman D and Bray F: Cancer
incidence and mortality worldwide: Sources, methods and major
patterns in GLOBOCAN 2012. Int J Cancer. 136:E359–E386. 2015.
View Article : Google Scholar
|
|
2
|
Dušek L, Mužík J, Malúšková D, Májek O,
Pavlík T, Koptíková J, Gregor J, Brabec P and Abrahámová J:
Epidemiology of screening-targeted cancers according to new data of
the Czech National Cancer Registry. Klin Onkol. 27(Suppl 2): 19–39.
2014.In Czech. View Article : Google Scholar
|
|
3
|
Rosenberg AR, Kroon L, Chen L, Li CI and
Jones B: Insurance status and risk of cancer mortality among
adolescents and young adults. Cancer. Dec 9–2014.Epub ahead of
print. PubMed/NCBI
|
|
4
|
Zhang X, Hu S, Zhang X, Wang L, Zhang X,
Yan B, Zhao J, Yang A and Zhang R: MicroRNA-7 arrests cell cycle in
G1 phase by directly targeting CCNE1 in human hepatocellular
carcinoma cells. Biochem Biophys Res Commun. 443:1078–1084. 2014.
View Article : Google Scholar
|
|
5
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zhang H, Cai K, Wang J, Wang X, Cheng K,
Shi F, Jiang L, Zhang Y and Dou J: MiR-7, inhibited indirectly by
lincRNA HOTAIR, directly inhibits SETDB1 and reverses the EMT of
breast cancer stem cells by downregulating the STAT3 pathway. Stem
Cells. 32:2858–2868. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jiang L, Liu X, Chen Z, Jin Y, Heidbreder
CE, Kolokythas A, Wang A, Dai Y and Zhou X: MicroRNA-7 targets
IGF1R (insulin-like growth factor 1 receptor) in tongue squamous
cell carcinoma cells. Biochem J. 432:199–205. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kong D, Piao YS, Yamashita S, Oshima H,
Oguma K, Fushida S, Fujimura T, Minamoto T, Seno H, Yamada Y, et
al: Inflammation-induced repression of tumor suppressor miR-7 in
gastric tumor cells. Oncogene. 31:3949–3960. 2012. View Article : Google Scholar
|
|
9
|
Li J, Zheng Y, Sun G and Xiong S:
Restoration of miR-7 expression suppresses the growth of Lewis lung
cancer cells by modulating epidermal growth factor receptor
signaling. Oncol Rep. 32:2511–2516. 2014.PubMed/NCBI
|
|
10
|
Fang Y, Xue JL, Shen Q, Chen J and Tian L:
MicroRNA-7 inhibits tumor growth and metastasis by targeting the
phosphoinositide 3-kinase/Akt pathway in hepatocellular carcinoma.
Hepatology. 55:1852–1862. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Saydam O, Senol O, Würdinger T, Mizrak A,
Ozdener GB, Stemmer-Rachamimov AO, Yi M, Stephens RM, Krichevsky
AM, Saydam N, et al: miRNA-7 attenuation in Schwannoma tumors
stimulates growth by upregulating three oncogenic signaling
pathways. Cancer Res. 71:852–861. 2011. View Article : Google Scholar :
|
|
12
|
Liu S, Zhang P, Chen Z, Liu M, Li X and
Tang H: MicroRNA-7 downregulates XIAP expression to suppress cell
growth and promote apoptosis in cervical cancer cells. FEBS Lett.
587:2247–2253. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Hansen TB, Jensen TI, Clausen BH, Bramsen
JB, Finsen B, Damgaard CK and Kjems J: Natural RNA circles function
as efficient microRNA sponges. Nature. 495:384–388. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Memczak S, Jens M, Elefsinioti A, Torti F,
Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer
M, et al: Circular RNAs are a large class of animal RNAs with
regulatory potency. Nature. 495:333–338. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Salzman J, Gawad C, Wang PL, Lacayo N and
Brown PO: Circular RNAs are the predominant transcript isoform from
hundreds of human genes in diverse cell types. PLoS One.
7:e307332012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Hansen TB, Wiklund ED, Bramsen JB,
Villadsen SB, Statha AL, Clark SJ and Kjems J: miRNA-dependent gene
silencing involving Ago2-mediated cleavage of a circular antisense
RNA. EMBO J. 30:4414–4422. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Salmena L, Poliseno L, Tay Y, Kats L and
Pandolfi PP: A ceRNA hypothesis: The Rosetta Stone of a hidden RNA
language? Cell. 146:353–358. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hsu MT and Coca-Prados M: Electron
microscopic evidence for the circular form of RNA in the cytoplasm
of eukaryotic cells. Nature. 280:339–340. 1979. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Cocquerelle C, Mascrez B, Hétuin D and
Bailleul B: Mis-splicing yields circular RNA molecules. FASEB J.
7:155–160. 1993.PubMed/NCBI
|
|
20
|
Jeck WR, Sorrentino JA, Wang K, Slevin MK,
Burd CE, Liu J, Marzluff WF and Sharpless NE: Circular RNAs are
abundant, conserved, and associated with ALU repeats. RNA.
19:141–157. 2013. View Article : Google Scholar :
|
|
21
|
Jeck WR and Sharpless NE: Detecting and
characterizing circular RNAs. Nat Biotechnol. 32:453–461. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL
and Yang L: Complementary sequence-mediated exon circularization.
Cell. 159:134–147. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Vicens Q and Westhof E: Biogenesis of
Circular RNAs. Cell. 159:13–14. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liang D and Wilusz JE: Short intronic
repeat sequences facilitate circular RNA production. Genes Dev.
28:2233–2247. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Danan M, Schwartz S, Edelheit S and Sorek
R: Transcriptome-wide discovery of circular RNAs in Archaea.
Nucleic Acids Res. 40:3131–3142. 2012. View Article : Google Scholar :
|
|
26
|
Wilusz JE and Sharp PA: Molecular biology.
A circuitous route to noncoding RNA. Science. 340:440–441. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Guo JU, Agarwal V, Guo H and Bartel DP:
Expanded identification and characterization of mammalian circular
RNAs. Genome Biol. 15:4092014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wang PL, Bao Y, Yee MC, Barrett SP, Hogan
GJ, Olsen MN, Dinneny JR, Brown PO and Salzman J: Circular RNA is
expressed across the eukaryotic tree of life. PLoS One.
9:e908592014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ghosal S, Das S, Sen R, Basak P and
Chakrabarti J: Circ2Traits: A comprehensive database for circular
RNA potentially associated with disease and traits. Front Genet.
4:2832013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Chen CY and Sarnow P: Initiation of
protein synthesis by the eukaryotic translational apparatus on
circular RNAs. Science. 268:415–417. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Kato M, Han TW, Xie S, Shi K, Du X, Wu LC,
Mirzaei H, Goldsmith EJ, Longgood J, Pei J, et al: Cell-free
formation of RNA granules: Low complexity sequence domains form
dynamic fibers within hydrogels. Cell. 149:753–767. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Ledford H: Circular RNAs throw genetics
for a loop. Nature. 494:4152013. View
Article : Google Scholar : PubMed/NCBI
|
|
33
|
Maqbool R, Ismail R and Hussain M:
Mutations in microRNA genes and their binding sites are
infrequently associated with human colorectal cancer in the
Kashmiri population. MicroRNA. 2:219–224. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Macha MA, Seshacharyulu P, Krishn SR, Pai
P, Rachagani S, Jain M and Batra SK: MicroRNAs (miRNAs) as
biomarker(s) for prognosis and diagnosis of gastrointestinal (GI)
cancers. Curr Pharm Des. 20:5287–5297. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Calin GA, Sevignani C, Dumitru CD, Hyslop
T, Noch E, Yendamuri S, Shimizu M, Rattan S, Bullrich F, Negrini M,
et al: Human microRNA genes are frequently located at fragile sites
and genomic regions involved in cancers. Proc Natl Acad Sci USA.
101:2999–3004. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Di Leva G, Garofalo M and Croce CM:
MicroRNAs in cancer. Annu Rev Pathol. 9:287–314. 2014. View Article : Google Scholar :
|
|
37
|
Yao J, Liang L, Huang S, Ding J, Tan N,
Zhao Y, Yan M, Ge C, Zhang Z, Chen T, et al: MicroRNA-30d promotes
tumor invasion and metastasis by targeting Galphai2 in
hepatocellular carcinoma. Hepatology. 51:846–856. 2010.PubMed/NCBI
|
|
38
|
Ma C, Qi Y, Shao L, Liu M, Li X and Tang
H: Downregulation of miR-7 upregulates Cullin 5 (CUL5) to
facilitate G1/S transition in human hepatocellular carcinoma cells.
IUBMB Life. 65:1026–1034. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
39
|
Stock AM, Hahn SA, Troost G, Niggemann B,
Zänker KS and Entschladen F: Induction of pancreatic cancer cell
migration by an autocrine epidermal growth factor receptor
activation. Exp Cell Res. 326:307–314. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Ribeiro FA, Noguti J, Oshima CT and
Ribeiro DA: Effective targeting of the epidermal growth factor
receptor (EGFR) for treating oral cancer: A promising approach.
Anticancer Res. 34:1547–1552. 2014.PubMed/NCBI
|
|
41
|
Tomao F, Di Tucci C, Imperiale L, Boccia
SM, Marchetti C, Palaia I, Muzii L and Panici PB: Cervical cancer:
Are there potential new targets? An update on preclinical and
clinical results. Curr Drug Targets. 15:1107–1120. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Ma D, Hovey RL, Zhang Z, Fye S, Huettner
PC, Borecki IB and Rader JS: Genetic variations in EGFR and ERBB4
increase susceptibility to cervical cancer. Gynecol Oncol.
131:445–450. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Nogueira-Rodrigues A, Moralez G,
Grazziotin R, Carmo CC, Small IA, Alves FV, Mamede M, Erlich F,
Viegas C, Triginelli SA, et al: Phase 2 trial of erlotinib combined
with cisplatin and radiotherapy in patients with locally advanced
cervical cancer. Cancer. 120:1187–1193. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Van Themsche C, Chaudhry P, Leblanc V,
Parent S and Asselin E: XIAP gene expression and function is
regulated by autocrine and paracrine TGF-beta signaling. Mol
Cancer. 9:2162010. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Gagnon V, Van Themsche C, Turner S,
Leblanc V and Asselin E: Akt and XIAP regulate the sensitivity of
human uterine cancer cells to cisplatin, doxorubicin and taxol.
Apoptosis. 13:259–271. 2008. View Article : Google Scholar
|
|
46
|
Yang WT and Zheng PS: Promoter
hypermethylation of KLF4 inactivates its tumor suppressor function
in cervical carcinogenesis. PLoS One. 9:e888272014. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Yang WT and Zheng PS: Krüppel-like factor
4 functions as a tumor suppressor in cervical carcinoma. Cancer.
118:3691–3702. 2012. View Article : Google Scholar
|
|
48
|
Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z
and Sharpless NE: Expression of linear and novel circular forms of
an INK4/ARF-associated non-coding RNA correlates with
atherosclerosis risk. PLoS Genet. 6:e10012332010. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Lukiw WJ: Circular RNA (circRNA) in
Alzheimer’s disease (AD). Front Genet. 4:3072013. View Article : Google Scholar
|
|
50
|
Hansen TB, Kjems J and Damgaard CK:
Circular RNA and miR-7 in cancer. Cancer Res. 73:5609–5612. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Kosik KS: Molecular biology: Circles
reshape the RNA world. Nature. 495:322–324. 2013.PubMed/NCBI
|
|
52
|
AbouHaidar MG, Venkataraman S, Golshani A,
Liu B and Ahmad T: Novel coding, translation, and gene expression
of a replicating covalently closed circular RNA of 220 nt. Proc
Natl Acad Sci USA. 111:14542–14547. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Ashwal-Fluss R, Meyer M, Pamudurti NR,
Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N and
Kadener S: circRNA biogenesis competes with pre-mRNA splicing. Mol
Cell. 56:55–66. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Perkel JM: Assume nothing: The tale of
circular RNA. Biotechniques. 55:55–57. 2013.PubMed/NCBI
|