Introduction
Colorectal cancer (CRC) is the third most common
cancer type worldwide (1) and the
second most common one in Korea (2). Early diagnosis of CRC is critical to
improve oncological outcomes and reduce medical expenses (3). In this study, identification of
CRC-specific biomarkers may be useful for diagnosis,
prognostication or prediction of therapeutic responses. The
biomarkers for CRC identified to date are limited to serum
carcinoembryonic antigen levels, mutations of KRAS, NRAS or BRAF
and microsatellite instability in CRC tissues. Previous studies by
our group reported fecal calgranulin B as a candidate maker for CRC
diagnosis. However, calgranulin B was additionally detected in the
stool of inflammatory bowel disease (IBD) patients (4,5). In a
subsequent study aimed at determining a complementary fecal marker
for CRC or IBD, deleted in malignant brain tumors 1 (DMBT1) was
incidentally identified in stools of CRC as well as IBD patients
[Yoo et al (unpublished data)].
DMBT1 is located on chromosome 10q25.3-q26.1
(6) and encodes three secretory
glycoproteins, specifically, DMBT1 protein, salivary agglutinin
(DMBT1SAG) and lung glycoprotein-340
(DMBT1gp340). DMBT1SAG and
DMBT1gp340 secreted from the salivary gland and
respiratory tract, respectively, share identical peptide sequences
(7) and monoclonal antibody
cross-reactivity (8). The DMBT1
protein is involved in mucosal innate immunity via binding to
various pathogens and host molecules, including surfactant protein,
immunoglobulin A and complement component 1q (9). However, upon secretion into the
extracellular matrix, DMBT1 triggers terminal differentiation of
epithelial cells (10,11). DMBT1 is mainly synthesized in the
trachea, lung, small intestine, salivary gland and stomach
(12). Expression of the protein is
increased in gastrointestinal epithelium of patients with IBD
(13) and Helicobacter
pylori-associated gastritis (14).
Due to its repetitive genomic structure, DMBT1 is
susceptible to genomic instability (15). Frequent homozygous deletion of DMBT1
was initially reported in medulloblastoma and glioblastoma
(6), as reflected by the designated
name ‘deleted in malignant brain tumor’. In subsequent studies,
deletion and/or loss of mRNA expression were observed in
esophageal, gastric, colorectal, lung and breast cancers (16–18),
supporting the theory that DMBT1 is a tumor suppressor gene.
Infrequent mutation of DMBT1 has been reported in astrocytic tumors
(19). In addition, upregulation of
DMBT1 has been reported in specific glioblastomas (20) and carcinomas of the stomach
(21) and lung (22). These diverse observations imply
complex roles of DMBT1 in carcinogenesis. To clarify the specific
functions of DMBT1 in various cancer types, studies focusing on its
clinicopathological impact are required. The present study
investigated the diagnostic utility of fecal and tissue expression
of DMBT1 in CRC along with its clinicopathological and prognostic
significance.
Materials and methods
Stool sample preparation and
patients
Initially, a liquid chromatography-mass spectrometry
(LC-MS) analysis of DMBT1 in stool from IBD patients (n=3) was
performed, which was extended to various groups of colorectal
disease (IBD, CRC and colorectal adenomas). The analysis was
performed using two sets: Development and validation. A total of
184 patients were enrolled in the developing (test) set (CRC, n=81;
adenoma, n=32; IBD, n=20; and control, n=51), while the validation
set included 256 individuals (CRC, n=93; adenoma, n=29; IBD, n=34;
and control, n=100) from May 2007 to December 2009. Colonoscopy was
performed with preparation and sedation according to individual
patient characteristics. CRC and colorectal adenoma were diagnosed
using colonoscopy and histopathology. Histopathological diagnosis
was performed by one pathologist (HJC) according to the World
Health Organization classification (23) and the 6th edition of the
Tumor-Nodes-Metastasis classification of the American Joint
Committee on Cancer (24). Advanced
adenoma (AA) were defined as polyps of >1 cm in size, villous
histopathology, high-grade dysplasia or carcinoma in situ
(25). IBD cases included
ulcerative colitis (n=35), Crohn's disease (n=12) and IBD,
unclassified (n=7) and those were diagnosed based on patient
symptoms, laboratory and colonoscopy parameters and
histopathological features (26–29).
Colonoscopy revealed negative findings in the control group. The
persons with an incomplete colonoscopy, or hyperplastic polyps, or
inflammatory polyps were excluded from this study.
Stool samples were collected
prospectively prior to bowel preparation
Proteins were extracted from 0.1 g of stool in 0.3
ml PBS containing protease inhibitors via vigorous vortexing,
followed by centrifugation at 12,000 × g for 10 min at room
temperature. The supernatant was collected without disturbing the
pellet and used for western blotting and LC-MS analyses, in
accordance with previous studies (4,5).
SDS-PAGE and LC-MS analysis
Stool proteins from IBD patients were measured using
Bicinchoninic Acid Protein Assay kit (Pierce; Thermo Fisher
Scientific, Inc., Waltham, MA, USA). Equivalent amounts of stool
protein (10 µg) were electrophoresed on 4–12% Bis-Tris Protein Gel
(Invitrogen; Thermo Fisher Scientific, Inc.), then fixed in a fixer
solution (40% methanol, 10% acetic acid) for 10 min and stained
using a Colloidal Blue Staining kit (Invitrogen; Thermo Fisher
Scientific, Inc.), according to the manufacturer's instructions.
Gel slices containing stool proteins from IBD patients were
excised, destained with 50% acetonitrile in 0.1 M ammonium
bicarbonate and dried in a SpeedVac concentrator (Eppendorf,
Hamburg, Germany) at room temperature. The dried gel pieces were
immersed in 30 µl sodium bicarbonate solution (25 mM, pH 8.8)
containing 50 ng trypsin (Promega Corp., Madison, WI, USA) at 37°C
overnight. Samples were desalted using Zip-Tips C18 (EMD Millipore,
Billerica, MA, USA) and dissolved in 10 µl 2% acetonitrile in 0.1%
formic acid. Analysis was performed using a linear ion trap mass
spectrometer system (LTQ-XL; Thermo Electron Corp., San Jose, CA,
USA) at the Facility of Omics Core Laboratory, National Cancer
Center (Goyang, Korea). Peptide separation was performed on an
Agilent 1100 system (Agilent Technologies, Santa Clara, CA, USA). A
3-µl sample of the peptide mixture was injected onto a C18-PepMap
column (150 mm; inner diameter, 75 µm; LC Packings, San Francisco,
CA, USA), and separated with a gradient of 5–50% acetonitrile in
0.1% formic acid over 60 min. The spray voltage was set at +1.7 kV
and capillary temperature at 200°C. The capillary voltage was set
at +20 V, the tube lens voltage at +100 V and the auxiliary gas at
zero. Full scan experiments were performed on the linear ion trap
mass spectrometer in the mass/charge range of 150-2,000. Systematic
MS/MS experiments were performed by changing the relative collision
energy and monitoring the intensities of the fragment ions. All
MS/MS samples were analyzed using the SEQUEST algorithm v.27, rev.
11 (Thermo Fisher Scientific, Inc.). SEQUEST was set up the
SwissProt (ftp://ftp.ebi.ac.uk/pub/databases/uniprot/knowledgebase)
and IPI human databases (ftp://ftp.ebi.ac.uk/pub/databases/IPI/current)
assuming trypsin as the digestion enzyme. A fragment ion mass
tolerance of 1.00 Da and parent ion tolerance of 1.2 Da were used
for the SEQUEST search.
Western blot analysis
Equivalent amounts of stool protein (10 µg) as
determined by a Bicinchoninic Acid Protein Assay kit (Pierce;
Thermo Fisher Scientific, Inc.) were subjected to SDS-PAGE (4–12%
Bis-Tris Protein Gel). After electrophoresis, the proteins were
transferred to polyvinylidene difluoride membranes (EMD Millipore),
followed by blocking via incubation in 1.5% non-fat dried milk
(Bio-Rad Laboratories, Inc., Hercules, CA, USA) and 1 mM
MgCl2 in Tris-buffered saline containing 1% Tween-20 for
2 h at 4°C. Membranes were incubated for 2 h at room temperature
with primary antibody against DMBT1 (1:1,000, cat. no. sc-80616;
Santa Cruz Biotechnology, Inc., Dallas, TX, USA), washed for 3×15
min with blocking solution, and incubated with diluted goat
anti-mouse Ig human ads-horseradish peroxidase (HRP)-conjugated
secondary antibody (1:10,000, cat. no. 1010-05; Southern Biotech,
Birmingham, UK) for 1 h at room temperature. This step was followed
by washing with blocking solution (3×15 min), incubation with
WEST-ZOL® plus chemiluminescence reagent (iNtRON
Biotechnology, Gyeonggi, Korea) for 1 min, at room temperature and
exposure to X-ray film (Kodak Blue XB-1; Eastman Kodak, Rochester,
NY, USA). The expression of total DMBT1 (Santa Cruz Biotechnology,
Inc.) in the stool samples of 325 subjects was measured via western
blot analysis and optical densities (ODs; arbitrary units of DMBT1
immunoreactive signal) of blots were quantified using TINA software
(version 2.10e; Raytest Isotopenmegerate GmbH, Straubenhardt,
Germany) after image scanning.
CRC tissue samples
Tissue microarray (TMA) blocks were constructed from
formalin-fixed paraffin-embedded tissues of 385 patients with CRC
who underwent surgical resection at the National Cancer Center of
Korea (Goyang-si, Korea) from January 2003 to December 2003. One
representative core tissue (2 mm in diameter) was taken from a
paraffin block of each case and arranged in a new recipient
paraffin block using a trephine apparatus (Superbiochips
Laboratories, Seoul, Korea). Each TMA block contained one core of
normal colorectal mucosal tissue (total number of normal
tissues=8). Sections of 4 µm in thickness were obtained from each
TMA block for immunohistochemical analysis. Clinical charts and
pathological reports of the 385 patients were reviewed. The tissue
study population included 152 women and 231 men with a mean age of
58 years (range, 25–86 years) and follow-up time of 64 months
(range, 2–74 months).
Immunohistochemistry
Immunohistochemical staining was performed using a
BenchMark XT automated slide stainer (Ventana Medical Systems,
Inc., Tucson, AZ, USA). De-waxing was implemented using the
BenchMark module (Ventana Medical Systems, Inc.). Antigen retrieval
was performed with Cell Conditioning solution 1/EDTA (pH 8.0) for
30 min at 98°C using the BenchMark staining module (Ventana Medical
Systems, Inc.). Sections were incubated with mouse monoclonal
anti-gp340 (5D7) antibody [conjugated with goat anti-mouse IgG
H&L HRP (cat. no. ab205719, 1:100, cat. no. ab17779; Abcam,
Cambridge, UK)] for 40 min at 42°C, and staining was performed on a
Ventana Bench mark XT autostainer and evaluated with an I-View
diaminobenzidine detection kit (both from Ventana Medical Systems,
Inc.). Slides were counterstained with hematoxylin II for 4 min at
room temperature and Bluing reagent (cat. no. 760-2037; Ventana
Medical Systems, Inc.) for 4 min at room temperature. For the
negative control, tissue sections were incubated with Tris-buffered
saline alone without the primary antibody. DMBT1 expression was
quantitatively evaluated using a double scoring system by
estimating the staining intensity and percentage of stained cancer
cells. The staining intensity was classified as 1 (weak), 2
(moderate) or 3 (strong). Immunoreactivity was scored as 0–300 by
multiplying the staining intensity by the percentage of cells
stained. All immunohistochemical staining results were evaluated
independently by two pathologists (HSP and HJC).
Statistical analysis
The OD values of DMBT1 expression levels in stool
are expressed as the mean ± standard deviation. One-way analysis of
variance (ANOVA) and the χ2 test were used to compare
DMBT1 levels among CRC, IBD, adenoma and control groups. In order
to compare stool DMBT levels between two individual groups, one-way
ANOVA was conducted, followed by a post-hoc Dunnett's test. The
diagnostic potential of fecal DMBT1 was evaluated using receiver
operator characteristics (ROC) curves, and the area under the ROC
curve (AUC) was calculated. Cut-off values for each marker were
determined from ROC and AUC values.
The χ2 test was additionally employed to
evaluate the association between the expression of DMBT1 determined
by immunohistochemical analysis and clinicopathological features.
Overall or disease-free survival was estimated using the
Kaplan-Meier method and compared with the log-rank test. The
prognostic value of protein expression was determined via
multivariate analysis using the Cox proportional hazards regression
model. P<0.05 was considered to indicate a statistically
significant difference. All statistical analyses were performed
using SPSS version 17.0 (SPSS Inc., Chicago, IL, USA).
Results
LC-MS analysis of DMBT1 in stool
samples of IBD and CRC patients
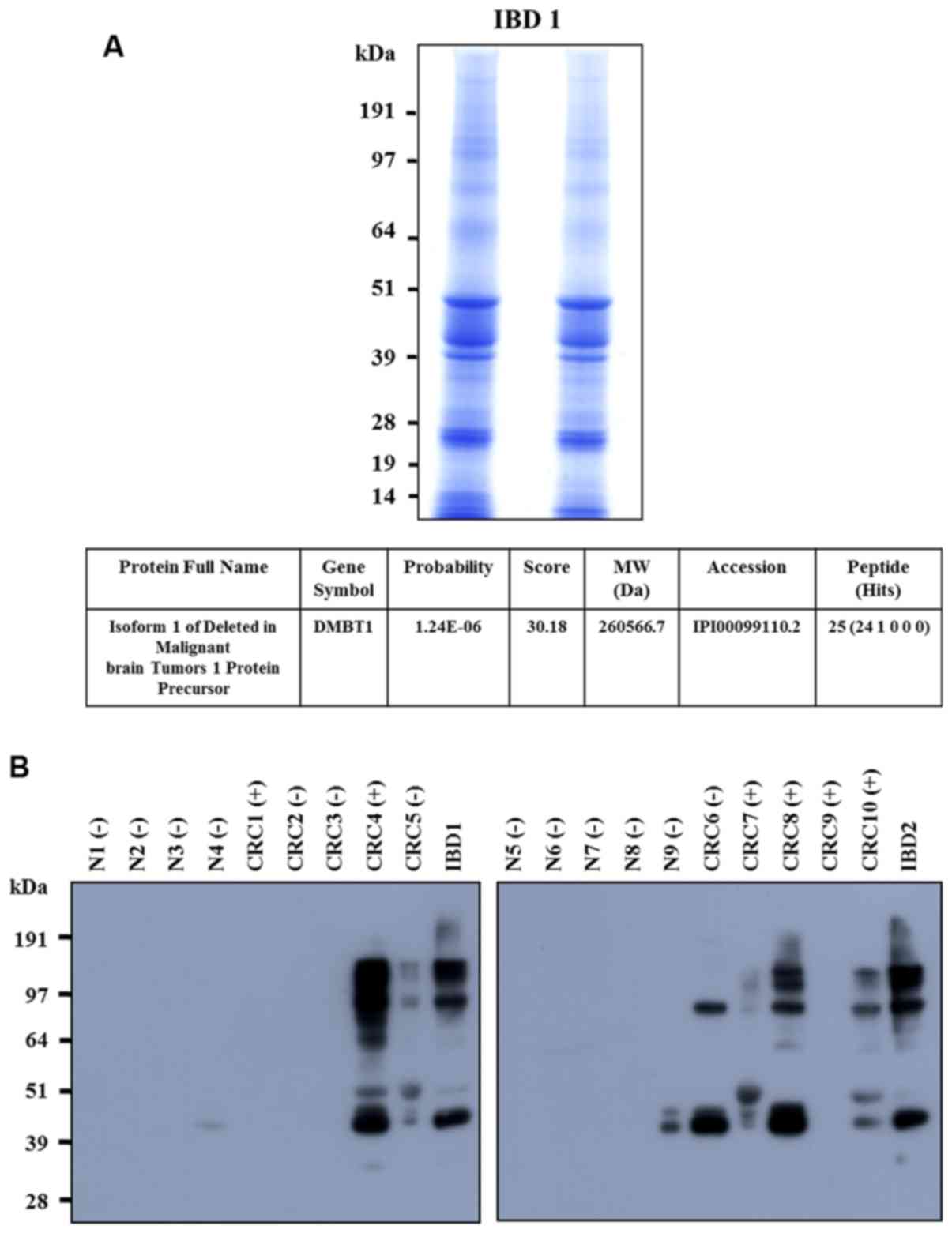
Stool extracts from IBD patients were separated via
SDS-PAGE. Gel slices were excised, digested with trypsin and
analyzed using LC-MS to determine the peptide mass. A SEQUEST
database search led to the identification of isoform 1 of DMBT1
(Fig. 1A). Upregulation of DMBT1
protein in stool samples of IBD and CRC patients relative to that
in stool samples from healthy individuals was confirmed based on
western blot analysis (Fig.
1B).
Fecal DMBT1 expression in CRC,
adenoma, IBD and control groups
The mean fecal DMBT1 protein expression was higher
in patients with CRC than that in the healthy control (P<0.0001)
and adenoma groups (P<0.0001) as well as higher in the IBD than
in the healthy control (P<0.01) and adenoma groups (P<0.05).
However, fecal DMBT1 levels were not significantly different
between patients with IBD and CRC or adenoma and healthy controls
in the test and validation sets (P>0.05). Similar results were
obtained with the combined set (Table
I).
 | Table I.OD values of fecal DMBT1 protein
precursor expression according to colorectal disease type. |
Table I.
OD values of fecal DMBT1 protein
precursor expression according to colorectal disease type.
| Groups | Test set | Validation set | Combined set |
|---|
| Cancer | (n=81)a,b,e | (n=93)a,b,e | (n=174)a,b,e |
|
| 945.48±1016.10 | 1086.94±1161.10 | 1021.09±1095.17 |
| Adenoma | (n=32)e | (n=29)e | (n=61)e |
|
| 310.79±603.17 | 409.26±678.90 | 357.60±636.80 |
| IBD | (n=20)c,d | (n=34)c,d | (n=54)c,d |
|
| 914.61±905.93 |
1079.90±1056.28 | 1018.68±997.70 |
| Control | (n=51) | (n=100) | (n=151) |
|
| 268.57±603.03 | 234.57±600.73 | 246.05±599.72 |
The AUC value for fecal DMBT1 compared between the
CRC and control groups was 0.684. The cut-off value for DMBT1 per
mg of stool protein was calculated as 1,224.0 OD. The proportion of
patients positive for fecal DMBT1 was not significantly different
among the subgroups of CRC patients according to CRC stage or site,
or between advanced and non-advanced adenoma (Table II). The expression of fecal DMBT1
was also not different among the subgroups of IBD (Table II). The sensitivity of the fecal
DMBT1 test for CRC, CRC+AA or CRC+AA+IBD was not high (48.9–50%),
while the specificity and positive predictive value (PPV) were
moderate (specificity, 86.8%; PPV, 81.3–85.3% compared with the
control; Table III).
 | Table II.Number of patients with fecal DMBT1
expression (cut-off value of OD >1,224.05) in each group of
combined-set diseases. |
Table II.
Number of patients with fecal DMBT1
expression (cut-off value of OD >1,224.05) in each group of
combined-set diseases.
| Pathology | n | Positive cases (n,
%) |
P-valueb |
P-valuef |
|---|
| Cancer | 174 | 87 (50.0) |
<0.0001c,e |
|
| pT
stage |
|
|
|
|
|
T1 | 24 | 14 (58.3) |
| 0.537 |
|
T2 | 17 | 7
(38.9) |
|
|
|
T3 | 105 | 51 (48.6) |
|
|
|
T4 | 27 | 15 (55.6) |
|
|
|
Location |
|
|
|
|
|
Right | 46 | 23 (50.0) |
| 1.000 |
|
Left | 128 | 64 (50.0) |
|
|
| IBDd | 54 | 27 (50.0) | 0.970d |
|
| UC | 35 | 19 (54.3) |
| 0.693 |
| CD | 12 | 5
(41.7) |
|
|
|
Unclassifieda |
7 | 3
(42.9) |
|
|
|
Adenomae | 61 | 14 (23.0) |
|
|
| AA |
8 | 2
(25.0) |
| 0.882 |
|
Non-AA | 53 | 12 (22.6) |
|
|
|
Controlsc | 151 | 20 (13.2) |
|
|
 | Table III.Sensitivity, specificity and
predictive value of fecal DMBT1 measurement for each group of
combined-set diseases. |
Table III.
Sensitivity, specificity and
predictive value of fecal DMBT1 measurement for each group of
combined-set diseases.
| Disease type of
each patient | Sensitivity | Specificity | PPV | NPV |
|---|
| CRC vs.
controls | 87/174
(50.0) | 131/151 (86.8) | 87/107
(81.3) | 131/218 (60.1) |
| CRC+AA vs.
controls | 89/182
(48.9) | 131/151 (86.8) | 89/109
(81.7) | 131/224 (58.5) |
| CRC+AA+IBD vs.
controls | 116/236 (49.1) | 131/151 (86.8) | 116/136 (85.3) | 131/251 (51.2) |
Tissue expression of DMBT1 protein and
clinicopathological correlation in CRC
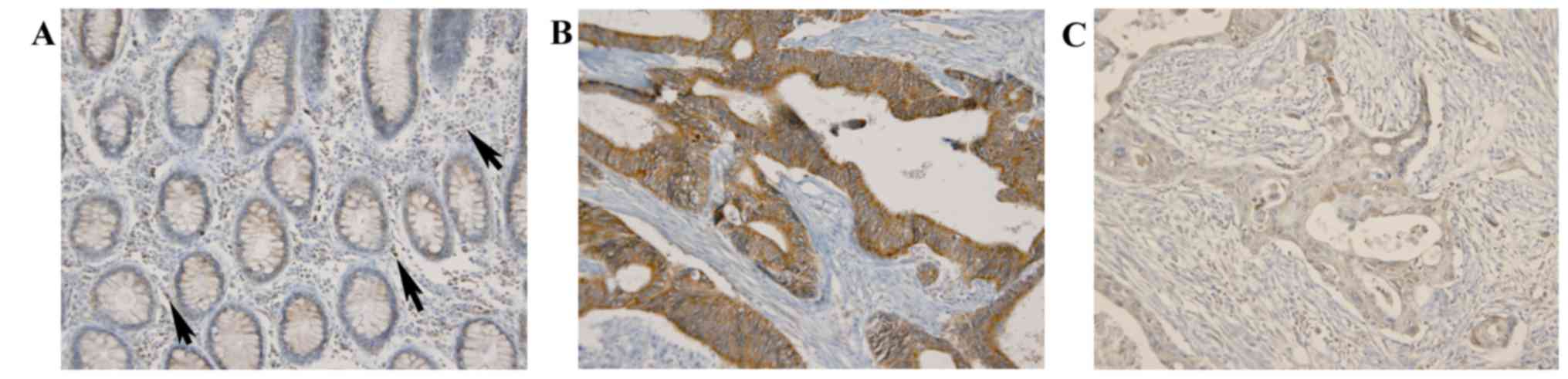
In normal colorectal crypt epithelium, DMBT1
exhibited diffuse mild cytoplasmic expression with an average
immunoreactive score (IR) of 90; DMBT1 was also expressed in
inflammatory cells infiltrating stroma (Fig. 2). In CRC cells, DMBT1 expression was
increased or decreased (Fig. 2)
with a mean IR value of 117.26±58.71 (range, 0–285). For the
χ2 test, DMBT1 tissue expression was classified as
follows: Group 1 (IR ≤10), group 2 (IR >10 and <150) and
group 3 (IR ≥150). A total of 18 out of 385 tumors (4.7%) were
classified as group 1, 255 (66.2%) as group 2 and 112 (29.1%) as
group 3. Group 1 (DMBT1 loss) was significantly associated with
lymph node metastasis (P=0.016), distant metastasis (P=0.013),
advanced stage (P=0.026) and higher histological grade (P=0.033).
Conversely, group 3 (DMBT1 overexpression) was significantly
associated with absence of lymph node metastasis (N0) or distant
metastasis (M0), lower stage and low histological grade (Table IV).
 | Table IV.DMBT1 expression in CRC tissues and
its association with clinical and pathological factors (n=385). |
Table IV.
DMBT1 expression in CRC tissues and
its association with clinical and pathological factors (n=385).
|
| DMBT1
expression |
|
|---|
|
|
|
|
|---|
| Characteristic | IRS≤10 |
10<IRS<150 | IRS≥150 | P-value |
|---|
| Sex |
|
|
| 0.764 |
|
Male | 11 (4.8) | 156 (67.5) | 64 (27.7) |
|
|
Female | 7 (4.5) | 99 (64.3) | 48 (31.2) |
|
| Age (years) |
|
|
| 0.090 |
|
<60 | 14 (6.8) | 137 (66.2) | 56 (27.1) |
|
|
≥60 | 4 (2.2) | 118 (66.3) | 56 (31.5) |
|
| T-stage |
|
|
| 0.839 |
| pT1,
2 | 2 (3.4) | 40 (69.0) | 16 (27.6) |
|
| pT3,
4 | 16 (4.9) | 215 (65.7) | 96 (29.4) |
|
| N-stage |
|
|
| 0.016 |
|
pN0 | 3 (1.9) | 101 (63.1) | 56 (35.0) |
|
| pN1,
2 | 15 (6.7) | 154 (68.4) | 56 (24.9) |
|
| M-stage |
|
|
| 0.013 |
| M0 | 8 (2.9) | 184 (65.9) | 87 (31.2) |
|
| M1 | 10 (9.4) | 71 (67.0) | 25 (23.6) |
|
| Stage |
|
|
| 0.026 |
| I,
II | 3 (2.0) | 95 (62.9) | 53 (35.1) |
|
| III,
IV | 15 (6.4) | 160 (68.4) | 59 (25.2) |
|
| Venous
invasion |
|
|
| 0.880 |
|
Absent | 11 (4.4) | 167 (67.1) | 71 (28.5) |
|
|
Present | 7 (5.1) | 88 (64.7) | 41 (30.1) |
|
| Lymphatic
invasion |
|
|
| 0.476 |
|
Absent | 7 (5.0) | 88 (62.4) | 46 (32.6) |
|
|
Present | 11 (4.5) | 167 (68.4) | 66 (27.0) |
|
| Perineural
invasion |
|
|
| 0.158 |
|
Absent | 10 (3.8) | 172 (64.7) | 84 (31.6) |
|
|
Present | 8 (6.7) | 83 (69.7) | 28 (23.5) |
|
| Histological
grade |
|
|
| 0.033 |
|
Low | 14 (4.1) | 222 (64.9) | 106 (31.0) |
|
|
High | 4 (9.3) | 33 (76.7) | 6 (14.0) |
|
Loss of DMBT1 expression and patient
outcomes
Since group 1 (DMBT1 loss) was associated with
adverse clinicopathological features, patients were subdivided into
two groups according to their DMBT1 status (loss vs. no loss) for
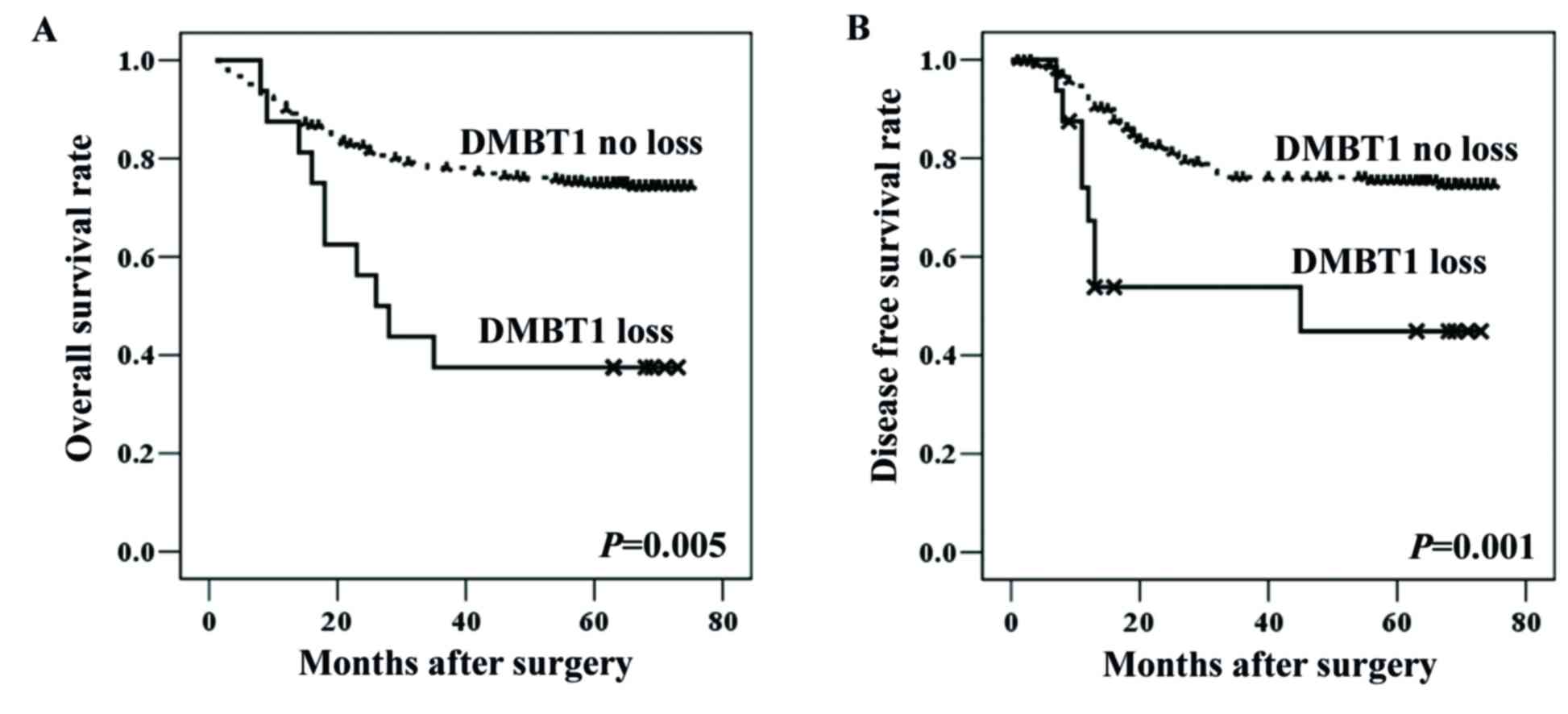
survival analysis. According to the univariate survival analysis,
CRC patients with DMBT1 loss (group 1) had significantly shorter
overall (P=0.005) and disease-free (P=0.001) survival than those
without DMBT1 loss (groups 2 and 3; Table V; Fig.
3). According to the multivariate survival analysis (Table VI), loss of DMBT1 was an
independent prognostic factor associated with poor overall survival
(hazard ratio, 2.272; 95% confidence interval, 1.175–4.391;
P=0.015) and disease recurrence (hazard ratio, 2.689; 95%
confidence interval, 1.275–5.670; P=0.009) in CRC patients after
adjusting for stage, as well as venous, lymphatic and perineural
invasion. Furthermore, the stage, as well as venous and perineural
invasion were independent prognostic factors associated with
overall and disease-free survival.
 | Table V.Univariate survival analysis in CRC
patients (n=385). |
Table V.
Univariate survival analysis in CRC
patients (n=385).
|
| Overall survival
(months) | Disease-free
survival (months) |
|---|
|
|
|
|
|---|
| Variables | Mean | 95% CI | P-value | Mean | 95% CI | P-value |
|---|
| Sex |
|
| 0.6842 |
|
| 0.0483 |
|
Male | 60 | 56–63 |
| 63 | 59–66 |
|
|
Female | 60 | 56–64 |
| 56 | 52–61 |
|
| Age (years) |
|
| 0.1964 |
|
| 0.6238 |
|
<60 | 61 | 58–64 |
| 59 | 55–63 |
|
|
≥60 | 58 | 54–62 |
| 61 | 57–65 |
|
| Stage |
|
| <0.0001 |
|
| <0.0001 |
| I and
II | 73 | 71–74 |
| 70 | 68–73 |
|
| III and
IV | 52 | 48–56 |
| 52 | 48–56 |
|
| Histological
grade |
|
| 0.0052 |
|
| 0.3271 |
|
Low | 61 | 59–64 |
| 61 | 58–63 |
|
|
High | 49 | 40–58 |
| 55 | 46–65 |
|
| Venous
invasion |
|
| <0.0001 |
|
| <0.0001 |
|
Absent | 67 | 65–70 |
| 68 | 65–70 |
|
|
Present | 46 | 41–51 |
| 42 | 36–48 |
|
| Lymphatic
invasion |
|
| <0.0001 |
|
| <0.0001 |
|
Absent | 70 | 67–72 |
| 68 | 65–71 |
|
|
Present | 55 | 51–58 |
| 55 | 51–59 |
|
| Perineural
invasion |
|
| <0.0001 |
|
| <0.0001 |
|
Absent | 66 | 63–68 |
| 66 | 64–69 |
|
|
Present | 48 | 42–53 |
| 43 | 37–49 |
|
| DMBT1
expression |
|
| 0.0050 |
|
| 0.0010 |
| No
loss | 61 | 58–64 |
| 61 | 58–64 |
|
|
Loss | 40 | 26–53 |
| 42 | 27–52 |
|
 | Table VI.Multivariate survival analysis in CRC
patients (n=385). |
Table VI.
Multivariate survival analysis in CRC
patients (n=385).
|
| Overall
survival | Disease-free
survival |
|---|
|
|
|
|
|---|
| Variables | HR | 95% CI | P-value | HR | 95% CI | P-value |
|---|
| Stagea |
|
|
|
|
|
|
| I, II
vs. III, IV | 5.604 | 2.480–12.664 | <0.0001 | 2.119 | 1.075–4.176 |
0.030 |
| Venous
invasion |
|
|
|
|
|
|
| Absent
vs. present | 2.220 | 1.426–3.455 | <0.0001 | 2.900 | 1.745–4.821 | <0.001 |
| Lymphatic
invasion |
|
|
|
|
|
|
| Absent
vs. present | 1.638 | 0.934–2.872 | 0.085 | 1.548 | 0.864–2.773 |
0.142 |
| Perineural
invasion |
|
|
|
|
|
|
| Absent
vs. present | 1.557 | 1.021–2.374 | 0.040 | 2.347 | 1.457–3.781 | <0.001 |
| DMBT1
expression |
|
|
|
|
|
|
| No loss
vs. loss | 2.272 | 1.175–4.391 | 0.015 | 2.689 | 1.275–5.670 |
0.009 |
Discussion
DMBT1, a multifunctional glycoprotein containing
multiple scavenger receptor cysteine-rich domains (9), is involved in the mucosal immune
defense, epithelial differentiation (9–11) and
tumor suppression (6). While loss
of DMBT1 mRNA expression has been detected in various cancer types
(16,17), conflicting results have been
reported by previous studies (20,21).
In an earlier study on colon cancer, a decrease in DMBT1 mRNA was
reported in 16.7% of cancer compared with that in normal tissue
samples (17). However, the
clinical validity of DMBT1 expression has not been evaluated in a
large population of CRC patients.
In the present study, fecal DMBT1 expression was
increased in patients with CRC, compared with that in the adenoma
and control groups, but the levels were not significantly different
from those of patients with IBD. In addition, DMBT1 expression was
reported to be upregulated in neutrophils, monocytes and epithelial
cells in IBD (30,31). Upregulation of fecal DMBT1 in CRC
patients may be induced via interactions with cancer cells and
microenvironments as well as immune reactions between epithelial
and inflammatory cells. However, low levels of fecal DMBT1 in
adenoma and control groups may be partly due to a less intense
inflammatory reaction in adenoma or normal colorectal mucosal
tissues, compared with that in CRC or IBD tissues. The results of
the present study demonstrate that fecal DMBT1 is not effective for
the differential diagnosis of CRC and IBD. However, a relatively
high specificity (86.8%) was observed for colorectal disease
(CRC+AA+IBD) despite the low sensitivity (48.9–50.0%).
Analysis of the clinical significance of DMBT1
expression in CRC tissues revealed that overexpression was more
frequent than loss (29.1 vs. 4.7%). Furthermore, expression of
DMBT1 was significantly associated with absence of lymph node
metastasis (N0) or distant metastasis (M0), lower stage and low
histologic grade in CRC patients. A previous study by Conde et
al (21) reported that DMBT1
protein expression is frequently decreased in well-differentiated
adenocarcinoma of the stomach. These conflicting results on the
expression patterns in gastric cancer vs. CRC tissues may be
attributed to differences in the applied antibody, criteria for
determining DMBT1 expression and organ specificity. In contrast to
the present result that loss of DMBT1 expression is an independent
prognostic factor associated with poor overall and disease-free
survival of CRC patients, an earlier study revealed no
clinicopathological significance of DMBT1 mRNA expression in
esophageal cancer (17). The
complex DMBT1 expression patterns in cancer are potentially
associated with functional dualism, i.e., epithelial protection and
differentiation (32). In early
carcinogenesis, DMBT1 upregulation may provide an advantage for
clonal expansion. At the late tumor stage, DMBT1 inactivation may
be beneficial, since differentiation counteracts tumor progression.
The present results suggest that loss of DMBT1 protein expression
represents an effective biomarker for tumor progression in CRC.
Consistent with the present observations, Goeppert et al
(33) reported that increased DMBT1
protein expression was associated with prolonged overall survival
in biliary tract cancer.
The present study has a number of limitations.
First, analysis of fecal DMBT1 was inefficient as a screening test
for CRC due to its low sensitivity. However, this drawback may be
overcome by combined application with another complementary
biomarker in the future. Furthermore, tissue and stool samples of
enrolled CRC patients were different. Fecal DMBT1 represents
protein secreted by cancer cells and the microenvironment.
Consequently, the fecal DMBT1 measured did not originate from
cancer cells alone but included a combination of proteins excreted
from cancer cells as well as inflammatory cells. However, the
tissue expression of DMBT1 was also measured in CRC tissues, which
did not include any stromal or inflammatory cell components. DMBT1
expression was additionally identified in inflammatory cells
infiltrating stroma (Fig. 2).
Despite the study limitations, the present results clearly
demonstrate that DMTB1 protein has a role in the progression of CRC
and loss of expression in tissue is a promising prognostic marker
for CRC patients.
In conclusion, the present study indicated that
DMBT1 is upregulated in stools of CRC patients, while exhibiting
variable expression in CRC tissues. Loss of DMBT1 protein in CRC
tissues is significantly associated with adverse
clinicopathological features (advanced stage, lymph node or distant
metastasis and high histological grade) and may be effectively
utilized as an independent poor prognostic factor for CRC.
Acknowledgements
Not applicable.
References
|
1
|
Ferlay J, Soerjomataram I, Dikshit R, Eser
S, Mathers C, Rebelo M, Parkin DM, Forman D and Bray F: Cancer
incidence and mortality worldwide: Sources, methods and major
patterns in GLOBOCAN 2012. Int J Cancer. 136:E359–E386. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Jung KW, Won YJ, Oh CM, Kong HJ, Lee DH
and Lee KH: Community of Population-Based Regional Cancer
Registries: Cancer Statistics in Korea: Incidence, mortality,
survival, and prevalence in 2014. Cancer Res Treat. 49:292–305.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Rhodes JM: Colorectal cancer screening in
the UK: Joint position statement by the British Society of
Gastroenterology, The Royal College of Physicians, and The
Association of Coloproctology of Great Britain and Ireland. Gut.
46:746–748. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kim BC, Joo J, Chang HJ, Yeo HY, Yoo BC,
Park B, Park JW, Sohn DK, Hong CW and Han KS: A predictive model
combining fecal calgranulin B and fecal occult blood tests can
improve the diagnosis of colorectal cancer. PLoS One.
9:e1061822014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yoo BC, Shin YK, Lim SB, Hong SH, Jeong SY
and Park JG: Evaluation of calgranulin B in stools from the
patients with colorectal cancer. Dis Colon Rectum. 51:1703–1709.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mollenhauer J, Wiemann S, Scheurlen W,
Korn B, Hayashi Y, Wilgenbus KK, von Deimling A and Poustka A:
DMBT1, a new member of the SRCR superfamily, on chromosome
10q25.3–26.1 is deleted in malignant brain tumours. Nat Genet.
17:32–39. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ligtenberg TJ, Bikker FJ, Groenink J,
Tornoe I, Leth-Larsen R, Veerman EC, Amerongen Nieuw AV and
Holmskov U: Human salivary agglutinin binds to lung surfactant
protein-D and is identical with scavenger receptor protein gp-340.
Biochem J. 359:243–248. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ligtenberg AJ, Karlsson NG and Veerman EC:
Deleted in malignant brain tumors-1 protein (DMBT1): A pattern
recognition receptor with multiple binding sites. Int J Mol Sci.
11:5212–5233. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Madsen J, Mollenhauer J and Holmskov U:
Review: Gp-340/DMBT1 in mucosal innate immunity. Innate Immun.
16:160–167. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Vijayakumar S, Takito J, Hikita C and
Al-Awqati Q: Hensin remodels the apical cytoskeleton and induces
columnarization of intercalated epithelial cells: Processes that
resemble terminal differentiation. J Cell Biol. 144:1057–1067.
1999. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Cheng H, Bjerknes M and Chen H:
CRP-ductin: A gene expressed in intestinal crypts and in pancreatic
and hepatic ducts. Anat Rec. 244:327–343. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Holmskov U, Mollenhauer J, Madsen J,
Vitved L, Gronlund J, Tornoe I, Kliem A, Reid KB, Poustka A and
Skjodt K: Cloning of gp-340, a putative opsonin receptor for lung
surfactant protein D. Proc Natl Acad Sci USA. 96:10794–10799. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Madsen J, Sorensen GL, Nielsen O, Tornøe
I, Thim L, Fenger C, Mollenhauer J and Holmskov U: A variant form
of the human deleted in malignant brain tumor 1 (DMBT1) gene shows
increased expression in inflammatory bowel diseases and interacts
with dimeric trefoil factor 3 (TFF3). PLoS One. 8:e644412013.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Garay J, Piazuelo MB, Lopez-Carrillo L,
Leal YA, Majumdar S, Li L, Cruz-Rodriguez N, Serrano-Gomez SJ,
Busso CS, Schneider BG, et al: Increased expression of deleted in
malignant brain tumors (DMBT1) gene in precancerous gastric
lesions: Findings from human and animal studies. Oncotarget.
8:47076–47089. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Mollenhauer J, Holmskov U, Wiemann S,
Krebs I, Herbertz S, Madsen J, Kioschis P, Coy JF and Poustka A:
The genomic structure of the DMBT1 gene: Evidence for a region with
susceptibility to genomic instability. Oncogene. 18:6233–6240.
1999. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wu W, Kemp BL, Proctor ML, Gazdar AF,
Minna JD, Hong WK and Mao L: Expression of DMBT1, a candidate tumor
suppressor gene, is frequently lost in lung cancer. Cancer Res.
59:1846–1851. 1999.PubMed/NCBI
|
|
17
|
Mori M, Shiraishi T, Tanaka S, Yamagata M,
Mafune K, Tanaka Y, Ueo H, Barnard GF and Sugimachi K: Lack of
DMBT1 expression in oesophageal, gastric and colon cancers. Br J
Cancer. 79:211–213. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Braidotti P, Nuciforo PG, Mollenhauer J,
Poustka A, Pellegrini C, Moro A, Bulfamante G, Coggi G, Bosari S
and Pietra GG: DMBT1 expression is down-regulated in breast cancer.
BMC Cancer. 4:462004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Mueller W, Mollenhauer J, Stockhammer F,
Poustka A and von Deimling A: Rare mutations of the DMBT1 gene in
human astrocytic gliomas. Oncogene. 21:5956–5959. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Mollenhauer J, Herbertz S, Holmskov U,
Tolnay M, Krebs I, Merlo A, Schrøder HD, Maier D, Breitling F,
Wiemann S, et al: DMBT1 encodes a protein involved in the immune
defense and in epithelial differentiation and is highly unstable in
cancer. Cancer Res. 60:1704–1710. 2000.PubMed/NCBI
|
|
21
|
Conde AR, Martins AP, Brito M, Manuel A,
Ramos S, Malta-Vacas J, Renner M, Poustka A, Mollenhauer J and
Monteiro C: DMBT1 is frequently downregulated in
well-differentiated gastric carcinoma but more frequently
upregulated across various gastric cancer types. Int J Oncol.
30:1441–1446. 2007.PubMed/NCBI
|
|
22
|
Mollenhauer J, Helmke B, Müller H,
Kollender G, Lyer S, Diedrichs L, Holmskov U, Ligtenberg T,
Herbertz S, Krebs I, et al: Sequential changes of the DMBT1
expression and location in normal lung tissue and lung carcinomas.
Genes Chromosomes Cancer. 35:164–169. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
World Health Organization, . WHO
Classification of Tumours of the Digestive SystemBosman FT,
Carneiro F, Hruban RH and Theise ND: 3. 4th. IARC; Lyon: 2010
|
|
24
|
American Joint Committee on Cancer: AJCC
Cancer Staging Atlas. Greene FL, Compton CC, Fritz AG, Shah JP and
Winchester DP: Springer; New York: 2006, doi:
org/10.1007/0-387-33126-3.
|
|
25
|
Lieberman DA, Weiss DG, Bond JH, Ahnen DJ,
Garewal H and Chejfec G: Use of colonoscopy to screen asymptomatic
adults for colorectal cancer. Veterans Affairs Cooperative Study
Group 380. N Engl J Med. 343:162–168. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yao T, Matsui T and Hiwatashi N: Crohn's
disease in Japan: Diagnostic criteria and epidemiology. Dis Colon
Rectum. 43 Suppl:S85–S93. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Choi CH, Jung SA, Lee BI, Lee KM, Kim JS
and Han DS: IBD Study Group of the Korean Association of the Study
of Intestinal Diseases: Diagnostic guideline of ulcerative colitis.
Korean J Gastroenterol. 53:145–160. 2009.(In Korean). PubMed/NCBI
|
|
28
|
Ye BD, Jang BI, Jeen YT, Lee KM, Kim JS
and Yang SK: IBD Study Group of the Korean Association of the Study
of Intestinal Diseases: Diagnostic guideline of Crohn's disease.
Korean J Gastroenterol. 53:161–176. 2009.(In Korean). PubMed/NCBI
|
|
29
|
Telakis E and Tsironi E: Indeterminate
colitis-definition, diagnosis, characteristics and management. Ann
Gastroenterol. 21:173–180. 2009.
|
|
30
|
Foell D, Wittkowski H and Roth J:
Monitoring disease activity by stool analyses: From occult blood to
molecular markers of intestinal inflammation and damage. Gut.
58:859–868. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Rosenstiel P, Sina C, End C, Renner M,
Lyer S, Till A, Hellmig S, Nikolaus S, Fölsch UR, Helmke B, et al:
Regulation of DMBT1 via NOD2 and TLR4 in intestinal epithelial
cells modulates bacterial recognition and invasion. J Immunol.
178:8203–8211. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Mollenhauer J, Helmke B, Müller H,
Kollender G, Krebs I, Wiemann S, Holmskov U, Madsen J, Otto HF and
Poustka A: An integrative model on the role of DMBT1 in epithelial
cancer. Cancer Detect Prev. 26:266–274. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Goeppert B, Roessler S, Becker N, Zucknick
M, Vogel MN, Warth A, Pathil-Warth A, Mehrabi A, Schirmacher P,
Mollenhauer J, et al: DMBT1 expression in biliary carcinogenesis
with correlation of clinicopathological data. Histopathology.
70:1064–1071. 2017. View Article : Google Scholar : PubMed/NCBI
|