Introduction
Prostate cancer (PCa) is one of the most commonly
diagnosed malignancies and the third most common lethal cancer in
men (1). Over 160,000 new cases of
prostate cancer were estimated to have been diagnosed in 2017 in
the US, which accounts for 19% of all cancer cases (2). Consequently, it is vitally important
and necessary to study the associated detailed mechanisms of this
disease.
TROAP, also named tastin and first cloned in 1995,
is a cytoplasmic protein that is necessary for the function of
trophinin as a cell adhesion molecule (3,4). The
associated adhesion molecule complex consists of bystin, trophinin
and TROAP, and bystin has been shown to be present in human PCa
(5). In addition to its function as
a component of the adhesion complex during embryo implantation,
TROAP has also been reported to be required for spindle assembly
during mitosis (6,7). It is also essential for centrosome
integrity and proper bipolar organisation of spindle assembly
during mitosis, playing an essential role in cell proliferation.
Levels of this protein peak during the G2/M phase and abruptly
decline after division. However, its biological function in
prostate cancer and cancer in general is still unclear. We found
that TROAP expression correlates with patient survival and
deduced that TROAP plays an important role in prostate cancer
progression.
In this study, we first examined the TROAP
expression profile using several online datasets of prostate
cancer; in addition, patient survival analysis with respect to the
target gene was performed using PROGgeneV2 Database. Next, the
regulatory effect of TROAP on cellular survival and apoptosis was
demonstrated by performing cell proliferation and flow cytometric
assays. Western blot analysis was used to determine the expression
of apoptosis and cell cycle-related biomarkers. Correlations
between WNT3 or survivin expression and TROAP transcripts in
prostate cancer tissues were also analysed. Finally, the regulation
of TROAP in PCa cell progression was established in vivo.
Collectively, our data suggest that TROAP could be an important
target for PCa diagnosis and treatment.
Materials and methods
Database analysis
To examine TROAP mRNA expression using microarray
data, the Oncomine database tool (www.oncomine.org) was used. Briefly, the TROAP
gene was queried in the database and the results were filtered by
selecting prostate cancer studies that reported TROAP
expression data (Reporter ID: 204649_at for Varambally Prostate,
Arredouani Prostate, Wallace Prostate and A_23_P150935 for Grasso
Prostate). Both probes were targeted to nucleotide sequence of
TROAP. P-values for each group were calculated using The Student's
t-test. Standardised normalisation techniques and statistical
calculations are provided by the Oncomine website. Additionally,
ProggeneV2 prognostic Database (http://watson.compbio.iupui.edu/chirayu) was used to
collect information about patient survival data with regards to
TROAP expression in prostate cancer (8).
Cell lines and culture conditions
Human prostate cell lines, LNCaP, WPMY-1, DU145,
22Rv1, PC-3 and C4-2 (all have been confirmed that they are not
misidentified or contaminated by checking established cell lines
with the list of known misidentified cell lines available from the
International Cell Line Authentication Committee http://iclac.org/databases/cross-contaminations) were
purchased from the Cell Bank of the Chinese Academy of Science
(Shanghai, China). LNCaP and WPMY-1 cells were cultured in DMEM
(Hyclone Laboratories; GE Healthcare Life Sciences, Logan, UT, USA;
SH30022.01B) with 10% fetal bovine serum (FBS). C4-2 and 22RV1
cells were cultured in RPMI-1640 medium (Hyclone Laboratories; GE
Healthcare Life Sciences, SH30809.01B) containing 10% FBS (Biowest,
Riverside, MO, USA; S1810). DU145 and PC-3 cells were cultured in
Ham's/F-12 (Gibco; Thermo Fisher Scientific, Inc., Waltham, MA,
USA; 11765054) with 10% FBS and 1% non-essential amino acids (NEAA,
Hyclone Laboratories; GE Healthcare Life Sciences; SH30238.01). The
culture medium was supplemented with 100 U/ml penicillin and 100
mg/ml streptomycin to avoid bacterial contamination. All cells were
cultured in humidified air at 37°C with 5% CO2.
Total RNA isolation and quantitative
real-time (qRT)-PCR analysis
Total RNA was prepared from several human prostate
cancer cell lines (WPMY-1, LNCap, 22Rv1, DU145, PC3 and C4-2) using
TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.)
following the manufacturer's protocol. Then, 1 µg of total RNA was
reverse transcribed in a final volume of 20 µl using random primers
and M-MLV Reverse Transcriptase (Promega, Madison, WI, USA; M170A)
and standard conditions. The reverse transcription reaction was
performed using the following conditions: 37°C for 30 min; 85°C for
5 sec and a hold at 4°C.
After reverse transcription, the qRT-PCR was
performed using the SYBR Green Master Mix Kit (Takara
Biotechnology, Co., Ltd., Dalian, China) and a BioRad CFX96
sequence detection system, according to the manufacturer's
instructions. The relative levels of TROAP were determined by
qRT-PCR using gene specific primers, and expression was normalised
to the internal control β-actin. The primer sequences used are as
follows: forward, 5′-GTGGACATCCGCAAAGAC-3′ and reverse,
5′-AAAGGGTGTAACGCAACTA-3′ for β-actin, and forward,
5′-GGTCAGGAGAAAAGCGGAGGAAG-3′ (sense) and reverse,
5′-AGGCGTGCGTTTCTGAGAGC-3′ for TROAP. The qRT-PCR was conducted
under the following conditions: 95°C for 60 sec, 40 cycles of 95°C
for 5 sec and 60°C for 20 sec. Each sample was run at least three
times. The relative expression level of TROAP was computed using
the comparative cycle threshold (CT) method 2−∆∆Cq
(9).
Lentivirus infection of prostate
cancer cells
The shRNA sequence targeting TROAP was synthesised
and ligated into pGreenPuro™ shRNA Cloning and Expression
Lentivector (#s SI505A-1; SBI System Biosciences, LLC, Palo Alto,
CA, USA), which contains green fluorescent protein (GFP) tag. The
shRNA sequence targeting human TROAP was:
5′-AGAACCAAGATCCAAGGAGATCTCGAGATCTCCTTGGATCTTGGTTCT-3′. The
non-targeting shCon sequence (control shRNA) used was
5′-TTCTCCGAACGTGTCACGTCTCGAGACGTGACACGTTCGGAGAA-3′. Lentiviral
particles were then packaged in the 93T cells and harvested using
ultra-centrifugation. DU145 and 22Rv1 cell lines cultured in 6-well
plates were infected with lentivirus containing targeting shRNA or
control shRNA. Cells were harvested after 3 days for qRT-PCR and
other experiments.
Cell proliferation assays
The effect of TROAP knockdown on cell proliferation
was determined using the
3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide
(MTT) method according to the manufacturer's (Sigma-Aldrich; Merck
KGaA) instructions. Briefly, DU145 and 22Rv1 cells infected with
shTROAP and shCon, separately, were seeded in 96-well plates
(3,000/well) and incubated for 5 days. At day 1 to day 5 after
seeding, MTT plus acidic isopropanol solution was added to every
well and samples were incubated at 37°C for 1 h. The absorbance
values were measured at 595 nm using a Biotek Epoch Microplate
Spectrophotometer (BioTek Instruments, Inc., Winooski, VT, USA).
Each group comprised five replicates.
Colony formation assays
For colony formation assays, DU145 and 22Rv1 cells
were seeded in 6-well plates at 500 cells/well and cultured in
medium containing 10% FBS, changing the medium every 3 days. After
14 days, the colonies were washed with PBS, fixed with 4%
paraformaldehyde, and stained with 0.1% freshly-prepared crystal
violet solution (Beyotime Unstitute of Biotechnology, Haimen,
China). Visible colonies containing 50 or more cells were observed
and manually counted using an inverted light microscope (Olympus
CKX41; Olympus Corp., Tokyo, Japan). Triplicate wells were measured
for each treatment group.
Flow cytometric analysis
For flow cytometric analysis, infected cells were
collected 6 days after inoculation by trypsinisation. Cells for
cell cycle analysis were stained using propidium iodide (PI) with
the Cycle TEST PLUS DNA Reagent kit (BD Biosciences, Franklin
Lakes, NJ, USA; NC9941088) according to the manufacturer's
protocol, and then subjected to FACScan. The percentage of the
cells in the G0/G1, S, and G2/M phases were computed and compared.
For apoptosis analysis, double staining was performed using the
FITC Annexin V Apoptosis Detection kit (Nanjing KeyGen Biotech,
Co., Ltd., Nanjing, China), according to the manufacturer's
instructions, and then the stained cells were analysed using
FACScan (BD Biosciences). The percentage of early apoptotic and
apoptotic cells were compared to that in the control groups for
each experiment.
Western blot assay
Cells were lysed using RIPA buffer containing 100 mM
Tris-HCl (pH 6.8), 10 mM EDTA, 4% SDS, and 10% glycine supplemented
with PMSF (Roche, Basel, Switzerland). Total protein was quantified
using the Coomassie brilliant blue method at 560 nm. The
supernatant (10 µg of protein) was denatured and separated on 12%
SDS polyacrylamide gel electrophoresis (SDS-PAGE), and then
transferred to 0.22-μm polyvinylidene difluoride (PVDF) membranes
(EMD Millipore, Bedford, MA, USA) and incubated with specific
antibodies as follows: anti-GAPDH (dilution 1:500,000; cat. no.
10494-1-AP), anti-TROAP (dilution 1:1,000; cat. no. 13634-1-AP),
anti-PSA (dilution 1:1,000; cat. no. 10679-1-AP), anti-cleaved
caspase 3 (dilution 1:1,000; cat. no. 25546-1-AP), anti-caspase 9
(dilution 1:1,000; cat. no. 10380-1-AP), anti-cyclin A2 (dilution
1:1,000; cat. no. 18202-1-AP), anti-WNT3 (dilution 1:500; cat. no.
17983-1-AP), anti-survivin (dilution 1:1,000; cat. no. 10508-1-AP)
and anti-CNNB1 (dilution 1:1,000; cat. no. 51067-2-AP) from
Proteintech Group Inc., (Rosemont, IL, USA); anti-Bcl-2 (dilution
1:1,000; cat. no. 2876) and PARP (dilution 1:1,000; cat. no. 9542)
from Cell Signaling Technology, Inc. (Danvers, MA, USA);
anti-cyclin B1 (dilution 1:2,000; cat. no. 21540) from SAB
(Nanjing, China). Bands were visualised using enhanced
chemiluminescence (ECL-PLUS/Kit, Amersham Pharmacia Biotech, Tokyo,
Japan) reagents.
Pairwise correlation analysis
WNT3 and survivin were previously reported to be
relevant for carcinogenesis. To determine whether there is a
correlation between the expression of WNT3 or survivin and
TROAP transcripts in PCa, pairwise gene correlation analysis
was performed using the web server GEPIA (http://gepia.cancer-pku.cn/) (10). P<0.05 was defined as
statistically significant. Correlation analysis results were then
further validated by western blot analysis.
Xenograft tumourigenicity and gene
expression assay in vivo
The prior approval of the Ethics Committee of the
Third Affiliated Hospital, The Second Military Medical University
was obtained for use of the animals in this study.
Twelve male athymic nude mice (nu/nu), 4 weeks old
(weight, 15–20 g), were housed under pathogen-free conditions in a
barrier animal facility with a 12-h light/12-h dark cycle and
humidity (45–55%) at 21–24°C, and food and water were available
ad libitum. For in vivo tumourigenicity experiments,
DU145 cells stably infected with lentivirus or negative control
were collected, resuspended in PBS, and injected subcutaneously
into the right subaxillary of each male 4-week-old BALB/c nude
mouse (2×104 per mouse; SLRC Laboratory Animal,
Shanghai, China). The tumour volumes and weights were measured
every 3 days and maximum allowable tumour size was controlled
according to the IACUC guidelines (diameter, 2.0 cm; volume 4.2
cm3); tumour volume (V) was measured as V = length ×
width2 × 0.5. Forty-five days after injection, mice were
sacrificed by cervical dislocation and tumours were dissected,
imaged, and weighed; the tumours were also collected and fixed for
further western blot analysis. The relative amount of protein was
calculated based on the integrated optical density reference value
using ImageJ optical density gel analysis software (National
Institutes of Health, Bethesda, MD, USA). GAPDH was used as the
control.
Statistical analysis
Statistical analysis was performed using GraphPad
Prism 5.0 software (GraphPad Software, Inc., La Jolla, CA, USA).
Student's t-test was used for two-group comparisons and ANOVA with
Sidak's multiple comparisons were used to test the significance of
multigroup data. Data are presented as mean ± SD. A P-value
<0.05 was considered to be indicative of statistical
significance.
Results
TROAP is overexpressed in PCa and
higher expression is associated with shorter overall survival
To achieve a profound understanding of the role of
TROAP in PCa, a publicly available array data source, the Oncomine
database tool (www.oncomine.org), was first used to investigate the
expression levels of TROAP. The Oncomine Cancer Microarray
database contains several datasets comparing human prostate
carcinoma to normal tissue. TROAP expression was commonly highly
expressed in several PCa groups compared to that in adjacent normal
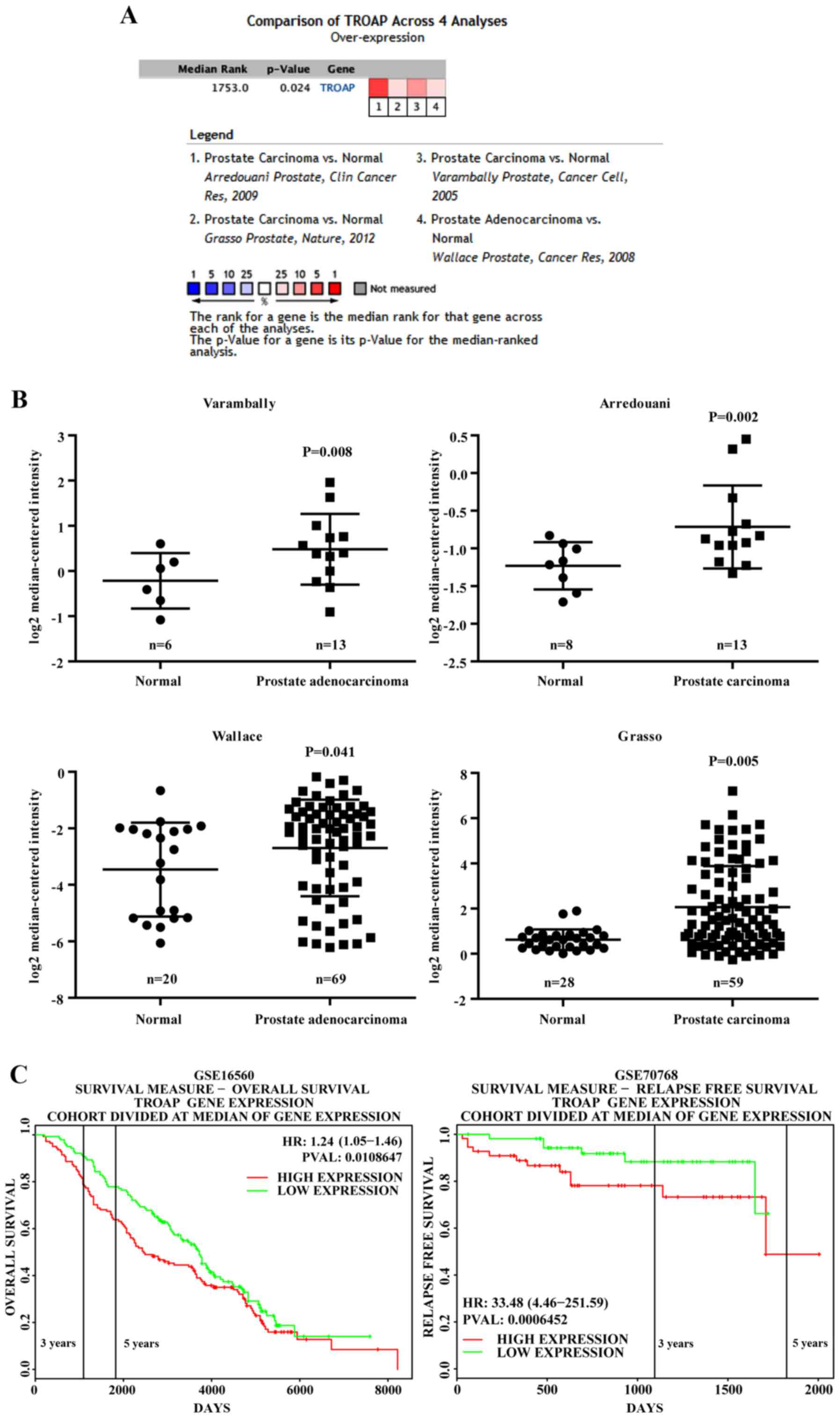
tissue. As shown in Fig. 1A and B,
four datasets [Arredouani et al (11), Grasso et al (12), Varambally et al (13) and Wallace et al (14)] showed that TROAP is significantly
overexpressed in PCa as compared to levels in adjacent normal
tissue, with fold changes ranging from 1.27 to 2.08 (11–14).
Additionally, by exploring the PROGgeneV2 Prognostic Database, we
found that PCa patients (GSE16560) (15) with TROAP levels that were higher
than the median expression level had significant shorter overall
survival than those with lower TROAP levels (Fig. 1C). Notably, another database
(GSE70768) (16) showed that higher
TROAP expression is associated with poor relapse-free survival
(Fig. 1C).
TROAP expression in prostate cells and
knockdown efficiency in PCa cell lines
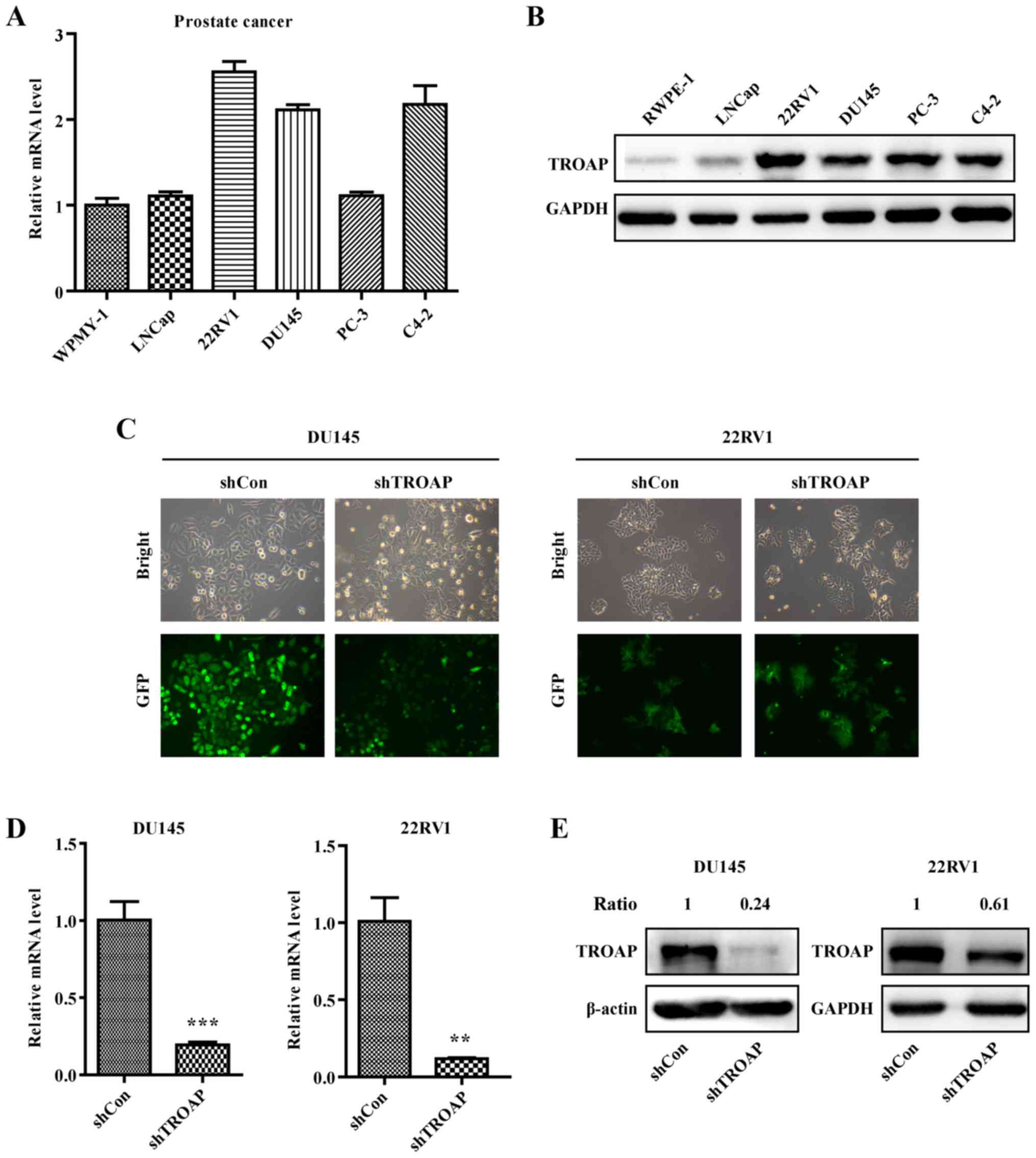
The expression of TROAP in different PCa cell lines
including WPMY-1, LNCaP, 22Rv1, DU145, PC3 and C4-2 was detected by
qRT-PCR and western blotting. The results showed that the
expression of TROAP is relatively high in PCa cells, especially
non-metastatic 22Rv1 and metastatic DU145 cells (Fig. 2A and B), which are
androgen-independent PCa cell lines. Accordingly, these two cell
lines were selected for further study. DU145 and 22Rv1 cells were
separately infected with the shTROAP and shCon lentiviruses. Based
on the observed expression of GFP using a microscope, the infection
efficiencies were adequate (Fig.
2C). Results of qRT-PCR illustrated that the mRNA levels of
TROAP were significantly suppressed in DU145 and 22Rv1 cells
(Fig. 2D). Accordingly, the protein
levels of TROAP were decreased obviously with shRNA targeting in
DU145 and 22Rv1 cells (Fig.
2E).
Inhibition of PCa cell proliferation
by shRNA-mediated TROAP knockdown
To determine the effect of different treatments on
PCa cell proliferation, prostate cancer cell numbers were assessed
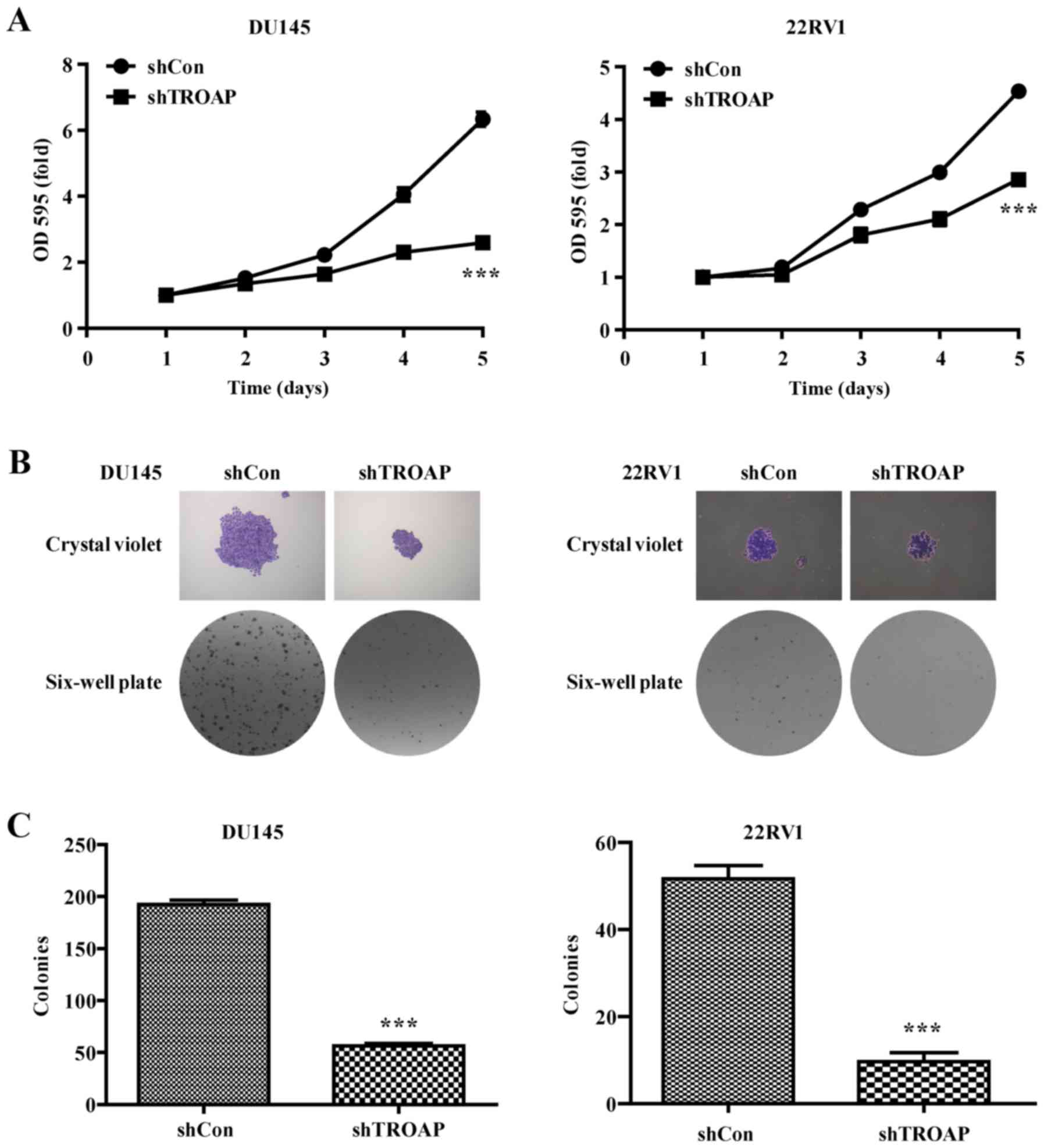
using MTT and colony formation assays. As shown by the cell growth
curve, growth of the shTROAP-treated groups was significantly
slower than that of the shCon groups for DU145 and 22Rv1 cells
(Fig. 3A). Furthermore, the sizes
of independent colonies were much smaller in DU145 and 22RV1 cells
infected with shTROAP, compared to that in cells infected with
shCon (Fig. 3B). Moreover, the
numbers of colonies formed in shTROAP-infected groups were
significantly lower than that in shCon-infected groups (P<0.001,
Fig. 3C). The data demonstrated a
significant reduction in cell expansion with reduced TROAP
expression.
Knockdown of TROAP leads to cell cycle
arrest and cell cycle modulation through cyclin A2/cyclinB1
Having established that TROAP expression is related
to PCa cell proliferation, we were next interested in whether the
cell cycle is influenced by TROAP. After performing cell cycle
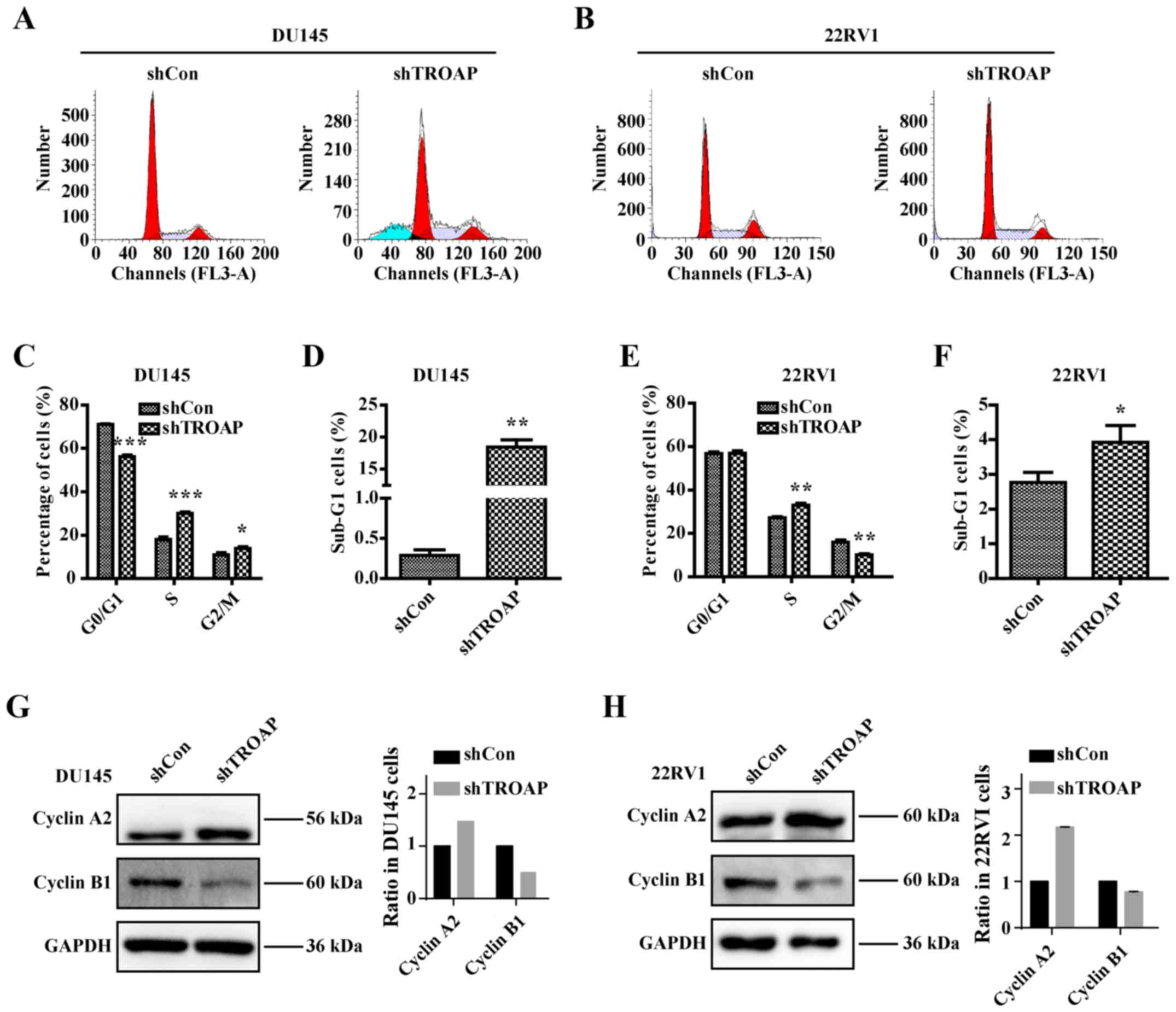
analysis by flow cytometry, we found that inhibition of TROAP
significantly decreased the number of cells in the G0/G1 phase and
increased the number of cells in the S phase in the DU145 and 22Rv1
cells (Fig. 4A and B). The S phase
cell percentage was increased from 18% in the shCon group to 30% in
the shTROAP group in DU145 cells and from 27% in the shCon group to
33% in the shTROAP group in the 22Rv1 cells (Fig. 4C and E). TROAP knockdown further led
to significant increases in the sub-G1-phase cell population in
both DU145 and 22Rv1 cells (Fig. 4D and
F). We then focused on the molecular mechanisms through which
TROAP knockdown delays S phase to mitosis transition by analysing
the expression of checkpoint proteins. Western blotting showed that
downregulation of TROAP increased the expression of cyclin A2 but
decreased the expression of cyclin B1, compared to that in
respective controls, for the DU145 and 22Rv1 cells (Fig. 4G and H). The results thus
corresponded to a cell cycle arrest at S phase.
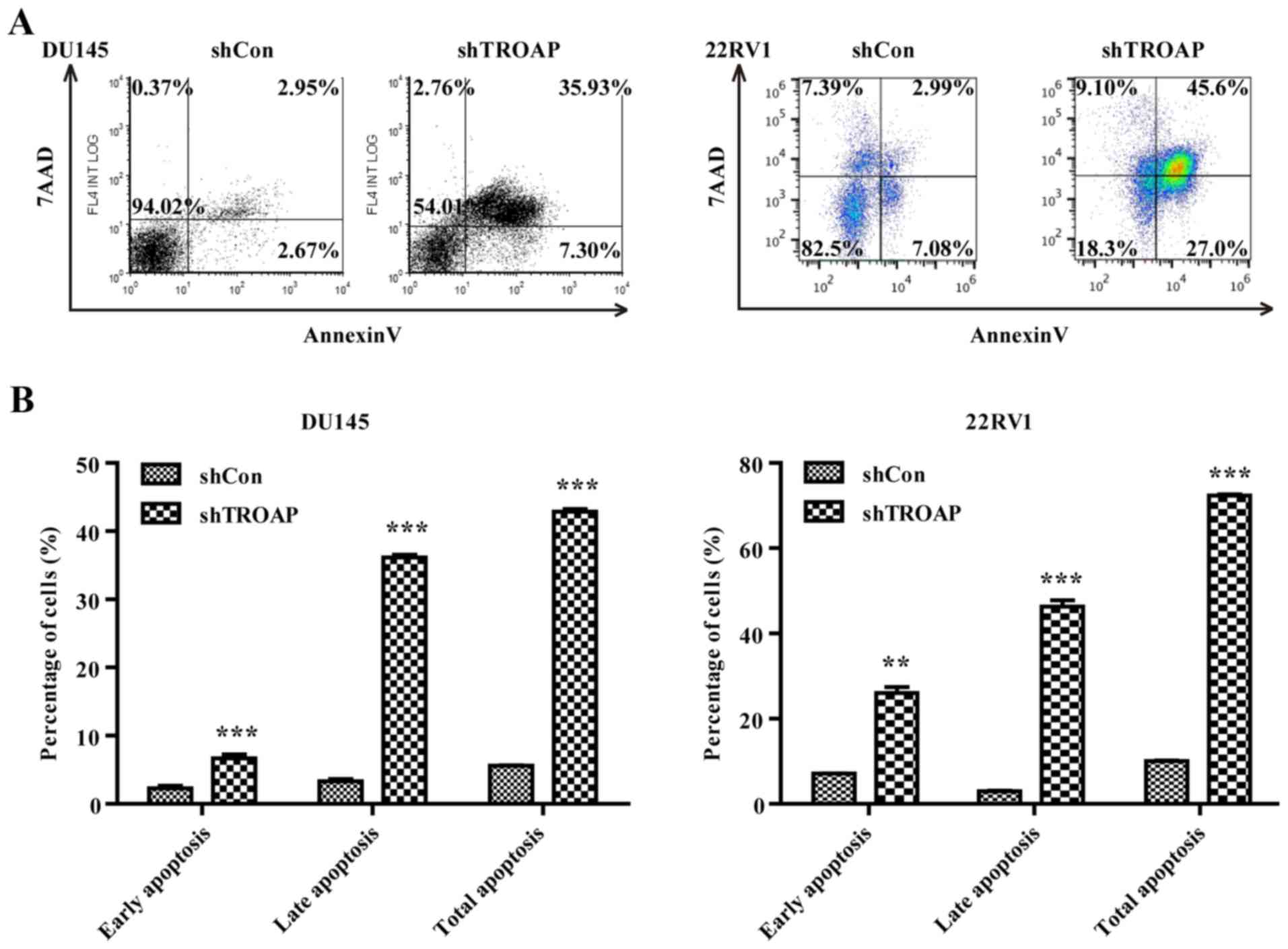
Knockdown of TROAP promotes apoptosis
in PCa cell lines through the mitochondrial apoptosis pathway
Based on the observed increases in the sub-G1-phase
cell population (Fig. 4D and F), we
attempted to determine whether the decrease in TROAP was associated
with cancer cell apoptosis by flow cytometry. TROAP silencing
significantly accelerated both DU145 and 22Rv1 cell apoptosis.
Lentivirus-mediated shRNA increased both early and late apoptosis
in the DU145 and 22Rv1 cells (Fig. 5A
and B). The total percentage of apoptotic cells increased from
6% in the shCon group to 43% in the shTROAP group for DU145 cells,
and from 10% in the shCon group to 72% in the shTROAP group for
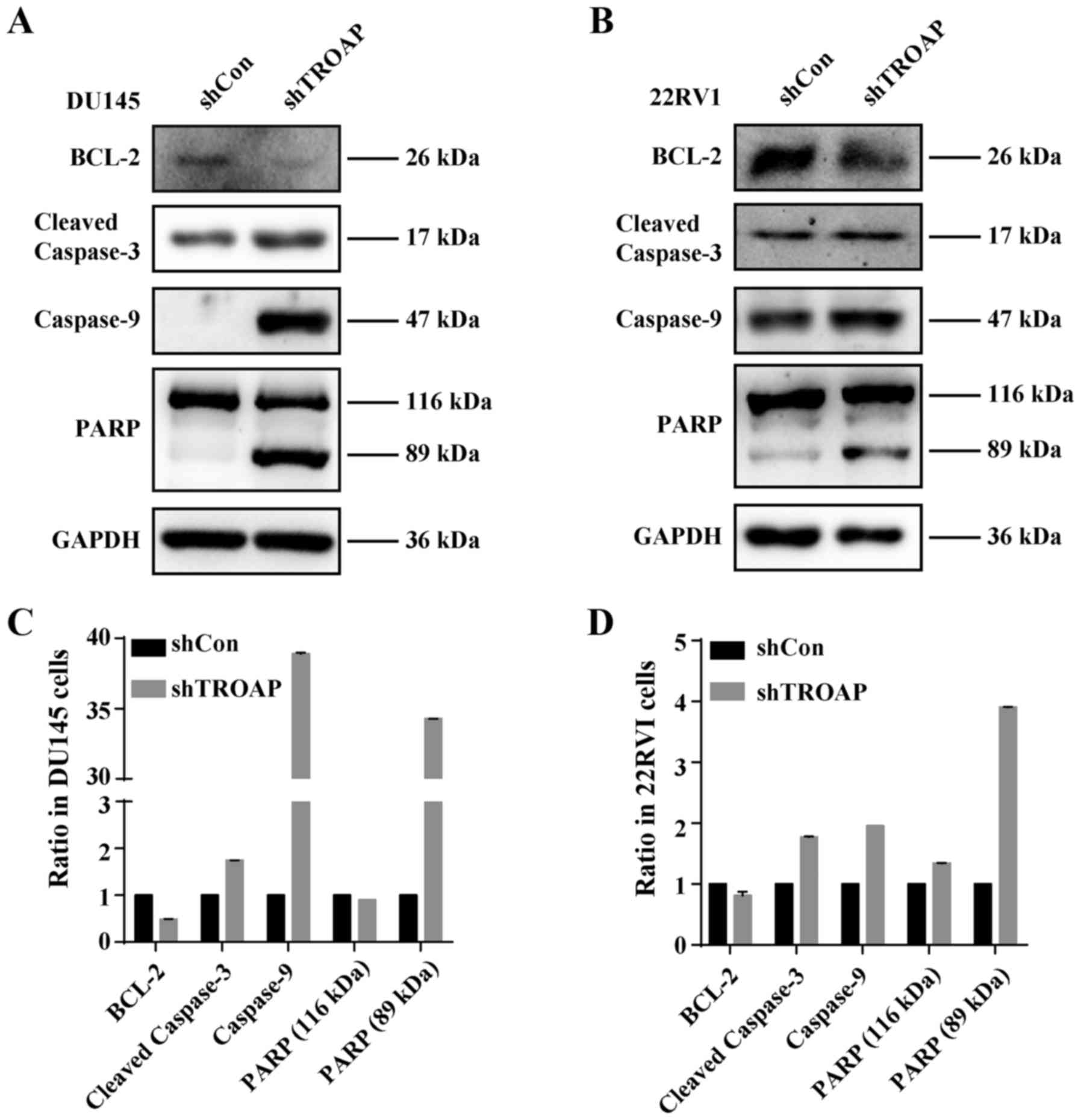
22Rv1 cells. To investigate the underlying mechanisms of TROAP
knockdown-induced apoptosis in these cells, the protein levels of
Bcl-2, caspase-9, cleaved-caspase-3 and PARP in PCa cells were
measured by western blot analysis after treatment with shTROAP.
Western blot analysis showed that depletion of TROAP decreased the
levels of Bcl-2, whereas levels of caspase-9, caspase-3 and PARP
increased (Fig. 6). The results
demonstrated that the mitochondrial apoptosis pathway was activated
by downregulation of TROAP in PCa cells.
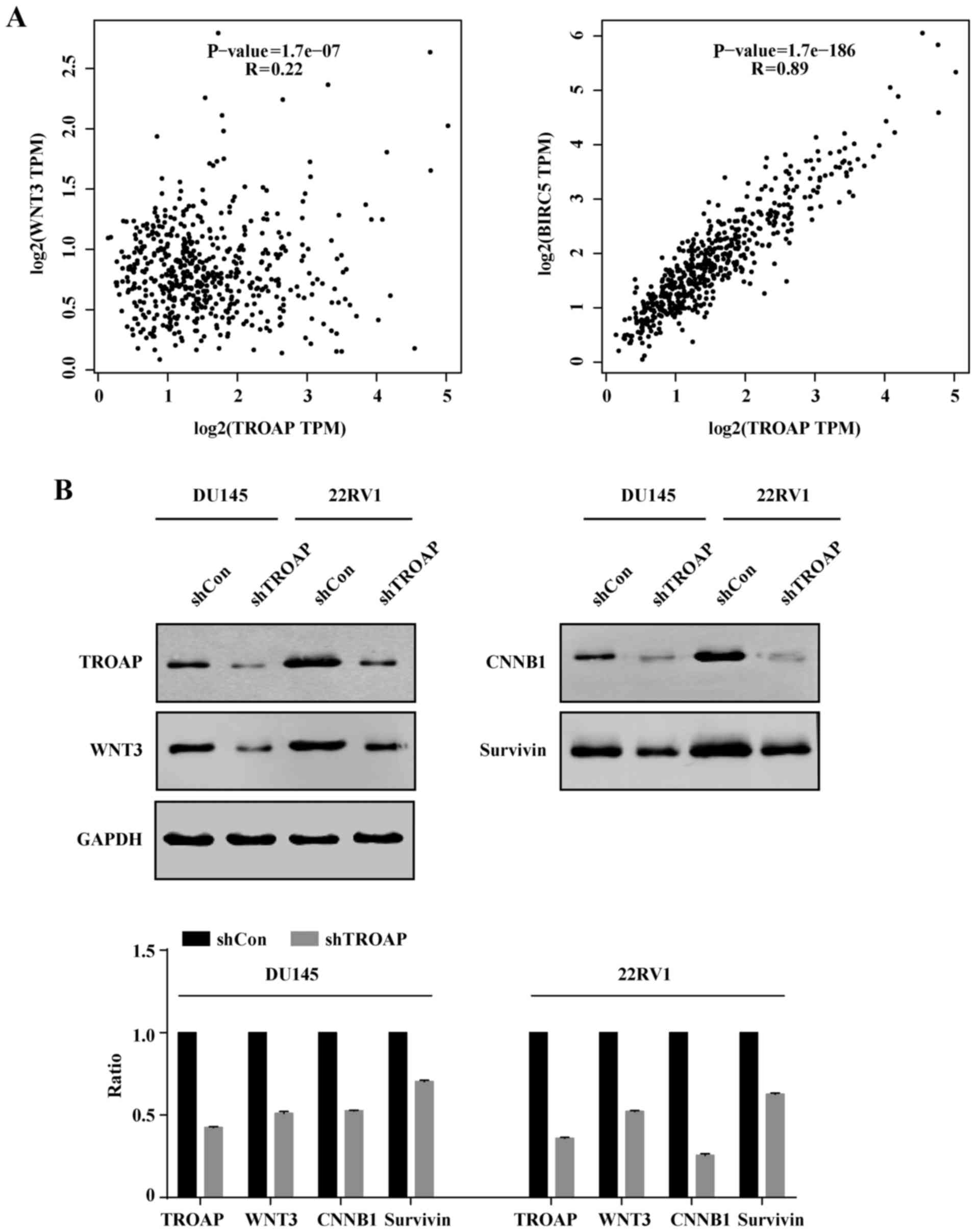
TROAP expression correlates with WNT3
and survivin levels in PCa
Survivin is demonstrated to protect cells from
caspase-induced apoptosis. Meanwhile, it was the downstream target
of wnt/β-catenin. Hence, we deduced that TROAP induced apoptosis by
interrupting the expression of surviving and wnt. Correlation
analysis indicated that WNT3 and survivin (BIRC5) levels were
positively correlated with TROAP expression (P<0.05; Fig. 7A). To further validate this
analysis, we performed western blotting. Consistently, it was
verified that protein levels of WNT3 and survivin decreased with
TROAP knockdown (Fig. 7B).
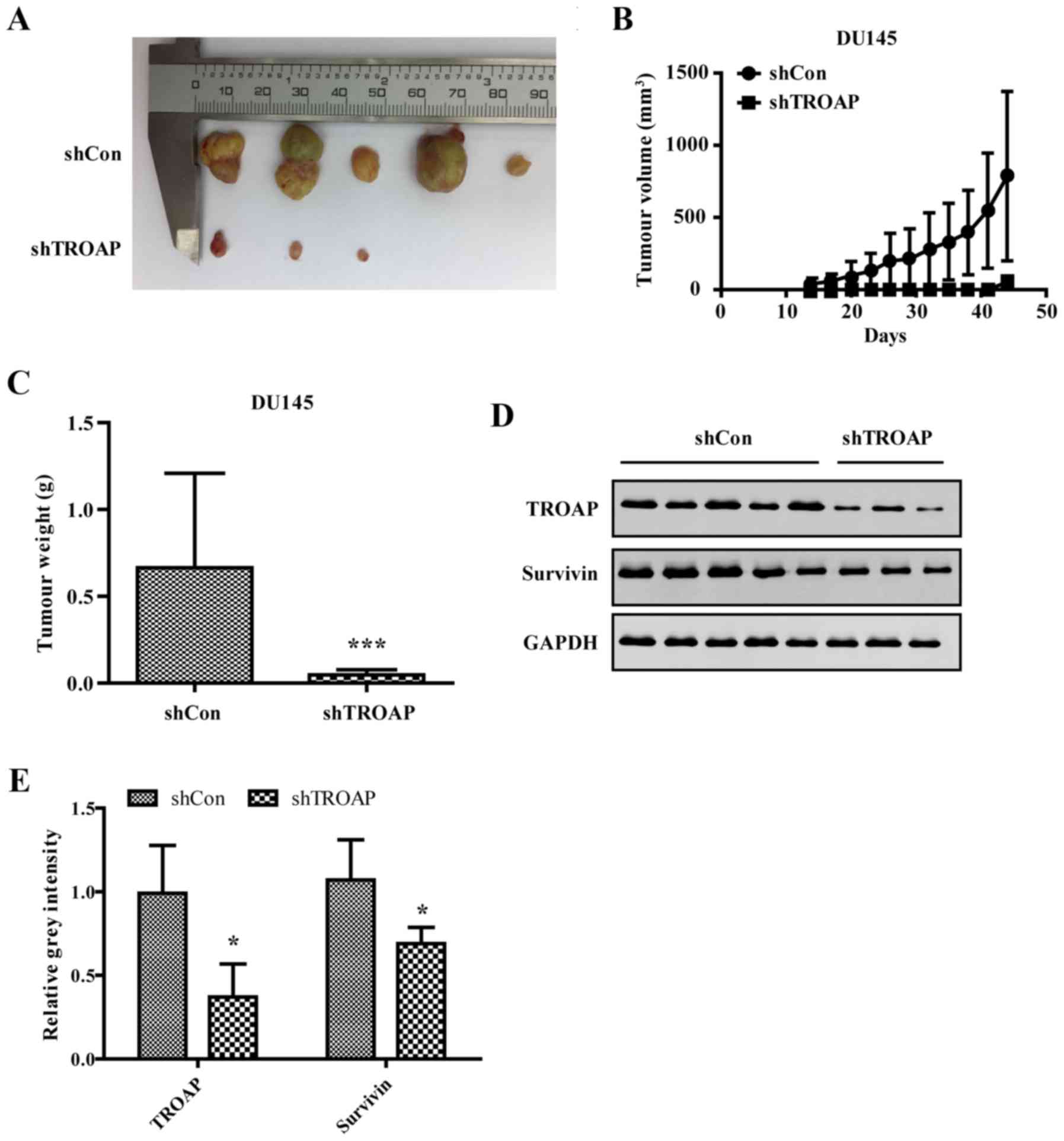
Knockdown of TROAP inhibits PCa growth
in nude mice
To investigate the effect of TROAP on PCa cell
proliferation in vivo, a xenograft model was utilised to
assess the tumorigenesis of DU145 cells after shTROAP or shCon
transduction. As shown, the largest diameter of the tumours was
approximately 10 mm in the shCon group. Tumours in mice injected
with shTROAP-transduced cells were significantly smaller than those
originating from shCon-infected cells (Fig. 8A). Both the volume and the weight of
tumours derived from shTROAP DU145 cells were significantly
decreased compared to those of the shCon group (Fig. 8B and C). Thus, the results confirmed
that TROAP knockdown can inhibit PCa tumour growth in
vivo.
To further illustrate that tumour suppression by
shTROAP is associated with the knockdown of TROAP and associated
downregulation of survivin in PCa cells, mice from the two groups
were sacrificed after the last measurement. Protein expression
levels of both TROAP and survivin were detected by western
blotting. As shown in Fig. 8D and
E, the levels of both were efficiently decreased in tumour
tissue of the shTROAP group compared to those in the shcontrol
group.
Discussion
In the present study, we initially demonstrated that
TROAP is upregulated in PCa tissues. Importantly, clinical data
have suggested that TROAP is associated with poor overall and
relapse-free survival in prostate cancer patients, underscoring the
oncogenic role of this marker, particularly in relapse. We also
demonstrated for the first time that TROAP protein expression is
relatively high in several PCa cell lines by using qRT-PCR and
western blot analysis.
Silencing of TROAP significantly inhibited the
growth and colony formation ability of PCa cells through cell cycle
arrest at the S phase and the induction of apoptosis. More
importantly, downregulation of TROAP dramatically decreased cyclin
B1 and Bcl-2 levels and increased cyclin A2, caspase-9 protein
levels and cleaved PARP.
Knockdown of TROAP caused cell cycle arrest and
apoptosis through the cyclin/Cdk pathway. Cyclin A2 is a core cell
cycle regulator that activates Cdk1 and Cdk2 (17). The expression of cyclin A2 and CDK2,
are known to promote S phase entry in mammals (18). Cyclin B1 is a key regulator of cell
cycle progression from the S phase to the G2/M phase and its
downregulation therefore results in S phase arrest (19). Therefore, upregulation of cyclin A2
and downregulation of cyclin B1, induced by TROAP silencing,
explains why PCa cells were arrested at the S phase. Furthermore,
it was found that in the shTROAP group, the G2/M phase cell ratio
was increased in DU145 cells and decreased G2/M in 22RV1 cells.
This may be owing to the different metastatic abilities in DU145
and 22RV1 cells as metastasis may relate to the cell cycle
(20,21). This needs further study. In
addition, more cell cycle markers are required to determine the
exact point at which the cell cycle is disrupted.
To further confirm that apoptosis induced by TROAP
depletion is associated with the mitochondrial pathway, the levels
of Bcl-2, cleaved caspase-3, caspase-9, and PARP were detected by
western blotting. The results demonstrated that the mitochondrial
pathway was activated by downregulation of TROAP in PCa cells.
Survivin is the smallest member of the human IAPs (inhibitor of
apoptosis proteins) and is the downstream target gene of the Wnt
signalling pathway. WNT3, a member of the Wnt family, was reported
to be associated with hepatic, lung, and colorectal carcinogenesis
(22–25). It can protect cells from
caspase-induced apoptosis, activated in mitosis, presumably by
binding caspase-3, −7, and −9 (26). The results of the correlation
analysis indicated that TROAP expression was correlated with WNT3
and survivin in PCa. Knockdown of TROAP could decrease the Wnt 3
and survivin expression, which confirms both the in vitro
and in vivo experiments. Hence, we deduced that TROAP may
affect proliferation through the WNT signalling pathway.
In conclusion, knockdown of TROAP decreased cell
proliferation and colony formation and induced cell cycle arrest
and apoptosis in vitro. Taken together, we concluded that
TROAP functions as a regulator of PCa to control tumour growth.
In conclusion, collectively, our observations
indicate that TROAP is one of the driving mechanisms of
Wnt/β-catenin signalling. Our findings provide new insights into
the mechanism of the progression of PCa.
Acknowledgements
Not applicable.
Funding
The present study was supported by grants from the
Youth Program of Shanghai Municipal Commission of Health and Family
Planning (no. 20164Y0040); the National Natural Science Foundation
of China (no. 81772747); the National Natural Science Foundation of
China for Youths (no. 81702515, 81702501) and the Shanghai Sailing
Program (no. 17YF1425400).
Availability of data and materials
The data used and/or analysed during the present
study can be obtained from the corresponding author on reasonable
request.
Authors' contributions
JY, CC, MC and ZS performed the vector construction
experiments and drafted the manuscript. SG and FQ participated in
the RT-PCR and western blot experiments. XP, QY and YT participated
in the cellular function experiments. LW and WY participated in the
animal experiments and were responsible for the data analysis and
for the figure formation. XC participated in the research design,
reviewed the literature and performed data examination. All authors
read and approved the manuscript and agree to be accountable for
all aspects of the research in ensuring that the accuracy or
integrity of any part of the work are appropriately investigated
and resolved.
Ethics approval and consent to
participate
Animal experiments in this study were approved by
the Ethics Committee of the Third Affiliated Hospital, The Second
Military Medical University.
Patient consent for publication
Not applicable.
Competing interests
The authors state that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2016. CA Cancer J Clin. 66:7–30. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Hoffman RM, Meisner AL, Arap W, Barry M,
Shah SK, Zeliadt SB and Wiggins CL: Trends in United States
prostate cancer incidence rates by age and stage, 1995–2012. Cancer
Epidemiol Biomarkers Prev. 25:259–263. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Fukuda MN, Sato T, Nakayama J, Klier G,
Mikami M, Aoki D and Nozawa S: Trophinin and tastin, a novel cell
adhesion molecule complex with potential involvement in embryo
implantation. Genes Dev. 9:1199–1210. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Nadano D, Nakayama J, Matsuzawa S, Sato
TA, Matsuda T and Fukuda MN: Human tastin, a proline-rich
cytoplasmic protein, associates with the microtubular cytoskeleton.
Biochem J. 364:669–677. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ayala GE, Dai H, Li R, Ittmann M, Thompson
TC, Rowley D and Wheeler TM: Bystin in perineural invasion of
prostate cancer. Prostate. 66:266–272. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Yang S, Liu X, Yin Y, Fukuda MN and Zhou
J: Tastin is required for bipolar spindle assembly and centrosome
integrity during mitosis. FASEB J. 22:1960–1972. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Li CW and Chen BS: Investigating core
genetic-and-epigenetic cell cycle networks for stemness and
carcinogenic mechanisms, and cancer drug design using big database
mining and genome-wide next-generation sequencing data. Cell Cycle.
15:2593–2607. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Goswami CP and Nakshatri H: PROGgeneV2:
Enhancements on the existing database. BMC Cancer. 14:9702014.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Tang Z, Li C, Kang B, Gao G, Li C and
Zhang Z: GEPIA: A web server for cancer and normal gene expression
profiling and interactive analyses. Nucleic Acids Res. 45:W98–W102.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Arredouani MS, Lu B, Bhasin M, Eljanne M,
Yue W, Mosquera JM, Bubley GJ, Li V, Rubin MA, Libermann TA, et al:
Identification of the transcription factor single-minded homologue
2 as a potential biomarker and immunotherapy target in prostate
cancer. Clin Cancer Res. 15:5794–5802. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Grasso CS, Wu YM, Robinson DR, Cao X,
Dhanasekaran SM, Khan AP, Quist MJ, Jing X, Lonigro RJ, Brenner JC,
et al: The mutational landscape of lethal castration-resistant
prostate cancer. Nature. 487:239–243. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Varambally S, Yu J, Laxman B, Rhodes DR,
Mehra R, Tomlins SA, Shah RB, Chandran U, Monzon FA, Becich MJ, et
al: Integrative genomic and proteomic analysis of prostate cancer
reveals signatures of metastatic progression. Cancer Cell.
8:393–406. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wallace TA, Prueitt RL, Yi M, Howe TM,
Gillespie JW, Yfantis HG, Stephens RM, Caporaso NE, Loffredo CA and
Ambs S: Tumor immunobiological differences in prostate cancer
between African-American and European-American men. Cancer Res.
68:927–936. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Sboner A, Demichelis F, Calza S, Pawitan
Y, Setlur SR, Hoshida Y, Perner S, Adami HO, Fall K, Mucci LA, et
al: Molecular sampling of prostate cancer: A dilemma for predicting
disease progression. BMC Med Genomics. 3:82010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ross-Adams H, Lamb AD, Dunning MJ, Halim
S, Lindberg J, Massie CM, Egevad LA, Russell R, Ramos-Montoya A,
Vowler SL, et al: CamCaP Study Group: Integration of copy number
and transcriptomics provides risk stratification in prostate
cancer: A discovery and validation cohort study. EBioMedicine.
2:1133–1144. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hochegger H, Takeda S and Hunt T:
Cyclin-dependent kinases and cell-cycle transitions: Does one fit
all? Nat Rev Mol Cell Biol. 9:910–916. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ding H, Han C, Guo D, Wang D, Chen CS and
D'Ambrosio SM: OSU03012 activates Erk1/2 and Cdks leading to the
accumulation of cells in the S-phase and apoptosis. Int J Cancer.
123:2923–2930. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Xu X, Zhang H, Zhang Q, Huang Y, Dong J,
Liang Y, Liu HJ and Tong D: Porcine epidemic diarrhea virus N
protein prolongs S-phase cell cycle, induces endoplasmic reticulum
stress, and up-regulates interleukin-8 expression. Vet Microbiol.
164:212–221. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Basak S, Jacobs SB, Krieg AJ, Pathak N,
Zeng Q, Kaldis P, Giaccia AJ and Attardi LD: The
metastasis-associated gene Prl-3 is a p53 target involved in
cell-cycle regulation. Mol Cell. 30:303–314. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Song H, Hur I, Park HJ, Nam J, Park GB,
Kong KH, Hwang YM, Kim YS, Cho DH, Lee WJ, et al: Selenium Inhibits
Metastasis of Murine Melanoma Cells through the Induction of Cell
Cycle Arrest and Cell Death. Immune Netw. 9:236–242. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Nambotin SB, Tomimaru Y, Merle P, Wands JR
and Kim M: Functional consequences of WNT3/Frizzled7-mediated
signaling in non-transformed hepatic cells. Oncogenesis. 1:e312012.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kato S, Hayakawa Y, Sakurai H, Saiki I and
Yokoyama S: Mesenchymal-transitioned cancer cells instigate the
invasion of epithelial cancer cells through secretion of WNT3 and
WNT5B. Cancer Sci. 105:281–289. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Voloshanenko O, Erdmann G, Dubash TD,
Augustin I, Metzig M, Moffa G, Hundsrucker C, Kerr G, Sandmann T,
Anchang B, et al: Wnt secretion is required to maintain high levels
of Wnt activity in colon cancer cells. Nat Commun. 4:26102013.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wang HS, Nie X, Wu RB, Yuan HW, Ma YH, Liu
XL, Zhang JY, Deng XL, Na Q, Jin HY, et al: Downregulation of human
Wnt3 in gastric cancer suppresses cell proliferation and induces
apoptosis. OncoTargets Ther. 9:3849–3860. 2016. View Article : Google Scholar
|
|
26
|
Li Z, Pei XH, Yan J, Yan F, Cappell KM,
Whitehurst AW and Xiong Y: CUL9 mediates the functions of the 3M
complex and ubiquitylates survivin to maintain genome integrity.
Mol Cell. 54:805–819. 2014. View Article : Google Scholar : PubMed/NCBI
|