Introduction
The two major forms of inflammatory bowel disease
(IBD) are ulcerative colitis and Crohn’s disease. The pathogenesis
of IBD is complex, and the most common treatment includes the use
of immunosuppressive agents, antibiotics and biological agents
(1,2). However, this approach is
symptom-oriented with harmful side effects; therefore, new insights
for novel therapeutic strategies are required. The fact that
intestinal bacteria have a role in the pathogenesis of IBD presents
a potential strategy for the reduction of bowel inflammation
(3). A number of previous studies
manipulated the intestinal microbiota via the administration of
probiotics in the treatment of IBD (4–6).
Bacteria-mediated gene delivery in IBD has previously been
investigated and found to be successful. Previous studies have
demonstrated that bactofection of the colonic mucosa using
Salmonella typhimurium SL7207 carrying genes encoding
antioxidant, anti-inflammatory and anti-angiogenic factors has been
effective in the treatment of dextran sulfate sodium (DSS)-induced
colitis in rodents (7–9). Several studies have also investigated
the effects of antibiotic treatment on the course of experimental
colitis in the view of bacterial (probiotic) therapy. Antibiotic
pretreatment may enable certain invasive bacterial strains to
better colonize the gut during DSS-induced colitis (10). A number of antibiotics, including
minocycline, exert intestinal anti-inflammatory effects and
attenuate the activation of experimental colitis. Furthermore,
these antibiotics show an additive effect on the recovery of
intestinal damage when used along with probiotic bacterial strains
(11).
Notably, previous studies have investigated therapy
based on stem cell administration for the treatment of IBD animal
models. Amelioration of experimental colitis has been shown using
hematopoietic stem cells (12),
mesenchymal stem cells (13) and
colonic stem cells (14). However,
despite the promising data on stem cell therapy of IBD, the use of
induced pluripotent stem (iPS) cells for the treatment of IBD has
not yet been assessed. In two recent reviews, we presented the
rationale behind in vivo reprogramming using bacterial
vectors for gene delivery into the colon tissue (15,16).
We hypothesized that reprogramming intestinal cells into a
pluripotent state could demonstrate potential for IBD therapy and
prevention. This theory was based on the presumption that adult
somatic cells with the capacity to return to the pluripotent state
have the ability to differentiate into the desired cell phenotype,
this phenotype exhibiting resistance to the IBD-inducing stimuli.
Several studies have shown that a pre-emptive stem cell therapy may
lead to a protective phenotype later when the damaging event occurs
(17,18). However, the concept of in
vivo reprogramming has yet to be verified experimentally.
The present study aimed to investigate the use of
bacterial vectors for oral delivery of genes encoding reprogramming
factors into the colon and the effects of this treatment on the
course of DSS-induced colitis in mice. The therapeutic and
preventive approaches were compared, and the role of antibiotic
pretreatment in this process was also investigated.
Materials and methods
Bacteria and plasmids
The bacterial strain Salmonella typhimurium
SL7207, which is suitable for bactofection of eukaryotic cells, was
transformed with eukaryotic expression plasmids pcDNA3-RFP
(Addgene, Cambridge, MA, USA) (8),
carrying the gene encoding red fluorescent protein (RFP), or
pCX-OKS-2A (Addgene), carrying genes encoding the reprogramming
factors Oct3/4, Klf4 and Sox2 (OKS). The bacterial strain possessed
a natural resistance to streptomycin and the plasmid encoded a gene
for resistance to ampicillin. SL7207 bacteria with the appropriate
plasmid were grown in standard Luria-Bertani (LB) medium in the
presence of ampicillin (100 μg/ml) and streptomycin (50 μg/ml). The
bacterial culture was incubated without agitation until an optical
density of 0.4 was reached, measured at 600 nm. The culture was
then centrifuged (10 min, 5,000 × g, 4°C) and the pellet washed
three times with 15% glycerol in phosphate-buffered saline (PBS).
Bacteria were subsequently resuspended in 15% glycerol in PBS to
achieve 4×109 colony forming units (CFU) per ml. The
final solution was divided into 1-ml aliquots and stored at −80°C
until use for oral gavage. The bacterial count was confirmed by
plating serial dilutions of bacterial stock onto LB plates
containing the appropriate antibiotic.
In vivo survival of SL7207 bacteria
In the single gavage experiment, mice were fed via
gastric gavage with 109 CFU SL7207-OKS or SL7207-RFP in
0.25 ml. At several different time-points (4, 8, 12, 24, 36 and 48
h) a stool sample was taken, serially diluted in PBS and plated
onto LB plates containing streptomycin and ampicillin. In the
multiple gavage experiment, mice were fed four times via gastric
gavage every other day and the stool samples were taken 4 h after
the first gavage and then every 24 h. The average number of
surviving bacteria was expressed as CFU per mg of stool.
Colitis model
Male C57BL/6 mice (aged 12–16 weeks) were obtained
from Charles River Laboratories (Prague, Czech Republic). All mice
were kept in a controlled environment with a 12/12-h light/dark
cycle with ad libitum access to water and food. Mice
received either water or 2% DSS (MP Biomedicals, Solon, OH, USA;
molecular weight, 36,000–50,000) for seven days ad libitum
in drinking water starting from day 0. From day 7 DSS was changed
back to water for a further three days. Body weight and stool
consistency (0, normal; 1, soft-formed; 2, watery; 3, watery with
blood) were monitored every day until the end of the experiment.
Weight loss was expressed as a percentage of the initial weight of
the animal at the beginning of the experiment. Mice were sacrificed
on day 10. The animal experiments were approved by the
institutional review board and Ethics Committee of Comenius
University Faculty of Medicine (Bratislava, Slovakia).
Therapeutic vs. preventive bacterial
treatment
For the therapeutic experiment, 48 male C57BL/6 mice
were divided into six groups (n=8 per group) as follows: (i) DSS
PBS; (ii) H2O PBS; (iii) DSS RFP; (iv) H2O
RFP; (v) DSS OKS; and (vi) H2O OKS. Colitis was induced
according to the aforementioned protocol. On days 0, 2, 4 and 6 of
the colitis experiment mice in all groups were administered 0.25 ml
of the respective bacterial strain (109 CFU; groups 3–6)
or 15% glycerol in PBS (groups 1 and 2) using a gastric gavage. For
the preventive treatment experiment, 24 male C57BL/6 mice were
divided into four groups (n=6 per group): (i) DSS RFP; (ii)
H2O RFP; (iii) DSS OKS; and (iv) H2O OKS. On
days −4, −2 and 0 of the colitis experiment mice in all groups were
administered 0.25 ml of the respective bacterial strain
(109 CFU) using a gastric gavage.
Antibiotic pretreatment
Male C57BL/6 mice (n=32) were divided into four
groups (n=8 per group): (i) H2O PBS (CTRL); (ii) DSS
PBS; (iii) DSS ATB; and (iv) DSS ATB OKS (ATB indicates antibiotic
treatment). Mice in groups 3 and 4 were administered metronidazole
(750 mg/l) and streptomycin (1 g/l) in drinking water for four days
prior to the start of the experiment (days −8 to −4). On days −4,
−2 and 0 of the colitis experiment mice were administered 0.25 ml
OKS bacterial strain (109 CFU; group 4) or 15% glycerol
in PBS (groups 1, 2 and 3) using a gastric gavage. Colitis was
induced according to the aforementioned protocol. On days −4, −3,
−1, 1 and 3 of the experiment, a stool sample was taken from four
mice from each group, serially diluted and plated onto LB medium
containing streptomycin and ampicillin. Samples were obtained
immediately prior to the first gavage (day −4), 24 h after the
first, second and third gavage (days −3, −1 and 1) and 48 h after
the third gavage (day 3). The average bacterial count in the stool
was calculated from the number of colonies on the plates.
Statistical analysis
Data were analyzed using one- and two-way analysis
of variance, where appropriate. The Bonferroni post hoc test was
used to evaluate the differences between groups. P<0.05 was
considered to indicate a statistically significant difference. The
analysis was performed using GraphPad Prism 5 software (GraphPad
Software, Inc., San Diego, CA, USA). Data are presented as the mean
± standard error of the mean.
Results
In vivo survival of SL7207 bacteria
In the in vivo survival and passage analysis,
the administered bacteria were recovered from the stool after 36 h.
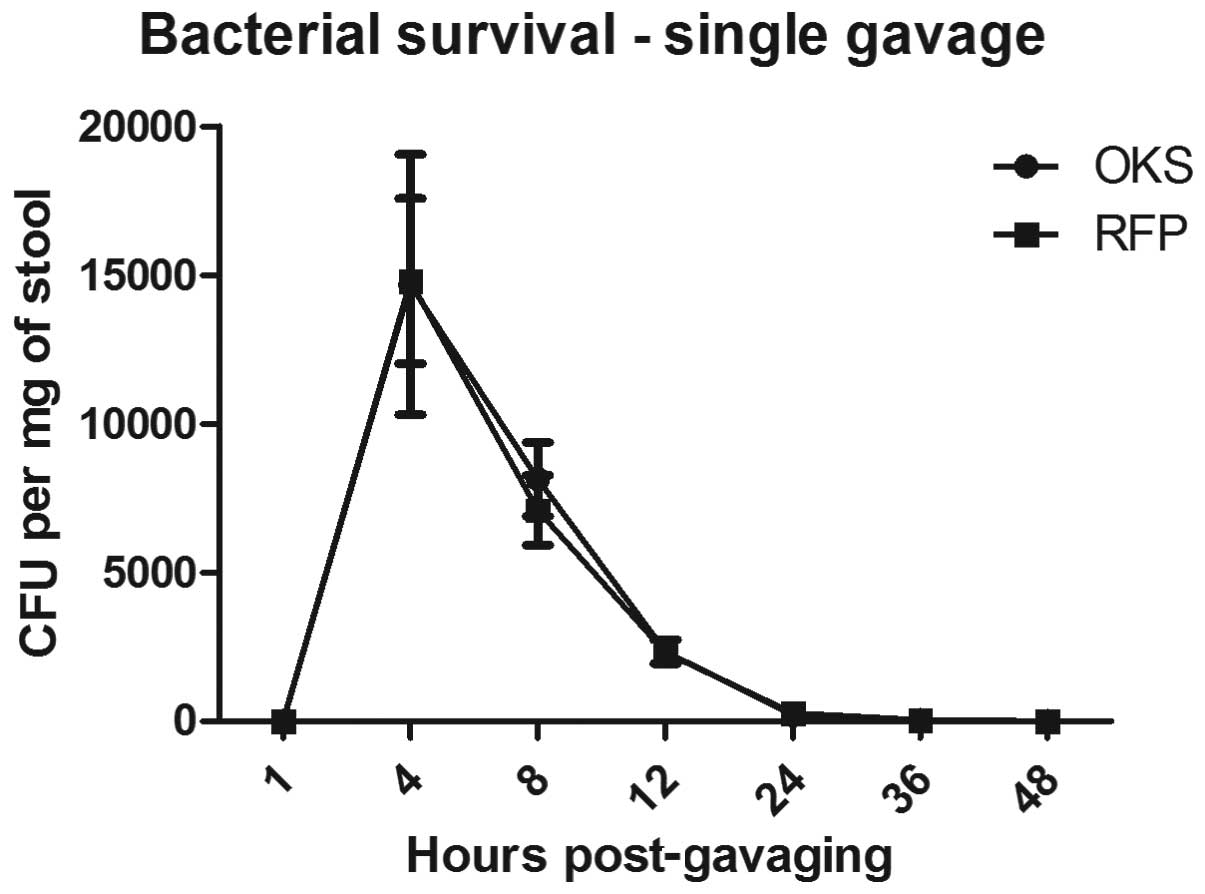
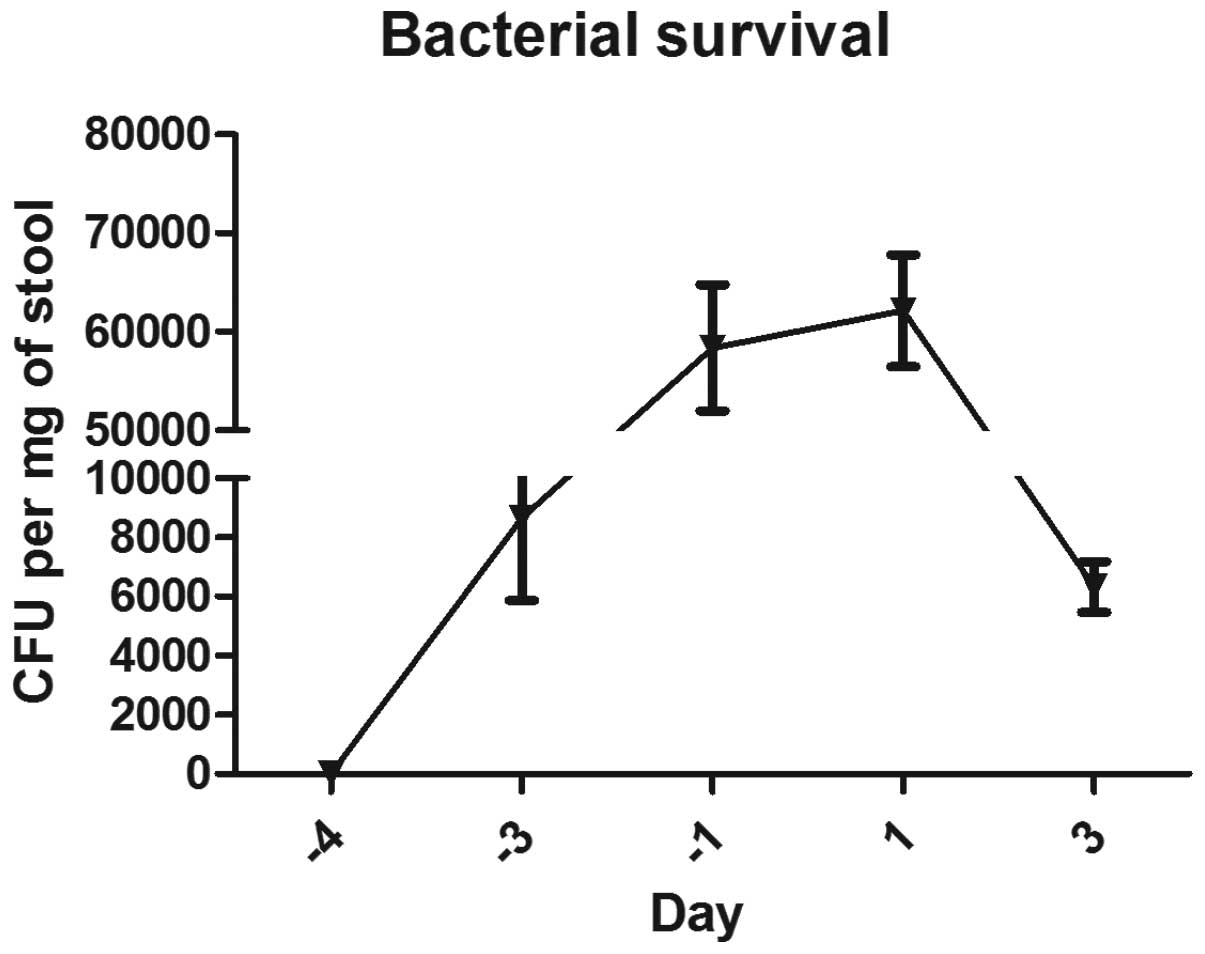
Fig. 1 shows the time course of
the bacterial recovery from the stool following a single
administration of 109 CFU SL7207-OKS or SL7207-RFP in
0.25 ml using gastric gavage. No bacteria were present in the stool
48 h after the single gavage.
To determine whether repeated administration of the
bacterial vector was likely to improve colonization of the colon,
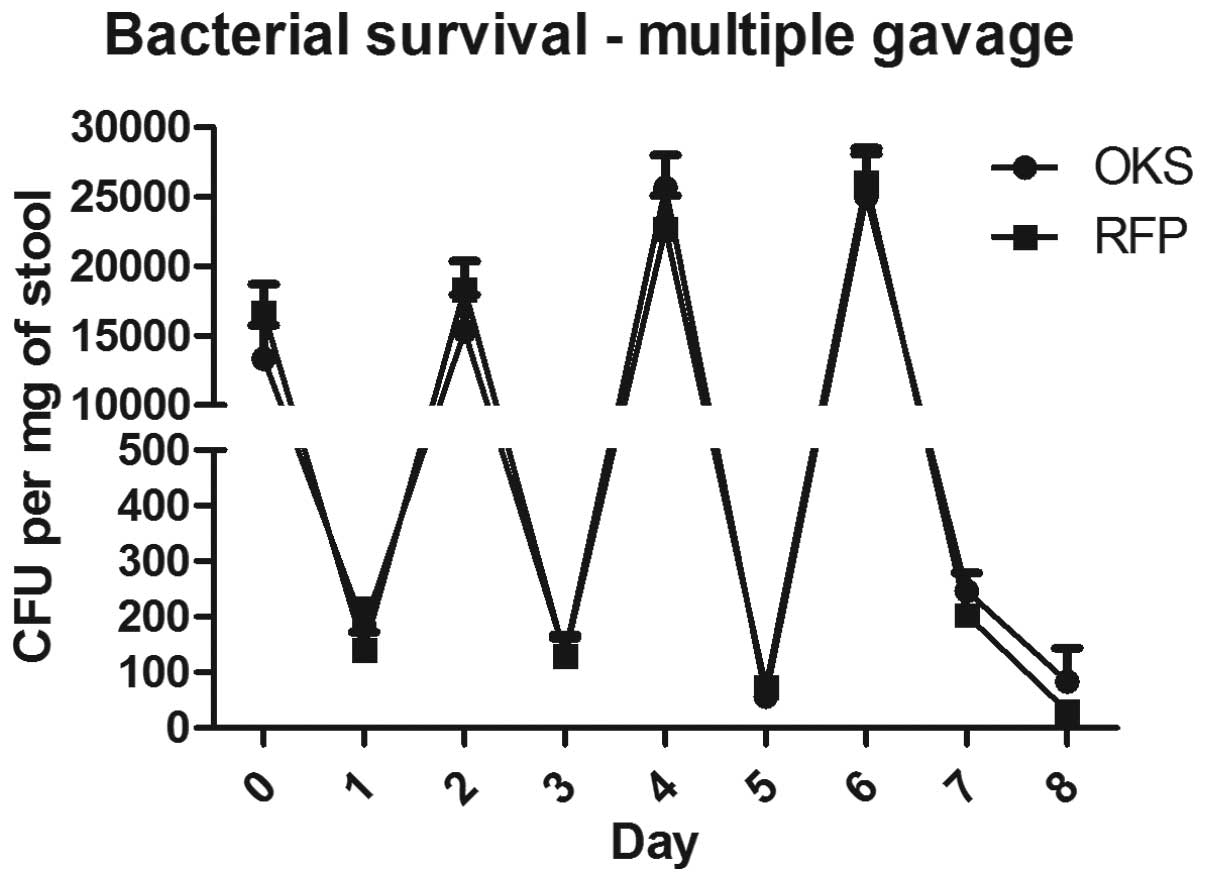
multiple gavages of the bacteria were analyzed. As shown in
Fig. 2, four consecutive
administrations of the bacterial vector every other day resulted in
detectable bacterial counts in the stool 48 h after the final
gavage (125 and 45 CFU/mg for OKS and RFP, respectively) compared
with no detectable bacteria following a single gavage.
Therapeutic vs. preventive bacterial
treatment of colitis
In the present study it was investigated whether the
bacterial strain SL7207 carrying the plasmid with genes encoding
three reprogramming factors (OKS) could have a positive effect on
the course of DSS-induced colitis. This was analyzed using basic
disease activity parameters, including weight loss, stool
consistency and colon length. In the first experiment, therapeutic
bacteria were administered concurrently with DSS treatment by four
gastric gavages every other day, starting from day 0. The weight
loss curves of all groups with colitis (DSS PBS, DSS RFP and DSS
OKS) indicated a similar course of ongoing colitis in all groups,
resulting in a significant weight loss at day 10 (75.8, 74.7 and
76.9%, respectively) compared with the control groups with no DSS
treatment. No significant differences were found among the DSS
groups throughout the course of colitis experiment (data not
shown). In addition, no significant differences among the DSS
groups were observed in stool consistency (mean score, 2.33, 2.11
and 2.00, respectively) and colon length (mean length, 6.7, 6.4 and
6.5 cm, respectively) on the day of sacrifice.
In the preventive bacterial treatment, bacterial
vectors were administered prior to induction of colitis by three
consecutive gastric gavages every other day, starting from day −4.
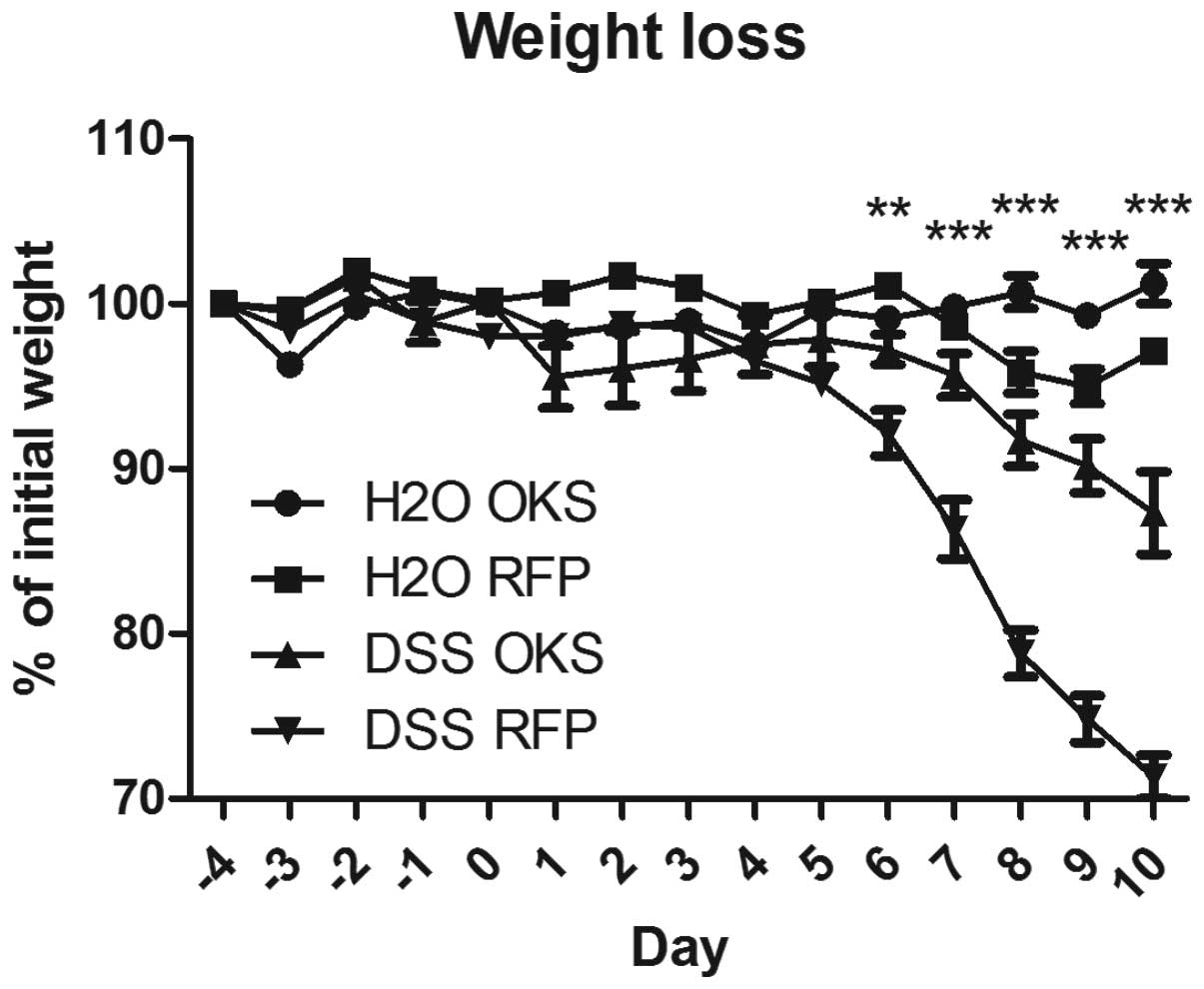
The group treated with bacteria carrying reprogramming genes (DSS
OKS) showed significantly improved weight loss compared with the
group receiving control bacteria (DSS RFP) (Fig. 3). This improvement was apparent
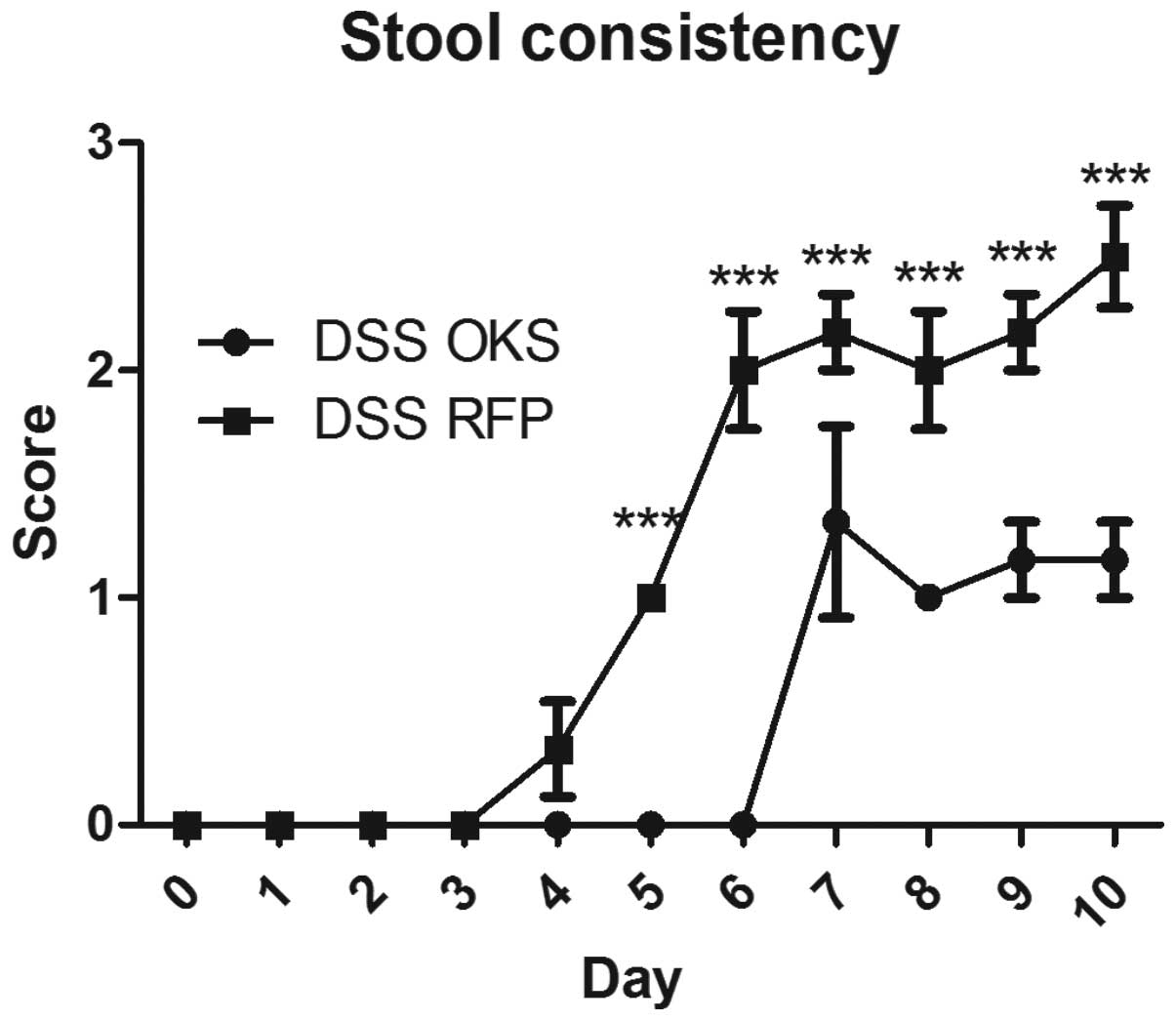
from day 6 to the end of the experiment. Similar differences
between the DSS OKS and DSS RFP groups were observed for other
parameters, including stool consistency (starting from day 5)
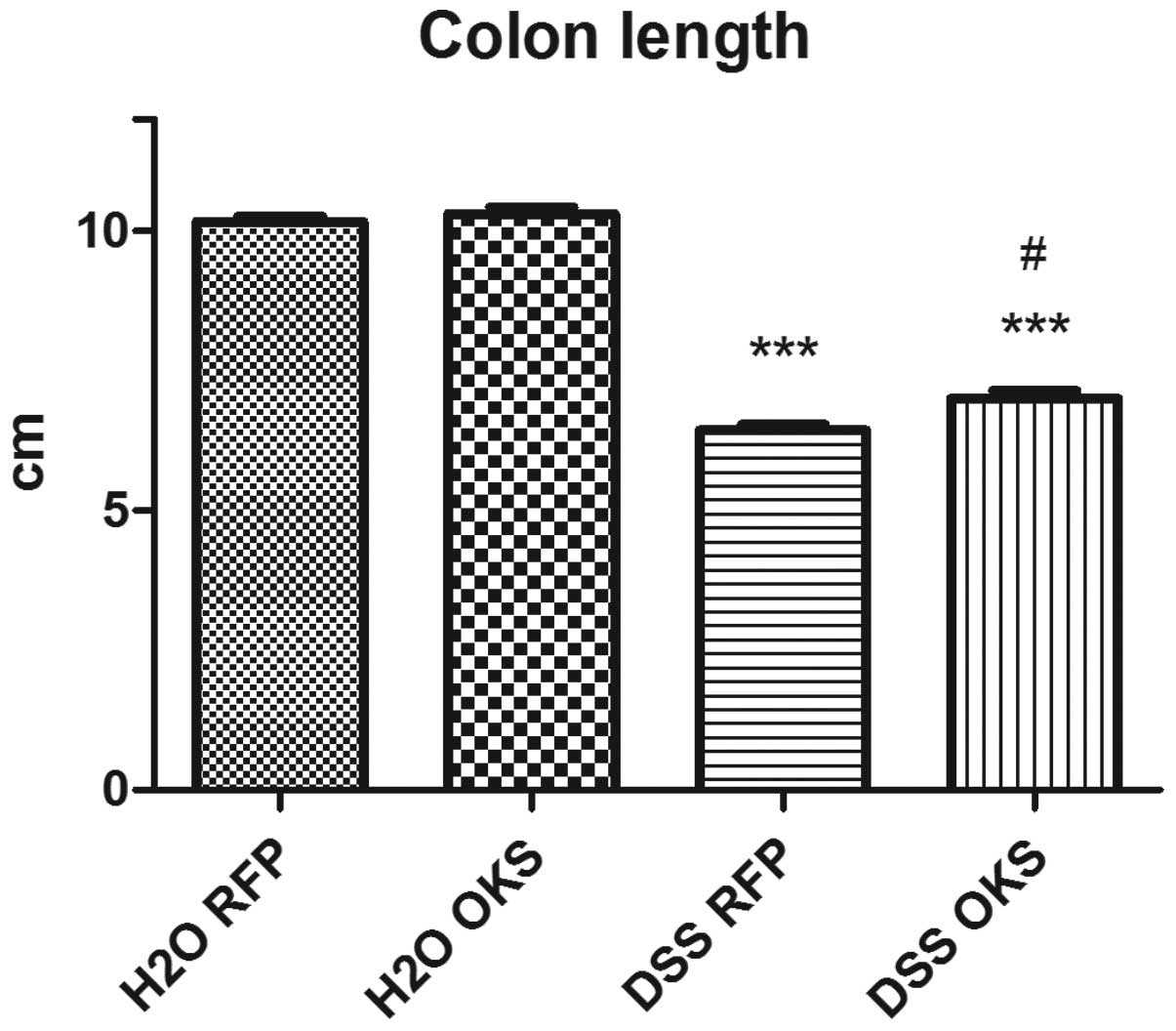
(Fig. 4) and colon length (on the
day of sacrifice) (Fig. 5).
Antibiotic pretreatment
Since the preventive approach was found to be more
beneficial than the therapeutic approach, the effect of antibiotic
pretreatment was then assessed. The same preventive treatment
protocol was performed, consisting of three gastric gavages of
bacterial vector or PBS every other day, starting from day −4, and
followed by the induction of colitis by DSS. Mice in the DSS ATB
and DSS ATB OKS groups received metronidazole and streptomycin in
their drinking water for four days prior to the first gavage.
Antibiotic pretreatment allowed the administered bacterial strain
SL7207-OKS to efficiently colonize the gut of mice in the DSS ATB
OKS group, as shown by plating the stool samples onto LB medium
containing streptomycin and ampicillin (Fig. 6). A significant number of resistant
bacteria were detected, even three days after the last gavage. No
bacteria resistant to both antibiotics were observed in the stools
of mice in other groups at any time-point (CTRL, DSS and DSS
ATB).
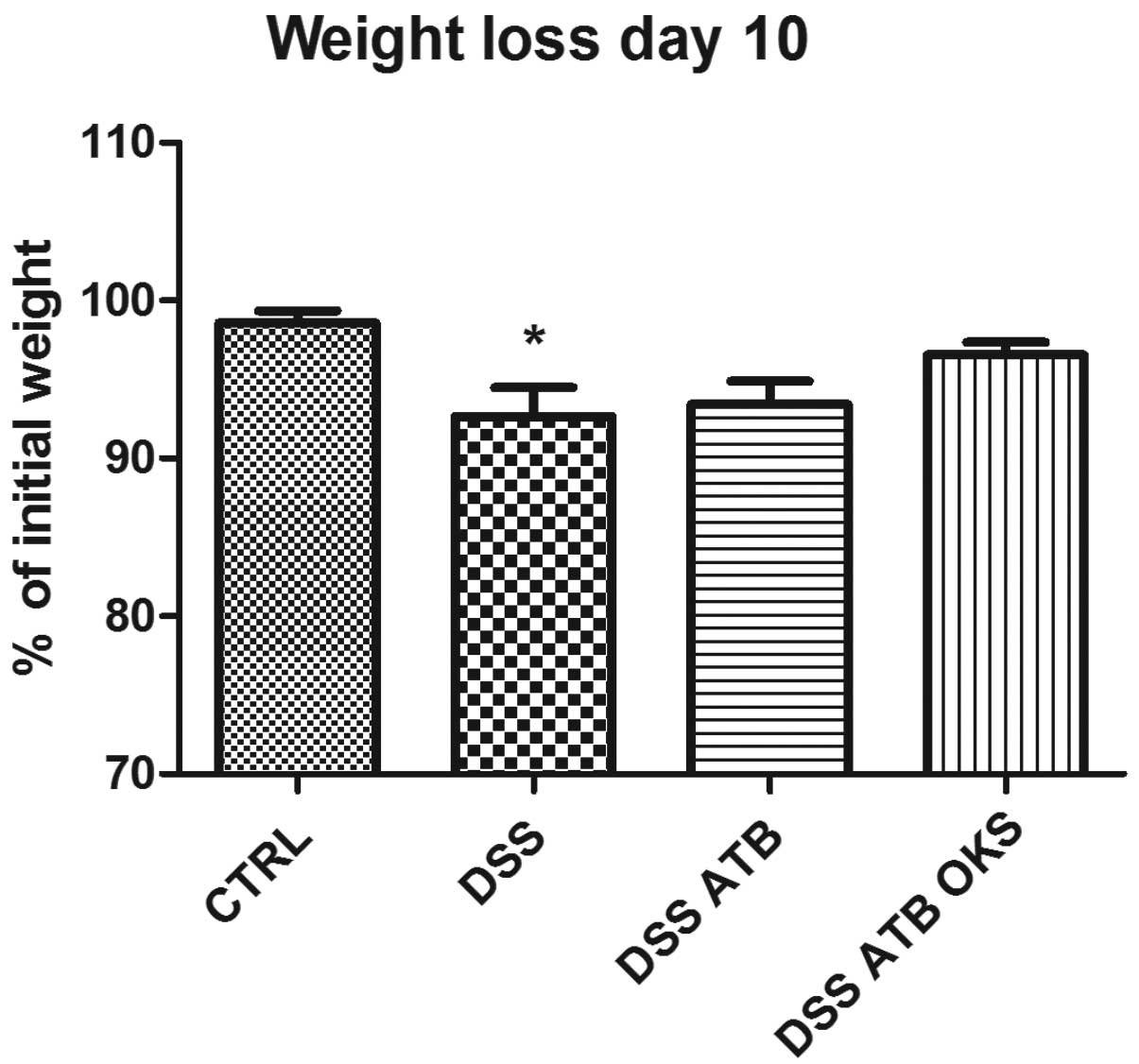
The DSS group with no treatment showed significant
weight reduction compared with the control group at the end of the
experiment (Fig. 7). Neither of
the groups that received antibiotic pretreatment (DSS ATB and DSS
ATB OKS) showed significant weight reduction, and the DSS ATB OKS
group showed only a slight weight loss (96.5%) compared with the
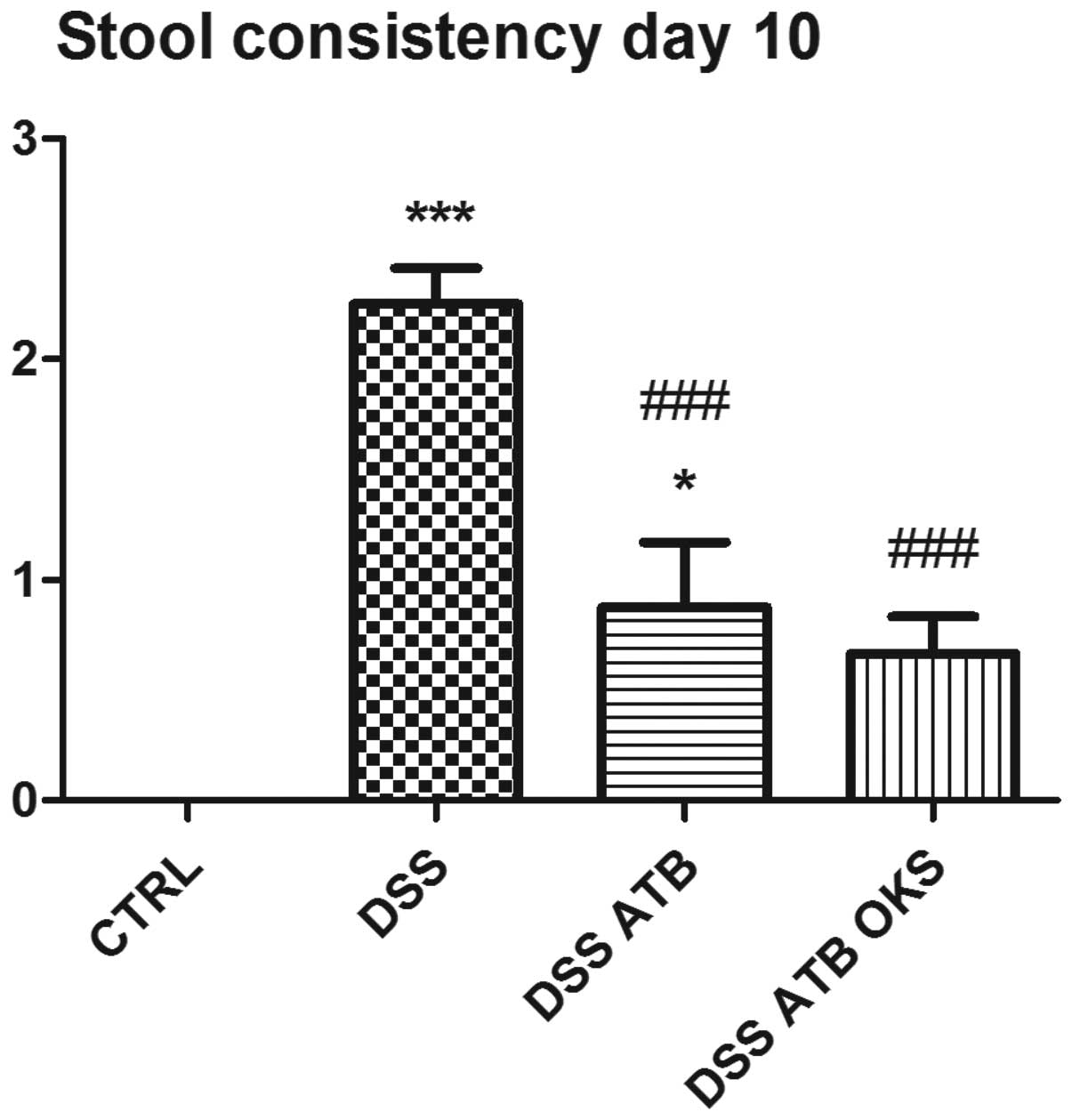
control group (98.6%). Furthermore, the DSS groups pretreated with
antibiotics exhibited significantly improved stool consistency at
the end of the experiment compared with the DSS group with no
antibiotic or bacterial treatment (Fig. 8). In addition, no significant
difference was observed between the DSS group receiving antibiotic
and bacterial treatment (DSS ATB OKS) and the control group without
colitis. The DSS and DSS ATB groups showed significantly lower
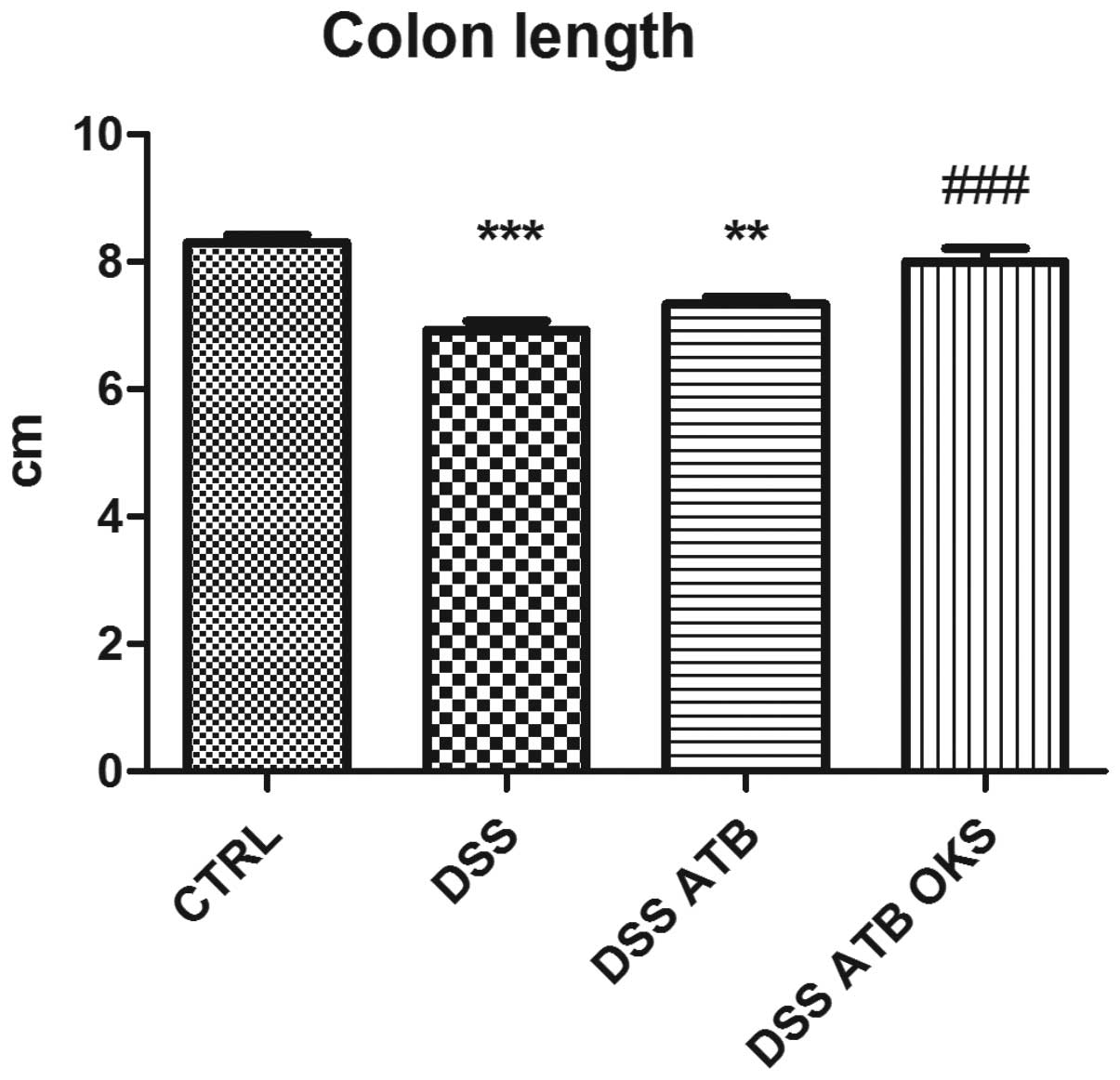
colon length compared with the control group (Fig. 9). This effect, however, was not
observed in the DSS group with both antibiotic and bacterial
treatment. Furthermore, the DSS ATB OKS group had significantly
improved colon length compared with the DSS group with no
treatment.
Discussion
The present study analyzed the effects of the
Salmonella-mediated delivery of reprogramming genes into the colon
and pretreatment with the antibiotics metronidazole and
streptomycin on chemically induced colitis. Two gavaging protocols,
therapeutic and preventive, were initially compared for the
administration of the bacterial vectors. It was shown that
pre-emptive bacterial treatment was superior to the therapeutic
treatment in terms of the basic parameters of colitis activity. The
lack of efficiency of the therapeutic approach may be due to the
requirement for healthy tissue for the bacterial vectors to enter
the host and release the expression plasmid into the host cells.
However, efficient delivery of the genes by SL7207 bacteria into
colonic cells, even during DSS-induced colitis, has been shown
previously (7,8). Assuming that reprogramming did occur
in the colon, the enhanced beneficial effects of the preventive
gene delivery of reprogramming factors are in accordance with the
preliminary data published on stem cell therapy that indicate that
a pre-emptive stem cell therapy may lead to a protective phenotype
later when the damaging event occurs (17,18).
Although there has been some investigation into the
differentiation of iPS cells into intestinal cells (19,20),
little is known regarding the specific conditions that are required
for the efficient reprogramming of intestinal cells into a
pluripotent state. To the best of our knowledge, the generation of
iPS cells from intestinal somatic cells has not yet been reported.
Given that the degeneration of intestinal mucosal cells has an
essential role in the pathogenesis of IBD, therapy based on their
reprogramming provides a rationale for the protection of these
cells against the harmful effects and thus prevention of damage in
IBD. It appears that the reprogramming processes could occur in the
intestine using in vivo gene delivery, despite the fact that
low transfection and reprogramming efficiencies are expected.
As a follow-on to our initial experiment, the
preventive administration protocol was employed and the effect of
antibiotic pretreatment was investigated. It was demonstrated that
the administered bacteria were present only in the stool of mice
with colitis that received both antibiotics and SL7207-OKS (the DSS
ATB OKS group). Notably, significant amounts of the administered
bacteria were detected in the stool even 72 h after the last
gavage. This indicates that antibiotic pretreatment markedly
improved the exposure time of the bacterial vector to the colonic
mucosa, thus increasing the efficiency of gene transfer. The actual
invasion of SL7207 bacteria into the colon tissue was shown in our
previous study (8).
Although neither antibiotics nor bacterial
reprogramming significantly altered the weight of the animals in
comparison with colitis control mice, the trend towards reducing
weight loss was clear. Colon length, as a measure of colitis
activity, was significantly improved by the combination of
antibiotic pretreatment and bacteria-mediated reprogramming
compared with the control colitis group. Similar results were found
for stool consistency, another measure of colitis activity.
Antibiotics alone and in combination with reprogramming genes led
to an improved stool consistency score, with the lowest score
observed in the combination group. These results indicate that
antibiotic pretreatment may enhance bacterial gene delivery into
the colon. Furthermore, in vivo reprogramming of colon cells
appears to have a preventive effect on the course of colitis.
It has been shown that gut inflammation stimulates
horizontal gene transfer between the members of the gut microbial
community, therefore spreading the determinants of fitness,
virulence and antibiotic resistance (21). The disruption of microbial flora
composition observed following antibiotic treatment may favor
several specific bacterial species that contain the necessary
factors, which are further spread across the community if the
inflammation takes place. This may explain the positive effect of
antibiotic pretreatment on the course of DSS colitis.
The present study does not provide direct evidence
of the expression of the reprogramming factors, nor is there
evidence of the effect of antibiotic pretreatment on the microbial
flora in the gut. In addition, the study lacks histological and
biochemical quantification of the inflammation. These limitations
make this study a preliminary investigation. However, this
experiment should be viewed as a first attempt to use bacterial
infection-mediated gene delivery in the context of a microflora
modified by antibiotics. It also shows for the first time, to the
best of our knowledge, the potential for therapeutic treatments for
IBD based on reprogramming the cells in vivo. The data in
the present study may therefore be used for further experiments
investigating the mechanism of in vivo reprogramming in
IBD.
Acknowledgements
This study was supported by the grant VEGA 1/0206/12
from The Ministry of Education, Science, Research and Sport of the
Slovak Republic.
References
|
1
|
Triantafillidis JK, Merikas E and
Georgopoulos F: Current and emerging drugs for the treatment of
inflammatory bowel disease. Drug Des Devel Ther. 5:185–210. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Blonski W, Buchner AM and Lichtenstein GR:
Inflammatory bowel disease therapy: current state-of-the-art. Curr
Opin Gastroenterol. 27:346–357. 2011.PubMed/NCBI
|
|
3
|
Thompson-Chagoyán OC, Maldonado J and Gil
A: Aetiology of inflammatory bowel disease (IBD): role of
intestinal microbiota and gut-associated lymphoid tissue immune
response. Clin Nutr. 24:339–352. 2005.PubMed/NCBI
|
|
4
|
Matsumoto S, Hara T, Hori T, et al:
Probiotic Lactobacillus-induced improvement in murine chronic
inflammatory bowel disease is associated with the down-regulation
of pro-inflammatory cytokines in lamina propria mononuclear cells.
Clin Exp Immunol. 140:417–426. 2005. View Article : Google Scholar
|
|
5
|
Steidler L, Hans W, Schotte L, et al:
Treatment of murine colitis by Lactococcus lactis secreting
interleukin-10. Science. 289:1352–1355. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bibiloni R, Fedorak RN, Tannock GW, et al:
VSL#3 probiotic-mixture induces remission in patients with active
ulcerative colitis. Am J Gastroenterol. 100:1539–1546. 2005.
|
|
7
|
Pálffy R, Behuliak M, Gardlík R, Jáni P,
Kádasi L, Turna J and Celec P: Oral in vivo bactofection in dextran
sulfate sodium treated female Wistar rats. Folia Biol (Krakow).
58:171–176. 2010.PubMed/NCBI
|
|
8
|
Palffy R, Gardlik R, Behuliak M, et al:
Salmonella-mediated gene therapy in experimental colitis in mice.
Exp Biol Med (Maywood). 236:177–183. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Gardlik R, Bartonova A and Celec P:
Therapeutic DNA vaccination and RNA interference in inflammatory
bowel disease. Int J Mol Med. 32:492–496. 2013.PubMed/NCBI
|
|
10
|
Drouet M, Vignal C, Singer E, et al: AIEC
colonization and pathogenicity: influence of previous antibiotic
treatment and preexisting inflammation. Inflamm Bowel Dis.
18:1923–1931. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Garrido-Mesa N, Utrilla P, Comalada M, et
al: The association of minocycline and the probiotic Escherichia
coli Nissle 1917 results in an additive beneficial effect in a
DSS model of reactivated colitis in mice. Biochem Pharmacol.
82:1891–1900. 2011.PubMed/NCBI
|
|
12
|
Khalil PN, Weiler V, Nelson PJ, et al:
Nonmyeloablative stem cell therapy enhances microcirculation and
tissue regeneration in murine inflammatory bowel disease.
Gastroenterology. 132:944–954. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
González MA, Gonzalez-Rey E, Rico L,
Büscher D and Delgado M: Adipose-derived mesenchymal stem cells
alleviate experimental colitis by inhibiting inflammatory and
autoimmune responses. Gastroenterology. 136:978–989.
2009.PubMed/NCBI
|
|
14
|
Zhou Q, Price DD, Dreher KL, Pronold B,
Callam CS, Sharma J and Verne GN: Localized colonic stem cell
transplantation enhances tissue regeneration in murine colitis. J
Cell Mol Med. 16:1900–1915. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wagnerova A and Gardlik R: In vivo
reprogramming in inflammatory bowel disease. Gene Ther.
20:1111–1118. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Gardlik R: Inducing pluripotency using in
vivo gene therapy. Med Hypotheses. 79:197–201. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Sheng X, Reppel M, Nguemo F, Mohammad FI,
Kuzmenkin A, Hescheler J and Pfannkuche K: Human pluripotent stem
cell-derived cardiomyocytes: response to TTX and lidocain reveals
strong cell to cell variability. PLoS One. 7:e459632012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yamada S, Nelson TJ, Behfar A, Crespo-Diaz
RJ, Fraidenraich D and Terzic A: Stem cell transplant into
preimplantation embryo yields myocardial infarction-resistant adult
phenotype. Stem Cells. 27:1697–1705. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Finkbeiner SR and Spence JR: A gutsy task:
generating intestinal tissue from human pluripotent stem cells. Dig
Dis Sci. 58:1176–1184. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Spence JR, Mayhew CN, Rankin SA, et al:
Directed differentiation of human pluripotent stem cells into
intestinal tissue in vitro. Nature. 470:105–109. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Stecher B, Denzler R, Maier L, et al: Gut
inflammation can boost horizontal gene transfer between pathogenic
and commensal Enterobacteriaceae. Proc Natl Acad Sci USA.
109:1269–1274. 2012. View Article : Google Scholar : PubMed/NCBI
|