Introduction
The 5α-dihydrotestosterone (DHT) androgen is
produced primarily by 5α-reductase in the testes (1). DHT regulates male reproductive
development, testes formation, growth of skeletal muscle and hair
growth, through activation of the androgen receptor (2). The affinity of DHT is 10-fold greater
than that of teststerone for the androgen receptor, and leads to
its hyperactivation, which induces shortening of the anagen phase
of hair follicle growth (3–5).
The hair growth cycle is modulated predominantly by
dermal papilla cells (DPCs), which are mesenchymal cells located at
the base of hair follicles, regulating formation of the hair
follicle and hair growth cycle through secretion of growth factors
and cytokines (6–11). Previous studies have demonstrated
that DHT inhibits protein kinase C, regulates of the expression of
B-cell lymphoma 2 (bcl-2)/blc-2-associated x protein (bax), and
upregulates the expression of dickkopf 1 in the DPCs, leading to
cell apoptosis, shortening of the hair cycle, a reduction in hair
growth, and hair loss (12–14).
MicroRNAs (miRNAs) are a class of small (~22 nt)
noncoding RNAs, which bind to mRNAs in a sequence-specific manner
to regulate the translation of target genes (15,16).
miRNAs are important in development, apoptosis and cell growth
(17). Various studies have been
performed to investigate the role of miRNAs in dermal papilla cells
from the balding and non balding scalp (14). In addition, investigations using
mice, in which Dicer, a key enzyme of miRNA metabolism, has been
knocked out, have revealed that miRNAs are essential for the
morphogenesis and maintenance of hair follicles (18).
However, although DHT is well known as a key
regulator of balding and hair follicle morphogenesis, DHT-dependent
alterations of the miRNA expression profile and putative mechanisms
remain to be elucidated. The present study investigated the
cellular effects of DHT and the miRNA expression prolife in normal
human DPCs (nHDPCs).
Materials and methods
Cells and culture conditions
The nHDPCs were purchased from Innoprot (Biscay,
Spain) and were cultured in Dulbecco’s modified Eagle’s medium
(DMEM; Gibco, Life Technologies, Grand Island, NY, USA),
supplemented with 10% fetal bovine serum (FBS; Sigma-Aldrich, St.
Louis, MO, USA) and 1% penicillin/streptomycin at 37°C in a
humidified atmosphere with 5% CO2.
Cell viability assay
The viability of the nHDPCs was measured using a
water-soluble tetrazolium salt (WST-1) assay (EZ-Cytox Cell
Viability Assay kit; Itsbio, Seoul, Korea). For the cell viability
assay, the nHDPCs were plated at a density of 5×103
cells/well in 96-well plates. After 24 h, the cells were treated
with doses of DHT between 0 and 1 mM at 37°C for 24, 48, or 72 h.
The cells were then incubated with WST-1 reagent at 37°C for 30
min, and the optical density was determined at 450 nm using a
microplate reader (iMark; Bio-Rad Laboratories, Inc., Hercules, CA,
USA).
Cell cycle assay
A propidium iodide (PI) staining based cell cycle
assay was performed using standard procedures, as described
previously (10). The nHDPCs
(2×106) were plated in 60 mm culture dishes and treated
with DHT for 24 h. The cells were then trypsinized with 0.25%
Trypsin-EDTA (Gibco Life Technologies) at 37°C, pelleted, washed
with phosphate-buffered saline (PBS), and fixed with 70% ethanol at
4°C for 3 h. The DNA in the fixed cells was stained using staining
solution containing 50 μg/ml PI (Sigma-Aldrich), 0.5% Triton
X-100 (Bioshop, Burlington, ON, Canada), and 100 μg/ml RNase
(Bioshop) at 37°C for 1 h. Following staining, the cells were
analyzed using a FL2 channel with an excitation wavelength of 488
nm and an emission wavelength of 578 nm, on a FACSCaliber flow
cytometer (BD Biosciences, San Jose, CA, USA).
Reactive oxygen species (ROS)
measurement
The measurement of ROS was performed, as previously
reported, using 2′,7′-dichlorofluorescein diacetate (DCF-DA)
(19). The nHDPCs
(2×106) were plated in 60 mm culture dishes and treated
with DHT at 37°C for 24 h. 2′, 7′-Dichlorodihydrofluorescin
diacetate (DCF-DA; 20 μM) was added to the culture medium,
and the cells were incubated at 37°C for 1 h. The cells were then
trypsinized with 0.25% Trypsin-EDTA at 37°C, pelleted, washed with
PBS, and analyzed using a FL1 channel with an excitation wavelength
of 488 nm and an emission wavelength of 530 nm on a FACSCaliber
flow cytometer (BD Biosciences).
Senescence-associated β-galactosidase
(SA-β-gal) assay
For the detection of senescent cells, an SA-β-gal
assay was performed, as previously described (20). Briefly, the nHDPCs
(2×106) were plated in 60 mm culture dishes and treated
with DHT at 37°C for 24 h. The cells were then fixed with Fixative
Solution (Senescence Detection kit; Biovision, Milpitas, CA, USA)
and stained using a Staining Solution mix (Senescence Detection
kit) supplemented with X-gal at 37°C for 24 h. Images of the
SA-β-gal stained cells were captured using a camera mounted to a
light microscope (CKX41; Olympus Corporation, Tokyo, Japan), and
the number of stained cells were counted in five randomly selected
microscopic fields from each condition.
miRNA microarray
The RNA in the cells was isolated using TRIzol
reagent (Gibco Life Technologies), according to the manufacturer’s
instructions. The RNA integrity was evaluated using an Agilent 2100
Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA), and the
RNA quality was evaluated using spectrophotometry at the 260/280 nm
ratio (Ultrospec 2100 Pro UV-Vis; Amersham Biosciences, GE
Healthcare Life Sciences, Piscataway, NJ, USA). Samples with an RNA
integrity score >7.8 and an RNA quality score >2.0 were used
for the microarray. A total of 100 ng RNA was labeled with cyanine
dye (Cy3) using an Agilent miRNA labeling kit (Agilent
Technologies). The labeled RNAs were purified using Micro Bio-Spin
P-6 columns (Bio-Rad Laboratories, Inc.) and hybridized using a
SurePrint G3 Human v16 miRNA Microarray kit (8×60 K; Release 16.0;
Agilent Technologies) at 65°C for 20 h. The microarray was scanned
using an Agilent microarray scanner (Agilent Technologies), and the
images were analyzed using Agilent Feature Extraction version 10.7
software (Agilent Technologies). The digitized data were analyzed
and the fold change was determined using GeneSpring GX version 11.5
software (Agilent Technologies).
miRNA target gene prediction and
biological function analysis
The putative target genes of significant miRNAs were
identified using the probability of interaction by target
accessibility (PITA; http://genie.weizmann.ac.il), microRNAorg (http://www.microrna.org) and TargetScan (http://www.targetscan.org) target prediction systems.
The Gene Ontologies (GOs) of the putative target genes were
analyzed using the Database for Annotation, Visualization and
Integrated Discovery (DAVID) Bioinformatics Resource 6.7
(http://david.abcc.ncifcrf.gov).
Statistical analysis
The data are presented as the mean ± standard
deviation. Statistical significance was calculated using Student’s
two-tailed t-test. Statistical analyses were conducted using
Microsoft Excel 2013 (Microsoft Corporation, Redmond, WA, USA).
P<0.01 was considered to indicate a statistically significant
difference, unless otherwise indicated.
Results
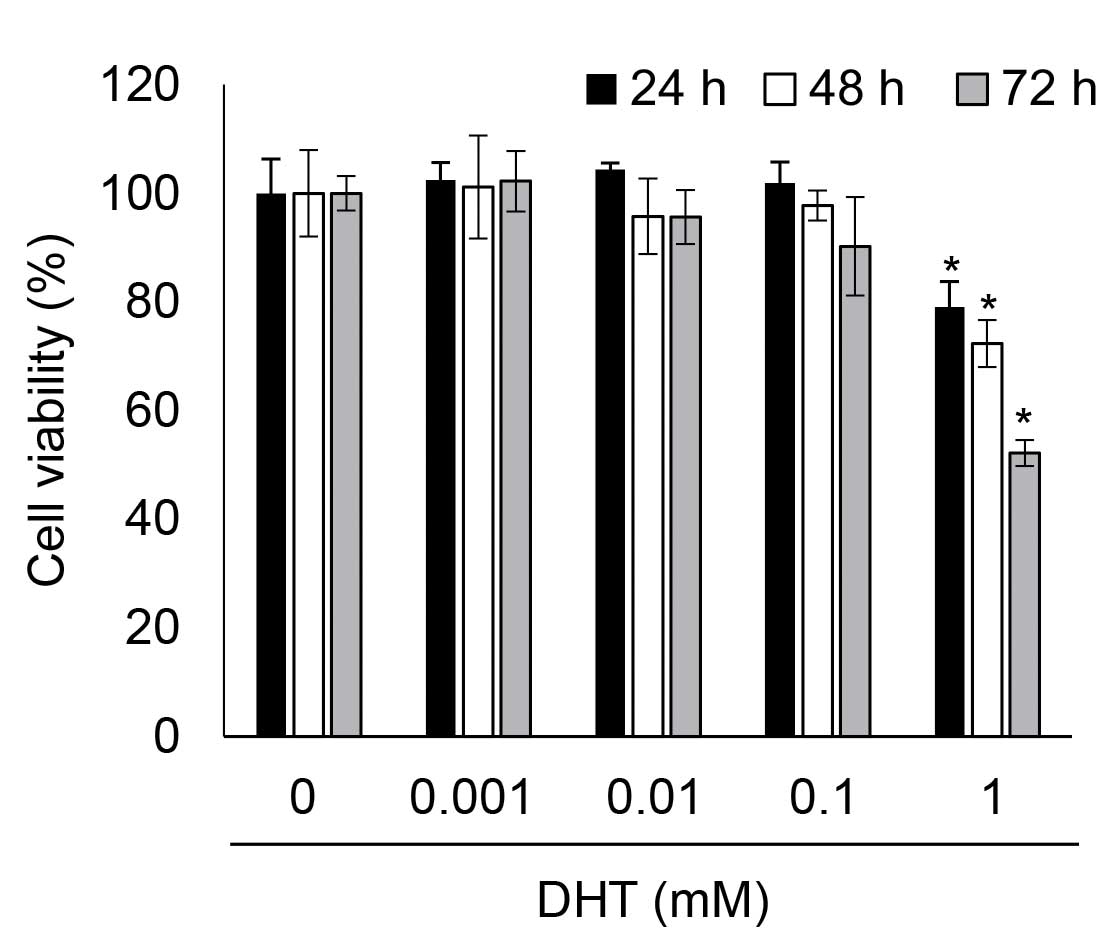
DHT induced cytotoxicity in nHDPCs
To determine whether DHT was associated with cell
viability in nHDPCs, the present study analyzed the viability of
DHT-treated nHDPCs after 24, 48, and 72 h using a WST-1 assay. Low
concentrations of DHT (<0.1 mM) demonstrated no significant
toxicity in the nHDPCs at any of the time-points assessed. However,
as shown in Fig. 1, cytotoxicity
was significantly increased by 1 mM DHT in the nHDPCs at every
time-point assessed. Thus, it was determined that 1 mM DHT-induced
cytotoxicity in the nHDPCs following exposure for ≥24 h, which led
to an exposure duration of 24 h being selected for use in the
subsequent experiments.
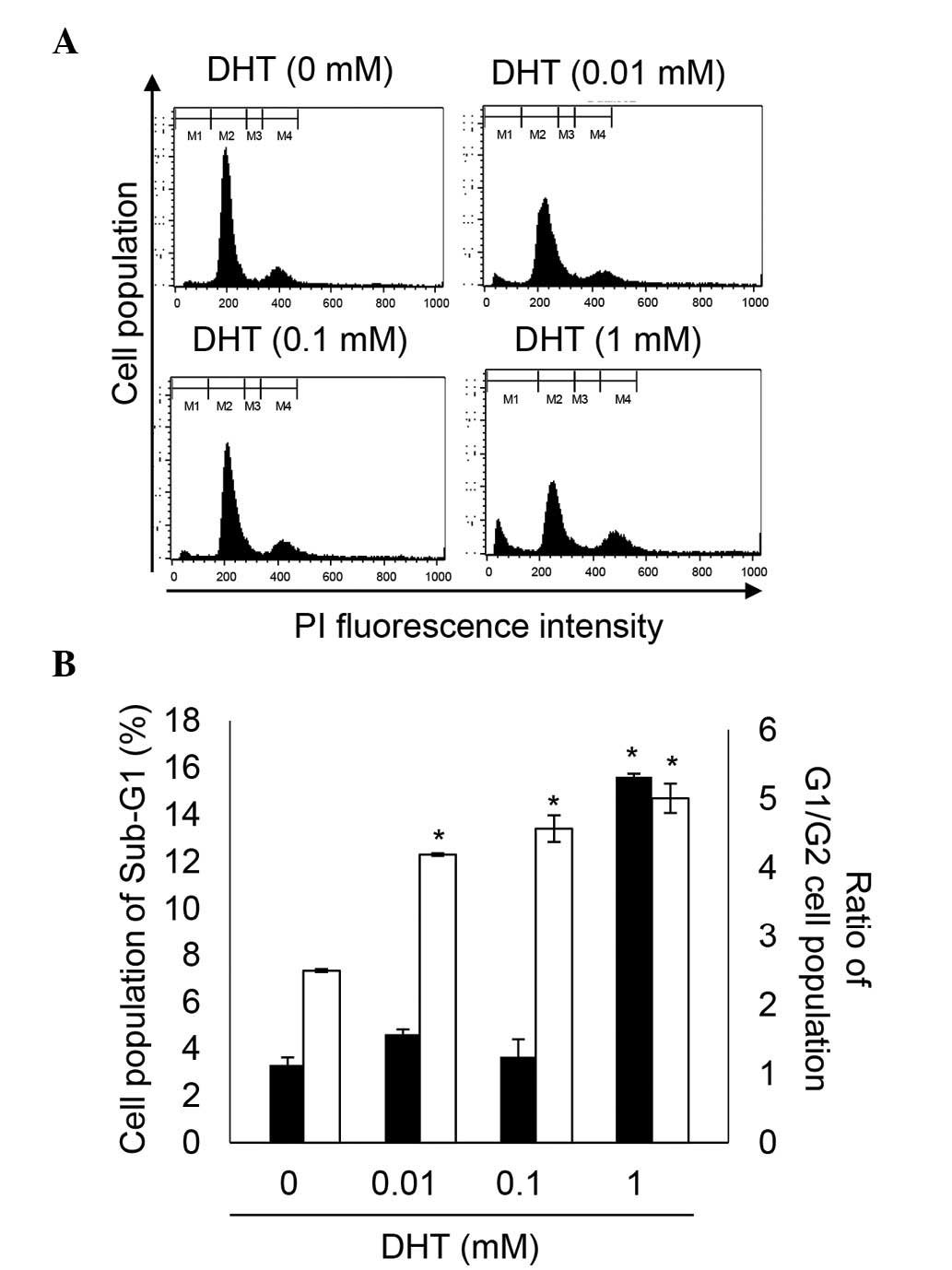
DHT induces cell death and cell cycle
arrest in nHDPCs
Previous experiments established that high levels of
DHT induce apoptosis (14,21). In agreement with the previous
experiments (Fig. 2), the present
study demonstrated that 1 mM DHT increased cell death between 3.36
and 15.62% in the nHDPCs. In addition, the G1/G2 ratio was
significantly increased by concentrations of DHT
>10−6 M, in a dose-dependent manner. The DHT-induced
increment in G1/G2 ratio indicated that DHT-induced G2 cell cycle
arrest. Therefore, high-doses of DHT reduced cell viability through
induction of cell death and G2 cell cycle arrest in the nHDPCs.
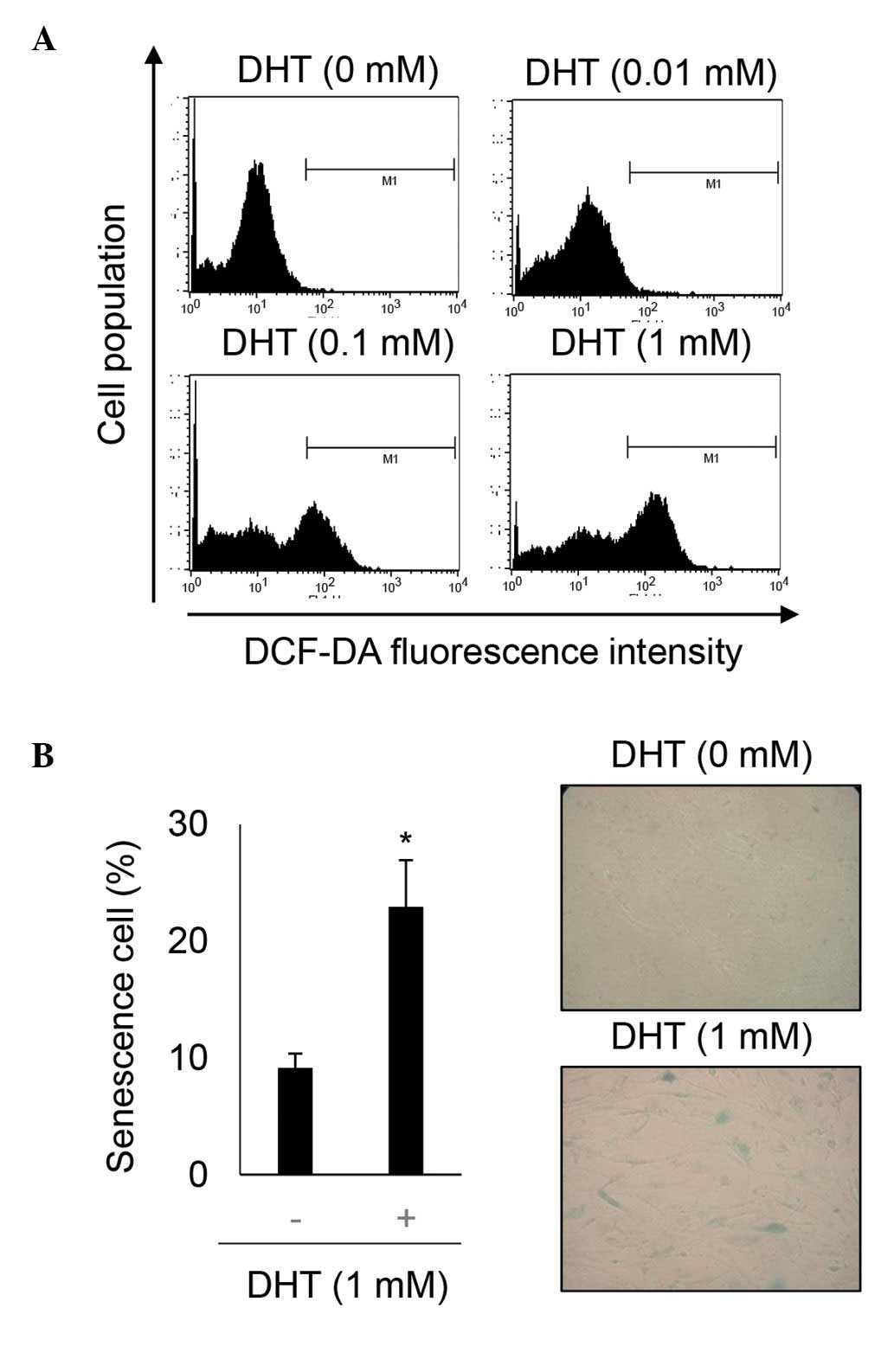
DHT increases ROS levels in nHDPCs
DHT can induce ROS in prostate cancer cell lines,
which express the androgen receptor at a high level (14,22–24).
Additionally, ROS are a key inducer of retinoblastoma-mediated
senescence (25). As nHPDCs also
express androgens at a high level (26), the present study investigated
whether 1 mM DHT-induced ROS in these cells. The levels of ROS were
determined using DCF-DA staining in untreated nHDPCs and in 1 mM
DHT-treated nHDPCs. As shown Fig.
3A, DHT significantly increased the level of ROS in the nHDPCs.
In addition, the cellular effect underlying the effect of 1 mM DHT
in enhancing ROS levels in the nHDPCs was investigated. As shown in
previous experiments in a prostate cell line (23), accumulated ROS induced senescence
in the nHDPCs, as assessed by SA-β-gal activity (Fig. 3B).
As DHT induced growth arrest, cell death, cell cycle
arrest, ROS production and senescence, comparative microarray
analysis of miRNAs was performed to identify the miRNA signatures
in the DHT-treated nHDPCs. Total RNA was extracted from the
untreated nHDPCs and nHDPCs treated with 1 mM DHT for 24 h. The
total RNA was labeled with Cy3 and hybridized to
microarray-containing probes for 1,205 annotated miRNAs. The
untreated cells were then compared with the 1 mM DHT-treated
nHDPCs, in which 55 miRNAs that were upregulated and 6 were
downregulated, by more than two-fold (Table I). Among the five miRNAs
significantly upregulated in the DHT-treated nHDPCs, the level of
miR-3663-3p increased by 219.04-fold, miR-485-3p by 200.81-fold,
miR-7 by 173.64-fold, miR-125a-3p by 154.55-fold, and miR-4271-by
108-fold. In addition, in the five miRNAs, which were significantly
downregulated in the DHT-treated nHDPCs, the level of miR-450a
decreased by 95.69-fold, miR-1181 by 93.76-fold, miR-3656 by
2.84-fold, miR-4286 by 2.29-fold and miR-370 by 2.24-fold.
 | Table ImiRNAs exhibiting a ≥2-fold change in
expression following treatment of the nHDPCs with DHT. |
Table I
miRNAs exhibiting a ≥2-fold change in
expression following treatment of the nHDPCs with DHT.
| miRNA | Fold change | Direction of
change | Chromosome |
|---|
| Has-let-7a* | 28.48 | Up | 9 |
| hsa-miR-1181 | −93.76 | Down | 19 |
|
hsa-miR-1207-5p | 5.23 | Up | 8 |
|
hsa-miR-1225-5p | 3.08 | Up | 16 |
| hsa-miR-1246 | 2.49 | Up | 2 |
| hsa-miR-1249 | 35.73 | Up | 22 |
|
hsa-miR-125a-3p | 154.55 | Up | 19 |
| hsa-miR-1268 | 2.38 | Up | 15 |
| hsa-miR-128 | 35.55 | Up | 2 |
| hsa-miR-1290 | 2.06 | Up | 1 |
| hsa-miR-132 | 40.88 | Up | 17 |
| hsa-miR-134 | 90.23 | Up | 14 |
| hsa-miR-135a* | 45.73 | Up | 3 |
| hsa-miR-138 2* | 52.92 | Up | 16 |
| hsa-miR-146a | −2.01 | Down | 5 |
| hsa-miR-148b | 50.96 | Up | 12 |
| hsa-miR-150* | 98.34 | Up | 19 |
| hsa-miR-1539 | 40.93 | Up | 18 |
| hsa-miR-154* | −2.24 | Down | 14 |
| hsa-miR-17* | 60.48 | Up | 13 |
| hsa-miR-1915 | 3.07 | Up | 10 |
| hsa-miR-197 | 84.06 | Up | 1 |
| hsa-miR-1973 | 2.20 | Up | 4 |
| hsa-miR-202 | 35.08 | Up | 10 |
| hsa-miR-28 5p | 36.51 | Up | 3 |
| hsa-miR-324-5p | 36.40 | Up | 17 |
|
hsa-miR-3613-3p | 78.20 | Up | 13 |
| hsa-miR-3646 | 50.21 | Up | 20 |
| hsa-miR-3651 | 2.98 | Up | 9 |
| hsa-miR-3656 | −2.84 | Down | 11 |
|
hsa-miR-3663-3p | 219.04 | Up | 10 |
| hsa-miR-369-3p | 36.36 | Up | 14 |
| hsa-miR-370 | 40.16 | Up | 14 |
| hsa-miR-371-5p | 78.83 | Up | 19 |
| hsa-miR-378 | 48.73 | Up | 5 |
| hsa-miR-409-5p | 44.22 | Up | 14 |
| hsa-miR-423-5p | 49.28 | Up | 17 |
| hsa-miR-4270 | 36.98 | Up | 3 |
| hsa-miR-4271 | 108.00 | Up | 3 |
| hsa-miR-4281 | 2.81 | Up | 5 |
| hsa-miR-4286 | −2.29 | Down | 8 |
| hsa-miR-4291 | 53.23 | Up | 9 |
| hsa-miR-4299 | 2.14 | Up | 11 |
| hsa-miR-431 | 35.70 | Up | 14 |
| hsa-miR-431* | 25.62 | Up | 14 |
| hsa-miR-4317 | 38.79 | Up | 18 |
| hsa-miR-4327 | 34.79 | Up | 21 |
| hsa-miR-450a | −95.69 | Down | X |
| hsa-miR-483-5p | 39.06 | Up | 11 |
| hsa-miR-485-3p | 200.81 | Up | 14 |
| hsa-miR-500a | 2.30 | Up | X |
|
hsa-miR-513a-5p | 2.68 | Up | X |
| hsa-miR-513b | 47.72 | Up | X |
| hsa-miR-550a | 18.95 | Up | 7 |
| hsa-miR-572 | 39.29 | Up | 4 |
| hsa-miR-630 | 3.53 | Up | 15 |
| hsa-miR-642b | 105.36 | Up | 19 |
| hsa-miR-7 | 173.64 | Up | 9 |
| hsa-miR-762 | 5.26 | Up | 16 |
| hsa-miR-770-5p | 56.99 | Up | 14 |
| hsa-miR-874 | 62.43 | Up | 5 |
Subsequently, the putative target genes of
DHT-regulated miRNAs were identified using the PITA, microRNAorg
and Targetscan target prediction systems (Table II). A total of 587 putative target
genes of the upregulated miRNAs and 140 putative target genes of
the downregulated miRNAs were identified in PITA. Using
microRNAorg, 488 putative target genes of upregulated miRNAs and
312 putative target genes of downregulated miRNAs were found, and
691 putative target genes of upregulated miRNAs and 219 putative
target genes of down regulated miRNAs were identified using
Targetscan. Of these, 339 were overlapping target genes of
upregulated miRNAs and 111 were overlapping target genes of
downregulated miRNAs in all three target prediction systems.
 | Table IINumber of significant miRNA targets
using three prediction databases. |
Table II
Number of significant miRNA targets
using three prediction databases.
| Database | Target miRNAs
(n) | Overlapping miRNAs
in all three databases (n) |
|---|
| Upregulated target
miRNAs | | 339 |
| Targetscan | 691 | |
| PITA | 587 | |
| microRNAorg | 488 | |
| Downregulated
target miRNAs | | 111 |
| Targetscan | 219 | |
| PITA | 140 | |
| microRNAorg | 312 | |
To investigate a association between the
aforementioned effects of DHT and the putative miRNA target genes,
GO analysis of each putative target gene was performed using DAVID.
The genes were classified according to GO terms associated with the
five effects of DHT and the number of putative target genes
associated with each GO term were counted. As shown in Table III, the putative target genes of
the uppregulated and downregulated miRNAs were associated with five
antioxidant-associated GO terms, 17 apoptosis and cell
death-associated terms, 11 proliferation and cell growth-associated
terms, 1 age associated term and 14 cell cycle-associated GO terms.
The miRNAs and their putative target genes are shown in Table IV. Overall, these results
demonstrated that DHT exerted negative effects, which were
associated with an alteration in cellular miRNA expression
profiles.
 | Table IIIGenes grouped according to the GO
terms, associated with the effects of 5α-dihydrotestosterone. |
Table III
Genes grouped according to the GO
terms, associated with the effects of 5α-dihydrotestosterone.
| A,
Antioxidant-associated genes |
|---|
|
|---|
| Accession No. | GO term | Upregulated
(n) | Downregulated
(n) |
|---|
| GO:0006733 | Oxidoreduction
coenzyme metabolic process | 3 | 0 |
| GO:0006979 | Response to
oxidative stress | 5 | 2 |
| GO:0042542 | Response to
hydrogen peroxide | 2 | 0 |
| GO:0015980 | Energy derivation
by oxidation of organic compounds | 0 | 3 |
| GO:0055114 | Oxidation
reduction | 6 | 6 |
| B, Apoptosis and
cell death-associated genes |
|---|
|
|---|
| Accession No. | GO term | Upregulated
(n) | Downregulated
(n) |
|---|
| GO:0006916 | Anti-apoptosis | 7 | 4 |
| GO:0008624 | Induction of
apoptosis by extracellular signals | 4 | 0 |
| GO:0042981 | Regulation of
apoptosis | 17 | 7 |
| GO:0043066 | Negative regulation
of apoptosis | 8 | 4 |
| GO:0043065 | Positive regulation
of apoptosis | 8 | 2 |
| GO:0006917 | Induction of
apoptosis | 6 | 0 |
| GO:0006915 | Apoptosis | 9 | 5 |
| GO:0043067 | Regulation of
programmed cell death | 18 | 4 |
| GO:0010941 | Regulation of cell
death | 18 | 4 |
| GO:0043069 | Negative regulation
of programmed cell death | 9 | 0 |
| GO:0060548 | Negative regulation
of cell death | 9 | 0 |
| GO:0043068 | Positive regulation
of programmed cell death | 8 | 2 |
| GO:0010942 | Positive regulation
of cell death | 8 | 2 |
| GO:0012502 | Induction of
programmed cell death | 6 | 0 |
| GO:0008219 | Cell death | 11 | 6 |
| GO:0016265 | Death | 11 | 6 |
| GO:0012501 | Programmed cell
death | 9 | 5 |
| C, Proliferation
and cell growth-associated genes |
|---|
|
|---|
| Accession No. | GO term | Upregulated
(n) | Downregulated
(n) |
|---|
| GO:0008283 | Cell
proliferation | 9 | 3 |
| GO:0008284 | Positive regulation
of cell proliferation | 8 | 5 |
| GO:0042127 | Regulation of cell
proliferation | 12 | 7 |
| GO:0008285 | Negative regulation
of cell proliferation | 4 | 3 |
| GO:0030308 | Negative regulation
of cell growth | 3 | 0 |
| GO:0040008 | Regulation of
growth | 7 | 4 |
| GO:0048638 | Regulation of
developmental growth | 2 | 0 |
| GO:0045926 | Negative regulation
of growth | 3 | 0 |
| GO:0001558 | Regulation of cell
growth | 4 | 2 |
| GO:0045927 | Positive regulation
of growth | | 2 |
| GO:0040007 | Growth | 3 | 0 |
| D, Aging-associated
genes |
|---|
|
|---|
| Accession No. | GO term | Upregulated
(n) | Downregulated
(n) |
|---|
| GO:0007568 | Aging | 3 | 0 |
| E, Cell
cycle-associated genes |
|---|
|
|---|
| Accession No. | GO term | Upregulated
(n) | Downregulated
(n) |
|---|
| GO:0051726 | Regulation of cell
cycle | 13 | 0 |
| GO:0045786 | Negative regulation
of cell cycle | 5 | 0 |
| GO:0051327 | M phase of meiotic
cell cycle | 4 | 0 |
| GO:0051321 | Meiotic cell
cycle | 4 | 2 |
| GO:0045930 | Negative regulation
of mitotic cell cycle | 2 | 0 |
| GO:0010948 | Negative regulation
of cell cycle process | 2 | 0 |
| GO:0007346 | Regulation of
mitotic cell cycle | 4 | 0 |
| GO:0022403 | Cell cycle
phase | 8 | 2 |
| GO:0010564 | Regulation of cell
cycle process | 3 | 0 |
| GO:0007049 | Cell cycle | 14 | 4 |
| GO:0022402 | Cell cycle
process | 9 | 4 |
| GO:0000278 | Mitotic cell
cycle | 6 | 0 |
| GO:0000075 | Cell cycle
checkpoint | 2 | 0 |
| GO:0000087 | M phase of mitotic
cell cycle | 3 | 0 |
 | Table IVTarget genes of significantly
regulated miRNAs in DHT-treated nHDPCs. |
Table IV
Target genes of significantly
regulated miRNAs in DHT-treated nHDPCs.
A, Targets of up
regulated miRNAs
|
|---|
| miRNA | Antioxidant | Apoptosis and cell
death | Proliferation and
cell growth | Aging | Cell cycle |
|---|
| a3663-3p | GAPDHS, NDUFA8,
GAPDH, DEGS2, DCXR | CARD9, ADA | FOXS1, ENO1 | – | – |
| a485-3p | APOA4, PRDX1,
NDUFAB1, NQO2 | GNRH1, PRDX1 | – | – | – |
| a7 | CYP11A1, UCP2, NQO2
NEIL1, BCKDHA, FADS3, ALKBH2 | DAPL1, CASP12,
DDX41, DAPK3, BCL2L12, CRYAA, CSTB, INHA | BMP10, LBX1, INHA,
IL34, CKLF, SLC3A2, ENO3, BDKRB1, OGFR, | – | RNF167, INHA,
CDC37, CRYAA |
| a125-3p | BCKDHA, NDUFS7,
FTMT, PLOD3, TH, COX6B1, HGD, AKR1C1 | PYCARD, LGALS12,
TGFB1, LRDD, GML, ADA | BDKRB1, SCGB3A1,
NPPA, TGFB1, ENO1, E4F1, FTMT, AGER, ADA, FGF6, PRG4, GML | AGER, ADA,
TGFB1 | TUBB2A, SPAG5,
PKMYT1, CDC20, TGFB1, E4F1, GML, CDK5RAP3 PARD6A, GPS2 |
| a4271 | BCKDHA, NDUFS7,
NDUFB11, NDUFB10, HAO2, NDUFS8, FADS3, FDX1L, ALOX12B, IL4I1,
NSDHL | GZMM, DAPL1, LRDD,
ATP2A1, MGC29506 | SSTR4, PRTN3, GHRH,
ILK, PYY, PRSS2, BARHL2, OGFR, ENO1 | – | BGLAP, PKMYT1,
ILK |
B, Targets of
down-regulated miRNAs
|
|---|
| miRNA | Antioxidant | Apoptosis and cell
death | Proliferation and
cell growth | Aging | Cell cycle |
|---|
| a450a | UQCRH, ALKBH2 | – | – | – | – |
| a1181 | – | – | – | – | – |
| a3656 | – | CARD9, INS,
TMEM102, SFN, ATP2A1 | INS, SFN, SCGB3A1,
VGF | – | INS, SFN |
| a428 | NDUFB11, NMRAL1,
FDX1L | AARS, MUC5AC,
DAPK3, CDK5, TGFB1, PROC, MIF, LRDD, TBRG4 | – | FANCG, CDK5,
SERTAD1, TGFB1, TBRG4, PARD6A | – |
| a154 | – | IFIH1, CASP12, PF4,
PRDX1 | VTI1B, PRDX1,
RARRES3, GNL3 | – | – |
Discussion
The results of the present study provided evidence
that DHT-induced growth arrest, cell death, cell cycle arrest, ROS
production and senescence in nHDPCs. In the hair follicle, DHT is
produced by 5α-reductase and it accumulates, which induces
androgenetic alopecia through DHT-mediated cell death and decreased
growth rate (27–29). As shown in Figs. 1 and 2, 1 mM
DHT repressed cell growth by inducing cell cycle arrest and cell
death. In a previous report, activation of the androgen receptor
provoked ROS-mediated senescence (30,31).
As shown in Fig. 3, measurement of
ROS revealed that 1 mM DHT significantly elevated the levels of ROS
in the nHDPCs. In the nHDPCs, which exhibited increased activity of
the androgen receptor by DHT, 1 mM DHT significantly increased the
percentage of senescent cells (Fig.
3B). Specifically, an association was observed between the
effects of DHT and the regulation of miRNAs by DHT. Using miRNA
microarray analysis, 61 miRNAs (55 upregulated and 6 downregulated)
were identified, in which the miRNA levels were increased of
decreased by more than two-fold by DHT in the nHDPCs (Table I). One of these, miRNA-125a-3p has
been demonstrated as a repressor of cell proliferation and
migration through targeting Fyn (32). In addition, miR-485-5p (39.06-fold
increase) inhibits cell growth and migration in breast cancer cell
lines (33), whereas miRNA-7
regulates the mammalian target of rapamycin and phosphoinositide
3-kinase/Akt pathways, and targets Bcl-2, X-linked inhibitor of
apoptosis protein and ETS2 repressor factor, which affect cell
growth and the repression of intrinsic apoptosis (34–39).
Furthermore, the present study predicted the target genes of
DHT-regulated miRNAs and performed GO analysis of potential target
genes using the DAVID bioinformatics resources. A correlation was
found between DHT-induced alterations in miRNA expression profiles
and DHT-induced cellular effects, by grouping the target genes,
according to GO terms, with five biological processes, which
impacted in DHT-treated cells (Tables
II and III). The results
revealed that the DHT-induced alteration of the miRNA profile was
associated with the aforementioned cellular effects of DHT, of
induced cell growth, cell cycle arrest, cell death, ROS induction
and senescence.
In conclusion, the present study demonstrated that
DHT-induced growth arrest, cell death, cell cycle arrest, ROS
production and senescence by upregulating and downregulating the
expression of DHT-specific miRNAs in nHDPCs. These findings support
the hypothesis that miRNA regulation is involved in DHT-induced
androgenetic alopecia.
Acknowledgments
This study was supported by Konkuk University in
2013.
References
|
1
|
Yazdan P: Update on the genetics of
androgenetic alopecia, female pattern hair loss, and alopecia
areata: Implications for molecular diagnostic testing. Semin Cutan
Med Surg. 31:258–266. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Alsantali A and Shapiro J: Androgens and
hair loss. Curr Opin Endocrinol Diabetes Obes. 16:246–253. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Canguven O and Burnett AL: The effect of 5
alpha-reductase inhibitors on erectile function. J Androl.
29:514–523. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Rove KO, Debruyne FM, Djavan B, Gomella
LG, Koul HK, Lucia MS, Petrylak DP, Shore ND, Stone NN and Crawford
ED: Role of testosterone in managing advanced prostate cancer.
Urology. 80:754–762. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Hillier SG and Tetsuka M: Role of
androgens in follicle maturation and atresia. Baillieres Clin
Obstet Gynaecol. 11:249–260. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
McElwee KJ, Kissling S, Wenzel E, Huth A
and Hoffmann R: Cultured peribulbar dermal sheath cells can induce
hair follicle development and contribute to the dermal sheath and
dermal papilla. J Invest Dermatol. 121:1267–1275. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Yang CC and Cotsarelis G: Review of hair
follicle dermal cells. J Dermatol Sci. 57:2–11. 2010. View Article : Google Scholar :
|
|
8
|
Tang L, Bernardo O, Bolduc C, Lui H,
Madani S and Shapiro J: The expression of insulin-like growth
factor 1 in follicular dermal papillae correlates with therapeutic
efficacy of finasteride in androgenetic alopecia. J Am Acad
Dermatol. 49:229–233. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Stenn KS, Combates NJ, Eilertsen KJ,
Gordon JS, Pardinas JR, Parimoo S and Prouty SM: Hair follicle
growth controls. Dermatol Clin. 14:543–558. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Peus D and Pittelkow MR: Growth factors in
hair organ development and the hair growth cycle. Dermatol Clin.
14:559–572. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Stenn KS and Paus R: Controls of hair
follicle cycling. Physiol Rev. 81:449–494. 2001.PubMed/NCBI
|
|
12
|
Ferraris C, Cooklis M, Polakowska RR and
Haake AR: Induction of apoptosis through the PKC pathway in
cultured dermal papilla fibroblasts. Exp Cell Res. 234:37–46. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kwack MH, Sung YK, Chung EJ, Im SU, Ahn
JS, Kim MK and Kim JC: Dihydrotestosterone-inducible dickkopf 1
from balding dermal papilla cells causes apoptosis in follicular
keratinocytes. J Invest Dermatol. 128:262–269. 2008.
|
|
14
|
Winiarska A, Mandt N, Kamp H, Hossini A,
Seltmann H, Zouboulis CC and Blume-Peytavi U: Effect of
5alpha-dihydrotestosterone and testosterone on apoptosis in human
dermal papilla cells. Skin Pharmacol Physiol. 19:311–321. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ambros V, Bartel B, Bartel DP, Burge CB,
Carrington JC, Chen X, Dreyfuss G, Eddy SR, Griffiths-Jones S,
Marshall M, et al: A uniform system for microRNA annotation. RNA.
9:277–279. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Valencia-Sanchez MA, Liu J, Hannon GJ and
Parker R: Control of translation and mRNA degradation by miRNAs and
siRNAs. Genes Dev. 20:515–524. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ha TY: MicroRNAs in Human Diseases: From
Cancer to Cardiovascular Disease. Immune Netw. 11:135–154. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Andl T, Murchison EP, Liu F, Zhang Y,
Yunta-Gonzalez M, Tobias JW, Andl CD, Seykora JT, Hannon GJ and
Millar SE: The miRNA-processing enzyme dicer is essential for the
morphogenesis and maintenance of hair follicles. Curr Biol.
16:1041–1049. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Bae S, Lee EJ, Lee JH, Park IC, Lee SJ,
Hahn HJ, Ahn KJ, An S, An IS and Cha HJ: Oridonin protects HaCaT
keratinocytes against hydrogen peroxide-induced oxidative stress by
altering microRNA expression. Int J Mol Med. 33:185–193. 2014.
|
|
20
|
Kim YJ, Cha HJ, Nam KH, Yoon Y, Lee H and
An S: Centella asiatica extracts modulate hydrogen peroxide-induced
senescence in human dermal fibroblasts. Exp Dermatol. 20:998–1003.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Simões VL, Alves MG, Martins AD, Dias TR,
Rato L, Socorro S and Oliveira PF: Regulation of apoptotic
signaling pathways by 5α-dihydrotestosterone and 17β-estradiol in
immature rat Sertoli cells. J Steroid Biochem Mol Biol. 135:15–23.
2013. View Article : Google Scholar
|
|
22
|
Mirochnik Y, Veliceasa D, Williams L,
Maxwell K, Yemelyanov A, Budunova I and Volpert OV: Androgen
receptor drives cellular senescence. PLoS One. 7:e310522012.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Mehraein-Ghomi F, Lee E, Church DR,
Thompson TA, Basu HS and Wilding G: JunD mediates androgen induced
oxidative stress in androgen-dependent LNCaP human prostate cancer
cells. Prostate. 68:924–934. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ruizeveld de Winter JA, Trapman J, Vermey
M, Mulder E, Zegers ND and van der Kwast TH: Androgen receptor
expression in human tissues: An immunohistochemical study. J
Histochem Cytochem. 39:927–936. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Takahashi A, Ohtani N, Yamakoshi K, Iida
S, Tahara H, Nakayama K, Nakayama KI, Ide T, Saya H and Hara E:
Mitogenic signalling and the p16INK4a-Rb pathway cooperate to
enforce irreversible cellular senescence. Nat Cell Biol.
8:1291–1297. 2006. View
Article : Google Scholar : PubMed/NCBI
|
|
26
|
Hodgins MB, Choudhry R, Parker G, Oliver
RF, Jahoda CA, Withers AP, Brinkmann AO, van der Kwast TH, Boersma
WJ, Lammers KM, et al: Androgen receptors in dermal papilla cells
of scalp hair follicles in male pattern baldness. Ann NY Acad Sci.
642:448–451. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Eicheler W, Happle R and Hoffmann R: 5
alpha-reductase activity in the human hair follicle concentrates in
the dermal papilla. Arch Dermatol Res. 290:126–132. 1998.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Trüeb RM: Molecular mechanisms of
androgenetic alopecia. Exp Gerontol. 37:981–990. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Inui S and Itami S: Molecular basis of
androgenetic alopecia: From androgen to paracrine mediators through
dermal papilla. J Dermatol Sci. 61:1–6. 2011. View Article : Google Scholar
|
|
30
|
Mirochnik Y, Veliceasa D, Williams L,
Maxwell K, Yemelyanov A, Budunova I and Volpert OV: Androgen
receptor drives cellular senescence. PLoS One. 7:e310522012.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Colavitti R and Finkel T: Reactive oxygen
species as mediators of cellular senescence. IUBMB Life.
57:277–281. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Ninio-Many L, Grossman H, Shomron N,
Chuderland D and Shalgi R: microRNA-125a-3p reduces cell
proliferation and migration by targeting Fyn. J Cell Sci.
126:2867–2876. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Anaya-Ruiz M, Bandala C and Perez-Santos
JL: miR-485 acts as a tumor suppressor by inhibiting cell growth
and migration in breast carcinoma T47D cells. Asian Pac J Cancer
Prev. 14:3757–3760. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang Y, Liu J, Liu C, Naji A and Stoffers
DA: MicroRNA-7 regulates the mTOR pathway and proliferation in
adult pancreatic β-cells. Diabetes. 62:887–895. 2013. View Article : Google Scholar :
|
|
35
|
Fang Y, Xue JL, Shen Q, Chen J and Tian L:
MicroRNA-7 inhibits tumor growth and metastasis by targeting the
phosphoinositide 3-kinase/Akt pathway in hepatocellular carcinoma.
Hepatology. 55:1852–1862. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Xiong S, Zheng Y, Jiang P, Liu R, Liu X
and Chu Y: MicroRNA-7 inhibits the growth of human non-small cell
lung cancer A549 cells through targeting BCL-2. Int J Biol Sci.
7:805–814. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Chou YT, Lin HH, Lien YC, Wang YH, Hong
CF, Kao YR, Lin SC, Chang YC, Lin SY, Chen SJ, et al: EGFR promotes
lung tumorigenesis by activating miR-7 through a Ras/ERK/Myc
pathway that targets the Ets2 transcriptional repressor ERF. Cancer
Res. 70:8822–8831. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Jiang L, Liu X, Chen Z, Jin Y, Heidbreder
CE, Kolokythas A, Wang A, Dai Y and Zhou X: MicroRNA-7 targets
IGF1R (insulin-like growth factor 1 receptor) in tongue squamous
cell carcinoma cells. Biochem J. 432:199–205. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Liu S, Zhang P, Chen Z, Liu M, Li X and
Tang H: MicroRNA-7 downregulates XIAP expression to suppress cell
growth and promote apoptosis in cervical cancer cells. FEBS Lett.
587:2247–2253. 2013. View Article : Google Scholar : PubMed/NCBI
|