Introduction
Lung cancer is the leading cause of
cancer-associated mortality worldwide (1). In the United States, it was estimated
that there would be 221,200 new cases and 158,040 mortalities
caused by lung cancer in 2015 (2).
The occurrence and progression of lung cancer is complex and
associated with various signaling pathways. Its incidence and
mortality are increasing each year as a result of environmental
pollution, particularly in China (3). Lung cancer can be divided into two
clinically relevant subtypes: Small cell lung cancer (SCLC) and
non-small cell lung cancer (NSCLC) (4). NSCLC, as the predominant form of lung
cancer, accounts for 80–85% of all cases of lung cancer (5). Large cell carcinoma, adenocarcinoma,
squamous carcinoma and adenosquamous carcinoma are types of NSCLC
(6). Patients with NSCLC are often
diagnosed in the late stages when it is locally advanced or has
metastasized, rendering it one of the most lethal forms of cancer
(7). Although there has been
progress regarding early detection and treatment by radical
surgical resection combined with chemotherapy and radiation
therapy, the prognosis of patients with NSCLC remains poor and the
5-year survival rate is ≤15% (8,9).
Thus, it is important to develop novel strategies and therapeutic
targets for the treatment of NSCLC.
Increasing evidence has indicated that microRNAs
(miRNAs) are deregulated in lung cancer (10–12).
miRNAs are a type of endogenous non-protein-coding short RNA (19–22
nucleotides in length) that are widely expressed in eukaryotes
(13). miRNAs are critical for the
regulation of various diverse physiological and pathological
processes, including proliferation, cell death, cell cycle,
migration, invasion, development and differentiation. miRNAs
function via complementary base pairing with target mRNAs in the 3′
untranslated region (3′ UTR), leading to cleavage or translation
repression of the mRNA (14–16).
Previous studies have demonstrated that more than half of miRNAs
are located in cancer-associated genomic regions, suggesting that
aberrant expression of miRNAs may be important during tumorigenesis
and progression (17). miRNAs are
stable molecules and may be useful for cancer diagnosis, treatment
and predicting prognosis (18,19).
miRNAs have previously been demonstrated to function as oncogenes
or tumor suppressors, and are aberrantly expressed in various types
of human malignancy (20). Thus,
investigating miRNAs may be critical to elucidate the prognostic
value and therapeutic potential of miRNAs in lung cancer.
miRNA (miR)-320 has previously been reported to be
frequently downregulated in multiple types of cancer. However, to
the best of our knowledge, the expression, biological functions and
molecular mechanisms of miR-320 in NSCLC have not been
investigated. Thus, the aim of the present study was to elucidate
the expression, biological functions and molecular mechanisms of
miR-320 in NSCLC.
Materials and methods
Clinical specimens
The procedures were approved by the Ethic Committee
on Human Experimentation of The First Affiliated Hospital of Dalian
Medical University (Dalian, China) and written informed consent was
also obtained from each patient. Samples of primary cancer tissue
(n=81) and paired normal adjacent tissue (NAT) were obtained from
patients who had undergone surgery at The First Affiliated Hospital
of Dalian Medical University. None of the patients had been treated
with radiotherapy or chemotherapy prior to surgery. All samples
were rapidly placed in liquid nitrogen and stored at −80°C until
use. The clinical data of NSCLC patients are presented in Table I.
 | Table IA comparison of microRNA-320
expression in non-small cell lung cancer and clinicopathological
features. |
Table I
A comparison of microRNA-320
expression in non-small cell lung cancer and clinicopathological
features.
| Clinical
features | Cases, n | miR-320 expression
| P-value |
|---|
| Low | High |
|---|
| Gender | | | | 0.812 |
| Male | 38 | 30 | 8 | |
| Female | 43 | 33 | 10 | |
| Age, years | | | | 0.544 |
| <60 | 49 | 37 | 12 | |
| ≥60 | 32 | 26 | 6 | |
| Smoking history,
years | | | | 0.434 |
| <10 | 34 | 25 | 9 | |
| ≥10 | 47 | 38 | 9 | |
| Tumor
differentiation, grade | | | | 0.591 |
| I–II | 45 | 34 | 11 | |
| III–IV | 36 | 29 | 7 | |
| TNM classification,
stage | | | | 0.026 |
| I | 22 | 13 | 9 | |
| II | 31 | 28 | 3 | |
| III+IV | 28 | 22 | 6 | |
| Metastasis | | | | 0.011 |
| No | 33 | 21 | 12 | |
| Yes | 48 | 42 | 6 | |
Cell culture and transfection
NSCLC cell lines H23, H522 and non-tumorigenic
bronchial epithelium BEAS-2B cells were obtained from American Type
Culture Collection (Manassas, VA, USA). H23 and H522 cells were
cultured in RPMI-1640 medium (Gibco; Thermo Fisher Scientific,
Inc., Waltham, MA, USA) supplemented with 10% fetal bovine serum
(FBS), 100 U/ml penicillin and 100 mg/ml streptomycin (Gibco;
Thermo Fisher Scientific, Inc.) in a cell incubator at 37°C with an
atmosphere of 5% CO2. BEAS-2B cells were cultured in
LHC-9 medium (Gibco; Thermo Fisher Scientific, Inc.) containing 10%
FBS.
Mature miR-320 mimics
(5′-AGCGGGAGAGUUGGGUCGAAAA-3′), miRNA mimics negative control (NC;
5′-UUCUCCGAACGUGUCACGUTT-3′). and the luciferase reporter plasmid
were obtained from Shanghai GenePharma Co., Ltd. (Shanghai, China).
H23 and H522 were transfected with miR-320 mimics, NC or
co-tranfected with luciferase reporter plasmid using Lipofectamine
2000 (Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's protocol.
RNA isolation and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
Total RNA was extracted from the tumor tissues, NATs
and cells using TRIzol reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) following the manufacturer's protocol. Then,
PrimeScript reverse transcription-PCR kit (Takara Biotechnology
Co., Ltd., Dalian, China) was used to perform RT to obtain cDNA.
For analysis of miR-320 expression levels, a SYBR PrimeScript miRNA
RT-PCR kit (Takara Biotechnology Co., Ltd.) was used with the ABI
7500 Real-Time PCR detection system (Applied Biosystems; Thermo
Fisher Scientific, Inc.), with U6 serving as the internal control.
Each sample was analyzed in triplicate. Primers were synthesized by
Guangzhou RiboBio Co., Ltd. (Guangzhou, China). Data were analyzed
using the ΔΔCq method (21)
Cell proliferation assay
The effect of miR-320 on NSCLC cell proliferation
was analyzed using a
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)
assay. Transfected cells (miR-320 or NC) were seeded in 96-well
plates at a density of 3,000 cells/well in 100 µl medium.
MTT assay was performed every 24 h for 120 h following
transfection. Briefly, 20 µl MTT (5 mg/ml; Sigma-Aldrich,
St. Louis, MO, USA) was added into each well and cells were
incubated for 4 h at 37°C. Subsequently, the formazan was dissolved
in 200 µl dimethyl sulfoxide. The absorbance at 490 nm was
measured using a microplate reader (ELx800; Bio-Tek Instruments,
Inc., Winooski, VT, USA). For each treatment group wells were
assessed in triplicate.
Cell migration and invasion assay
The migration and invasion ability of NSCLC cells
transfected with miR-320 mimics and NC was analyzed using Transwell
chambers with an 8-µm pore polycarbonate membrane (EMD
Millipore, Billerica, MA, USA). For cell migration assays,
2×104 transfected cells in 100 µl serum-free
RPMI-1640 medium were placed into the upper chambers. A volume of
500 µl RPMI-1640 medium supplemented with 20% FBS was added
into the lower chambers. The cell invasion assays were performed
according to the same procedure, although the Transwell chambers
were pre-coated with Matrigel (BD Biosciences, San Jose, CA).
Subsequent to incubation (12 h for migration assay and 24 h for
invasion assay), cells that did not migrate or invade through the
pores were carefully wiped away with cotton wool. Subsequently, the
inserts were fixed with 100% methanol for 10 min, and stained with
0.5% crystal violet (Beyotime Institute of Biotechnology, Haimen,
China) and imaged with an inverted microscope (IX71; Olympus
Corporation, Tokyo, Japan) at x200 magnification.
Bioinformatics analysis
The target gene information of miR-320 was analyzed
using Targetscan (www.targetscan.org/).
Western blotting
To determine the expression level of fatty acid
synthase (FAS) at the protein level, western blot analysis was
performed. Protein samples of the transfected cell lines were
harvested using radioimmunoprecipitation assay lysis buffer
(Beyotime Institute of Biotechnology) containing protease
inhibitors and phosphatase inhibitors (Roche Diagnostics, Basel,
Switzerland). Protein concentrations (30 µg) were measured
by a bicinchoninic acid assay kit (Nanjing KeyGen Biotech Co.,
Ltd., Nanjing, China). Briefly, equal quantities of protein lysates
were separated by 10% SDS-polyacrylamide gel electrophoresis for 20
min at 70 V and transferred to polyvinylidene fluoride membrane
(EMD Millipore). After blocking with 5% non-fat milk Tris-buffered
saline solution containing 0.1% Tween 20, and incubated with
primary antibodies overnight at 4°C.
Primary antibodies in the current study included
mouse anti-FAS monoclonal antibody (1:1,000 dilution; cat no.
ab106062; Abcam, Cambridge, MA, USA) and mouse anti-GAPDH
monoclonal antibody (1:1,000 dilution; cat no. ab125247; Abcam).
The membranes were then incubated with goat anti-mouse IgG
horseradish peroxidase-conjugated secondary antibody (1:5,000
dilution; cat. no. ab6789; Abcam) was applied to the membranes for
1 h at room temperature. The protein blots were visualized with ECL
reagents (Pierce; Thermo Fisher Scientific, Inc.), imaged using a
FluorChem imaging system (ProteinSimple, San Jose, CA, USA) and
normalized to GADPH. Protein bands were normalized to GAPDH and
analyzed using the AlphaEase FC2 software (Alpha Innotech, San
Leandro, CA, USA).
Luciferase assay
The luciferase assays were performed to determine
whether FAS was a direct target of miR-320. NSCLC cells were plated
in a 24-well plate at 40–50% confluency and were transfected with
the luciferase reporter plasmid (synthesized by GenePharma Co.,
Ltd.), miR-320 mimics or NC in a 12-well plate using Lipofectamine
2000 according to the manufacturer's protocol. After 48 h, the
activities of firefly and Renilla luciferases were measured
with the Dual-Lucif-erase Reporter Assay system (Promega
Corporation, Madison, WI, USA). The firefly luciferase activity was
normalized to the Renilla luciferase activity. The wells
were assessed in triplicate for each treatment group.
Statistical analysis
Data are presented as the mean ± standard deviation,
and compared using SPSS software (version 11.0; SPSS, Inc.,
Chicago, IL, USA). One-way analysis of variance followed by a least
significant differences test was conducted to analyze the data.
P<0.05 was considered to indicate a statistically significant
difference.
Results
miR-320 expression in human NSCLC tissues
and cell lines
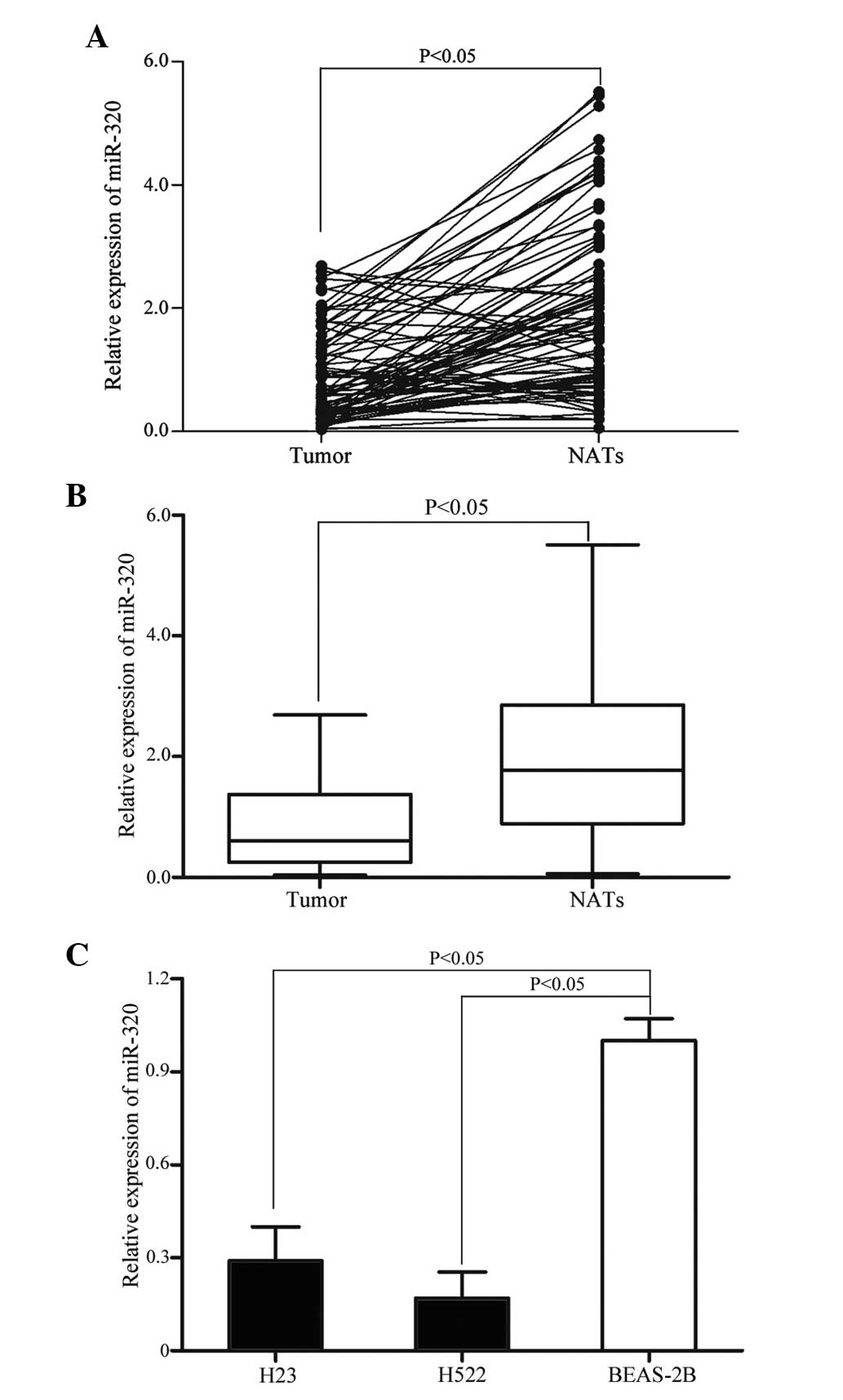
To examine the function of miR-320 in NSCLC, miR-320
expression was analyzed by RT-qPCR. As demonstrated in Fig. 1A and B, the level of miR-320 was
significantly down-regulated in NSCLC tissue samples when compared
with matched NATs (P=0.002). The expression level of miR-320 in
NSCLC cell lines and non-tumorigenic bronchial epithelium BEAS-2B
cells was also detected. As demonstrated in Fig. 1C, miR-320 expression level was also
significantly decreased in H23 and H522 cells compared with BEAS-2B
cells (P=0.001). These results indicate that miR-320 may be
important during lung carcinogenesis.
Association between miR-320 expression
level and clinicopathological features of patients with NSCLC
Statistical analysis was performed to determine
whether the expression level of miR-320 was associated with the
clinicopathological features of patients with NSCLC. The results
demonstrated that the expression level of miR-320 was significantly
associated with the TNM classification (P=0.028) and metastasis
(P=0.016). However, no correlation was identified between the
miR-320 expression level and the other clinicopathological features
(gender, age, smoking history and tumor differentiation).
miR-320 suppresses cell proliferation in
H23 and H522 cells
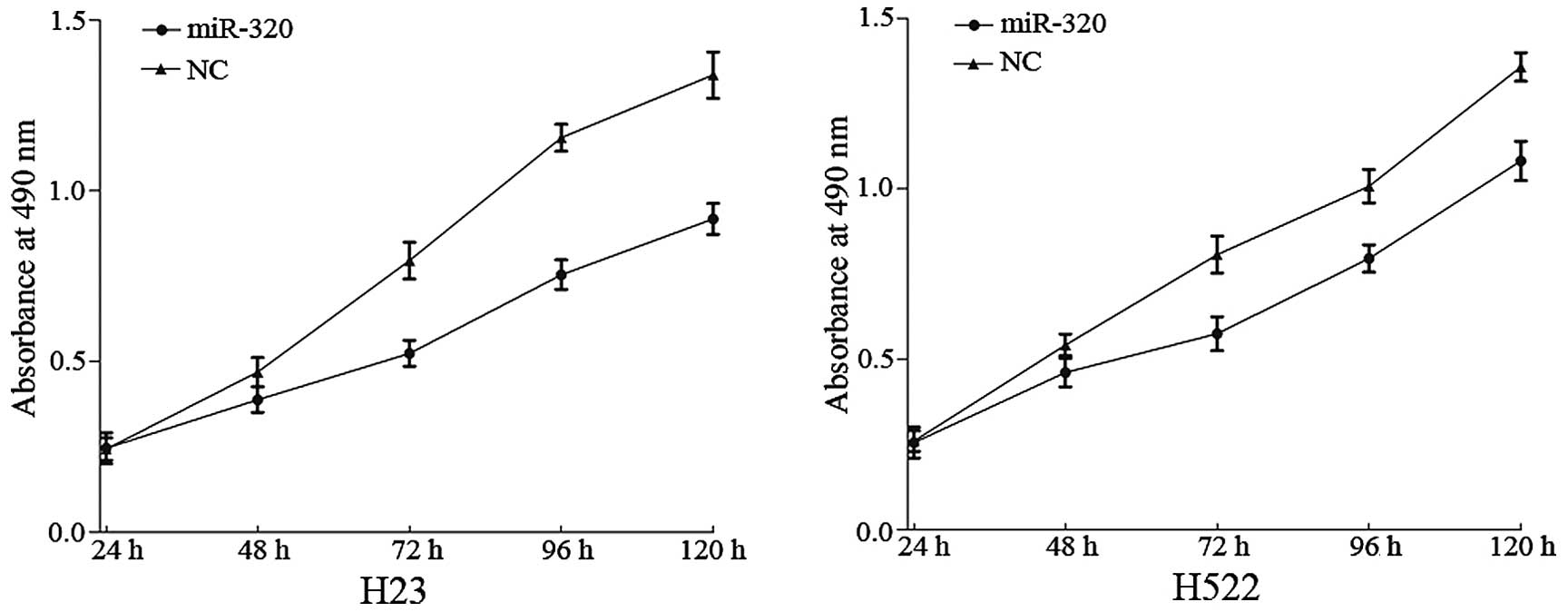
The effect of miR-320 on NSCLC cell proliferation
was measured by MTT assay. It was demonstrated that the absorbance
in H23 (P=0.008) and H522 (P=0.019) cells transfected with miR-320
was significantly decreased when compared with the NC group
(P<0.05; Fig. 2). This verified
that miR-320 inhibits the proliferation of H23 and H522 cells.
miR-320 inhibits cell migration and
invasion in H23 and H522 cells
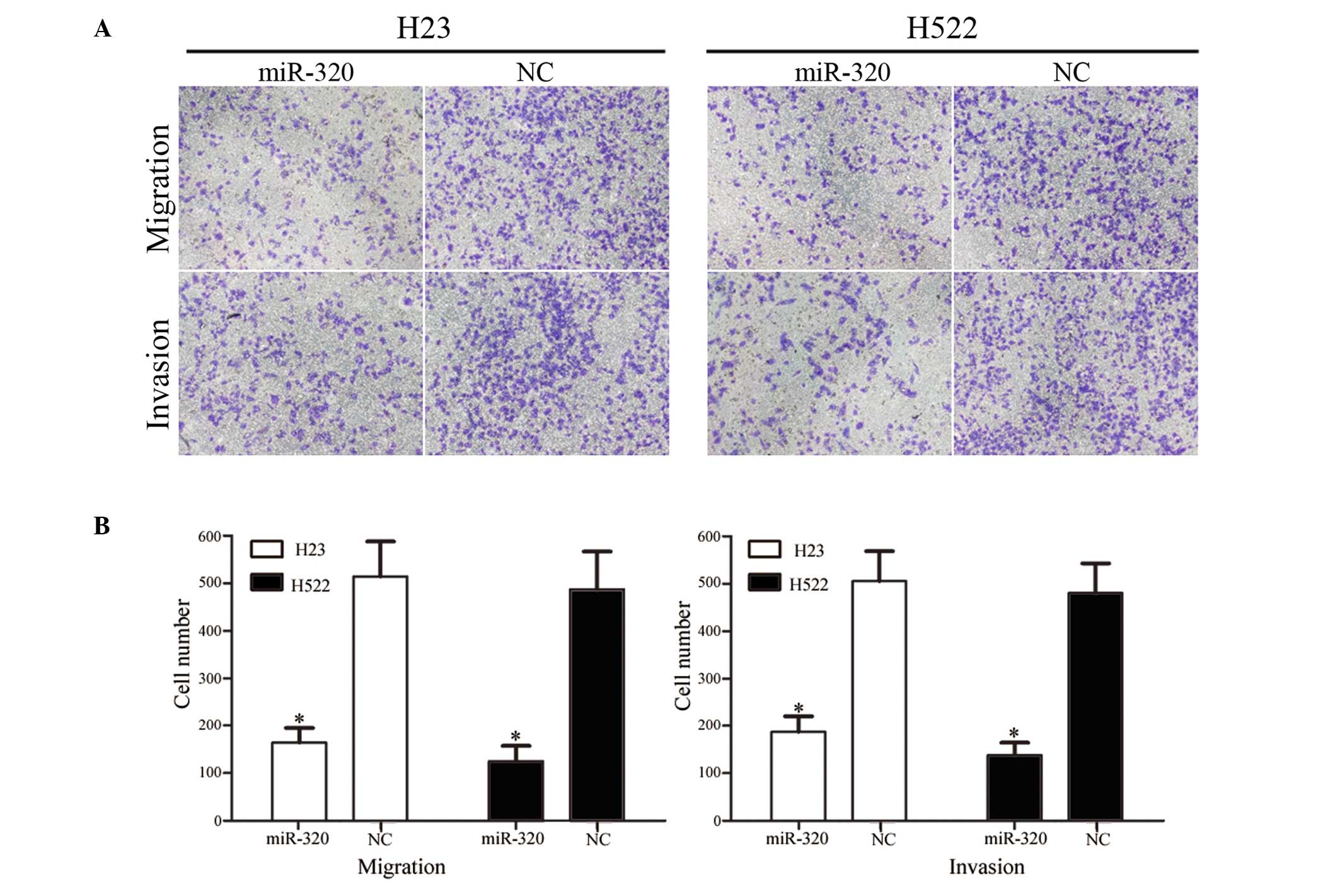
The effect of miR-320 on NSCLC cell migration and
invasion was measured using Transwell apparatus. As demonstrated in
Fig. 3, overexpression of miR-320
significantly reduced the migration and invasion ability of H23
(P=0.021 for migration, 0.025 for invasion) and H522 (P=0.015 for
migration; P=0.020 for invasion) cells compared with the NC group
(P<0.05). These results indicate that miR-320 may be involved in
reducing the migration and invasion potential of NSCLC cells in
vitro.
miR-320 directly targets FAS
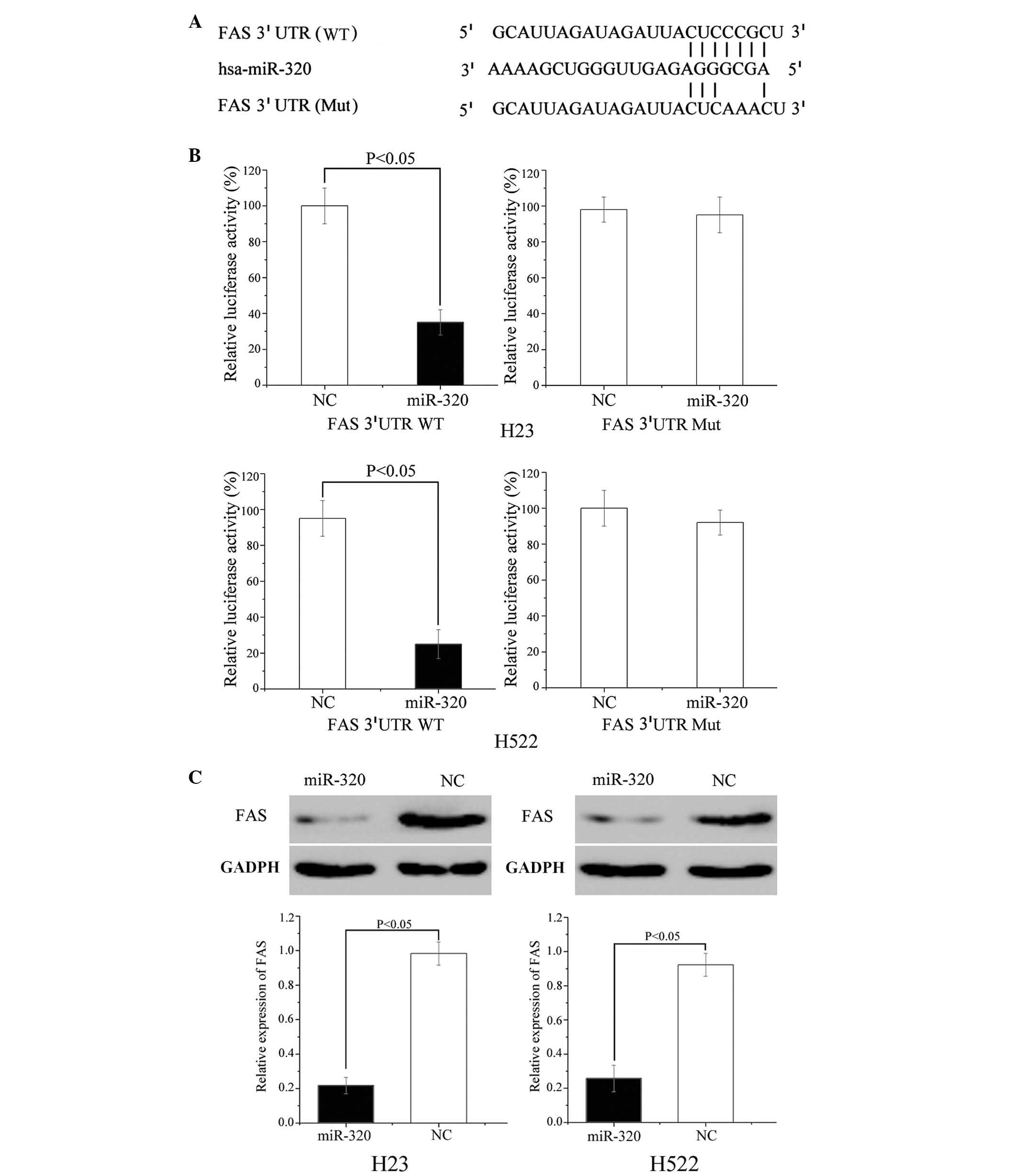
FAS protein is important for cancer proliferation
and metastasis, and was predicted to be a direct target gene of
miR-320 (Fig. 4A); luciferase
assays were performed to verify this. As demonstrated in Fig. 4B, miR-320 mimics significantly
inhibited the luciferase activity of the wild type but not the
mutant FAS 3′ UTR constructs in H23 (P=0.034) and H522 cells
(P=0.018).
Furthermore, western blot analysis was performed to
determine the protein expression level of FAS in H23 and H522 cells
following transfection with miR-320. The results of the current
study demonstrate that the expression level of FAS was
significantly inhibited in miR-320 mimic-transfected H23 (P=0.021)
and H522 (P=0.028) cells compared with cells transfected with NC
(P<0.05; Fig. 4C). These
results demonstrate that FAS is a direct target gene of miR-320
in vitro.
Discussion
Increasing evidence indicates that the aberrant
expression of miRNAs is a characteristic of multiple types of
malignancy, including lung cancer (13,22).
The expression of miR-320 has been observed to be downregulated in
various types of human cancer, including breast cancer, oral
squamous cell carcinoma (OSCC), colon cancer, acute myelogenous
leukemia, osteosarcoma and glioma (23–28).
However, to the best of our knowledge, no previous studies have
investigated the expression of miR-320 in lung cancer. The present
study demonstrated that miR-320 was significantly downregulated in
NSCLC tissue samples and cell lines. Furthermore, the expression of
miR-320 was significantly associated with the TNM classification
and presence of metastasis. These results indicate that miR-320 may
be important in NSCLC.
miR-320 has previously been demonstrated to be a
tumor suppressor during tumorigenesis and progression in multiple
types of cancer. In colon cancer, upregulation of miR-320 was
demonstrated to inhibit cell growth, cell cycle, migration and
invasion, whereas downregulation of miR-320 had the opposite effect
on these biological processes (24). Additionally, it was demonstrated
that overexpression of miR-320 enhanced the sensitivity of human
colon cancer cells to fluorouracil and oxaliplatin by targeting
forkhead box M1 in vitro (24). In OSCC, miR-320 was demonstrated to
suppress tumor angiogenesis via downregulation of neuropilin 1
(26). Cheng et al
(27) also reported that miR-320
inhibits osteosarcoma cell proliferation by targeting FAS. In human
glioma, miR-320 decreased cell growth by directly targeting E2F
transcription factor 1 (28).
However, to the best of our knowledge, no previous studies have
investigated the function of miR-320 in lung cancer. The present
study demonstrated that miR-320 inhibits NSCLC cell proliferation,
migration and invasion. The current study increased the knowledge
of the functions of miR-320 in cancer. These findings indicate that
miR-320 performs an essential function in these forms of cancer,
and may have the potential to be developed as an anticancer
therapeutic agent.
Identification of miR-320 target genes is essential
for understanding its mechanism in lung carcinogenesis and for the
development of novel targeted therapies. The current study
successfully demonstrated that FAS is a direct target gene of
miR-320. Bioinformatics software (TargetScan) predicted that FAS
was a direct target mRNA of miR-320 and luciferase assays
demonstrated that miR-320 directly interacts with FAS 3′ UTR.
Furthermore, manipulation of miR-320 expression in NSCLC cell lines
decreased the expression of FAS at the protein level. These
findings suggest that miR-320 acts as a tumor suppressor in NSCLC
by targeting FAS.
FAS, a multifunctional enzymatic complex, catalyzes
the formation of palmitate from acetyl-coenzyme A and
malonylcoenzyme A (29). The
endogenous FAS is expressed at very low levels in normal human
tissues (30). However, increasing
studies have demonstrated that the expression level of FAS is
highly upregulated and involved in cancer proliferation and
metastasis in various types of human tumor, including breast,
colorectal, prostate, bladder, ovarian, esophageal, gastric and
lung cancer, oral carcinoma, thyroid cancer and endometrial
carcinoma, and also in mesothelioma, nephroblastoma,
retinoblastoma, soft tissue sarcomas, melanoma and hepatocellular
carcinoma (31–33). FAS was also previously reported to
be correlated with various clinicopathological features of cancer.
For example, overexpression of FAS in NSCLC was reported to be
significantly associated with bone metastasis (34). These studies suggested that
targeting FAS may present as a therapeutic approach.
FAS has previously been demonstrated to be regulated
by multiple miRNAs in multiple types of cancer, including lung
cancer. miR-601, miR-10b and miR-663 are involved in the biology of
lung cancer via direct or indirect regulation of FAS (35–37).
In osteosarcoma, miR-142-3p and miR-320 have been reported to act
as tumor suppressors by regulation of FAS (27,38).
Previous studies demonstrated that miR-424 and miR-195 regulate FAS
to inhibit osteosarcoma cell migration and invasion (39,40).
In colorectal and breast cancer, miR-196b and miR-21 modulate cell
apoptosis by repressing FAS expression (41,42).
In gastric cancer, altered miR-106a expression levels exert
oncogenic effects in gastric carcinogenesis by targeting FAS
(43). In prostate cancer, miR-185
and miR-342 have been demonstrated to decrease cell proliferation,
migration and invasion by inhibiting FAS indirectly (44). In hepatocellular carcinoma,
restoration of miR-449 suppresses cell proliferation by
downregulating FAS (45). In the
present study, overexpression of miR-320 in NSCLC cell lines
demonstrated that miR-320 inhibits cell proliferation, migration
and invasion via downregulation of FAS. Taken together, these
results indicate that miR-320 may act as a tumor suppressor by
inhibiting the oncogenic activity of FAS.
In conclusion, this was the first study, to the best
of our knowledge, to demonstrate that miR-320 is downregulated in
NSCLC, and is significantly associated with the TNM classification
and metastasis. It was also demonstrated that miR-320 contributes
to cell proliferation, migration and invasion by directly targeting
FAS in NSCLC. The identification of FAS as a candidate target gene
of miR-320 may provide an understanding of the potential
carcinogenic mechanisms in NSCLC. These findings have therapeutic
implications and may be exploited for developing treatment
strategies for NSCLC.
References
|
1
|
Siegel R, Ma J, Zou Z and Jemal A: Cancer
statistics, 2014. CA Cancer J Clin. 64:9–29. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Yang X, Chen BB, Zhang MH and Wang XR:
MicroRNA-126 inhibits the proliferation of lung cancer cell line
A549. Asian Pac J Trop Med. 8:239–242. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ni T, Mao G, Xue Q, Liu Y, Chen B, Cui X,
Lv L, Jia L, Wang Y and Ji L: Upregulated expression of ILF2 in
non-small cell lung cancer is associated with tumor cell
proliferation and poor prognosis. J Mol Histol. 46:325–335. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Peters S, Adjei AA, Gridelli C, Reck M,
Kerr K and Felip E; ESMO Guidelines Working Group: Metastatic
non-small-cell lung cancer (NSCLC): ESMO Clinical Practice
Guidelines for diagnosis, treatment and follow-up. Ann Oncol.
23(Suppl 7): vii56–vii64. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zhang T, Zhang DM, Zhao D, Hou XM, Liu XJ,
Ling XL and Ma SC: The prognostic value of osteopontin expression
in non-small cell lung cancer: A meta-analysis. J Mol Histol.
45:533–540. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Leidinger P, Galata V, Backes C, Stähler
C, Rheinheimer S, Huwer H, Meese E and Keller A: Longitudinal study
on circulating miRNAs in patients after lung cancer resection.
Oncotarget. 6:16674–16685. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Cai J, Fang L, Huang Y, Li R, Yuan J, Yang
Y, Zhu X, Chen B, Wu J and Li M: miR-205 targets PTEN and PHLPP2 to
augment AKT signaling and drive malignant phenotypes in non-small
cell lung cancer. Cancer Res. 73:5402–5415. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Pisters KM, Evans WK, Azzoli CG, Kris MG,
Smith CA, Desch CE, Somerfield MR, Brouwers MC, Darling G, Ellis
PM, et al: Cancer care Ontario and American society of clinical
oncology adjuvant chemotherapy and adjuvant radiation therapy for
stages I–IIIA resectable non small-cell lung cancer guideline. J
Clin Oncol. 25:5506–5518. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Fiori ME, Barbini C, Haas TL, Marroncelli
N, Patrizii M, Biffoni M and De Maria R: Antitumor effect of
miR-197 targeting in p53 wild-type lung cancer. Cell Death Differ.
21:774–782. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hatley ME, Patrick DM, Garcia MR,
Richardson JA, Bassel-Duby R, van Rooij E and Olson EN: Modulation
of K-Ras-dependent lung tumorigenesis by MicroRNA-21. Cancer Cell.
18:282–293. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Seike M, Goto A, Okano T, Bowman ED,
Schetter AJ, Horikawa I, Mathe EA, Jen J, Yang P, Sugimura H, et
al: MiR-21 is an EGFR-regulated anti-apoptotic factor in lung
cancer in never-smokers. Proc Natl Acad Sci USA. 106:12085–12090.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chen C, Zhao Z, Liu Y and Mu D:
microRNA-99a is down-regulated and promotes proliferation,
migration and invasion in non-small cell lung cancer A549 and H1299
cells. Oncol Lett. 9:1128–1134. 2015.PubMed/NCBI
|
|
14
|
Ambros V: The functions of animal
microRNAs. Nature. 431:350–355. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Broderick JA and Zamore PD: MicroRNA
therapeutics. Gene Ther. 18:1104–1110. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Guo H, Ingolia NT, Weissman JS and Bartel
DP: Mammalian microRNAs predominantly act to decrease target mRNA
levels. Nature. 466:835–840. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Mitchell PS, Parkin RK, Kroh EM, Fritz BR,
Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O'Briant
KC, Allen A, et al: Circulating microRNAs as stable blood-based
markers for cancer detection. Proc Natl Acad Sci USA.
105:10513–10518. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K,
Guo J, Zhang Y, Chen J, Guo X, et al: Characterization of microRNAs
in serum: A novel class of biomarkers for diagnosis of cancer and
other diseases. Cell Res. 18:997–1006. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kent OA and Mendell JT: A small piece in
the cancer puzzle: microRNAs as tumor suppressors and oncogenes.
Oncogene. 25:6188–6196. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
22
|
Ling DJ, Chen ZS, Zhang YD, Liao QD, Feng
JX, Zhang XY and Shi TS: MicroRNA-145 inhibits lung cancer cell
metastasis. Mol Med Rep. 11:3108–3114. 2015.
|
|
23
|
Yan LX, Huang XF, Shao Q, Huang MY, Deng
L, Wu QL, Zeng YX and Shao JY: MicroRNA miR-21 overexpression in
human breast cancer is associated with advanced clinical stage,
lymph node metastasis and patient poor prognosis. RNA.
14:2348–2360. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wan LY, Deng J, Xiang XJ, Zhang L, Yu F,
Chen J, Sun Z, Feng M and Xiong JP: miR-320 enhances the
sensitivity of human colon cancer cells to chemoradiotherapy in
vitro by targeting FOXM1. Biochem Biophys Res Commun. 457:125–132.
2015. View Article : Google Scholar
|
|
25
|
Schaar DG, Medina DJ, Moore DF, Strair RK
and Ting Y: miR-320 targets transferrin receptor 1 (CD71) and
inhibits cell proliferation. Exp Hematol. 37:245–255. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wu YY, Chen YL, Jao YC, Hsieh IS, Chang KC
and Hong TM: miR-320 regulates tumor angiogenesis driven by
vascular endothelial cells in oral cancer by silencing neuropilin
1. Angiogenesis. 17:247–260. 2014. View Article : Google Scholar
|
|
27
|
Cheng C, Chen ZQ and Shi XT: MicroRNA-320
inhibits osteosarcoma cells proliferation by directly targeting
fatty acid synthase. Tumour Biol. 35:4177–4183. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sun JY, Xiao WZ, Wang F, Wang YQ, Zhu YH,
Wu YF, Miao ZL and Lin YC: MicroRNA-320 inhibits cell proliferation
in glioma by targeting E2F1. Mol Med Rep. 12:2355–2359.
2015.PubMed/NCBI
|
|
29
|
Lupu R and Menendez JA: Targeting fatty
acid synthase in breast and endometrial cancer: An alternative to
selective estrogen receptor modulators? Endocrinology.
147:4056–4066. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Weiss L, Hoffmann GE, Schreiber R, Andres
H, Fuchs E, Körber E and Kolb HJ: Fatty-acid biosynthesis in man, a
pathway of minor importance. Purification, optimal assay
conditions, and organ distribution of fatty-acid synthase. Biol
Chem Hoppe Seyler. 367:905–912. 1986. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chen J, Zhuang D, Cai W, Xu L, Li E, Wu Y
and Sugiyama K: Inhibitory effects of four plants flavonoids
extracts on fatty acid synthase. J Environ Sci (China). 21(Suppl
1): S131–S134. 2009. View Article : Google Scholar
|
|
32
|
Liu ZL, Wang G, Peng AF, Luo QF, Zhou Y
and Huang SH: Fatty acid synthase expression in osteosarcoma and
its correlation with pulmonary metastasis. Oncol Lett. 4:878–882.
2012.PubMed/NCBI
|
|
33
|
Menendez JA and Lupu R: Fatty acid
synthase and the lipogenic phenotype in cancer pathogenesis. Nat
Rev Cancer. 7:763–777. 2007. View
Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang Y, Zhang XR, Fu J, Tan W and Zhang W:
Prognostic value of expression of FASE, HER-2/neu, bcl-2 and p53 in
stage I non-small cell lung cancer. Zhonghua Zhong Liu Za Zhi.
26:369–372. 2004.In Chinese. PubMed/NCBI
|
|
35
|
Liu ZY, Zhang GL, Wang MM, Xiong YN and
Cui HQ: MicroRNA-663 targets TGFB1 and regulates lung cancer
proliferation. Asian Pac J Cancer Prev. 12:2819–2823.
2011.PubMed/NCBI
|
|
36
|
Ohdaira H, Nakagawa H and Yoshida K:
Profiling of molecular pathways regulated by microRNA 601. Comput
Biol Chem. 33:429–433. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Huang J, Sun C, Wang S, He Q and Li D:
microRNA miR-10b inhibition reduces cell proliferation and promotes
apoptosis in non-small cell lung cancer (NSCLC) cells. Mol Biosyst.
11:2051–2059. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Yang YQ, Qi J, Xu JQ and Hao P:
MicroRNA-142-3p, a novel target of tumor suppressor menin, inhibits
osteosarcoma cell proliferation by down-regulation of FASN. Tumour
Biol. 35:10287–10293. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Mao JH, Zhou RP, Peng AF, Liu ZL, Huang
SH, Long XH and Shu Y: microRNA-195 suppresses osteosarcoma cell
invasion and migration in vitro by targeting FASN. Oncol Lett.
4:1125–1129. 2012.PubMed/NCBI
|
|
40
|
Long XH, Mao JH, Peng AF, Zhou Y, Huang SH
and Liu ZL: Tumor suppressive microRNA-424 inhibits osteosarcoma
cell migration and invasion via targeting fatty acid synthase. Exp
Ther Med. 5:1048–1052. 2013.PubMed/NCBI
|
|
41
|
Wu MF, Yang J, Xiang T, Shi YY and Liu LJ:
miR-21 targets Fas ligand-mediated apoptosis in breast cancer cell
line MCF-7. J Huazhong Univ Sci Technolog Med Sci. 34:190–194.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Mo JS, Alam KJ, Kang IH, Park WC, Seo GS,
Choi SC, Kim HS, Moon HB, Yun KJ and Chae SC: MicroRNA 196B
regulates FAS-mediated apoptosis in colorectal cancer cells.
Oncotarget. 6:2843–2855. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Wang Z, Liu M, Zhu H, Zhang W, He S, Hu C,
Quan L, Bai J and Xu N: miR-106a is frequently upregulated in
gastric cancer and inhibits the extrinsic apoptotic pathway by
targeting FAS. Mol Carcinog. 52:634–646. 2013. View Article : Google Scholar
|
|
44
|
Li X, Chen YT, Josson S, Mukhopadhyay NK,
Kim J, Freeman MR and Huang WC: MicroRNA-185 and 342 inhibit
tumorigenicity and induce apoptosis through blockade of the SREBP
metabolic pathway in prostate cancer cells. PLoS One. 8:e709872013.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhang H, Feng Z, Huang R, Xia Z, Xiang G
and Zhang J: MicroRNA-449 suppresses proliferation of hepatoma cell
lines through blockade lipid metabolic pathway related to SIRT1.
Int J Oncol. 45:2143–2152. 2014.PubMed/NCBI
|