Introduction
Liver transplantation is the primary treatment for
patients with acute liver failure, end-stage liver disease and
inherited liver-based metabolic disorders. However, the demand for
suitable organs for transplantation far exceeds the available donor
organs (1). Tissue engineering and
regenerative medicine-based strategies are a promising alternative
to organ transplantation (2–4).
Tissue-specific cells and scaffolding biomaterials
are essential for liver tissue engineering (5). As primary hepatocytes exhibit poor
proliferative potential in vitro, it may be more feasible to
generate hepatocytes via the differentiation of mesenchymal stem
cells (MSCs). MSCs are commonly cultured as 2D monolayers using
conventional tissue culture techniques, which may result over time
in loss of replicative ability, reduced colony-forming efficiency
and poor differentiation capacity (6,7). The
microenvironment has a crucial influence on stem cell biology.
Therefore, the present study investigated spheroid culture, which
has been reported to improve cell-cell contact and interactions of
cells with the extracellular matrix (ECM) compared with traditional
monolayer methods (8). As cells
exist in their native morphology, significant differences in
phenotype and responses have been observed between monolayer and
spheroid cultures (9,10). Our previous study revealed that 3D
spheroid cultures of MSCs enhanced cell yield and maintained
stemness, in addition to osteogenetic and adipogenetic
differentiation efficiencies (11).
In the last decade, advances in organ and tissue
decellularization have made it possible to obtain tissue-specific
ECM from whole organs via the perfusion of the organs with various
detergents (12–15). Whole organ decellularization
represents a potential strategy for the fabrication of scaffolds
for the engineering of tissues and organs, as the decellularized
scaffolds maintain their microarchitecture and retain numerous
bioactive signals that are difficult to replicate artificially
(16). Decellularized liver
scaffolds may act as anchors for hepatocyte-like cells derived from
stem cells, and aid their attachment, proliferation and
organization (17–19). In addition, decellularized liver
scaffolds may be an alternative option for heterotopic hepatocyte
transplantation (20).
In the present study, spheroid culture and
decellularized liver scaffolds (DLSs) were utilized to establish a
novel 3D culture system to promote maturation of hepatocyte-like
cells from mouse bone marrow (BM)-derived MSCs. The Albp-ZsGreen
adenoviral vector, which is driven by the albumin (ALB) promoter,
was utilized for real-time monitoring of the differentiation status
of hepatocytes from stem cells. The findings of the present study
may be useful for cell transplantation purposes.
Materials and methods
Animals
The study was approved by the Ethics Committee of
Sichuan University (Chengdu, China). Three livers were isolated
from 6-month-old male Bama miniature pigs weighing 10–15 kg for
perfusion decellularization. Male C57BL/6 mice (n=3; age, 8 weeks;
weight, 20–25 g) were used for hepatocyte isolation. All animals
were obtained from the Animal Experiment Center of Sichuan
University (Chengdu, China). The mice and Bama miniature pigs were
maintained on an alternating 12-h light/dark cycle, fed regular
chow, and given water ad libitum.
The surgeries were performed under ketamine (6 mg/kg
body weight, administered intramuscular; Kelun, Chengdu, China) and
xylazine (10 mg/kg intramuscular; Kelun) anesthesia. Under deep
anesthesia, a laparotomy was performed and the liver was exposed.
After systemic heparinization through the inferior vena cava, the
hepatogastric ligament was carefully dissected. The proximal PV was
catheterized. The hepatic artery and common bile duct were ligated
and transected. All perihepatic ligaments were severed.
Simultaneously, the liver was slowly perfused with 2 l deionized
water containing 0.1% EDTA (Kelun) through a cannula in the PV, and
the SHIVC was transected, allowing outflow of the perfusate.
Following blanching, the liver was stored at −80°C overnight. The
Bama miniature pigs were sacrificed during the perfusion process
due to an excessive amount of blood loss.
Cultivation of mouse BM-MSCs
Commercial mouse BM-MSCs were purchased from Cyagen
Biosciences Inc. (Guangzhou, China). C57BL/6 mouse MSC growth
medium (cat. no. MUBMX-90011; Cyagen Biosciences Inc.) was utilized
to culture cells and was replaced at least every 2 days. Cells at
passage 4–6 were used for subsequent experiments.
Formation of BM-MSCs spheroids
For spheroid cultures, the harvested BM-MSCs were
suspended in 10 ml serum-free medium at 1×106 cells/ml
and cultured in glass spheroid dishes (13×8×4 cm), which were
coated with Sigmacote® (Sigma-Aldrich; Merck KGaA,
Darmstadt, Germany). Spheroid dishes were incubated with continuous
rocking at 0.167 Hz for 24 h to induce spheroid formation. BM-MSC
spheroids were stained with 4′,6-diamidino-2-phenylindole (DAPI;
Santa Cruz Biotechnology, Inc., Dallas, TX, USA). The viability of
BM-MSC spheroids was assessed using the FluoroQuench™ fluorescent
stain (One Lambda; Thermo Fisher Scientific, Inc., Waltham, MA,
USA) according to the manufacturer's protocol. Samples were imaged
using a Leica DFC495 fluorescence microscope (Leica Microsystems
GmbH, Wetzlar, Germany).
Evaluation of decellularized porcine
liver
DLSs were obtained as previously described (21). DLS samples were fixed in 4%
paraformaldehyde, and stained with hematoxylin and eosin and 2.5%
glutaraldehyde, prior to observation under a scanning electron
microscope (22).
Cell seeding and hepatic
differentiation
Two culturing methods for differentiation [single
cell (2D) and spheroids + DLS (3D)] were studied. DLSs were
incubated in culture medium at 37°C overnight. Following aspiration
of the medium, 100 µl cell suspension of harvested BM-MSC spheroids
was pipetted onto the center of the DLS via a negative pressure
suction device. Spheroids were allowed to settle and attach to the
scaffold for 4 h. Subsequently, 2 ml medium of stage one was added
slowly to the spheroids. To induce hepatic differentiation,
serum-free Iscove's modified Dulbecco's medium (HyClone; GE
Healthcare Life Sciences, Logan, UT, USA) supplemented with growth
factors was utilized, as described previously (23): i) 10 ng/ml basic fibroblast growth
factor (bFGF) and 20 ng/ml epidermal growth factor (EGF) for 2
days; ii) 20 ng/ml hepatocyte growth factor (HGF), 10 mg/ml bFGF,
and 0.61 mg/m; nicotinamide (NAM) for 7 days; and iii) insulin
transferrin selenium (ITS) premix solution (10 µg/ml insulin, 5.5
µg/ml transferrin, 5 ng/ml selenium), 1 µmol/l dexamethasone (DXM)
sodium phosphate, and 20 ng/l oncostatin M (OSM) for 14 days.
Supplements were all purchased from Sigma-Aldrich; Merck KGaA. The
culture medium was replaced every 3 days during the differentiation
period.
Albp-ZsGreen adenovirus
transduction
To monitor the differentiation of hepatocytes from
BM-MSCs, the Albp-ZsGreen adenoviral vector containing the ALB
promoter was designed and constructed as previously described
(24). 2D and 3D hepatocyte-like
cells (1×106 cells/well) were incubated with the
Albp-ZsGreen adenoviral vector (10 µl; 1×108 plaque
formation units; multiplicity of infection, 100) in 6-well tissue
culture plates for 2 h prior to examining ALB expression at 48 h
using a Leica DM40000B microscope (Leica Microsystems GmbH).
Following DAPI staining, the percentage of ZsGreen-positive cells
was determined using ImageJ software version 1.48 (National
Institutes of Health, Bethesda, MD, USA). Three samples of both 2D
and 3D undifferented cells were incubated with adenovirus as
controls.
Western blot analysis
Differentiated cells in 2D and 3D culture systems
were homogenized to generate protein lysates using
radioimmunoprecipitation assay lysis buffer (Beyotime Institute of
Biotechnology, Haimen, China) with protease inhibitors (Beyotime
Institute of Biotechnology). Equal quantities of protein (80 µg)
were separated by 10% SDS-PAGE. Proteins were transferred onto
polyvinylidene difluoride membranes (EMD Millipore, Billerica, MA,
USA). Membranes were blocked in 5–10% non-fat milk in Tris-buffered
saline containing 0.1% Tween-20 for 1 h at room temperature and
then incubated overnight with anti-ALB (1:1,000; rabbit monoclonal;
cat. no. ab207327; Abcam, Cambridge, UK) and anti-GAPDH (1:8,000;
mouse monoclonal; cat. no. MAB374-AF647; EMD Millipore) primary
antibodies at 4°C. Following this, the membranes were washed twice
with TBST and incubated with horseradish peroxidase-conjugated goat
anti-rabbit (cat. no. ab6721; 1:1,000) and goat anti-mouse (cat.
no. ab6789; 1:1,000) secondary antibodies (both from Abcam) for 2 h
at room temperature. Protein bands were visualized using an
enhanced ehemiluminescence kit (Thermo Scientific, Inc., Waltham,
MA, USA).
ALB and urea production
Conditioned media from the differentiated BM-MSCs of
2D and 3D culture systems was collected on day 21 and ALB levels
were measured using a mouse ALB ELISA kit (cat. no. E90-134; Bethyl
Laboratories, Inc., Montgomery, TX, USA) according to the
manufacturer's protocol. A total of 2 mM heavy, diazonium-enriched
ammonium chloride (Cambridge Isotope Laboratories, Inc., Tewksbury,
MA, USA) was added to the medium to determine the metabolic ability
of differentiated cells. The total urea concentration and
proportion of diazonium-enriched urea and natural urea in the
medium were measured to determine the source of urea synthesis.
Supernatants were quantified by capillary gas chromatography and
mass spectrometry, as previously reported (25). Freshly isolated mouse primary
hepatocytes were included as a control. Hepatocytes were isolated
using a two-step perfusion method as previously described (25).
Immunofluorescence analysis
Differentiated cells from 2D and 3D groups were
fixed in 4% formaldehyde in PBS and permeabilized with 0.1% Triton
X-100 for 15 min at room temperature. Following permeabilization,
samples were blocked with 2% bovine serum albumin (Sigma-Aldrich;
Merck KGaA) in PBS (blocking buffer) for 1 h and subsequently
treated with primary antibodies diluted in blocking buffer
overnight at 4°C. The antibodies utilized were sheep anti-ALB
(1:1,000; cat. no. ab8940; Abcam), rabbit anti-α-fetoprotein
(1:1,000; AFP; cat. no. AF5134; Affinity Biosciences, Cell Signal
Transduction, Cambridge, UK) and rabbit anti-cytokeratin-19 (CK19;
1:1,000; cat. no. AF0192; Affinity Biosceinces, Cell Signal
Transduction). Alexa Fluor 488-conjugated rabbit anti-sheep (1:400;
cat. no. ab150181; Abcam) and goat anti-rabbit (1:400; cat. no.
ab150077; Abcam) secondary antibodies were incubated with samples
at room temperature for 1 h in the dark. Following nuclear staining
with DAPI, slides were mounted and observed under a Leica DMI6000
fluorescence microscope (Leica Microsystems GmbH).
Gene array analysis
Sample labeling and array hybridization were
performed with Whole Mouse Genome Oligo Microarray (cat. no.
G4122F; 4×44K; Agilent Technologies, Inc., Santa Clara, CA, USA).
Briefly, total RNA was extracted from BM-MSCs, primary mouse
hepatocytes and 3D hepatocyte-like cells using TRIzol reagent
(Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's protocol, and used for synthesis of cRNAs, which
were labeled with Cyanine 3-UTP. The concentration of cRNA was
measured using a NanoDrop ND-1000 (Thermo Fisher Scientific, Inc.,
Wilmington, DE, USA). The hybridized arrays were performed and
scanned using the Agilent DNA Microarray Scanner (Agilent
Technologies, Inc.). Data were normalized and analyzed using the
TIGR MultiExperiment Viewer version 4.8.1 (Institute of Genomic
Research, Rockville, MD, USA). Gene ontology (GO) analysis was
performed using the Database for Annotation, Visualization and
Integrated Discovery (DAVID; https://david.ncifcrf.gov) to determine the functions
of the predicted target genes and to uncover the miRNA-target gene
regulatory network based on the predicted biological processes and
molecular functions; P<0.01 was used as the threshold.
Statistical analysis
Data are expressed as the mean ± standard error of
three independent experiments. Data were analyzed using SPSS
software version 17.0 (SPSS, Inc., Chicago, IL, USA). A one-way
analysis of variance followed by the Dunnett's post hoc test was
performed to compare groups. P<0.05 was considered to indicate a
statistically significant difference.
Results
Characterization of BM-MSC
spheroids
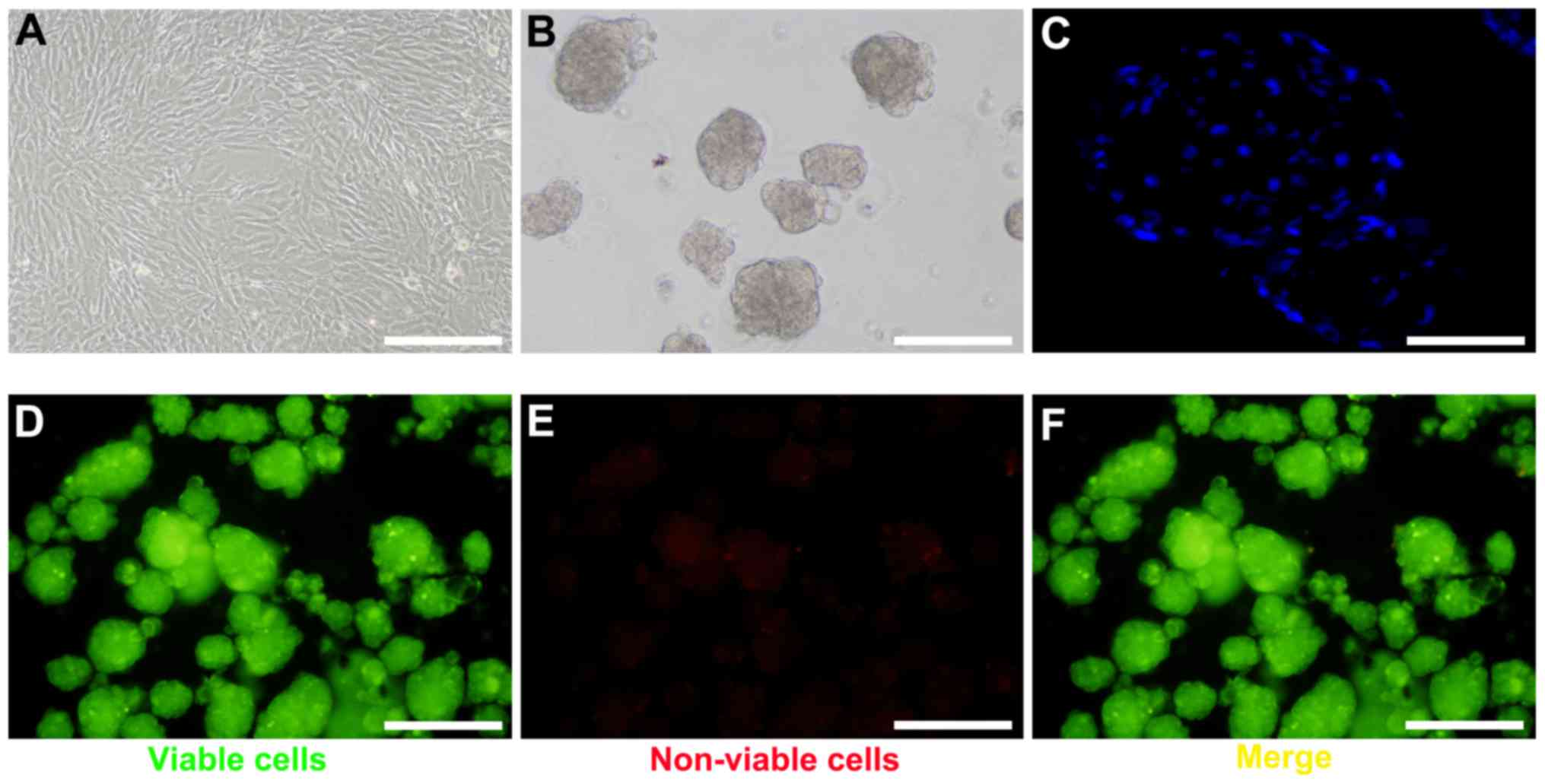
To generate BM-MSC spheroids, BM-MSCs were harvested
at passage 4–6 (Fig. 1A).
Following optimization of cell number and growth conditions,
spheroid formation was observed under a scanning electron
microscope within 24 h (Fig. 1B).
Based on DAPI staining, the cells within the spheroids were in
close proximity (Fig. 1C). The
FluoroQuench™ fluorescent staining assay revealed that the
viability of BM-MSCs in spheroids remained >95% in 3D culture
(Fig. 1D-F).
Characterization of DLSs
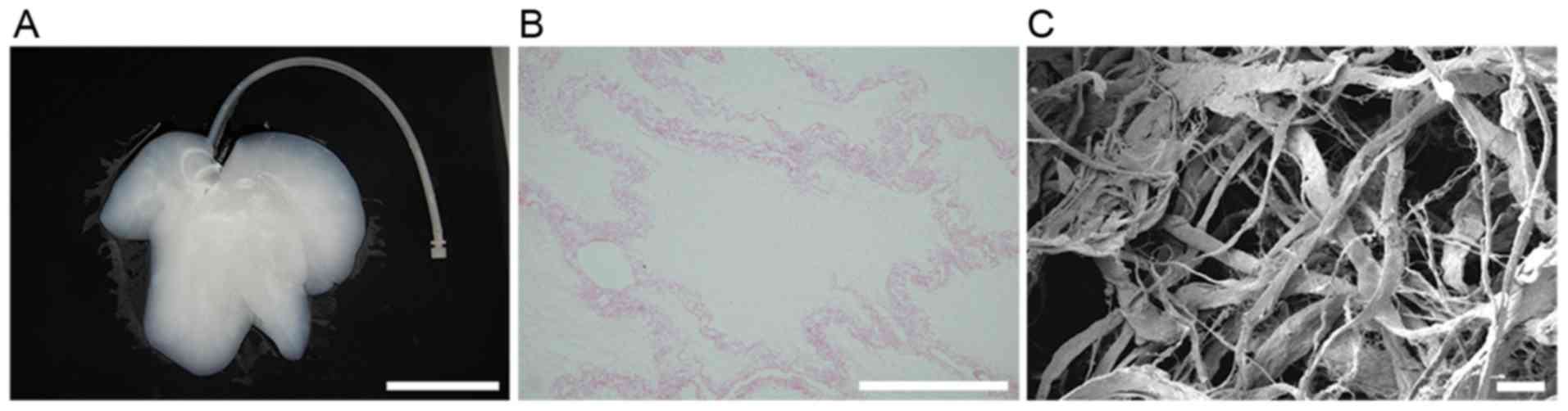
Whole-organ decellularization was achieved by portal
perfusion using sodium dodecyl sulfate and Triton X-100. Following
decellularization, the porcine liver parenchyma became
semi-transparent (Fig. 2A).
Hematoxylin and eosin staining revealed no visible cell nuclei and
cellular material in the decellularized liver scaffolds (Fig. 2B). Decellularized tissue sections
were observed under a scanning electron microscope to determine
whether the structure of the bio-scaffold was preserved (Fig. 2C). Reticular collagen fibers, which
provide support for the hepatic tissue, were evident.
Identification of hepatocyte-like
cells
The hepatocyte-like cells were generated from mouse
BM-MSCs as shown in Fig. 3A. The
expression of the Albp-ZsGreen adenovirus was not observed in
non-hepatic cells (Fig. 3B);
however, ALB synthesis was observed in mature hepatocytes, which
suggested that hepatic differentiation had occurred in 2D and 3D
cells (Fig. 3C). A greater
percentage of ZsGreen-positive cells was observed in the 3D group,
which suggested that the 3D culture system provided an improved
external microenvironment for differentiation (Fig. 3D). Similarly, the expression levels
of ALB in 3D cells were greater compared with the 2D cells
following induction (Fig. 3E). In
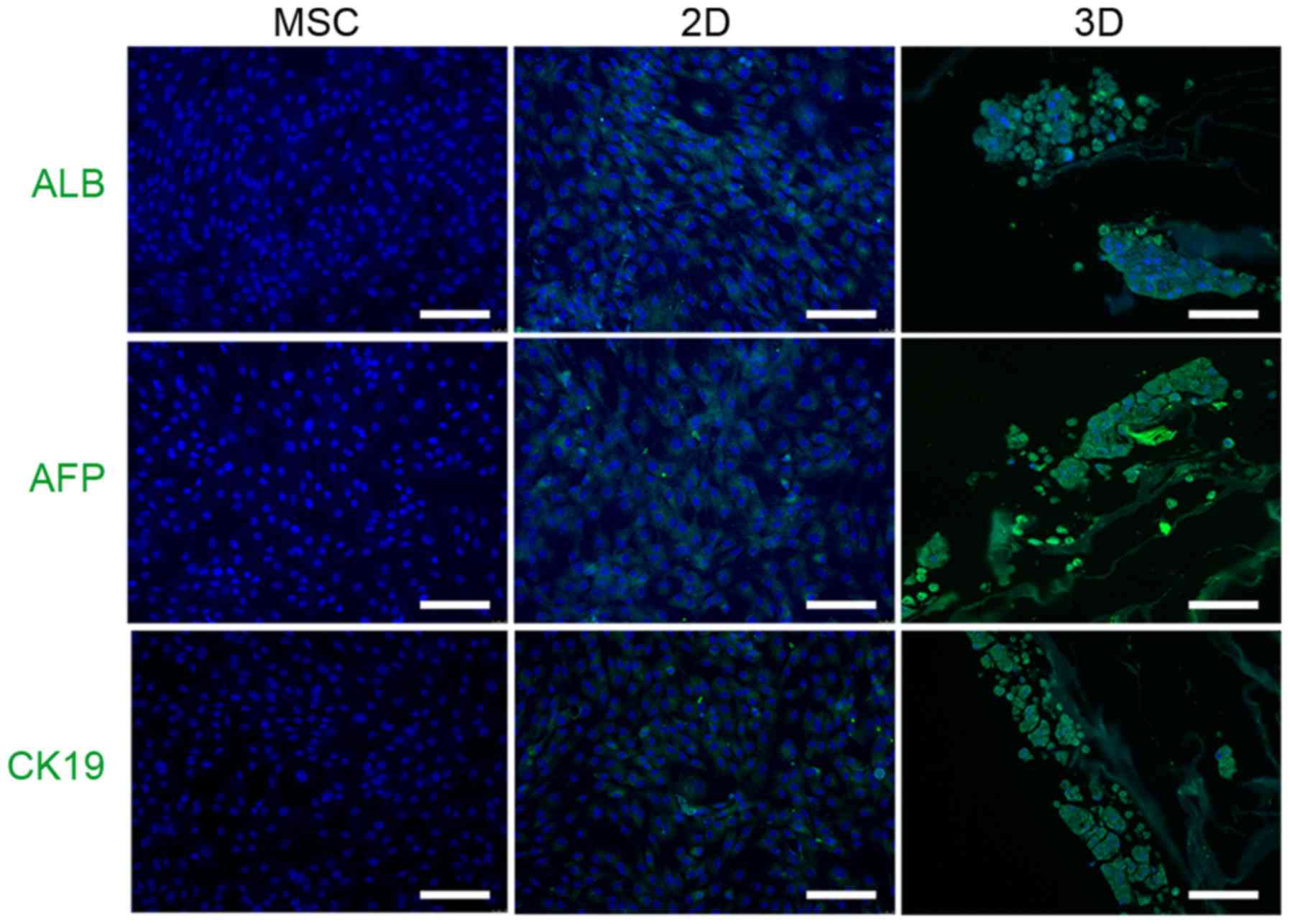
addition, immunocytochemistry demonstrated that 3D cells expressed
hepatocyte-like cell markers, including ALB, AFP and CK19, to a
greater extent than 2D cells (Fig.
4).
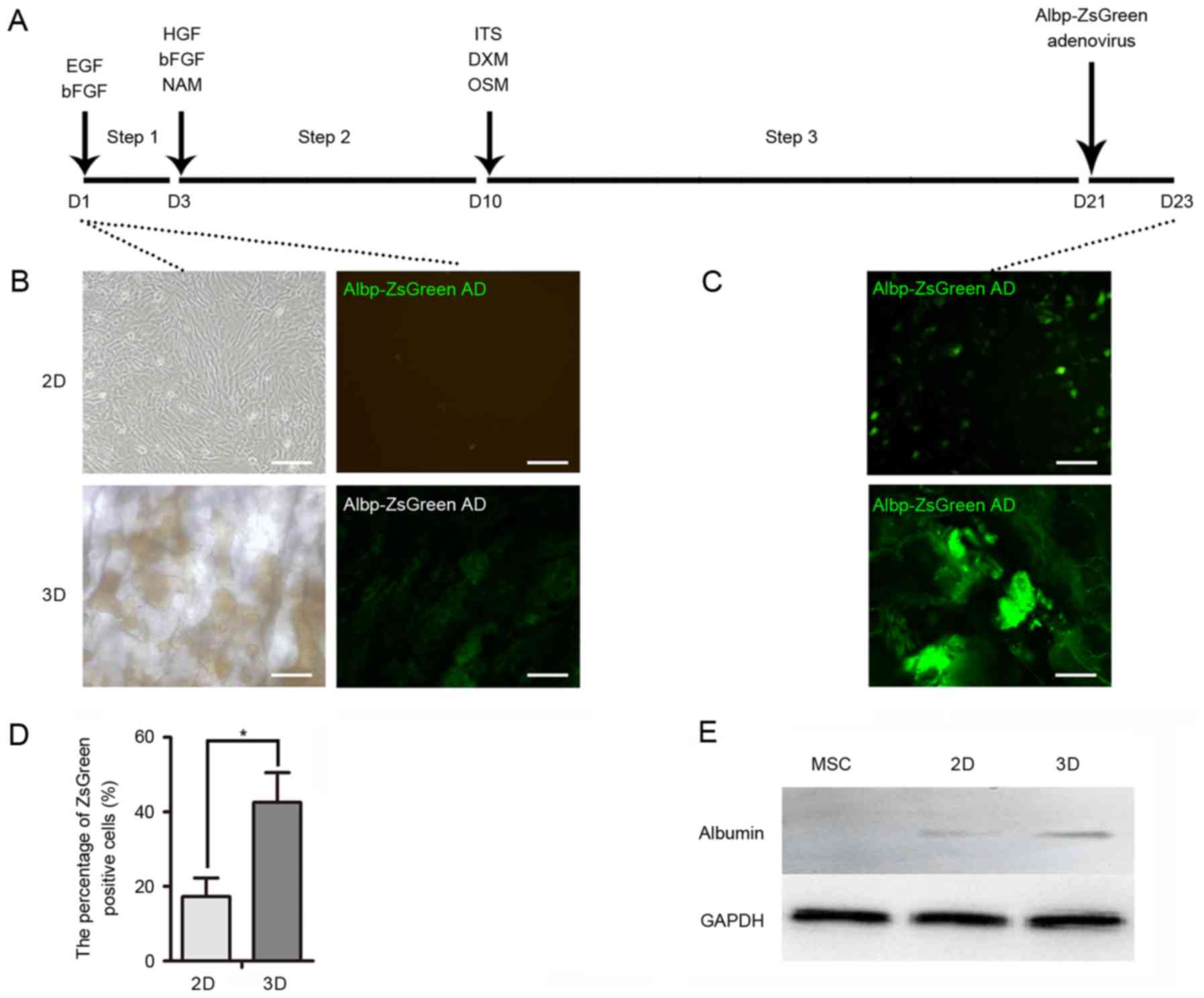 | Figure 3.Hepatocyte-like cell induction. (A)
Timeline of hepatic induction of mouse MSCs for 2D and 3D cultures.
Cells were incubated in medium containing various growth factors
for 23 days. Albp-ZsGreen adenovirus was added to induce expression
of ALB. (B) Green fluorescence of Albp-ZsGreen adenovirus indicated
ALB synthesis in non-hepatic or (C) hepatic differentiated MSCs.
(D) Percentage of Albp-ZsGreen-positive hepatocyte-like cells in
each group. (E) Western blot analysis demonstrated the expression
of ALB in undifferentiated MSCs and following hepatic
differentiation using 2D or 3D models. Data represent the mean ±
standard deviation of three independent experiments. *P<0.05.
Scale bars, 100 µm. MSCs, mesenchymal stem cells; ALB, albumin;
EGF, epidermal growth factor; bFGF, basic fibroblast growth factor;
HGF, hepatocyte growth factor; NAM, nicotinamide; DXM,
dextromethorphan; OSM, oncostatin M; ITS, insulin transferrin
selenium. |
Metabolic activity of differentiated
cells in 2D or 3D culture
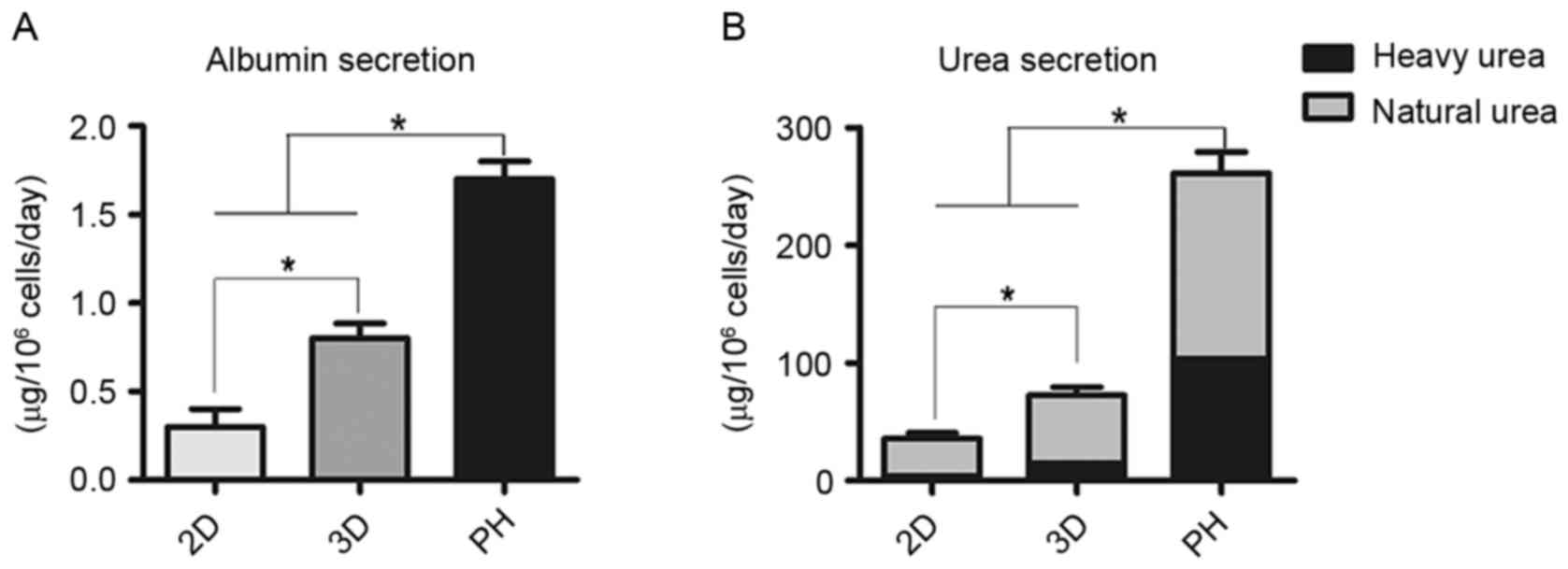
Compared with the 2D group, cumulative ALB secretion
by differentiated cells in the 3D group was significantly greater
compared with the 2D group (P<0.05; Fig. 5A). Clearance of heavy urea, which
contained ammonium chloride, was greater in the 3D group compared
with the 2D group (P<0.05; Fig.
5B). The proportion of heavy urea to total urea produced by the
2D group, 3D group and primary hepatocytes fluctuated from 9.6 to
11.8%, 19.8 to 22.1% and 39.8 to 42.1%, respectively (Fig. 5B).
Gene expression in 3D hepatocyte-like
cells
The results of the present study demonstrated the
advantage of 3D culture in promoting hepatic differentiation of
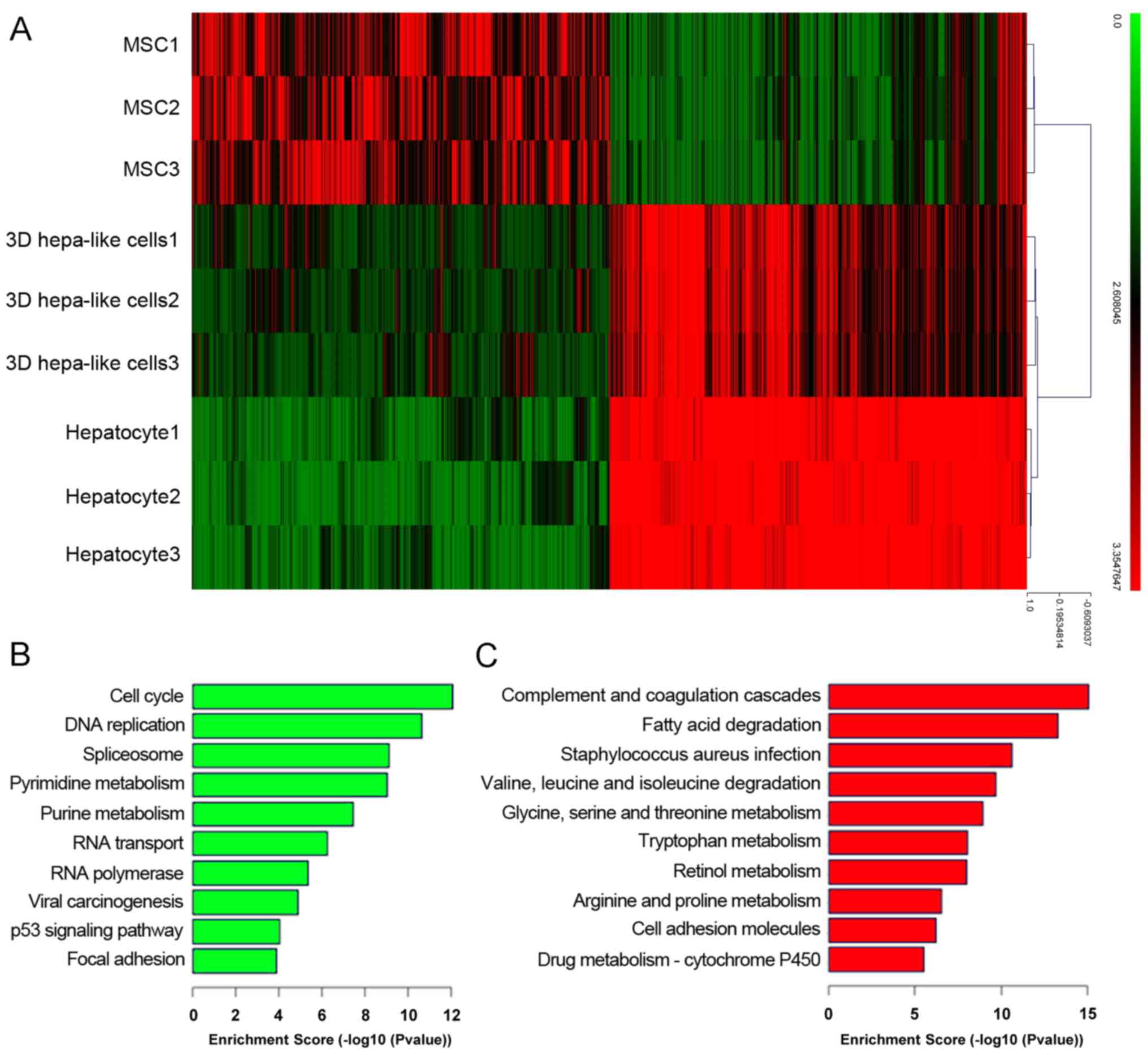
mouse BM-MSCs. The cDNA microarray showed that differentiated 3D
hepatocyte-like cells resembled mouse primary hepatocytes more than
mouse BM-MSCs (Fig. 6A). GO
enrichment analysis revealed that signaling pathways that
associated with liver function were significantly upregulated,
including those involved in fat metabolism, amino acid metabolism
and drug metabolism (Fig. 6C),
whereas signaling pathways associated with the cell cycle were
significant downregulated in the 3D group compared with BM-MSCs
(Fig. 6B).
Discussion
A limitation of traditional methods for the
induction of differentiation of adherent monolayer hepatocyte-like
cells is low differentiation efficiency (26). In the present study, a novel 3D
culture system was established to obtain differentiated hepatocyte
cells from spheroid cultures and DLSs.
Using a simplified portal vein perfusion procedure,
porcine liver was effectively decellularized and examined under a
scanning electron microscope. Subsequently, the 3D model was
utilized. In our previous study, spheroids were pipetted onto the
center of the DLS to allow spontaneous attachment. However, certain
spheroids detached from the DLS in the early stages of culture
(8,22). In the present study, a negative
pressure suction device was utilized to ensure the efficient
attachment of spheroids to the DLS.
A simple, efficient tool for real-time monitoring of
hepatocyte differentiation from stem cells is useful. As the
adenoviral vector is commonly utilized for gene expression studies
due to its efficient transduction, an Albp-ZsGreen adenoviral
vector was constructed to detect hepatocyte-like cells. Following
the second induction period, green fluorescence was detected at an
earlier stage in 3D cells compared with 2D cells and the
fluorescence intensity in 3D cells was significantly greater in a
time-dependent manner (data not shown).
Ammonia is a product of protein metabolism and
requires clearance by the liver. A functional urea cycle is an
important characteristic of mature hepatocytes. To assess urea
production and the detoxification of ammonia via the urea cycle,
heavy ammonium chloride was added to the culture medium. The
concentration of urea produced by differentiated cells in the 3D
group was greater compared with the 2D group. In addition, the
results of the present study demonstrated that ~20% of urea was
produced by urea cycle activity in 3D cells, which was
significantly greater compared with 2D cells.
The liver is the largest internal organ providing
essential metabolic, exocrine and endocrine functions. These
functions include production of bile, metabolism of dietary
compounds, detoxification, regulation of glucose levels via
glycogen storage and control of blood homeostasis by secretion of
clotting factors and serum proteins, including ALB. In the present
study, GO analysis revealed that the majority of upregulated genes
were associated with liver function, whereas cell cycle-associated
pathways were significantly downregulated.
In conclusion, the results of the present study
suggested that a 3D culture system may promote hepatic
differentiation of mouse MSCs, to generate high yields of mature
hepatocytes. This miniaturized culture system may possess unique
advantages over previous methods and may provide a potential
strategy for cell transplantation and drug research.
Acknowledgements
The present study was supported by the National Key
Clinical Project, the National Natural Scientific Foundations of
China (grant no. 81200315) and the Doctoral Program of Colleges and
Universities Specialized Research Foundation (grant no.
20120181110090).
References
|
1
|
Brown RS Jr: Live donors in liver
transplantation. Gastroenterology. 134:1802–1813. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Fox IJ, Daley GQ, Goldman SA, Huard J,
Kamp TJ and Trucco M: Use of differentiated pluripotent stem cells
in replacement therapy for treating disease. Science.
345:12473912014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Chistiakov DA: Liver regenerative
medicine: Advances and challenges. Cells Tissues Organs.
196:291–312. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Forbes SJ and Newsome PN: New horizons for
stem cell therapy in liver disease. J Hepatol. 56:496–499. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Badylak SF, Doris T and Korkut U:
Whole-organ tissue engineering: Decellularization and
recellularization of three-dimensional matrix scaffolds. Annu Rev
Biomed Eng. 13:27–53. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Baer PC, Griesche N, Luttm W, Schubert R,
Luttm A and Geiger H: Human adipose-derived mesenchymal stem cells
in vitro: Evaluation of an optimal expansion medium preserving
stemness. Cytotherapy. 12:96–106. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Park E and Patel AN: Changes in the
expression pattern of mesenchymal and pluripotent markers in human
adipose-derived stem cells. Cell Biol Int. 34:979–984. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Li Y, Wu Q, Wang Y, Li L, Chen F, Shi Y,
Bao J and Bu H: Construction of bioengineered hepatic tissue
derived from human umbilical cord mesenchymal stem cells via
aggregation culture in porcine decellularized liver scaffolds.
Xenotransplantation. 24:e122852017. View Article : Google Scholar
|
|
9
|
Lin SJ, Jee SH, Hsiao WC, Yu HS, Tsai TF,
Chen JS, Hsu CJ and Young TH: Enhanced cell survival of melanocyte
spheroids in serum starvation condition. Biomaterials.
27:1462–1469. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Frith JE, Thomson B and Genever PG:
Dynamic three-dimensional culture methods enhance mesenchymal stem
cell properties and increase therapeutic potential. Tissue
Engineering Part C Methods. 16:735–749. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Li Y, Guo G, Li L, Chen F, Bao J, Shi YJ
and Bu H: Three-dimensional spheroid culture of human umbilical
cord mesenchymal stem cells promotes cell yield and stemness
maintenance. Cell Tissue Res. 360:297–307. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Uygun BE, Soto-Gutierrez A, Yagi H, Izamis
ML, Guzzardi MA, Shulman C, Milwid J, Kobayashi N, Tilles A,
Berthiaume F, et al: Organ reengineering through development of a
transplantable recellularized liver graft using decellularized
liver matrix. Nat Med. 16:814–820. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ott HC, Matthiesen TS, Goh SK, Black LD,
Kren SM, Netoff TI and Taylor DA: Perfusion-decellularized matrix:
Using nature's platform to engineer a bioartificial heart. Nat Med.
14:213–221. 2008. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Song JJ, Guyette JP, Gilpin SE, Gabriel G,
Vacanti JP and Ott HC: Regeneration and experimental orthotopic
transplantation of a bioengineered kidney. Nat Med. 19:646–651.
2013. View
Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ott HC, Clippinger BC, Schuetz C,
Pomerantseva I, Ikonomou L, Kotton D and Vacanti JP: Regeneration
and orthotopic transplantation of a bioartificial lung. Nat Med.
16:927–933. 2010. View
Article : Google Scholar : PubMed/NCBI
|
|
16
|
Crapo PM, Gilbert TW and Badylak SF: An
overview of tissue and whole organ decellularization processes.
Biomaterials. 32:3233–3243. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lang R, Stern MM, Smith L, Liu Y,
Bharadwaj S, Liu G, Baptista PM, Bergman CR, Soker S, Yoo JJ, et
al: Three-dimensional culture of hepatocytes on porcine liver
tissue-derived extracellular matrix. Biomaterials. 32:7042–7052.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ji R, Zhang N, You N, Li Q, Liu W, Jiang
N, Liu J, Zhang H, Wang D, Tao K and Dou K: The differentiation of
MSCs into functional hepatocyte-like cells in a liver biomatrix
scaffold and their transplantation into liver-fibrotic mice.
Biomaterials. 33:8995–9008. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Jiang WC, Cheng YH, Yen MH, Chang Y, Yang
VW and Lee OK: Cryo-chemical decellularization of the whole liver
for mesenchymal stem cells-based functional hepatic tissue
engineering. Biomaterials. 35:3607–3617. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhou P, Lessa N, Estrada DC, Severson EB,
Lingala S, Zern MA, Nolta JA and Wu J: Decellularized liver matrix
as a carrier for the transplantation of human fetal and primary
hepatocytes in mice. Liver Transpl. 17:418–427. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wu Q, Bao J, Zhou YJ, Wang YJ, Du ZG, Shi
YJ, Li L and Bu H: Optimizing perfusion-decellularization methods
of porcine livers for clinical-scale whole-organ bioengineering.
Biomed Res Int. 2015:7854742015.PubMed/NCBI
|
|
22
|
Bao J, Wu Q, Wang Y, Li Y, Li L, Chen F,
Wu X, Xie M and Bu H: Enhanced hepatic differentiation of rat bone
marrow-derived mesenchymal stem cells in spheroidal aggregate
culture on a decellularized liver scaffold. Int J Mol Med.
38:457–465. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Schwartz RE, Reyes M, Koodie L, Jiang Y,
Blackstad M, Lund T, Lenvik T, Johnson S, Hu WS and Verfaillie CM:
Multipotent adult progenitor cells from bone marrow differentiate
into functional hepatocyte-like cells. J Clin Invest.
109:1291–1302. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Tang J, Wu Q, Li Y, Wu X, Wang Y, Zhu L,
Shi Y, Bu H, Bao J and Xie M: Construction of a general albumin
promoter reporter system for real-time monitoring of the
differentiation status of functional hepatocytes from stem cells in
mouse, rat and human. Biomed Rep. 6:627–632. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Bao J, Fisher JE, Lillegard JB, Wang W,
Amiot B, Yu Y, Dietz AB, Nahmias Y and Nyberg SL: Serum-free medium
and mesenchymal stromal cells enhance functionality and stabilize
integrity of rat hepatocyte spheroids. Cell Transplant. 22:299–308.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Książek K: A Comprehensive review on
mesenchymal stem cell growth and senescence. Rejuvenation Res.
12:105–116. 2009. View Article : Google Scholar : PubMed/NCBI
|