Introduction
Prostate cancer is one of the most malignant types
of tumour in developed countries, and prostate cancer-associated
mortality has recently increased. According to statistics from the
American Cancer Society in 2016 (1), >1.8 million new cases of prostate
cancer occur each year. In addition, >26,000 patients die from
prostate cancer per year, accounting for 27% of the total number of
patients. Due to diverse dietary habits and more advanced medical
equipment, early and small prostate cancer lesions are increasingly
detected. The appropriate treatment for prostate cancer depends on
the disease stage. Radical prostatectomy is the primary choice for
the treatment of early stage prostate cancer. However, endocrine
therapy may be more effective for the treatment of prostate tumours
that have progressed to an advanced stage.
Gene therapy is a novel treatment approach that is
primarily in an experimental stage. The androgen-androgen receptor
(AR) signalling pathway serves an important role during all stages
of prostate cancer. Following binding to the AR in prostate
epithelial cells, androgens translocate to the cell nucleus. Once
in the nucleus, androgens bind to androgen response elements
upstream of the target genes, which leads to DNA transcription,
promotes the abnormal proliferation of prostate epithelial cells
and enables carcinogenesis (2).
During the androgen-sensitive stage of prostate cancer, the
blockade or removal of androgens prevents the binding of androgens
to the AR, which may delay the development of prostate cancer.
However, at 18 months, prostate cancer becomes
androgen-independent. Therefore, the tumour growth may not be
inhibited using androgen suppressors. However, numerous studies
have demonstrated that the AR retains a role in
androgen-independent prostate cancer via AR gene mutation and gene
amplification, and acts together with co-modulating factors to
stimulate the abnormal activation of other signalling pathways
(3–5). Therefore, the development of prostate
cancer may be suppressed by inhibiting AR.
The clustered regularly interspaced short
palindromic repeats-associated protein (CRISPR/Cas) system has
recently become an extensively used gene-editing technology
(6,7). Compared with the previous zinc-finger
nuclease (ZFN) and transcription activator-like effector nuclease
(TALEN) technologies, the CRISPR/Cas system is more widely used in
basic and in certain clinical studies (8). A number of researchers have begun to
apply the CRISPR/Cas system in studies investigating inherited
genetic diseases, and certain studies have achieved relevant
results (9–13). For example, Kawamura et al
(14) established homeobox protein
NANOG (NANOG)- and NANOGP8-knockout DU145 cell lines, which
exhibited a significantly decreased malignant potential compared
with control cells.
In the present study, a site-specific CRISPR/Cas
system was designed to cleave the AR gene in androgen-positive
prostate cancer cell lines and decrease the growth of
androgen-dependent prostate cancer in vitro. Stable LNCaP
cell lines harbouring Cas9 and single-guide RNAs (sgRNAs) were
constructed to knock out AR, and the knockout activity and
efficiency of CRISPR was investigated. The results of the present
study demonstrated that the treatment with CRISPR/Cas significantly
inhibited the growth of LNCaP cells. These results suggested that
the CRISPR/Cas system may be a potential therapeutic strategy for
the treatment of prostate cancer.
Materials and methods
Cell culture
The human LNCaP cell line was purchased from the
Type Culture Collection of the Chinese Academy of Sciences
(Shanghai, China) and preserved in the Laboratory of the Second
Affiliated Hospital of Soochow University. All cells were cultured
in RPMI 1640 medium (HyClone; GE Healthcare Life Sciences, Logan,
UT, USA) supplemented with 100 U/ml penicillin, 100 µg/ml
streptomycin and 10% fetal bovine serum (Gibco; Thermo Fisher
Scientific, Inc., Waltham, MA, USA). The cell culture plates were
maintained in humidified incubators at 37°C with 5%
CO2.
Construction of AR-Cas9-sgRNA stable
cell lines
Lentiviral vectors (LVs) harbouring Cas9 and sgRNA
(Shanghai GeneChem Co., Ltd., Shanghai, China) were constructed for
transfection of the LNCaP cells. According to the manufacturer's
protocol, the LVs harbouring Cas9 were transfected into LNCaP cells
using polybrene (Shanghai GeneChem Co., Ltd.), and the LNCaP-Cas9
stable cell lines were isolated. In addition, three different
sgRNAs were designed according to the three different target sites
in the AR gene and packaged into the LVs to form LV-sgRNA-AR230,
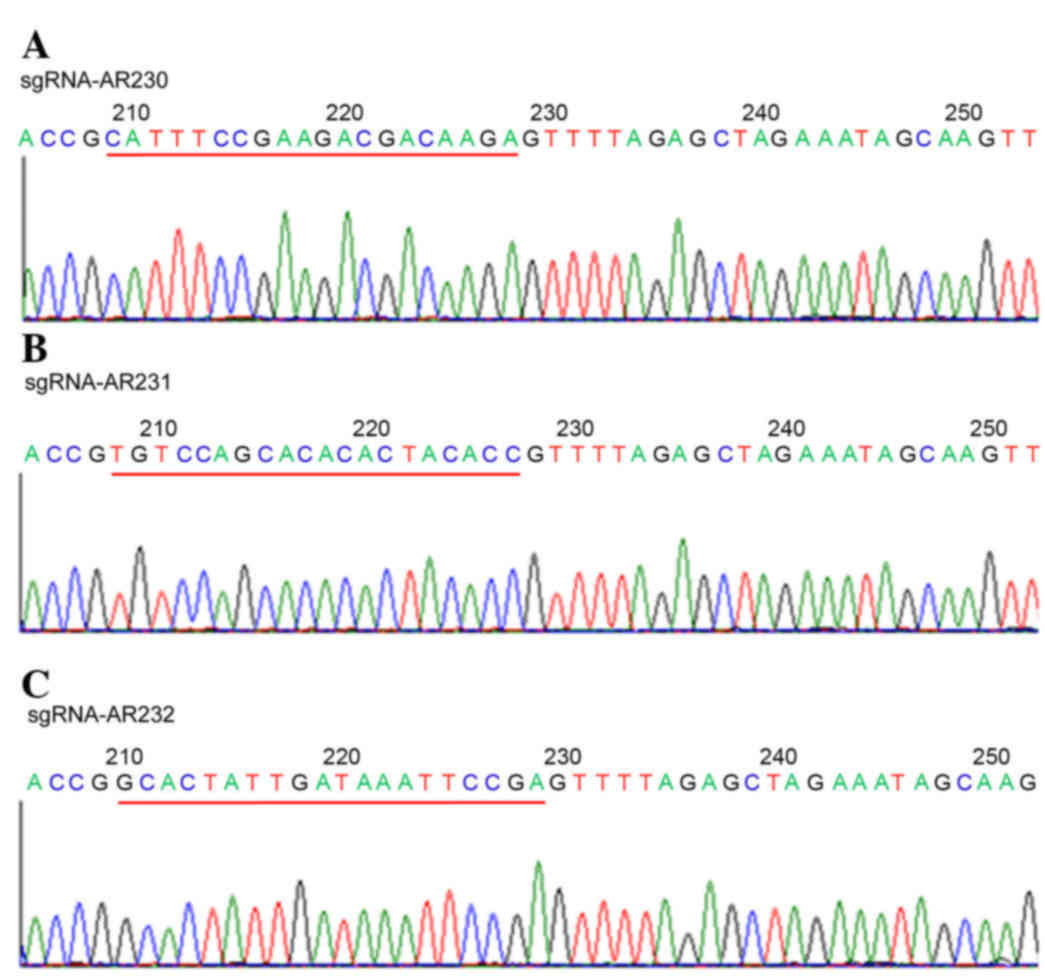
LV-sgRNA-AR231 and LV-sgRNA-AR232. Sequencing was performed using
the Sanger method (Shanghai GeneChem Co., Ltd.) to confirm the
sequence of the three sgRNAs. The sequences of the sgRNAs were as
follows: sgRNA-AR230, CATTTCCGAAGACGACAAGA; sgRNA-AR231,
TGTCCAGCACACACTACACC; and sgRNA-AR232, GCACTATTGATAAATTCCGA. The
three LV-sgRNA-ARs were transfected into the LNCaP cells to
eventually generate the LV-Cas9-sgRNA-AR stable cell lines, and 48
h following LV transfection, subsequent experiments were performed.
The LV transfection altered the growth of the LNCaP cells to a
certain degree.
Polymerase chain reaction (PCR)
detection of the mutation
The LV-Cas9-sgRNA-AR-infected and
LV-Cas9-control-infected LNCaP cells (LNCaP-Cas9-sgRNA-AR and
LNCaP-Cas9-control) were collected into Eppendorf tubes for the
detection of the mutations. The mutations were detected using the
Knockout and Mutation Detection kit (Shanghai GeneChem Co., Ltd.),
which may efficiently recognize and detect specific double-stranded
DNA mutations. DNA was extracted from the samples using a TIANamp
Genomic DNA kit (Tiangen Biotech Co., Ltd., Beijing, China). The
specific sequences of the three primers were as follows: sgRNA-230
sense, 5′-AAGACTGGGGGTATGATCACC-3′ and sgRNA-230 antisense,
5′-AGGCCAGTATCATTAAGTCCC-3′; sgRNA-231 sense,
5′-ATCCAAGGATATGCTAGGTTGG-3′ and sgRNA-231 antisense,
5′-GAGACTTGTAACAATCCCTCTC-3′; sgRNA-232 sense,
5′-AAGACTGGGGGTATGATCACC-3′ and sgRNA-232 antisense,
5′-AGGCCAGTATCATTAAGTCCC-3′.
The PCR reactions were performed according to the
manufacturer's protocol, using the following cycling conditions: 35
cycles of degradation at 95°C for 20 sec, annealing at 55°C for 20
sec, and extension at 72°C for 30 sec. The DNA product was digested
using a T7E1 enzyme digestion reaction system at 45°C for 20 min,
and a stop buffer was added to terminate the reaction. The samples
were stained with GelRed dye (Biotium, Inc., Freemont, CA, USA),
and detected following separation using 2% agarose gel
electrophoresis at 105 V for 30 min.
Cell proliferation assay
A Cell Counting kit (CCK)-8 assay (Dojindo Molecular
Technologies, Inc., Kumamoto, Japan) was used to assess the
proliferation of the transfected cells. The cells were seeded at an
initial density of 5×104 cells/ml in 96-well plates (5
wells/group) and incubated for 24, 48, 72, 96 and 120 h
post-transfection with LV-sgRNA-AR230. For the analysis, 10 µl
CCK-8 reagent was added to each well, and the plates were incubated
for 2 h at 37°C in a 5% CO2 incubator. The optical
density at 450 nm was then measured. All experiments were performed
in triplicate, and the mean results were calculated.
Measurement of cellular apoptosis and
the cell cycle
Cellular apoptosis was detected using an Annexin
V-allophycocyanin (APC)/7-amino-actinomycin D (7-ADD) kit
(MultiSciences Biotech Co., Ltd., Hangzhou, China). 7-AAD is a
nucleic-acid dye that is superior to propidium iodide (PI) for
multicoloured fluorescence analyses. In the present study, flow
cytometry was used to analyse cellular apoptosis. The
LNCaP-Ca9-sgRNA-AR230 and LNCaP-Cas9-control cell samples were
trypsinized and centrifuged at 300 × g for 5 min at room
temperature. Following two washes with PBS buffer at 300 × g for 5
min at room temperature, the cells were re-suspended with 500 µl 1X
binding buffer at a concentration of 1–3×106 cells/ml.
Subsequently, 5 µl Annexin V-APC and 10 µl 7-AAD was added to the
cells and incubated at room temperature in the dark for 15 min.
Using a CytoFLEX flow cytometer (Beckman Coulter, Inc., Brea, CA,
USA), the cells were divided into four regions (Q1-Q4). Region Q1
was representative of mechanical error; region Q2 was
representative of late apoptotic or necrotic cells; region Q3 was
representative of early apoptotic cells; and region Q4 was
representative of living cells. Data analysis was performed using
FlowJo software (version 7.6.1; BD Biosciences, Franklin Lakes, NJ,
USA).
For the cell cycle assay, the LNCaP-Cas9-sgRNA-AR230
and LNCaP-Cas9-control cell samples were washed twice with 1 ml
cold PBS buffer, and the cells were fixed in 70% ethanol for 12–24
h at −20°C. The cells were washed with cold PBS buffer and stained
with 500 µl PI at 37°C in the dark for 30 min. The analyses were
performed using a CytoFLEX flow cytometer (Beckman Coulter, Inc.).
The percentage of cellular apoptosis was calculated according to
the percentage of the sub-G1 peak in the cell cycle profile
following the PI staining (15).
Statistical analysis
All data are presented as the mean ± standard
deviation. The statistical analyses were performed using SPSS 19.0
software (IBM Corp., Armonk, NY, USA). Significant differences
between the groups were analysed using one-way analysis of variance
followed by a Newman-Keuls post hoc test and t-tests. P<0.05 was
considered to indicate a statistically significant difference.
Results
Construction of sgRNAs targeting the
AR gene
A total of three AR targets were selected in the
present study. All three sgRNAs were expressed using the following
lentiviral preparations: LV-sgRNA-AR230, LV-sgRNA-AR231, and
LV-sgRNA-AR232. These viral packaged sgRNAs were confirmed by
sequencing, as presented in Fig.
1.
Confirmation of gene editing activity
by PCR detection of mutations following viral infection
The viral preparations were used to infect LNCaP
cells that permanently express Cas9. The cells were harvested 7
days subsequent to the viral infection for DNA extraction and
mutation analysis. As presented in Fig. 2A, the expected sizes of the
PCR-amplified products were obtained for the three sgRNA-associated
CRISPR/Cas9 preparations as follows: 983, 702 and 983 bp for
sgRNA-AR230, sgRNA-AR231 and sgRNA-AR232, respectively.
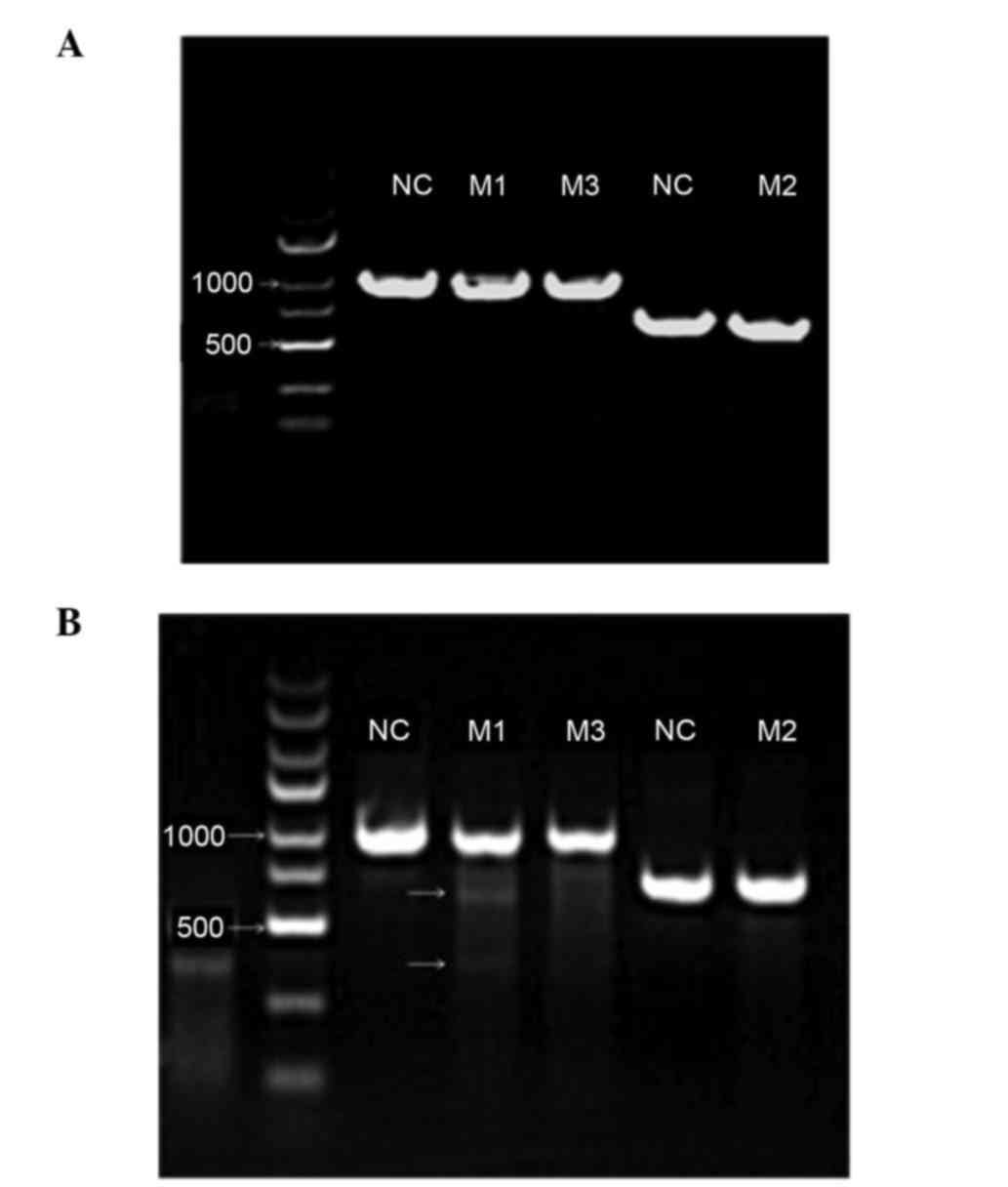 | Figure 2.Enzymatic electrophoresis of three
target sites determined using T7E1 endonuclease. (A) Images of
agarose gel demonstrating that the correct fragments of the three
different androgen receptor target sites relevant to LV-230, LV-231
and LV-232 were amplified. Lanes M1, M2, and M3 exhibit the
amplified products at the expected sizes (LV230, 983 bp; LV231, 702
bp; and LV-232, 983 bp). The two NC lanes exhibit the corresponding
products in the NC groups. The same primers were used in lanes M1
and M3 and, therefore, the NC was shared. (B) Agarose gel image
exhibiting the mismatch-associated short fragments in the M1 lane,
indicating the gene-editing effect of the single-guide
RNA-associated clustered regularly interspaced short palindromic
repeats-associated protein system. No such obvious gene editing
effect was detected in the M2 and M3 lanes. NC, negative
control. |
These three viral preparations exhibited different
gene editing effects in the LNCaP cells. Using T7E1 endonuclease
(Fig. 2B), gene editing was
observed in the cells infected with the virus expressing
sgRNA-AR230, and not in the cells infected with the other two
viruses. Therefore, the sgRNA-AR230 viral preparation was used in
the following cell proliferation and apoptosis analyses.
Effects of AR-knockout on tumour cell
proliferation
Since androgens may promote prostate cancer cell
proliferation via the AR, the present study assessed whether
prostate cancer cell proliferation was affected by knocking out AR.
At 24 h subsequent to the viral infection, green fluorescent
protein was observed in ~80% of the LNCaP cells. The CCK-8 assays
demonstrated that the cells in the control group
(LNCaP-Cas9-control) proliferated significantly more rapidly
compared with those in the experimental group
(LNCaP-Cas9-sgRNA-AR230). Notably, the partial knockout of AR using
the CRISPR/Cas9 system suppressed cell proliferation, and the most
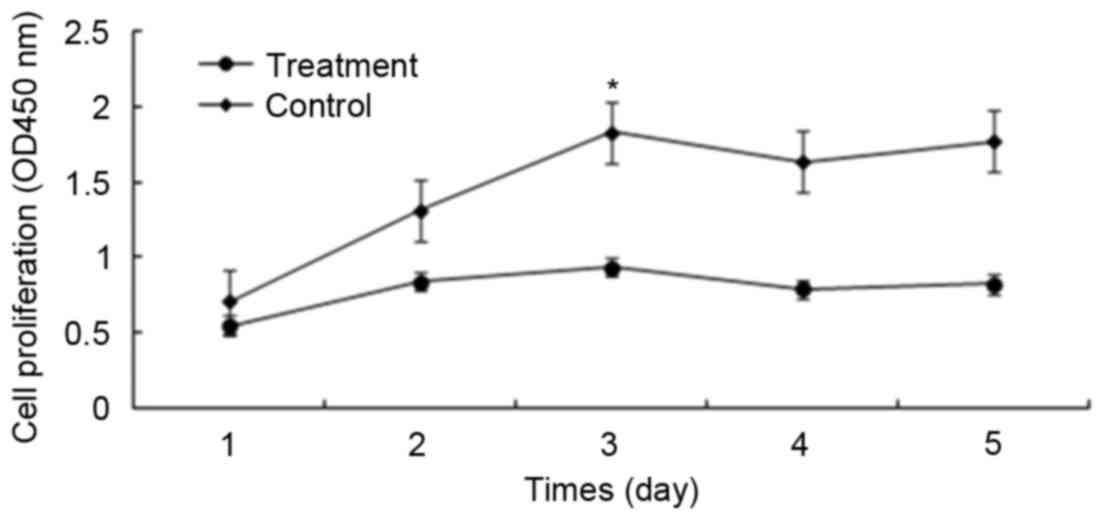
significant alterations were observed at 48–72 h. As presented in
Fig. 3, the difference in cell
proliferation between the two groups was significant (P<0.05).
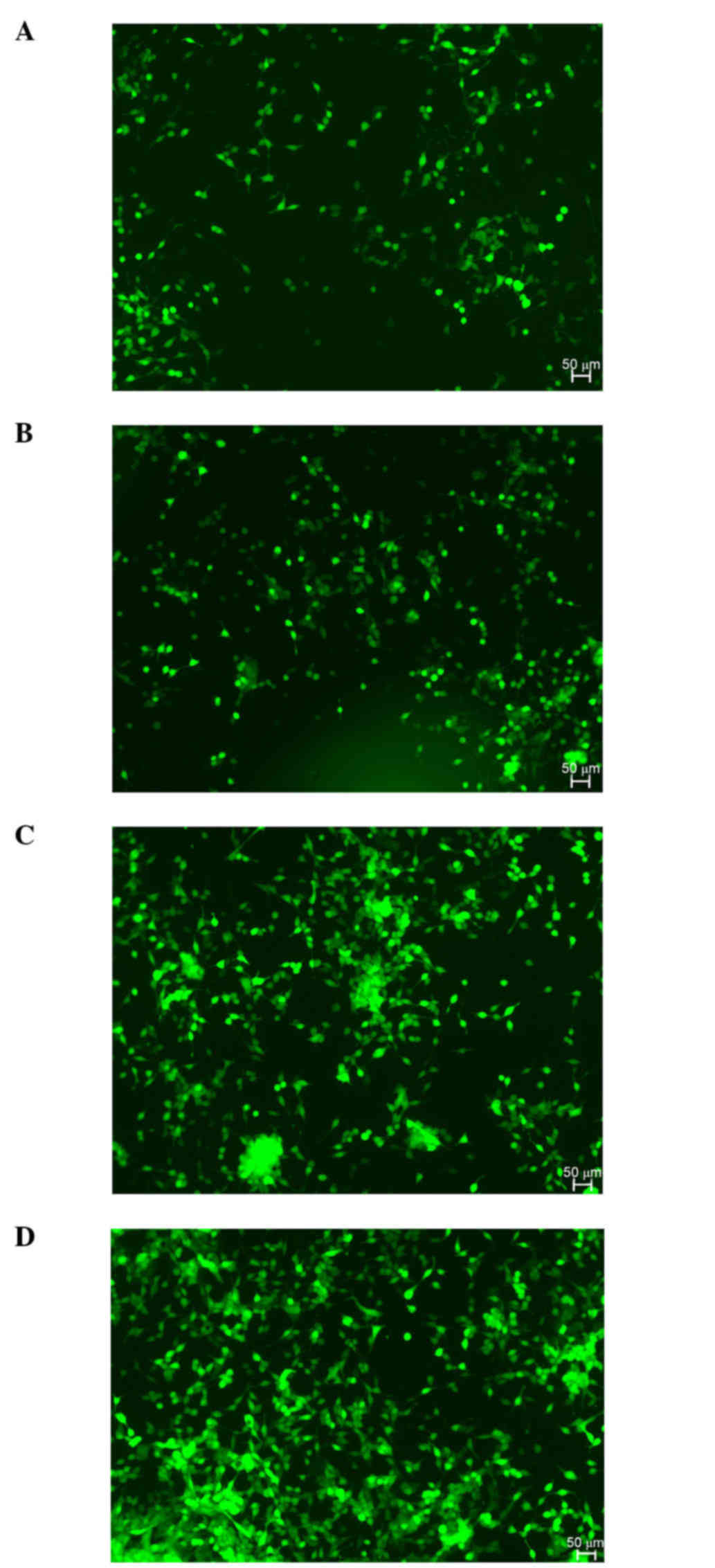
Fig. 4 illustrates the cell
proliferation in the experimental and control groups 24 and 72 h
post-transfection. The cell proliferation rate in the experimental
group was decreased compared with that in the control group 72 h
post-transfection.
Effects of AR knockdown on tumour cell
apoptosis
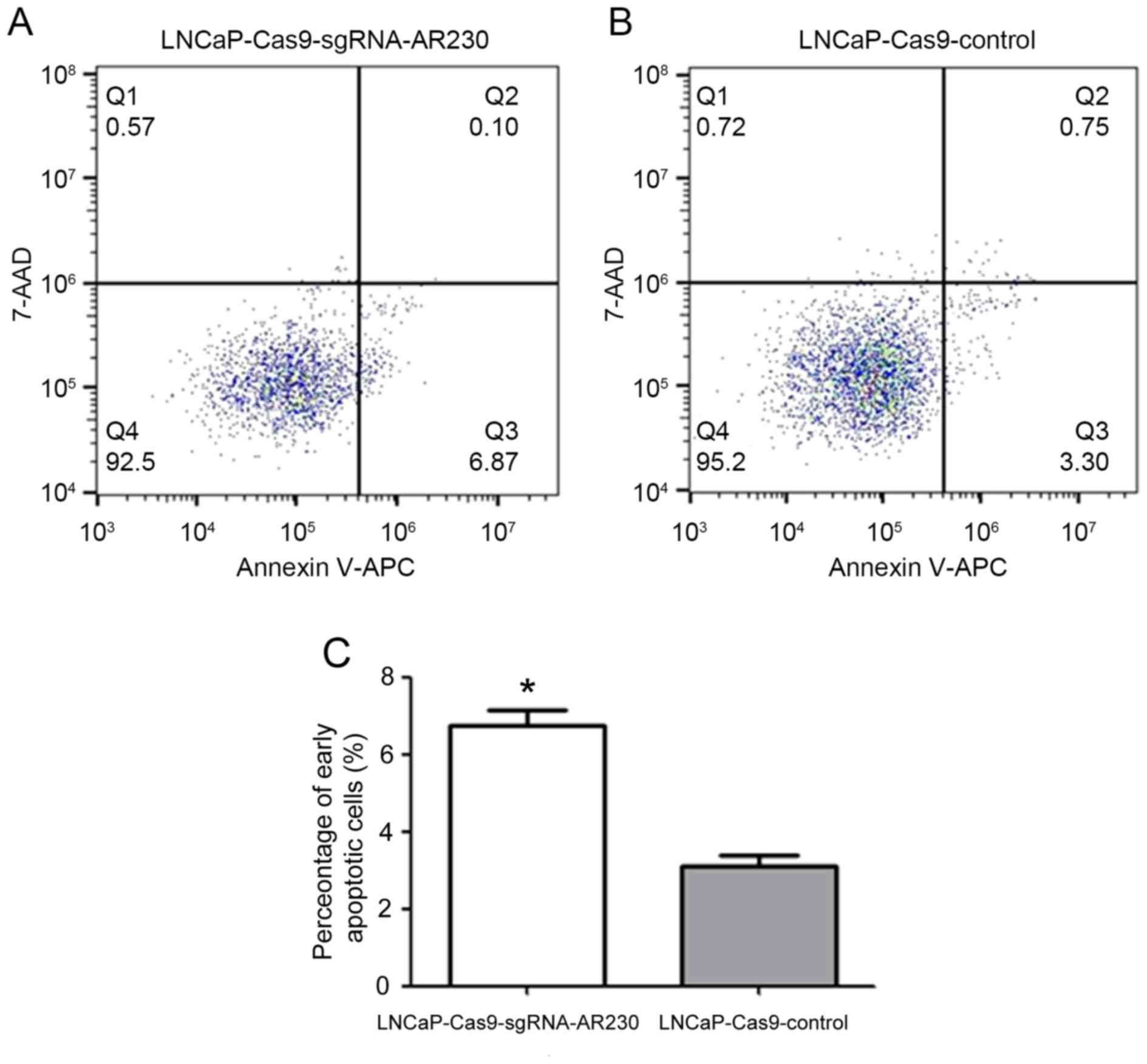
Cellular apoptosis was analysed by flow cytometry as
presented in Fig. 5. To
investigate whether the decreased cell proliferation was due to
cellular apoptosis, flow cytometry was used to evaluate the effect
of AR knockdown on cellular apoptosis. It was observed that the
proportion of early apoptosis (region Q3) in the
LNCaP-Cas9-sgRNA-AR 230-treated cells was increased compared with
that induced by the LNCaP-Cas9-control treatment (6.87% vs. 3.30%).
The difference between the two groups was statistically significant
(P<0.05) according to the results of a two-tailed Student's
t-test.
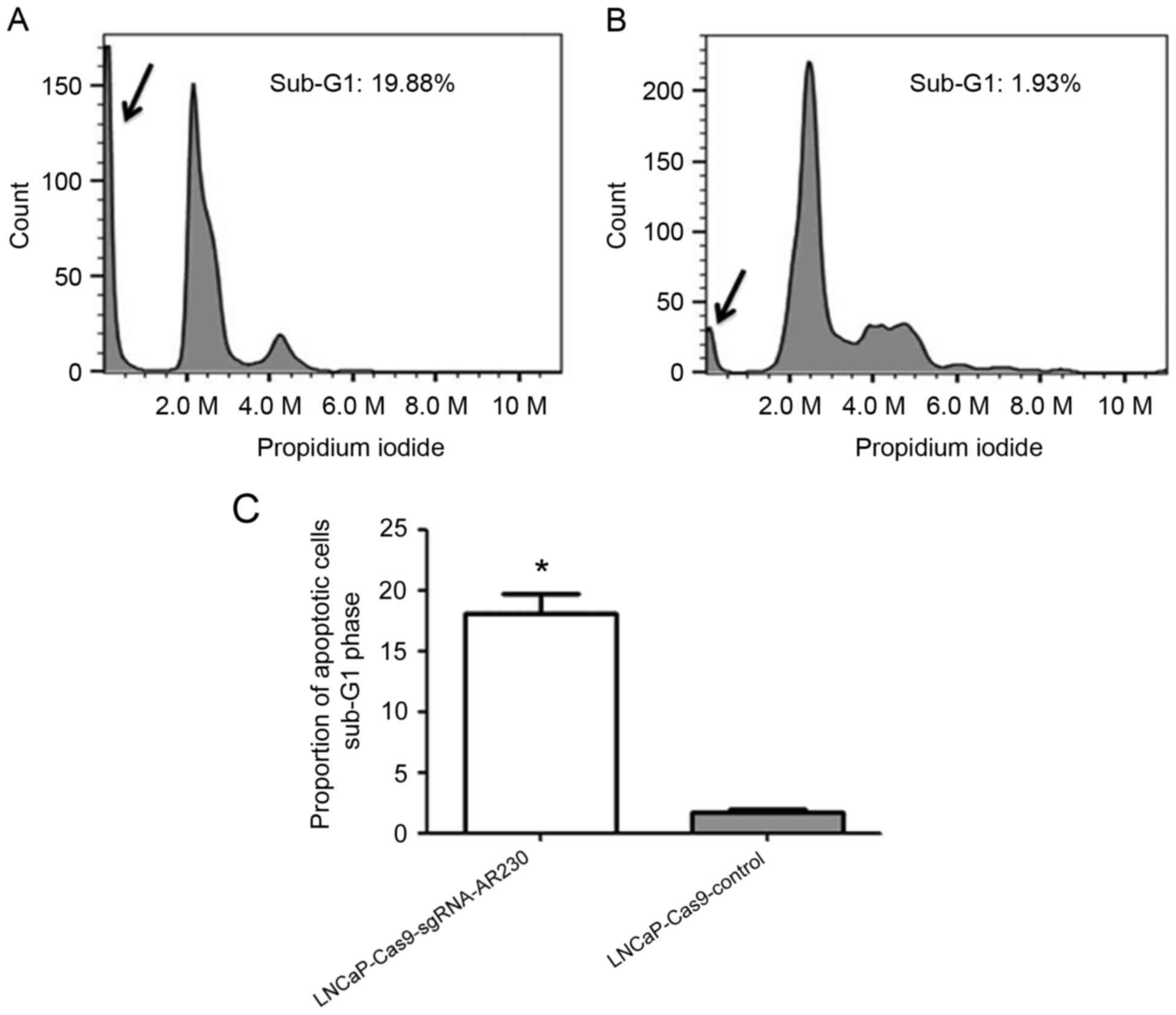
The sub-G1 peak in the cell cycle profile was
detected in the two groups, as presented in Fig. 6. The proportion of sub-G1 phase
cells was 19.88% in the LNCaP-Cas9-sgRNA-AR230 group and 1.93% in
the LNCaP-Cas9-control group, (Fig.
6). A significant increase in the number of cells in the sub-G1
phase was observed in the LNCaP-Cas9-sgRNA-AR230 group compared
with that in the LNCaP-Cas9-control group (P<0.05).
Discussion
The A-AR signalling pathway serves a significant
role during all stages of prostate cancer. During the early stages,
resecting the prostate gland may yield a positive therapeutic
outcome. During the middle stages of prostate carcinogenesis,
decreasing the levels of androgen via surgery or anti-androgenic
drugs may be effective. Following a certain period (~18 months),
prostate cancer frequently becomes insensitive to androgens and
continues to grow under androgen-depleted conditions. Nevertheless,
the AR continues to serve important roles during the androgen
refractory stage of prostate cancer via multiple mechanisms,
including gene mutations, gene amplification and the abnormal
activation of other relevant signalling pathways (16,17).
Consequently, editing the expression of ARs may be a reliable
treatment for prostate cancer.
The present study partially knocked out the AR using
gene-editing technology and observed its effects on the
proliferation and apoptosis of LNCaP cancer cells. The CRISPR/Cas9
system was used to successfully edit the AR gene. The partial AR
knockout significantly inhibited the growth of LNCaP cells by
inhibiting cell proliferation, and the blockade was implemented by
promoting cellular apoptosis. According to the cellular apoptosis
and cell cycle analysis using flow cytometry, apoptosis was
significantly increased in the experimental group compared with
that in the control group. This observation is consistent with
previous studies (18,19).
The CRISPR/Cas9 system has recently become a hot
topic in the field of gene editing. Compared with ZFN and TALEN
technologies, the CRISPR/Cas9 system has advantages in operability
and repeatability (20,21). The CRISPR/Cas9 system has been
widely used in studies investigating various tumours, including
cervical cancer, liver cancer, lymphoma and prostate cancer. The
present study used the CRISPR/Cas9 system to edit the AR gene,
leading to a partial knockout in androgen-dependent LNCaP cells.
Stable LNCaP cells containing sgRNA and Cas9 were constructed.
These cells represented a solid foundation for further studies. The
influence of AR gene editing by CRISPR/Cas9 was observed in the
LNCaP cells, although a statistically significant difference was
not noted.
However, the present study has certain limitations.
The scope of this study was limited to androgen-dependent prostate
cancer cells due to their hormone dependence. In future studies,
androgen-independent prostate cancer cells may be used to further
analyse the CRISPR/Cas9 system. The present study confirmed that
CRISPR/Cas9 was able to edit the AR gene, although the quantitative
accuracy of the knockout of the AR gene was not investigated. In
addition, the CRISPR/Cas9 system has off-target effects. Therefore,
future studies may investigate methods to avoid these off-target
effects.
In conclusion, the CRISPR/Cas9 system was able to
edit the expression of AR and restrain the growth of
androgen-dependent prostate cancer cells in vitro,
suggesting the potential of the CRISPR/Cas9 system in future cancer
therapy.
Acknowledgements
The authors were financially supported by the Suzhou
Science and Technology Bureau Development Plan (grant no.
SYS201475), the Jiangsu Provincial Special Program of Clinical
Medical Science (grant no. BL2014040), the Suzhou Clinical Special
Disease Diagnosis and Treatment Program (grant no. LCZX201406), the
Suzhou Science and Technology Development Plan (grant no.
SS201534), and the Suzhou Science and Technology Development Plan
Guidance Project (grant no. SYSD2015091). This project additionally
received the Second Affiliated Hospital of Soochow University
Preponderant Clinic Discipline Group project funding (grant no.
XKQ2015009).
References
|
1
|
DeSantis CE, Siegel RL, Sauer AG, Miller
KD, Fedewa SA, Alcaraz KI and Jemal A: Cancer statistics for
African Americans, 2016: Progress and opportunities in reducing
racial disparities. CA Cancer J Clin. 66:290–308. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Feldman BJ and Feldman D: The development
of androgen-independent prostate cancer. Nat Rev Cancer. 1:34–45.
2001. View
Article : Google Scholar : PubMed/NCBI
|
|
3
|
Saraon P, Jarvi K and Diamandis EP:
Molecular alterations during progression of prostate cancer to
androgen independence. Clin Chem. 57:1366–1375. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Yuan X and Balk SP: Mechanisms mediating
androgen receptor reactivation after castration. Urol Oncol.
27:36–41. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Cohen MB and Rokhlin OW: Mechanisms of
prostate cancer cell survival after inhibition of AR expression. J
Cell Biochem. 106:363–371. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Horii T and Hatada I: Genome engineering
using the CRISPR/Cas system. World J Med Genet. 4:69–76. 2014.
View Article : Google Scholar
|
|
7
|
Mali P, Yang L, Esvelt KM, Aach J, Guell
M, DiCarlo JE, Norville JE and Church GM: RNA-guided human genome
engineering via Cas9. Science. 339:823–826. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Pu J, Frescas D, Zhang B and Feng J:
Utilization of TALEN and CRISPR/Cas9 technologies for gene
targeting and modification. Exp Biol Med (Maywood). 240:1065–1070.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Seeger C and Sohn JA: Targeting hepatitis
B virus with CRISPR/Cas9. Mol Ther Nucleic Acids. 3:e2162014.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Lu XJ, Qi X, Zheng DH and Ji LJ: Modeling
cancer processes with CRISPR-Cas9. Trends Biotechnol. 33:317–319.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Tang H and Shrager JB: CRISPR/Cas-mediated
genome editing to treat EGFR-mutant lung cancer: A personalized
molecular surgical therapy. EMBO Mol Med. 8:83–85. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Matano M, Date S, Shimokawa M, Takano A,
Fujii M, Ohta Y, Watanabe T, Kanai T and Sato T: Modeling
colorectal cancer using CRISPR-Cas9-mediated engineering of human
intestinal organoids. Nat Med. 21:256–262. 2015.PubMed/NCBI
|
|
13
|
Wang G, Zhao N, Berkhout B and Das AT:
CRISPR-Cas9 can inhibit HIV-1 replication but NHEJ repair
facilitates virus escape. Mol Ther. 24:522–526. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kawamura N, Nimura K, Nagano H, Yamaguchi
S, Nonomura N and Kaneda Y: CRISPR/Cas9-mediated gene knockout of
NANOG and NANOGP8 decreases the malignant potential of prostate
cancer cells. Oncotarget. 6:22361–22374. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhang HT, Jun WU, Zhang HF and Zhu QF:
Efflux of potassium ion is an important reason of HL-60 cells
apoptosis induced by tachyplesin. Acta Pharmacol Sin. 27:1367–1374.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Heinlein CA and Chang C: Androgen receptor
in prostate cancer. Endocr Rev. 25:276–308. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Shore N: Management of early-stage
prostate cancer. Am J Manag Care. 20 12 Suppl:S260–S272.
2014.PubMed/NCBI
|
|
18
|
Yu L, Wang X, Zhu D, Ding W, Wang L, Zhang
C, Jiang X, Shen H, Liao S, Ma D, et al: Disruption of human
papillomavirus 16 E6 gene by clustered regularly interspaced short
palindromic repeat/Cas system in human cervical cancer cells. Onco
Targets Ther. 8:37–44. 2014.PubMed/NCBI
|
|
19
|
Hu Z, Yu L, Zhu D, Ding W, Wang X, Zhang
C, Wang L, Jiang X, Shen H, He D, et al: Disruption of HPV16-E7 by
CRISPR/Cas system induces apoptosis and growth inhibition in HPV16
positive human cervical cancer cells. Biomed Res Int 2014.
6128232014.
|
|
20
|
Ding Q, Regan SN, Xia Y, Oostrom LA, Cowan
CA and Musunuru K: Enhanced efficiency of human pluripotent stem
cell genome editing through replacing TALENs with CRISPRs. Cell
Stem Cell. 12:393–394. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Pattanayak V, Guilinger JP and Liu DR:
Determining the specificities of TALENs, Cas9, and other genome
editing enzymes. Methods Enzymol. 546:47–78. 2014. View Article : Google Scholar : PubMed/NCBI
|