Introduction
Diabetic retinopathy (DR), one of the most common
complications of diabetes, is the leading cause of blindness and
visual impairment in the working population (25–65 years old) in
developing countries (1). As a
kind of fundus lesions with specific changes, the pathology of DR
is retinal micro-vascular dysfunction, and its clinical features
are mainly manifestations of capillary periocular cell reduction,
basement membrane thickening, abnormal proliferation of endothelial
cells and angiogenesis, leading to macular edema, vitreous
hemorrhage, retinal detachment and other serious injuries (2). At present, the treatment of DR is
mainly through retinal photocoagulation and vitreous surgery, but
the visual function of most patients has been already irreversibly
damaged before the treatment. So far, DR is still lack of effective
prevention and treatment. Therefore, seeking for new effective
diagnosis and treatment methods has been a hot issue in the study
of DR.
MicroRNAs (MiRNAs), a class of non-coding
single-stranded small RNA, about 18–24 nucleotides in length, can
transcriptionally or post-transcriptionally regulate the expression
of the target genes by specific binding to the 3′-UTR of target
mRNAs (3–5). MiRNAs play an important regulatory
role in the process of visual function formation via involving in
regulation of the physiological processes such as ocular
development, neural differentiation, functional maintenance and
apoptosis. Kovacs et al (6), have confirmed for the first time that
the miRNAs expression profile has changed during DR. MiR-29 has
been found to play important roles in the process of high glucose
(HG) induced apoptosis in RPE cells, thus involved in DR progress
(7). MiR-20b plays critical role
in DR by targeting protein kinase B-3 (AKT3) (8). MiRNA-29a could down-regulate the
expression of angiotensinogen (AGT), thereby repressing the
development of DR (9). Moreover,
miR-200b has been reported might alleviate DR development via the
down-regulation of its target gene vascular endothelial growth
factor A (VEGFA) (10).
MiR-219-5p, a tumor suppressor in a variety of
cancers, may play an important role in regulating cell growth and
apoptosis (11–14). So far, the expression and role of
miR-219-5p in DR remain unclear. Therefore, the present study aimed
to investigate the role of miR-219-5p in the development of DR, and
to explore the underlying molecular mechanism.
Materials and methods
Cell culture
The human retinal pigment epithelial (RPE) cell line
ARPE-19 was obtained from the American Type Culture Collection
(ATCC, Manassas, VA). Cells were cultured in DMEM/F-12 medium
(Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA)
containing HG (25 mM D-glucose) or normal glucose (control, 5 mM
D-glucose) (15,16), 10% fetal bovine serum (FBS; Gibco;
Thermo Fisher Scientific, Inc.), 2 mM L-glutamine (Sigma-Aldrich;
Merck KGaA, Darmstadt, Germany), 100 µg/ml streptomycin, and 100
units/ml penicillin (Sigma-Aldrich; Merck KGaA), and incubated at
37°C with 5% CO2. The cells were passaged until reaching
80~90% confluence.
Cell transfection
The day before cell transfection, ARPE-19 cells were
seeded into a 6-well plate at the concentration of 5×104
cells per well and cultured in a incubator at 37°C with 5%
CO2. Then the ARPE-19 cells were transfected with
miR-219-5p inhibitor, its negative control, or miR-219-5p
inhibitor+LRH-1-siRNA with 30 µl Lipofectamine 2000 (Invitrogen;
Thermo Fisher Scientific, Inc.) according to the manufacturer's
protocol. Cells without any treatment were recognized as the
control group. These cells were initially transfected for 6 h, then
the culture medium was replaced with DMEM/F-12 medium containing 50
mM HG (7). After incubation for 24
h, the cells were subjected to following experiments.
Cell viability assay
Twenty four hours after treatment, cell viability
assay was used to measure the cell viability. Briefly, ARPE-19
cells were trypsinized, re-suspended, and then reseeded into
96-well plates and incubated at 37°C for 24, 48 and 72 h
respectively. Subsequently, 20 mg/ml MTT solution (Sigma-Aldrich;
Merck KGaA) was added into each well and then incubated for another
4 h. At the end of the experiment, the optical density (OD) value
at 570 nm was detected by using a SynergyTM 2 Multi-function
Microplate Reader (Bio-Tek Instruments, Inc., Winooski, VT, USA).
Tests were repeated at least for three times.
Cell apoptosis assay
To analyze cell apoptosis, cell apoptosis assay was
performed using flow cytometry (BD Biosciences, Franklin Lakes, NJ,
USA). 24 h after treatment, ARPE-19 cells were stained with annexin
V-FITC and propidium iodide (PI) (CST Biological Reagents Co.,
Ltd., Shanghai, China) in line with the manufacturer's
instructions. Finally, we used flow cytometry to analyze cell
apoptosis. Tests were repeated at least for three times.
RNA extraction and quantitative
RT-PCR
Total RNA was extracted from ARPE-19 cells by using
TRIZOL reagent (Invitrogen; Thermo Fisher Scientific, Inc.)
following the manufacturer's protocol. The total RNA was reversely
transcripted into cDNA using a PrimeScript reverse transcription
reagent kit (Takara Biotechnology Co., Ltd., Dalian, China).
Subsequently, TaqMan Universal PCR Master Mix kit (Thermo Fisher
Scientific, Inc.) was performed to analyze the synthesized cDNAs.
The amplification conditions were as follows: 95°C for 10 min,
followed by 40 cycles of 95°C for 10 sec and 57°C for 60 sec. All
primer sequences were obtained from Genscript (Nanjing, China), and
the relative gene expression was analyzed by using the
2−ΔΔCq method (17).
Western blot analysis
After treatment for 24 h, ARPE-19 cells were
harvested by microcentrifugation. Total cell protein was extracted
using the lysis buffer (Promega Corporation, Madison, WI, USA).
Equal amount of the samples were separated by SDS-polyacrylamide
gel electrophoresis and then transferred onto an enhanced
nitrocellulose membrane. Blots were blocked with 5% skim milk at
room temperature for 2 h, incubated with the primary antibody
against LRH-1 (cat no. 12800), β-catenin (cat no. 8480), Cyclin D1
(cat no. 2978), c-Myc (cat no. 13978) (all dilution: 1:1,000) and
β-actin (cat no. 4970; dilution: 1:5,000) (dilution: 1:1,000; all
from Cell Signaling Technology, Danvers, MA, USA) at 4°C overnight,
and then incubated with a secondary antibody at room temperature
for 2 h. At the end of the test, blots were observed by using the
enhanced chemiluminescence detection system (Super Signal West Dura
Extended Duration Substrate; Pierce; Thermo Fisher Scientific,
Inc.).
Dual-luciferase reporter assay
We used bioinformatics software (http://www.microrna.Org/microrna/home.do) to predict
the promising targets of miR-219-5p, and we found that LRH-1 was
potentially targeted by miR-219-5p. In order to confirm our
prediction, dual-luciferase reporter assay was applied. In brief,
the 3′UTR of LRH-1 gene, which includes the miR-219-5p binding
domain, was sub-cloned into the XhoI and NotI sites of a psiCHECK2
luciferase vector (Promega Corporation) (LRH-1 WT). To point-mutate
the miR-219-5p binding domain on the 3′UTR of LRH-1, a QuikChange
Site-Directed Mutagenesis kit (Agilent Technologies, Inc., Santa
Clara, CA, USA) was performed according to the manufacturer's
instructions. Then the mutated LRH-1 3′UTR was sub-cloned into
psiCHECK2 luciferase vector (LRH-1 MUT). ARPE-19 cells were
co-transfected with LRH-1 WT or LRH-1 MUT and miR-219-5p mimic or
miR-control (control of miR-219-5p mimic). 48 h later, the
dual-luciferase reporter assay system (Promega Corporation) was
carried out to determine the luciferase activity.
Statistical analysis
SPSS v16.0 statistical software (SPSS, Inc.,
Chicago, IL, United States) was applied for all statistical
analyses. Data were presented as the mean ± SD. Comparison between
two groups were made by Student's t-test, between multiple groups
by one-way ANOVA followed by Tukey's test. P<0.05 was considered
to indicate a statistically significant difference.
Results
MiR-219-5p was up-regulated in HG
treated ARPE-19 cells
In current study, we firstly treated ARPE-19 cells
with 50 mM D-glucose or 0 mM D-glucose (control). Consistent with a
previous study, significant apoptosis in ARPE-19 cells was induced
after treatment with 50 mM D-glucose, as compared to the control
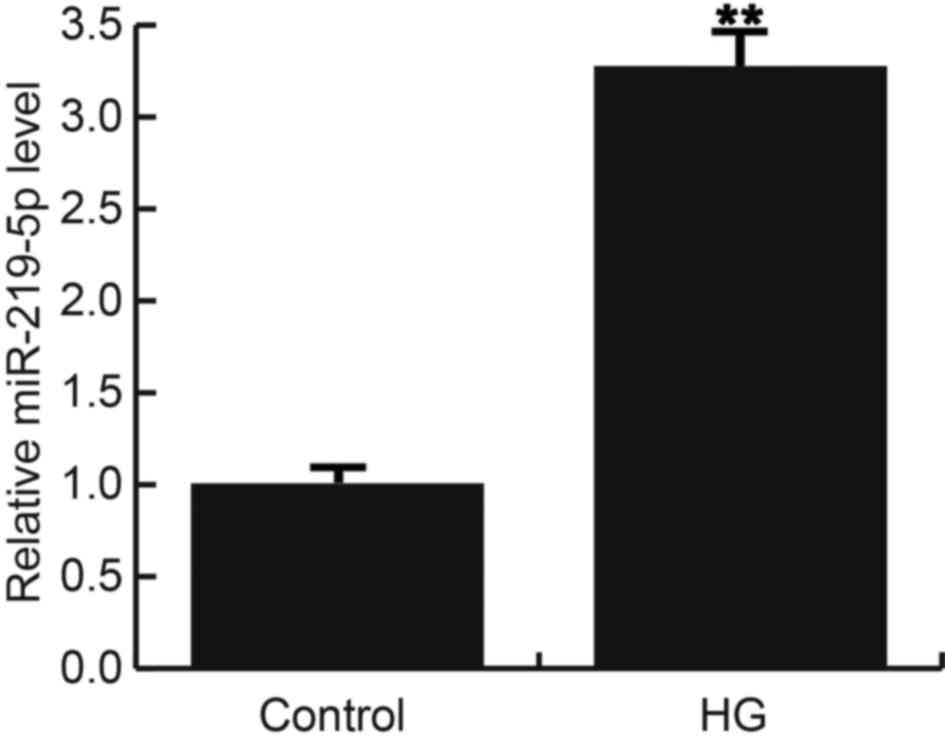
group with 0 mM D-glucose. 24 h after treatment, miR-219-5p was
detected using qRT-PCR, and the results indicated that compared
with the control cells, miR-219-5p expression was significantly
up-regulated in HG treated ARPE-19 cells (Fig. 1).
MiR-219-5p directly targets LRH-1
Bioinformatics software (http://www.microrna.Org/microrna/home.do) was
performed to predict the targets of miR-219-5p. About 4000 targets
were found to be potentially targeted by miR-219-5p, including
LRH-1 (Fig. 2A). LRH-1 has been
revealed to play an important role in the regulation of cell
proliferation and apoptosis via controlling the Wnt/β-catenin
signaling pathways (12,18). However, whether LRH-1 functions its
role in DR is still unclear. Therefore, we choose LRH-1 to make
further investigation. And the results of dual-luciferase reporter
assay indicated that miR-219-5p directly targets LRH-1 at 3′UTR
domain (Fig. 2B). And the lower
expression of LRH-1 in HG treated ARPE-19 cells was observed in the
current study (Fig. 2C and D).
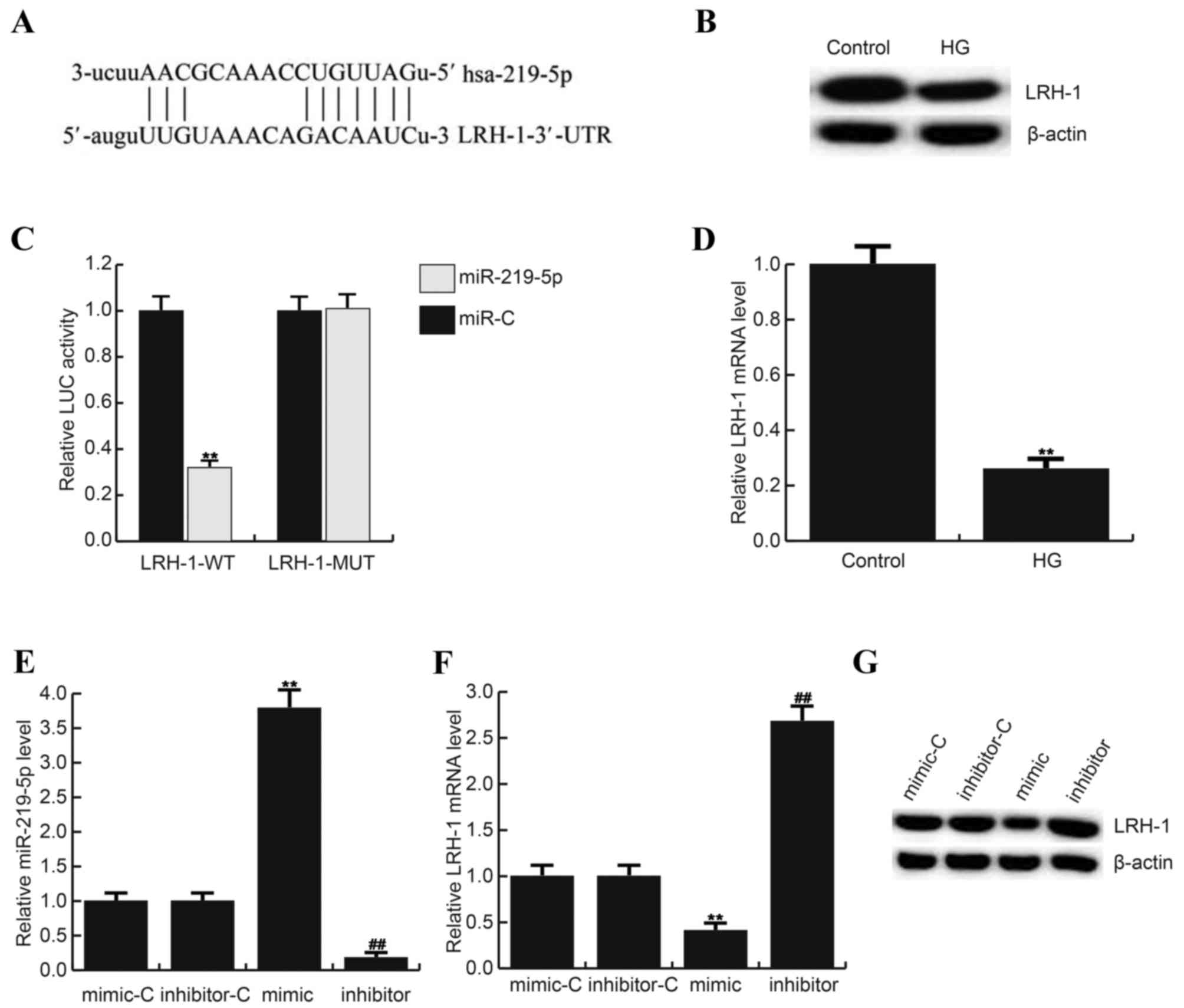 | Figure 2.LRH-1 is a direct target of
miR-219-5p. (A) Bioinformatics software (http://www.microrna.Org/microrna/home.do) was carried
out to predict the interaction between miR-219-5p and LRH-1 3′UTR.
(B) Luciferase activity was detected by Dual luciferase assay.
Here, (C) protein level of LRH-1 in ARPE-19 cells after treatment
with 0 mM D-glucose (Control) or 50 mM D-glucose. D: mRNA level of
LRH-1 in ARPE-19 cells after treatment with 0 mM D-glucose
(Control) or 50 mM D-glucose. (D) Effect of miR-219-5p inhibitor on
LRH-1 mRNA expression in ARPE-19 cells. Relative expression of (E)
miR-219-5p and (F) LRH-1 in different groups. (G) Effect of
miR-219-5p inhibitor on LRH-1 protein expression in in ARPE-19
cells. **P<0.01 vs Control; ##P<0.01 vs. Control.
Data are displayed as mean ± standard deviation. UTR, untranslated
region; HG, high glucose, cells treated with 50 mM D-glucose;
inhibitor, cells transfected with miR-219-5p inhibitors and treated
with 50 mM D-glucose; inhibitor+si-LRH-1, cells co-transfected with
miR-219-5p inhibitors and LRH-1-siRNA, and treated with 50 mM
D-glucose. miR, microRNA; LRH, liver receptor homolog; MUT, LRH-1
3′ UTR with a mutation in the miR-219-5p binding site; WT, wild
type; LUC, luciferase. |
Then we determined whether miR-219-5p regulates
LRH-1 expression in ARPE-19 cells, miR-219-5p mimic, mimic control,
miR-219-5p inhibitor or inhibitor control was transfected into
ARPE-19 cells, and the transfection efficiency was assessed by
qRT-PCR (Fig. 2E). The findings
suggested that miR-219-5p mimic significantly decreased both mRNA
(Fig. 2F) and protein (Fig. 2G) level of LRH-1 (55 kD) in ARPE-19
cells, while miR-219-5p inhibitor presented the opposite
effects.
MiR-219-5p inhibitor promoted ARPE-19
cell viability
Influence of miR-219-5p inhibitor on ARPE-19 cell
activity was detected using MTT assay. Compared with the control
group, the viability of ARPE-19 cell seriously reduced in the HG
group, and the declined cell viability was rescued by miR-219-5p
inhibitor. In addition, LRH-1-siRNA eliminated the enhanced cell
viability caused by miR-219-5p inhibitor (Fig. 3).
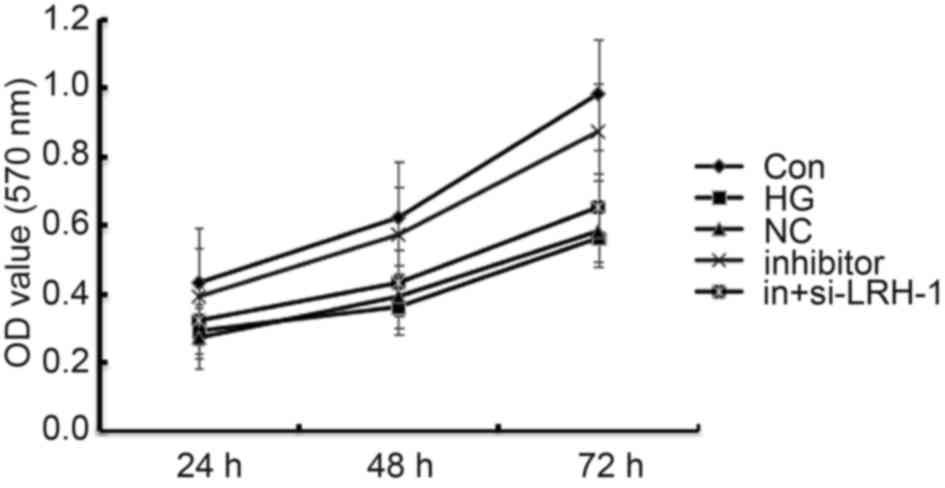 | Figure 3.Effects of miR-219-5p inhibitor on
ARPE-19 cell viability. ARPE-19 cell viability was measured by an
MTT assay 24, 48 and 72 h post treatment. Data are displayed as the
mean ± standard deviation. Con, control group; HG, high glucose,
cells treated with 50 mM D-glucose; NC, negative control, cells
transfected with the negative control of miR-219-5p inhibitor and
treated with 50 mM D-glucose; inhibitor, cells transfected with
miR-219-5p inhibitors and treated with 50 mM D-glucose;
in+si-LRH-1, cells co-transfected with miR-219-5p inhibitors and
liver receptor homolog-1-siRNA, and treated with 50 mM D-glucose;
OD, optical density. |
MiR-219-5p inhibitor repressed HG
induced ARPE-19 cell apoptosis
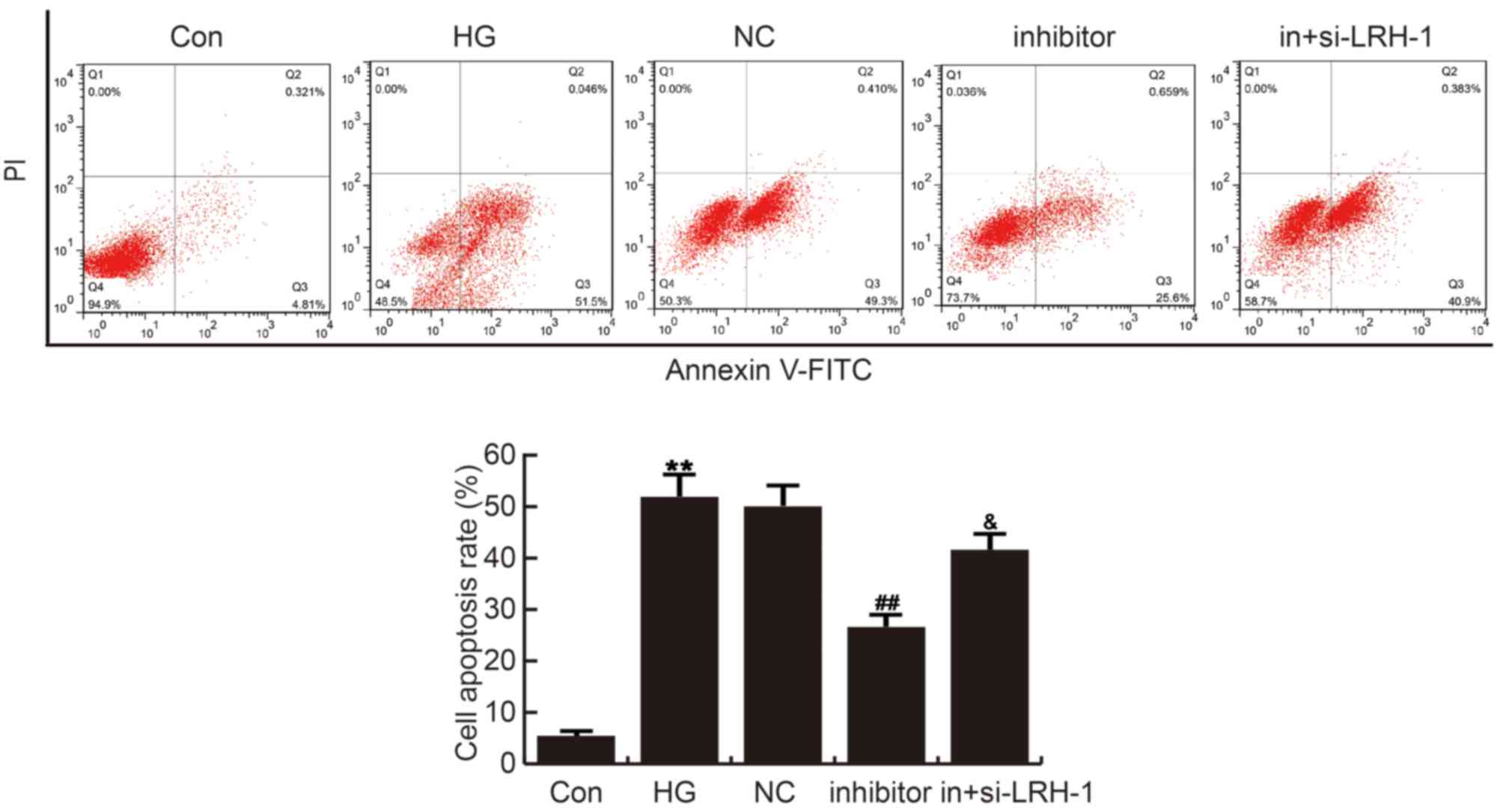
The effects of miR-219-5p inhibitor on ARPE-19 cell
apoptosis was analyzed by cell apoptosis assay with FCM. Compared
with the control group, ARPE-19 cell apoptosis was significantly
induced in HG group, which was repressed by miR-219-5p inhibitor
administration. These changes caused by miR-219-5p inhibitor were
eliminated by LRH-1-siRNA (Fig.
4).
MiR-219-5p inhibitor rescued the
LRH-1/Wnt/β-Catenin inhibition caused by HG
Finally, in order to explore the underlying
molecular mechanism of the role miR-219-5p played in DR process,
LRH-1/Wnt/β-Catenin pathway was analyzed, and western blotting and
qRT-PCR was performed respectively. As shown in Fig. 5, compared with the control group,
the level of LRH-1 (55 kD), β-Catenin (92 kD), Cyclin D1 (36 kD)
and c-Myc (62 kD) significantly reduced in HG treated ARPE-19
cells, and miR-219-5p inhibitor markedly enhanced LRH-1, β-Catenin,
Cyclin D1 and c-Myc expression when compared to the HG group. LRH-1
gene silencing eliminated the effects caused by miR-219-5p
inhibitor (Fig. 5).
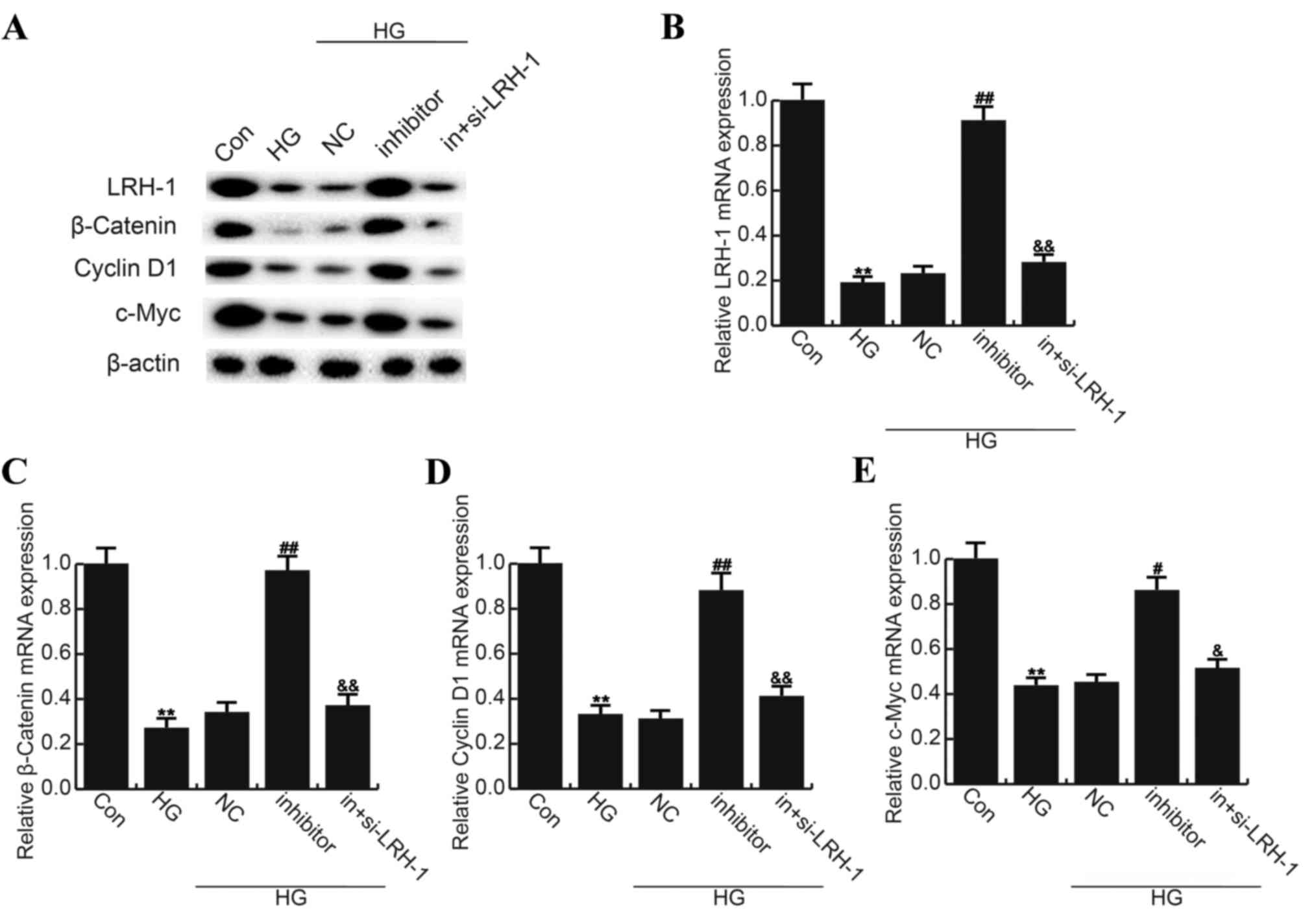 | Figure 5.Effects of miR-219-5p inhibitor on
Wnt/β-Catenin signaling pathway. (A) At 24 h post treatment, the
protein expression of LRH-1, β-Catenin, Cyclin D1 and c-Myc was
determined by western blot analysis. The mRNA level of (B) LRH-1,
(C) β-Catenin, (D) Cyclin D1 and (E) c-Myc was determined by
reverse transcription-quantitative polymerase chain reaction. Data
are displayed as the mean ± standard deviation. **P<0.01 vs.
Con; #P<0.05 and ##P<0.01 vs. HG;
&P<0.05 and &&P<0.01 vs.
inhibitor. Con, control group; HG, high glucose, cells treated with
50 mM D-glucose; NC, negative control, cells transfected with the
negative control of miR-219-5p inhibitor and treated with 50 mM
D-glucose; inhibitor, cells transfected with miR-219-5p inhibitors
and treated with 50 mM D-glucose; in+si-LRH-1, cells co-transfected
with miR-219-5p inhibitors and LRH-1-siRNA, and treated with 50 mM
D-glucose; LRH, liver receptor homolog. |
Discussion
Irreversible damage of the blood-retina barrier
(BRB) may caused by hyperglycemia or HG, resulting in loss of
vision or blindness, during DR (19). Hyperglycemia or HG induced
apoptosis in RPE cells, the essential elements of BRB, is a
characteristic process in DR (20). Increasing evidences have suggested
a regulatory function and the potential influences of microRNAs in
DR treatment. MiR-219-5p has been reported as a tumor suppressor in
a large number of cancers via the regulation of cell migration,
invasion, proliferation and apoptosis (11–14,21).
However, the regulation of cell apoptosis by miR-219-5p on human
RPE cells remains unclear. The present study aimed to investigate
the expression of miR-219-5p in human RPE cells, and to further
explore its role in regulating human RPE cell apoptosis under HG
conditions.
According a precious study, 50 mM D-glucose could
significantly induced human RPE cell apoptosis (7). Firstly, we examined the level of
miR-219-5p in human RPE cells after treatment with 50 mM D-glucose
for 24 h, and the findings suggested that miR-219-5p was
significantly up-regulated, indicating miR-219-5p may be involved
in the process of DR. Then we found that about 4000 targets may be
targeted by miR-219-5p, including LRH-1. LRH-1 (liver receptor
homologue-1), also called NR5A2, is a member of the nuclear
receptor subfamily which is involved in various biological
processes, including differentiation, steroid synthesis, and
bile-acid homeostasis, etc (22–24).
Recently, a large number of studies have revealed that LRH-1 plays
a key role in regulating cell growth, including apoptosis (12,18,25–28).
Due to its unclear role in DR, LRH-1 was further explored in the
present study. Our results indicated that LRH-1 was a direct target
of miR-219-5p, and LRH-1 was lower expressed in HG treated human
RPE cells compared to the control cells. Moreover, our findings
showed that LRH-1 was negatively regulated by miR-219-5p in human
RPE cells. These findings suggested that miR-219-5p may participate
in the development of DR by targeting LRH-1.
To study the role and molecular mechanism of
miR-219-5p in DR, we determined the effect of miR-219-5p inhibitor
on human RPE cell viability and cell apoptosis, and we found that
miR-219-5p inhibitor notably promoted human RPE cell viability
which was inhibited by HG treatment, and repressed HG induced human
RPE cell apoptosis. LRH-1/Wnt/β-Catenin pathway was inhibited in
human RPE cells after HG treatment, and this inhibition was rescued
by miR-219-5p inhibition. In addition, we revealed that the effects
of miR-219-5p inhibitor on human RPE cells could be eliminated by
LRH-1-siRNA under HG condition.
In summary, our study demonstrated that HG resulted
in the up-regulation of miR-219-5p expression in human RPE cells.
MiR-219-5p down-regulation promoted human RPE cell viability and
inhibited cell apoptosis through regulating LRH-1/Wnt/β-Catenin
pathway by targeting LRH-1. MiR-219-5p/LRH-1 may be promising
therapeutic targets for DR.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ding J and Wong TY: Current epidemiology
of diabetic retinopathy and diabetic macular edema. Curr Diab Rep.
12:346–354. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Yau JW, Rogers SL, Kawasaki R, Lamoureux
EL, Kowalski JW, Bek T, Chen SJ, Dekker JM, Fletcher A, Grauslund
J, et al: Global prevalence and major risk facers of diabetic
retinopathy. Diabetes Care. 35:556–564. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kim VN, Han J and Siomi MC: Biogenesis of
small RNAs in animals. Nat Rev Mol Cell Biol. 10:126–139. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Bartel DP: MicroRNAs: Target recognition
and regulatory functions. Cell. 136:215–233. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Valencia-Sanchez MA, Liu J, Hannon GJ and
Parker R: Control of translation and mRNA degradation by miRNAs and
siRNAs. Genes Dev. 20:515–524. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kovacs B, Lumayag S, Cowan C and Xu S:
MicroRNAs in early diabetic retinopathy in streptozotocin-induced
diabetic rats. Invest Ophthalmol Vis Sci. 52:4402–4409. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lin X, Zhou X, Liu D, Yun L, Zhang L, Chen
X, Chai Q and Li L: MicroRNA-29 regulates high-glucose-induced
apoptosis in human retinal pigment epithelial cells through PTEN.
In Vitro Cell Dev Biol Anim. 52:419–426. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Qin B, Liu J, Liu S, Li B and Ren J:
MiR-20b targets AKT3 and modulates vascular endothelial growth
factor-mediated changes in diabetic retinopathy. Acta Biochim
Biophys Sin (Shanghai). 48:732–740. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhang LQ, Cui H, Wang L, Fang X and Su S:
Role of microRNA-29a in the development of diabetic retinopathy by
targeting AGT gene in a rat model. Exp Mol Pathol. 102:296–302.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li EH, Huang QZ, Li GC, Xiang ZY and Zhang
X: Effects of miRNA-200b on the development of diabetic retinopathy
by targeting VEGFA gene. Biosci Rep. 37:pii: BSR20160572. 2017.
View Article : Google Scholar
|
|
11
|
Wang Q, Zhu L, Jiang Y, Xu J, Wang F and
He Z: miR-219-5p suppresses the proliferation and invasion of
colorectal cancer cells by targeting calcyphosin. Oncol Lett.
13:1319–1324. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Li C, Dong J, Han Z and Zhang K:
MicroRNA-219-5p represses the proliferation, migration, and
invasion of gastric cancer cells by targeting the
LRH-1/Wnt/β-catenin signaling pathway. Oncol Res. 25:617–627. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Huang C, Cai Z, Huang M, Mao C, Zhang Q,
Lin Y, Zhang X, Tang B, Chen Y, Wang X, et al: miR-219-5p modulates
cell growth of papillary thyroid carcinoma by targeting estrogen
receptor α. J Clin Endocrinol Metab. 100:E204–E213. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Huang N, Lin J, Ruan J, Su N, Qing R, Liu
F, He B, Lv C, Zheng D and Luo R: MiR-219-5p inhibits
hepatocellular carcinoma cell proliferation by targeting
glypican-3. FEBS Lett. 586:884–891. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ruiz MA, Feng B and Chakrabarti S:
Polycomb repressive complex 2 regulates MiR-200b in retinal
endothelial cells: Potential relevance in diabetic retinopathy.
PLoS One. 10:e01239872015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhao S, Li T, Li J, Lu Q, Han C, Wang N,
Qiu Q, Cao H, Xu X, Chen H and Zheng Z: miR-23b 3p induces the
cellular metabolic memory of high glucose in diabetic retinopathy
through a SIRT1-dependent signalling pathway. Diabetologia.
59:644–654. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhai G, Song J, Shu T, Yan J, Jin X, He J
and Yin Z: LRH-1 senses signaling from phosphatidylcholine to
regulate the expansion growth of digestive organs via synergy with
Wnt/β-catenin signaling in zebrafish. J Genet Genomics. 44:307–317.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ahsan H: Diabetic retinopathy-biomolecules
and multiple pathophysiology. Diabetes Metab Syndr. 9:51–54. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Simó R, Villarroel M, Corraliza L,
Hernández C and Garcia-Ramírez M: The retinal pigment epithelium:
Something more than a constituent of the blood-retinal
barrier-implications for the pathogenesis of diabetic retinopathy.
J Biomed Biotechnol. 2010:1907242010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Cheng J, Deng R, Zhang P, Wu C, Wu K, Shi
L, Liu X, Bai J, Deng M, Shuai X, et al: miR-219-5p plays a tumor
suppressive role in colon cancer by targeting oncogene Sall4. Oncol
Rep. 34:1923–1932. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Sablin EP, Blind RD, Uthayaruban R, Chiu
HJ, Deacon AM, Das D, Ingraham HA and Fletterick RJ: Structure of
liver receptor homolog-1 (NR5A2) with PIP3 hormone bound in the
ligand binding pocket. J Struct Biol. 192:342–348. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Fayard E, Auwerx J and Schoonjans K:
LRH-1: An orphan nuclear receptor involved in development,
metabolism and steroidogenesis. Trends Cell Biol. 14:250–260. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Stein S and Schoonjans K: Molecular basis
for the regulation of the nuclear receptor LRH-1. Curr Opin Cell
Biol. 33:26–34. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang Q, Zhao S, Pang X and Chi B:
MicroRNA-381 suppresses cell growth and invasion by targeting the
liver receptor homolog-1 in hepatocellular carcinoma. Oncol Rep.
35:1831–1840. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Jiang W, Tian Y, Jiang S, Liu S, Zhao X
and Tian D: MicroRNA-376c suppresses non-small-cell lung cancer
cell growth and invasion by targeting LRH-1-mediated Wnt signaling
pathway. Biochem Biophys Res Commun. 473:980–986. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Baquié M, St-Onge L, Kerr-Conte J,
Cobo-Vuilleumier N, Lorenzo PI, Moreno Jimenez CM, Cederroth CR,
Nef S, Borot S, Bosco D, et al: The liver receptor homolog-1
(LRH-1) is expressed in human islets and protects {beta}-cells
against stress-induced apoptosis. Hum Mol Genet. 20:2823–2833.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wang S, Lan F, Huang L, Dong L, Zhu Z, Li
Z, Xie Y and Fu J: Suppression of hLRH-1 mediated by a DNA
vector-based RNA interference results in cell cycle arrest and
induction of apoptosis in hepatocellular carcinoma cell BEL-7402.
Biochem Biophys Res Commun. 333:917–924. 2005. View Article : Google Scholar : PubMed/NCBI
|