Introduction
Non-small cell lung cancer (NSCLC) is a common
pathological type of lung cancers (1). NSCLC exerts rapid growth and
aggressive features during its progression. Even though marked
improvement in the diagnosis and treatment of NSCLC has been
achieved, the prognosis of NSCLC patients particularly for those in
advanced stages is still poor (2).
Thus, it is imperative to investigate new biomarkers and treatment
regimens for NSCLC.
MicroRNAs (miRNAs) are a group of small, non-protein
coding, endogenous and single-stranded RNAs (3). miRNAs have been confirmed to play
vital roles in many biological processes including cell growth
(4), metastasis (5) and angiogenesis (6) by negatively regulating target mRNAs to
either translation or degradation.
Recently, microRNA-149 (miR-149) was identified as a
tumor inhibitor in human cancers. In hepatocellular carcinoma, low
expression of miR-149 was found to be correlated with larger tumor
mass size, capsular and vascular invasion, lymph node metastasis,
high histological grade, advanced tumor-node-metastasis (TNM) stage
and poor prognosis (7). In
vitro, miR-149 inhibited cell proliferation, migration and
invasion by targeting PPM1F (8). In
gastric cancer, miR-149 was upregulated by a chemical compound
named 18β-glycyrrhetinic acid (GRA) and inhibited the progression
of gastric tumors by targeting the Wnt-1 signaling pathway
(9). However, the expression and
functions of miR-149 in NSCLC are still unclear.
In the present study, we investigated the expression
and biological function of miR-149 in NSCLC progression.
Downregulation of miR-149 was confirmed to be correlated with large
tumor size, metastasis and poor prognosis in NSCLC patients.
miR-149 suppressed NSCLC tumor growth and metastasis by targeting
oncogenic transcription factor FOXM1 by gain- and loss-of-function
experiments.
Materials and methods
Clinical samples and cell lines
Forty-five paired non-small cell lung cancer and
matched tumor-adjacent tissues were collected at Linyi Central
Hospital (Linyi, China). The patients included 31 males and 14
females who received surgical resection between March 2011 and
January 2013. The age range was between 37 and 69 years. 293T
cells, normal lung epithelium BEAS-2B and 4 NSCLC cell lines (A549,
H1299, H1975 and HCC827) were purchased from the American Type
Culture Collection (ATCC; Manassas, VA, USA) and cultured in
Dulbecco's modified Eagle's medium (DMEM; Invitrogen, Waltham, MA,
USA) supplemented with 10% fetal bovine serum (FBS; Gibco, Grand
Island, NY, USA), 1% penicillin and 1% streptomycin.
Real-time quantitative reverse
transcription-PCR (qRT-PCR)
Total RNA from tissues and cells was isolated by
RNeasy Plus Micro and Mini kits (cat no. 74034; Qiagen, Venlo, The
Netherlands) according to the manufacturers instructions. One-Step
Perfect Real-Time RT-qPCR (SYBR-Green I) kit (Takara Biotechnology
Co., Ltd., Dalian, China) was used to detected the expression
levels of miR-149 and FOXM1 mRNA. miR-149 and RNU6 Bulge-Loop™
primers were purchased from RiboBio Co., Ltd. (Guangzhou, China).
FOXM1 and GAPDH primers were synthesized by Sangon Co., Ltd.
(Shanghai, China). Data were analyzed using the 2−ΔΔCt
method.
Cell transfection
miR-149 and negative control (miR-NC) lentivirus
particles were purchased from GeneCopoeia (Guangzhou, PR China).
miR-NC or miR-149 and package plasmids were co-transfected into
293T cells using Lipofectamine™ 2000 (Invitrogen) reagent according
to the manufacturers instructions. The supernatant including the
lentivirus was collected twice at 48 and 72 h after transfection.
Cells (15×104/well) were seeded into a 6-well plate
before the infection day. For lentivirus infection, 1 ml lentivirus
supernatant (miR-NC or miR-149) combined with 2 ml growth medium
and Polybrene (5 µg/ml) were added into one well. Puromycin was
used to select the successfully infected cells. The effect of
transfection was determined by qRT-PCR.
Anti-BrdU cell proliferation
assay
For cell proliferation detection, we used an
anti-BrdU assay according to the manufacturer's recommendation (BD
Biosciences, San Jose, CA, USA). Cells were cultured into a 24-well
plate. BrdU solution (0.1 mg/ml) was added to each well. Cells were
incubated at 37°C for 4 h. The incorporated BrdU was stained with
Alexa Fluor® 488 anti-BrdU monoclonal antibody (BD
Biosciences). Cell nuclei were stained with DAPI. Positive stained
cells were observed and counted under a fluorescence
microscope.
Cell apoptosis assay
To evaluate the apoptosis of NSCLC cells, the flow
cytometric assay was performed to evaluate the percentage of
apoptotic cells. In brief, cells were harvested and re-suspended in
phosphate-buffered saline (PBS) solution, and then stained with
Annexin V-FITC/PI detection kit (556547; BD Biosciences) and
subjected to FACS analysis.
Wound healing assay
Cells were seeded into a 6-well plate until
achieving 90% confluency. A pippette tip (100 µl) was used to make
a wound in the middle of the well. Remnant cells were washed away
using PBS. Basal DMEM was used to culture cells for 48 h. The
healing of the wound was observed under an inverted microscope.
Transwell assay
The upper face of 8-µm pore Transwell inserts (Nalge
Nunc International, Penfield, NY, USA) was coated with Matrigel (BD
Biosciences, Franklin Lakes, NJ, USA) for the invasion assay. In
brief, cells suspended with basal DMEM were seeded onto the upper
chamber. DMEM (750 µl) with 10% FBS was added into the lower
chambers, and. 24 h later, cells that had invaded the membranes and
stayed on the lower surface were stained with crystal violet. The
invaded cell number was counted under a microscope.
In vivo experiments
Genetic modified and wild-type tumor cells were
subcutaneously injected into nude mice to investigate the functions
of miR-149 on tumor growth or injected into the left pleural cavity
of the mice for tumor metastasis. Mice were sacrificed after 4
weeks. Final tumor volumes (V) were calculated based on the
following formula: V = (length × width2)/2. Mouse livers
were harvested to make paraffin-embedded tissue sections for
H&E staining.
Western blotting
Total cell proteins were extracted with RIPA buffer,
separated by 10% SDS-PAGE gel and transferred onto a nitrocellulose
membrane (Invitrogen). The membranes were incubated with the
following primary rabbit anti-human antibodies purchased from Cell
Signaling Technology Inc. (CST; Danvers, MA, USA) at 4°C overnight:
FOXM1 (5436; dilution at 1:1,000), cyclin D1 (2978; dilution at
1:1,000), MMP2 (87809; dilution at 1:1,000) and GAPDH (5174;
dilution at 1:1,000). Then, the membranes were incubated with
HRP-linked goat anti-rabbit antibodies (7074; dilution at 1:5,000;
CST), and the signals were detected using the Bio-Rad Gel imaging
system (Bio-Rad, Hercules, CA, USA).
Luciferase reporter assay
Stable miR-149 or miR-NC infected cells were grown
in a 24-well plate until achieving 50% confluency. Cells were
transfected with 50 ng of wild-type (WT) or mutant (MT)
3-untranslated region (3′-UTR) vectors of FOXM1 (RiboBio Co., Ltd.)
in a 24-well plate using Lipofectamine™ 2000 (Invitrogen) reagent
according to the manufacturers instructions for 48 h. The hLuc
fluorescence intensity and hRluc fluorescence intensity were
detected by the Dual-Luciferase Reporter Assay System (Promega,
Madison, WI, USA) following the manufacturer's instructions. The
ratio of Rluc/Luc was used to measure the effects of combination
between miR-149 and 3′-UTR of FOXM1.
Statistical analysis
All quantitative data are expressed as mean ± SD.
SPSS v21.0 software (IBM, Armonk, NY, USA) was used to perform
statistical analysis. The Pearson's Chi-square test was used for
enumeration data and the Student's t-test was performed for
measurement of data in the present study. Overall survival and
progression-free survival analysis were performed using the
Kaplan-Meier method for plotting and the log-rank test for
comparison. P<0.05 was indicative of a statistically significant
difference.
Results
miR-149 is downregulated in NSCLC
tissues and cell lines
qRT-PCR was used to detect the expression levels of
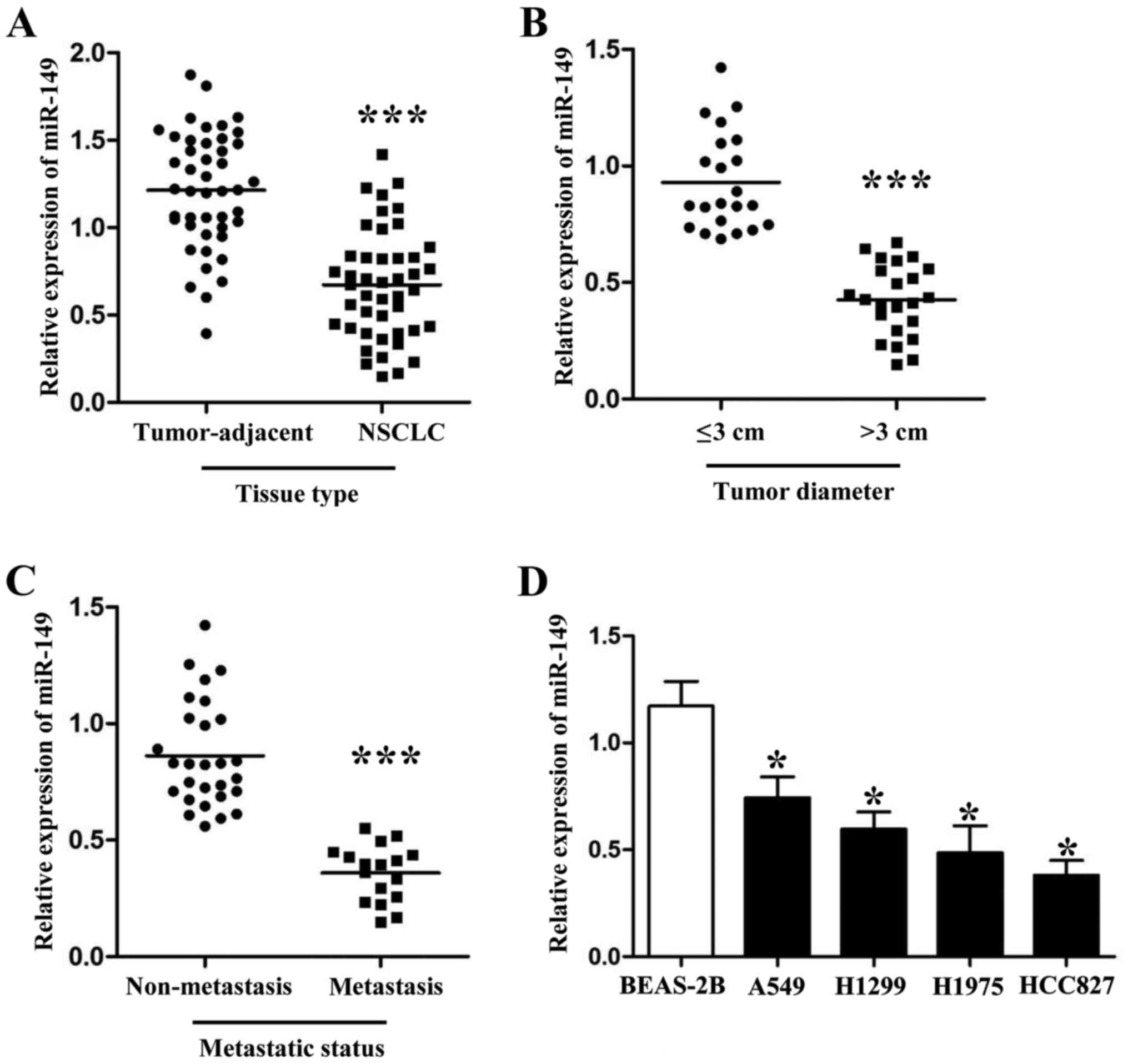
miR-149 in cancer tissues and different NSCLC cell lines. As shown
in Fig. 1A, the expression levels
of miR-149 were downregulated in the NSCLC tissues (Fig. 1A; P<0.001). Furthermore, we also
divided these 45 patients into different subgroups according to
their clinical features. As shown in Fig. 1B and C, PCR results revealed that
patients with large tumor size and metastasis had lower miR-149
expression levels than those who did not have these malignant
features (P<0.001, respectively). Next, compared with normal
lung epithelium cells, we also found that miR-149 was downregulated
in 4 NSCLC cell lines (A549, H1299, H1975 and HCC827; Fig. 1D; P<0.05, respectively).
Clinical significance of miR-149 in
NSCLC
To further investigate the clinical significance of
miR-149, we used the mean expression level of miR-149 (x=0.672) as
a cut-off value to divide 45 NSCLC tissues into 2 groups: miR-149
high expression group (n=20) and miR-149 low expression group
(n=25). As shown in Table I, low
expression of miR-149 was associated with large tumor size
(P=0.003), tumor metastasis (P=0.006) and advanced TNM stage
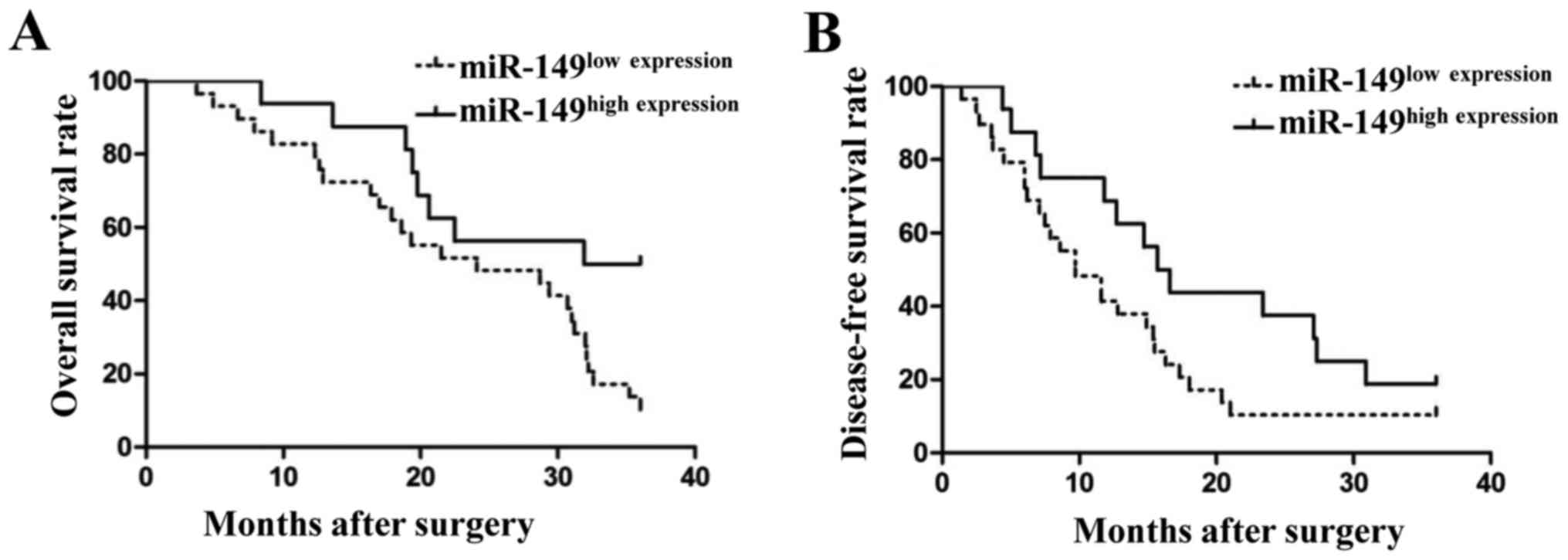
(P=0.038). We also compared the differences in the 3-year survival
rates between these 2 groups. As shown in Fig. 2, patients with low levels of miR-149
had a poor 3-year overall survival rate (Fig. 2A; P<0.05) and disease-free
survival rate (Fig. 2B; P<0.05).
These findings indicated that miR-149 could serve as a biomarker
for NSCLC patients.
 | Table I.Clinical significance of miR-149
expression in NSCLC (N=45). |
Table I.
Clinical significance of miR-149
expression in NSCLC (N=45).
|
| miR-149 expression
level |
|
|
|---|
|
|
|
|
|
|---|
| Clinical
characteristics | High n=20 | Low n=25 | χ2 | P-value |
|---|
| Sex |
|
|
|
|
| Male | 15 | 16 | 0.627 | 0.525 |
|
Female | 5 | 9 |
|
|
| Age (years) |
|
|
|
|
|
<50 | 9 | 17 | 2.409 | 0.142 |
| ≥50 | 11 | 8 |
|
|
| Smoking |
|
|
|
|
| Yes | 8 | 15 | 1.779 | 0.236 |
| No | 12 | 10 |
|
|
| Tumor size (cm) |
|
|
|
|
| ≥3 | 5 | 18 | 9.823 | 0.003a |
|
<3 | 15 | 7 |
|
|
| Metastasis |
|
|
|
|
| Yes | 3 | 14 | 7.946 | 0.006a |
| No | 17 | 11 |
|
|
| TNM stage |
|
|
|
|
| I+II | 13 | 8 | 4.862 | 0.038a |
|
III+IV | 7 | 17 |
|
|
miR-149 inhibits the proliferation and
induces apoptosis of NSCLC cells
Following the analysis of expression levels of
miR-149 in the 4 NSCLC cell lines, we used miR-149 lentivirus to
overexpress its expression level in HCC827 cells and the
anti-miR-149 lentivirus to knock down its expression level in A549
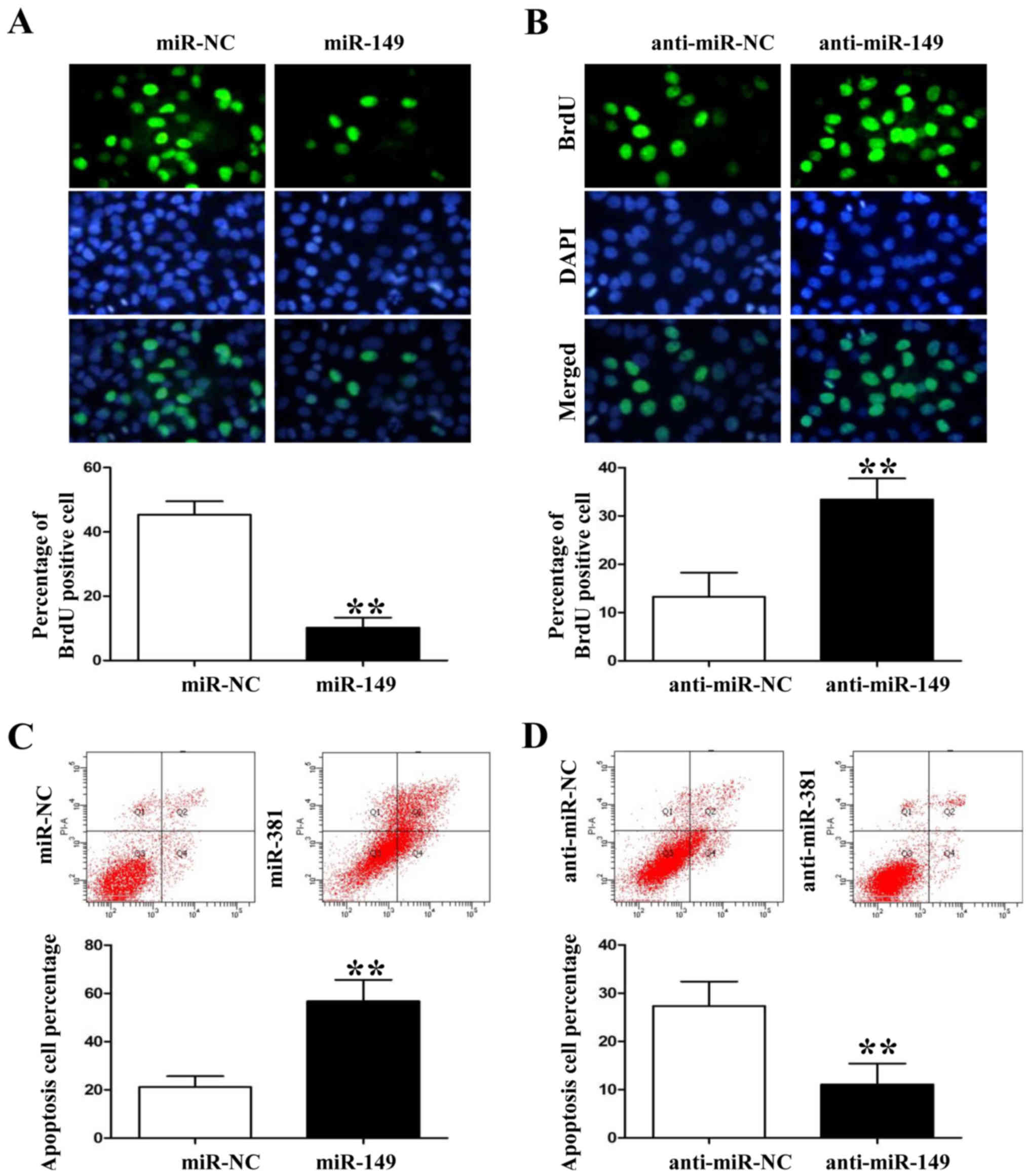
cells (P<0.05, data not shown). Anti-BrdU assay showed that
miR-149 overexpression significantly decreased the number of
BrdU-positive staining HCC827 cells (Fig. 3A; P<0.01). However, miR-149
knockdown induced A549 cell proliferation based on the BrdU
immunofluorescence results (Fig.
3B; P<0.01). In contrast, flow cytometric assay showed that
miR-149 overexpression increased the percentage of apoptotic HCC827
cells whereas miR-149 knockdown decreased A549 cell apoptosis
(Fig. 3C and D; P<0.01,
respectively).
miR-149 inhibits the migration and
invasion of NSCLC cells
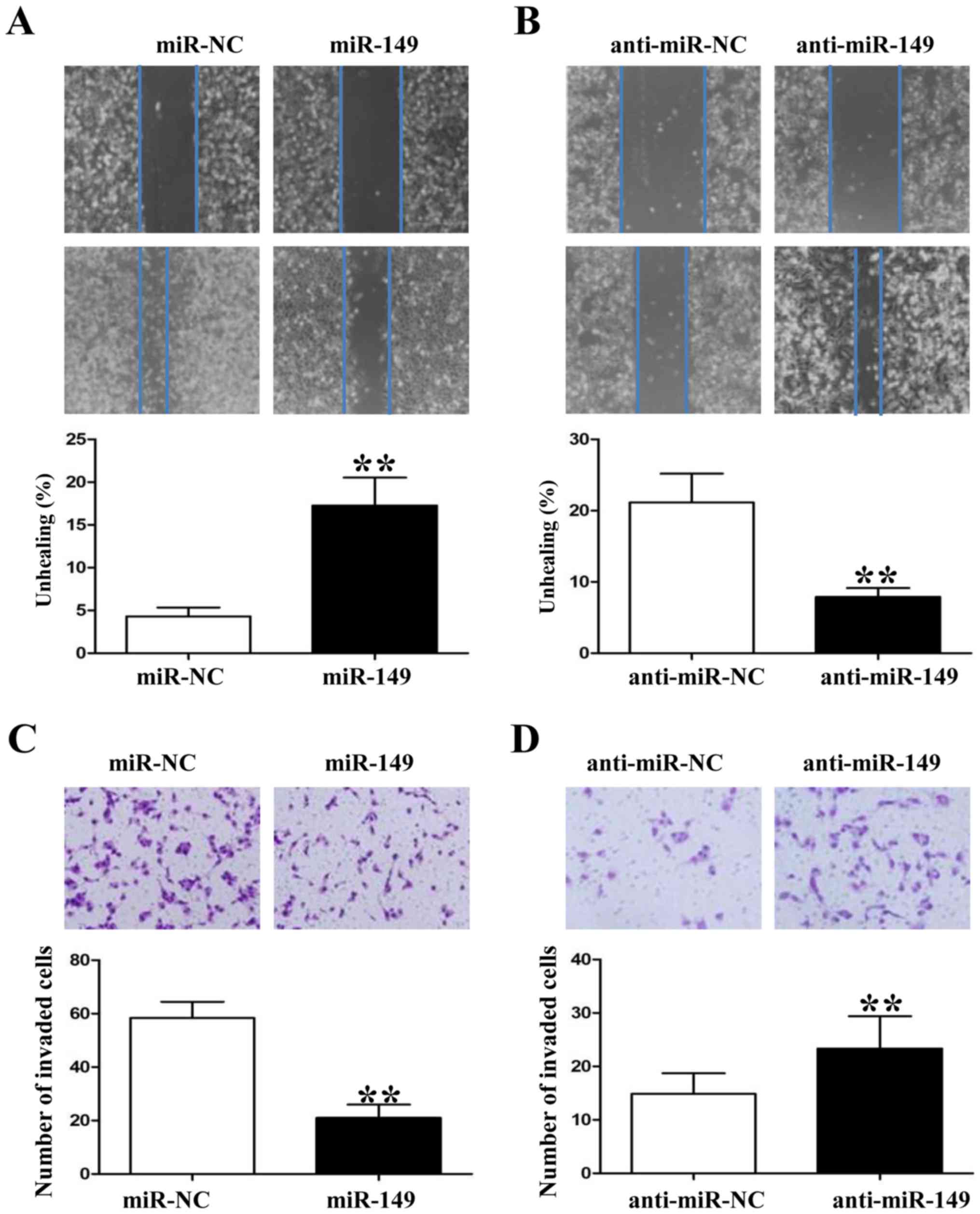
Since miR-149 was found to be associated with tumor
metastasis in clinical samples, we also used wound healing and
Transwell assays to demonstrate that upregulation of miR-149
impaired the migration and invasion of HCC827 cells (Fig. 4A and C; P<0.01, respectively). In
contrast, downregulation of miR-149 in A549 cells enhanced their
migration and invasion markedly (Fig.
4B and D; P<0.01, respectively). In summary, these data
suggest that miR-149 attenuates the metastatic ability of NSCLC
cells.
miR-149 inhibits NSCLC tumor growth
and metastasis in vivo
When we demonstrated that miR-149 inhibited NSCLC
growth and metastasis in vitro, we also established a tumor
subcutaneous growth model and liver metastasis model to further
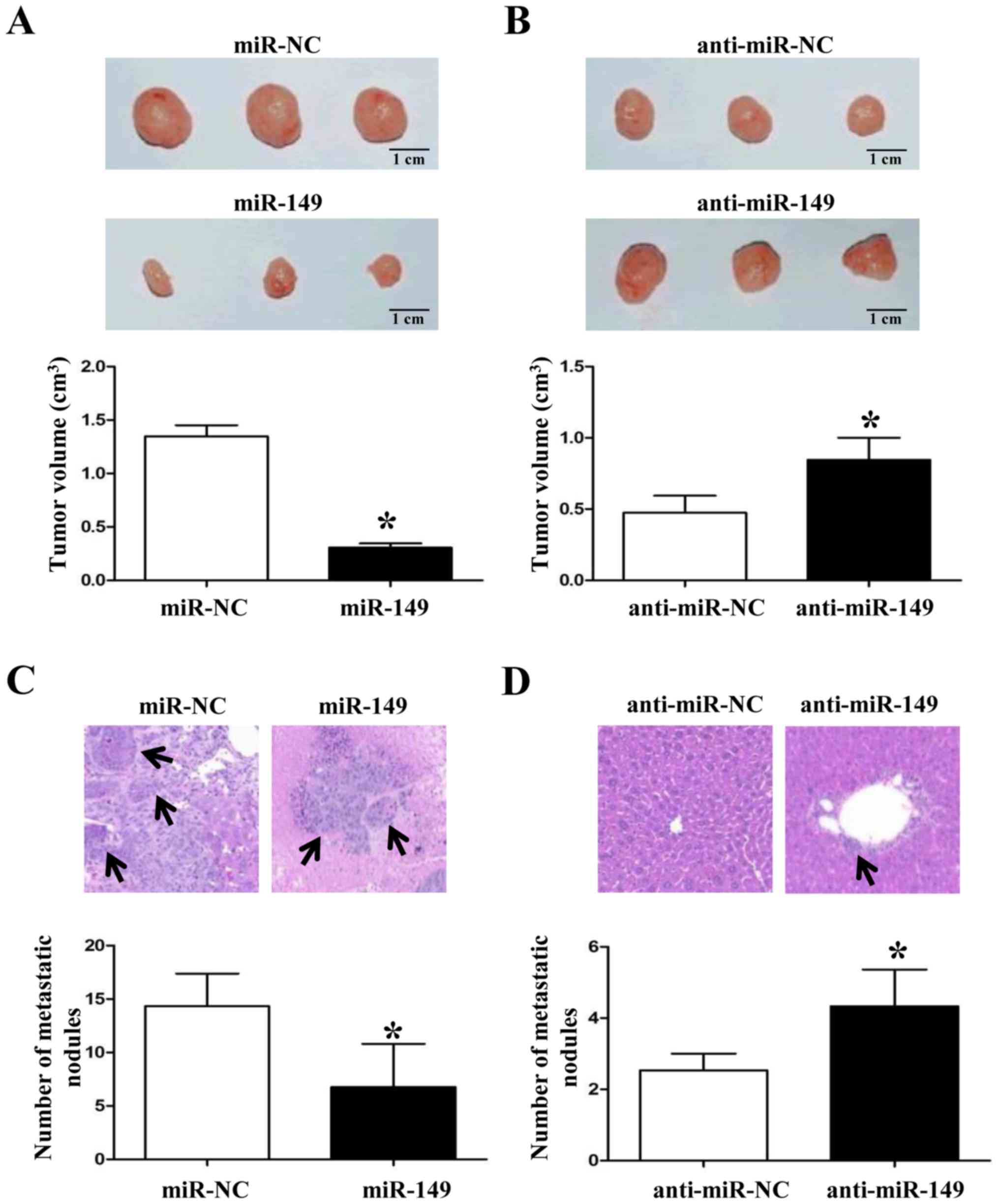
confirm the anticancer roles of miR-149 in vivo. As shown in
Fig. 5A and B, forced expression of
miR-149 in NSCLC cells inhibited the tumor growth in nude mice
(P<0.05, respectively). Subsequently, overexpression of miR-149
significantly reduced the liver metastasis of NSCLC cells (Fig. 5C and D; P<0.05, respectively).
These data suggest that miR-149 plays a key role in the growth and
metastasis of NSCLC cells in vitro and in vivo.
miR-149 exerts its anticancer
functions by inhibiting FOXM1 in NSCLC cells
FOXM1 was predicted as a potential downstream target
of miR-149 according to bioinformatic analysis by online database
screening (TargetScan: http://www.targetscan.org/ and miRanda: http://www.microrna.org/). To confirm the relationship
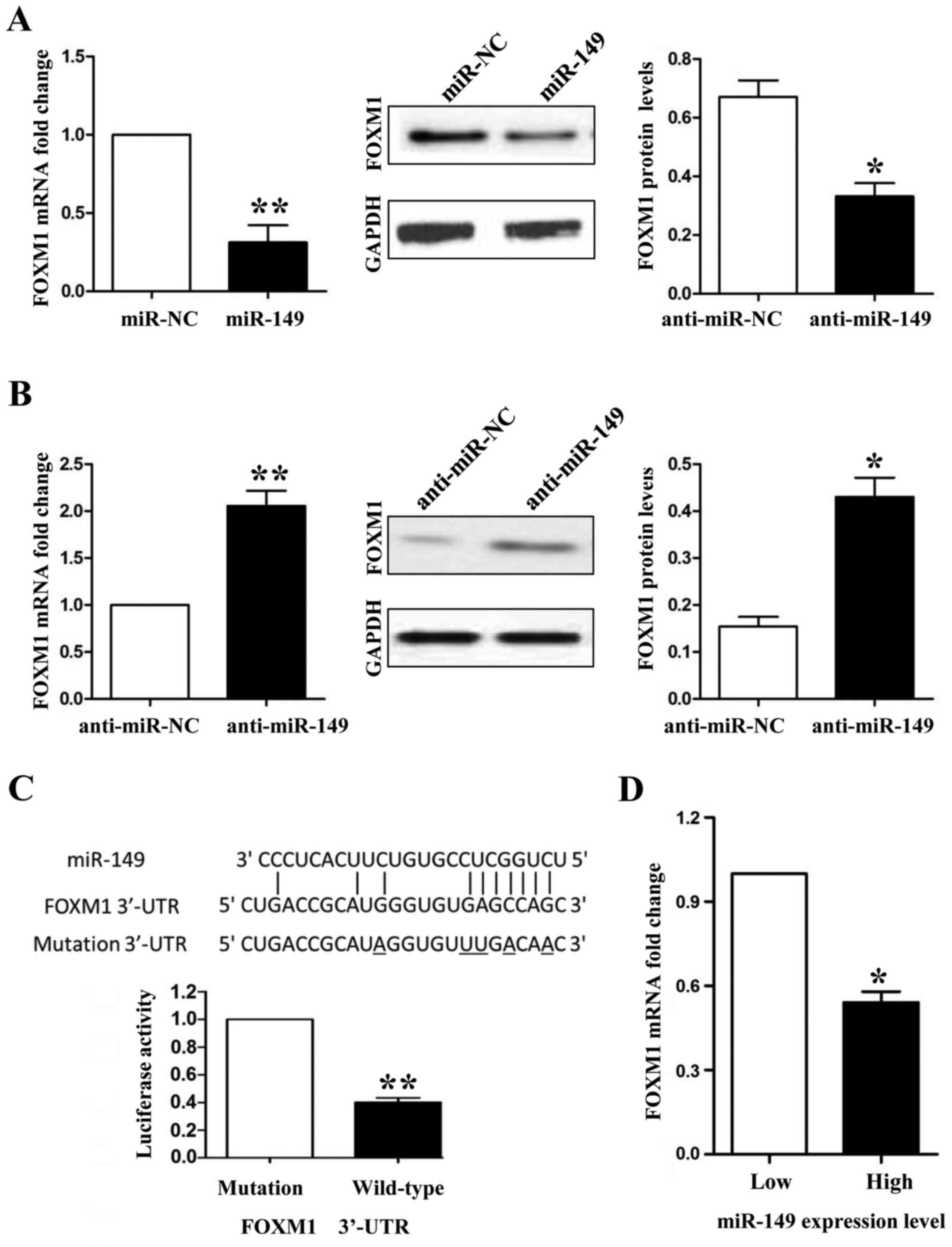
between miR-149 and FOXM1, first, we detected the expression of
FOXM1 in miR-194-overexpressing HCC827 and miR-194-knockdown A549
cells. Both the mRNA and protein expression levels of FOXM1 were
downregulated when we overexpressed miR-149 in the HCC827 cells
(Fig. 6A; P<0.01 and P<0.05,
respectively), whereas FOXM1 was upregulated when miR-149 was
inhibited in the A549 cells (Fig.
6B; P<0.01 and P<0.05, respectively). Secondly, we used a
Dual-Luciferase® Reporter Assay System to confirm that
only the luciferase activity of wild-type FOXM1 3′-UTR was
downregulated by miR-149 (Fig. 6C;
P<0.01). Finally, we detected the mRNA levels of FOXM1 in NSCLC
tissues by qRT-PCR and demonstrated that the mRNA expression of
FOXM1 was lower in the miR-149 high expression group than that
noted in the miR-149 low expression group (Fig. 6D; P<0.05). In the present study,
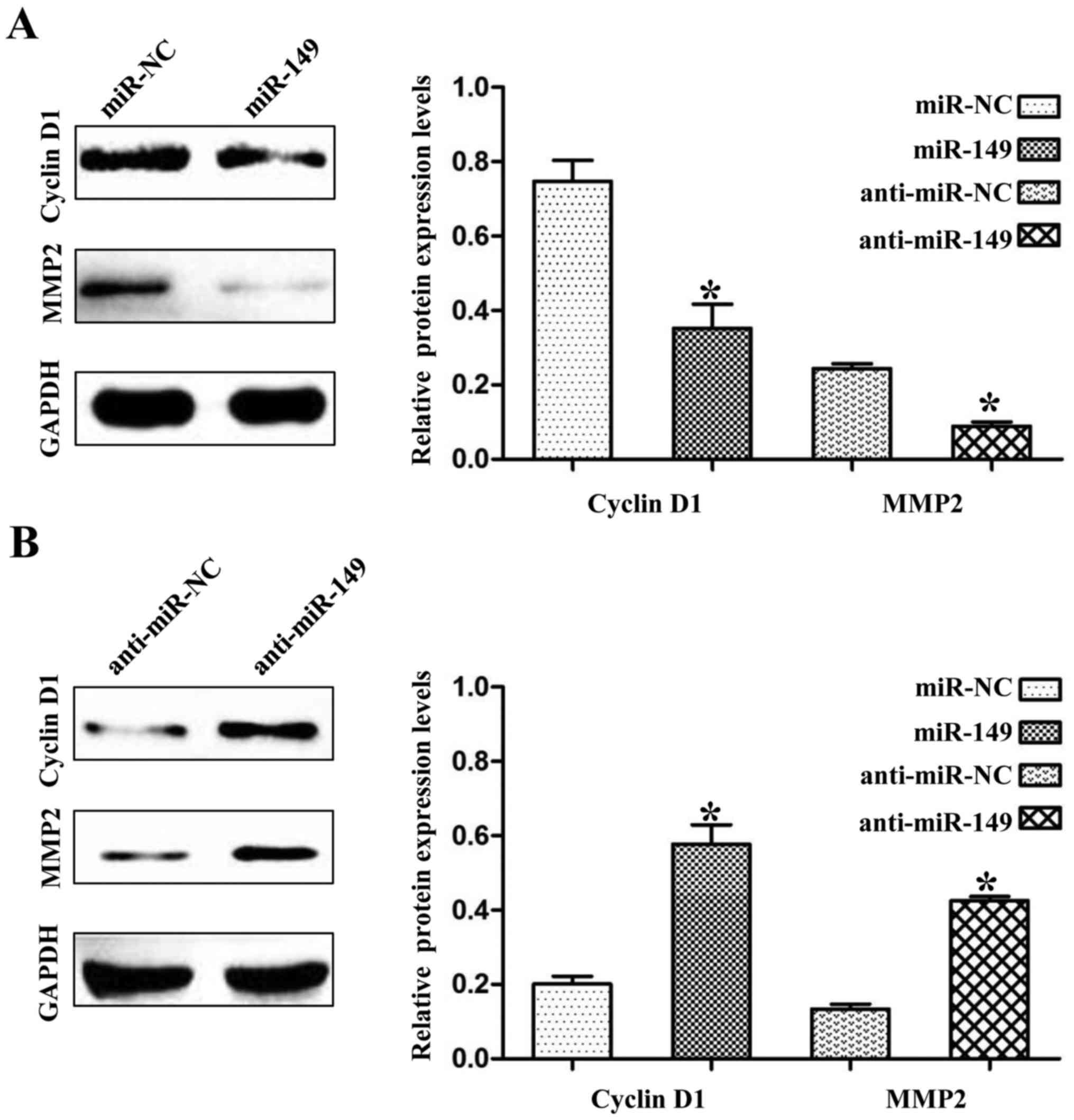
we found that the expression levels of cyclin D1 and MMP2 were also
negatively regulated by miR-149 in the NSCLC cells (Fig. 7A and B; P<0.01). Cyclin D1 and
MMP2 are not targets of miR-149, but they have been demonstrated to
be regulated by FOXM1 (10,11). Thus, these results may partly
explain the anticancer mechanisms of miR-149 in NSCLC.
Discussion
Tumor growth and metastasis are two vital risk
factors for the poor prognosis of NSCLC patients (12). Numerous studies have demonstrated
that microRNAs (miRNAs) are critical players in the genesis and
progression of human cancers, particularly in NSCLC. miR-502e was
found to serve as a tumor suppressor and inhibit NSCLC cell growth,
invasion and migration by targeting Zbtb7a partly depending on the
Wnt signaling pathway (13).
However, miR-30a reduced radiation-induced G2/M cell cycle arrest
and may also affect radiation-induced apoptosis by targeting ATF1
(14). This accumulating evidence
shows that miRNAs are promising biomarkers and therapeutic targets
in NSCLC.
In the present study, we revealed that miR-149 was
significantly decreased in NSCLC cancer tissues and cell lines.
Clinical data showed that low expression of miR-149 was associated
with large tumor size, tumor metastasis and advanced TNM stage. In
addition, low expression levels of miR-149 predicted a reduced
3-year overall and disease-free survival rates of NSCLC patients.
Taken together, these data indicated that miR-149 may play a
critical role in the development of NSCLC.
Functionally, we used gain- and loss-of-function
experiments to confirm that miR-149 inhibited NSCLC cell
proliferation, migration and invasion in vitro and it also
inhibited subcutaneous xenograft growth and liver metastasis in
vivo. These results were consistent with the expression levels
of miR-149 in the clinical samples.
Forkhead box protein M1 (FOXM1) is a member of the
FOX family of transcription factors. In NSCLC, FOXM1 has been
demonstrated as a tumor inducer due to its function in tumor
proliferation (15), invasion
(16) and chemoresistance (17). Moreover, FOXM1 was found to be a
target of several miRNAs in NSCLC such as miR-134 (18) and miR-509 (19). In the present study, we found that
FOXM1 may be a target of miR-149. To confirm their relationship, we
used PCR, western blot and luciferase reporter assays to
demonstrate that miR-149 inhibited FOXM1 mRNA and protein
expression levels by binding to its 3′-UTR in NSCLC cells.
Moreover, patients with low expression levels of miR-149 exerted
high FOXM1 mRNA levels. As a classical oncogenic transcription
factor in NSCLC, FOXM1 could enhance the transcription of many
oncogenes including cyclin D1 and MMP2 in lung cancer (11,20,21).
In addition, we also found that miR-149 decreased the expression
levels of cyclin D1 and MMP2 in NSCLC cells.
In conclusion, we demonstrated that miR-491 was
downregulated in NSCLC tissues and cell lines, and its reduced
expression was associated with malignant clinical features. miR-149
can inhibit NSCLC growth and metastatic behaviors by inhibiting
FOXM1. These results suggest that miR-149 is a potential
tumor-suppressor in NSCLC.
References
|
1
|
Gadgeel SM: Personalized therapy of
non-small cell lung cancer (NSCLC). Adv Exp Med Biol. 890:203–222.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Califano R, Romanidou O, Mountzios G,
Landi L, Cappuzzo F and Blackhall F: Management of NSCLC disease
progression after first-line EGFR tyrosine kinase inhibitors: What
are the issues and potential therapies? Drugs. 76:831–840. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ameis D, Khoshgoo N, Iwasiow BM, Snarr P
and Keijzer R: MicroRNAs in lung development and Disease. Paediatr
Respir Rev. 22:38–43. 2017.PubMed/NCBI
|
|
4
|
Singh S, Zheng Y, Jagadeeswaran G, Ebron
JS, Sikand K, Gupta S, Sunker R and Shukla GC: Deep sequencing of
small RNA libraries from human prostate epithelial and stromal
cells reveal distinct pattern of microRNAs primarily predicted to
target growth factors. Cancer Lett. 371:262–273. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zoni E and van der Pluijm G: The role of
microRNAs in bone metastasis. J Bone Oncol. 5:104–108. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Masotti A, Miller MR, Celluzzi A, Rose L,
Micciulla F, Hadoke PW, Bellucci S and Caporali A: Regulation of
angiogenesis through the efficient delivery of microRNAs into
endothelial cells using polyamine-coated carbon nanotubes.
Nanomedicine. 12:1511–1522. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lin L, Zhang YD, Chen ZY, Chen Y and Ren
CP: The clinicopathological significance of miR-149 and PARP-2 in
hepatocellular carcinoma and their roles in chemo/radiotherapy.
Tumour Biol. 37:12339–12346. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Luo G, Chao YL, Tang B, Li BS, Xiao YF,
Xie R, Wang SM, Wu YY, Dong H, Liu XD, et al: miR-149 represses
metastasis of hepatocellular carcinoma by targeting
actin-regulatory proteins PPM1F. Oncotarget. 6:37808–37823. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cao D, Jia Z, You L, Wu Y, Hou Z, Suo Y,
Zhang H, Wen S, Tsukamoto T, Oshima M, et al: 18β-glycyrrhetinic
acid suppresses gastric cancer by activation of miR-149-3p-Wnt-1
signaling. Oncotarget. 7:71960–71973. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Gao F, Bian F, Ma X, Kalinichenko VV and
Das SK: Control of regional decidualization in implantation: Role
of FoxM1 downstream of Hoxa10 and cyclin D3. Sci Rep. 5:138632015.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chen PM, Wu TC, Shieh SH, Wu YH, Li MC,
Sheu GT, Cheng YW, Chen CY and Lee H: MnSOD promotes tumor invasion
via upregulation of FoxM1-MMP2 axis and related with poor survival
and relapse in lung adenocarcinomas. Mol Cancer Res. 11:261–271.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chen YY, Huang TW, Tsai WC, Lin LF, Cheng
JB, Chang H and Lee SC: Risk factors of postoperative recurrences
in patients with clinical stage I NSCLC. World J Surg Oncol.
12:102014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhijun Z and Jingkang H: MicroRNA-520e
suppresses non-small-cell lung cancer cell growth by targeting
Zbtb7a-mediated Wnt signaling pathway. Biochem Biophys Res Commun.
486:49–56. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Guo Y, Sun W, Gong T, Chai Y, Wang J, Hui
B, Li Y, Song L and Gao Y: miR-30a radiosensitizes non-small cell
lung cancer by targeting ATF1 that is involved in the
phosphorylation of ATM. Oncol Rep. 37:1980–1988. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Kim IM, Ackerson T, Ramakrishna S,
Tretiakova M, Wang IC, Kalin TV, Major ml, Gusarova GA, Yoder HM,
Costa RH, et al: The Forkhead Box m1 transcription factor
stimulates the proliferation of tumor cells during development of
lung cancer. Cancer Res. 66:2153–2161. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Kong FF, Qu ZQ, Yuan HH, Wang JY, Zhao M,
Guo YH, Shi J, Gong XD, Zhu YL, Liu F, et al: Overexpression of
FOXM1 is associated with EMT and is a predictor of poor prognosis
in non-small cell lung cancer. Oncol Rep. 31:2660–2668. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang K, Zhu X, Zhang K, Zhu L and Zhou F:
FoxM1 inhibition enhances chemosensitivity of docetaxel-resistant
A549 cells to docetaxel via activation of JNK/mitochondrial
pathway. Acta Biochim Biophys Sin. 48:804–809. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li J, Wang Y, Luo J, Fu Z, Ying J, Yu Y
and Yu W: miR-134 inhibits epithelial to mesenchymal transition by
targeting FOXM1 in non-small cell lung cancer cells. FEBS Lett.
586:3761–3765. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ma N, Zhang W, Qiao C, Luo H, Zhang X, Liu
D, Zang S, Zhang L and Bai J: The tumor suppressive role of
miRNA-509-5p by targeting FOXM1 in non-small cell lung cancer. Cell
Physiol Biochem. 38:1435–1446. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Liu Y, Hock JM, Van Beneden RJ and Li X:
Aberrant overexpression of FOXM1 transcription factor plays a
critical role in lung carcinogenesis induced by low doses of
arsenic. Mol Carcinog. 53:380–391. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Balli D, Zhang Y, Snyder J, Kalinichenko
VV and Kalin TV: Endothelial cell-specific deletion of
transcription factor FoxM1 increases urethane-induced lung
carcinogenesis. Cancer Res. 71:40–50. 2011. View Article : Google Scholar : PubMed/NCBI
|