Introduction
Mesenchymal stem cells (MSCs) are adult stem cells
with the ability to differentiate into mesenchymal tissue cells
while retaining self-renewal and migration abilities (1). Many recent studies have revealed that
MSCs possess immunomodulatory functions that they exert through
cell-to-cell contacts, as well as by secreting growth factors,
cytokines, and chemokines (2,3). The
effect of immunosuppression with MSCs has been reported in
graft-versus-host disease (4) and
multiple system atrophy (5). MSCs
have the ability to migrate to damaged tissue by inducing
peripheral tolerance by inhibiting the release of pro-inflammatory
cytokines (2). The advantages of
MSC-based cell therapy have been demonstrated in acute lung injury
(6), myocardial infarction
(7), acute renal failure (8), cerebral ischemia (9) and Alzheimer's disease (10). At the cellular level, it has been
shown that MSCs can directly inhibit both T lymphocyte and
microglial cell proliferation and can negatively modulate the
cytokine-secretion profile of dendritic cells and
monocytes/macrophages (11–14).
In our recent study, we identified the scrapie
responsive gene 1 (SCRG1) secreted from MSCs and its receptor
complex bone marrow stromal cell antigen 1 (BST1)/β1 integrin, as
positive regulators of stem cell qualities (15). SCRG1, which was identified by Dron
et al as a protein that increased expression in the brain of
scrapie infected mice, was shown to be associated with
neurodegenerative changes in transmissible spongiform
encephalopathy, as well as in brain injury, and is associated with
autophagy (16–18). The SCRG1 gene encodes a 98-amino
acid, cytokine-like peptide with an N-terminal signal peptide
(19,20). Intriguingly, in stem cells SCRG1
was shown to maintain octamer-binding transcription factor 4
(Oct-4) and CD271/low-affinity nerve growth factor receptor (LNGFR)
expression and thereby maintain the MSC's potential for
self-renewal, migration abilities, as well as osteogenic
differentiation potential, even at high stem cell passage numbers
(15). Other cytokines and
chemokines secreted from MSCs have been implicated in
immunosuppression and repair of damaged tissues (21–24).
MSC differentiation along different pathways is regulated by
stimulation with various growth factors, cytokines, or chemokines,
as has been demonstrated in the differentiation of bone
marrow-derived MSCs (25,26).
SCRG1 secreted from MSCs is predicted to affect a
variety of cell types in vivo. In this study, we
hypothesized that SCRG1 secreted into the extracellular space by
MSCs exhibits paracrine activity. To explore this possibility, we
examined the paracrine effect of MSC-derived SCRG1 on the immune
response of Raw264.7 macrophages. In particular, as a readout of
macrophage function, we focused on macrophage production of
CC-chemokine ligand 22 [CCL22; also known as MDC
(macrophage-derived chemokine)], which is known to display
chemotactic activity for monocytes, dendritic cells, natural killer
cells, and chronically activated T lymphocytes (27–30).
Materials and methods
Reagents
Recombinant mouse SCRG1 (rmSCRG1), expressed in
yeast, was purchased from MyBiosource, Inc. (MBS1177239, San Diego,
CA, USA). The MAPK/ERK kinase inhibitor U0126 was purchased from
Calbiochem (Merck Millipore, Darmstadt, Germany).
Lipopolysaccharide (LPS) derived from Escherichia coli
0111:B4 was purchased from Sigma-Aldrich (St. Louis, MO, USA).
Cell culture
Mouse macrophage-like Raw264.7 cells (American Type
Culture Collection, Manassas, VA, USA) were maintained in minimum
essential medium Eagle's α-modification (αMEM) (Sigma-Aldrich)
supplemented with 10% fetal bovine serum (FBS) (HyClone, GE
Healthcare Life Sciences, Logan, UT, USA) under the condition of 5%
CO2 at 37°C.
Proliferation assay
Cell proliferation was analyzed by WST-1 assay
reagent (Roche Diagnostics, Basel, Switzerland) according to the
manufacturer's instructions. Raw264.7 cells were cultured on
96-well plates (Nunc; Thermo Fisher Scientific, Waltham, MA, USA)
in 100 µl complete medium containing with or without 100 ng/ml
rmSCRG1. After five days, the cells were added with 10 µl WST-1
reagent and incubated for 1 h. The absorbance was measured using an
MPR-A4i microplate reader (Tosoh Corp., Tokyo, Japan) at 450
nm.
Migration assay
The migration assay was performed using 8-µm pore
sized Transwell cell culture inserts (BD Biosciences, Franklin
Lakes, NJ, USA). Raw264.7 cells (1.0×105) were seeded on
the upper well in 350 µl serum-free αMEM containing 0.1% BSA
(Sigma-Aldrich). The lower well was filled in 600 µl complete
medium containing with or without 100 ng/ml rmSCRG1. After
incubation for 6 h, cells that had not migrated were scraped off
with a cotton swab. The number of cells migrated to the lower side
of the filter was stained with Diff-Quik Three-Step Stain Set
(Sysmex, Kobe, Japan), and then counted using a microscope (Olympus
IX70; Olympus Corp., Tokyo, Japan) under five high-power fields
(x400 magnification).
Adhesion assay
Raw264.7 cells (1.0×105) were seeded onto
a fibronectin-coated culture dish (BD Biosciences) and cultured in
complete medium with or without 100 ng/ml rmSCRG1. After 6 h,
non-adhered cells on the bottom of the dish were removed by washing
twice with phosphate-buffered saline (PBS). The number of cells
adhered to the culture dish were measured by the WST-1 assay
described above. The absorbance of the dye in the culture directly
correlates with the number of live cells.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Raw264.7 cells were either left unstimulated or
stimulated with 10 ng/ml LPS, 100 ng/ml rmSCRG1, or 10 ng/ml LPS
plus 100 ng/ml rmSCRG1 in the absence and presence of 1 mM U0126,
for 6 h. Total RNA extraction, cDNA synthesis, RT-qPCR were
performed with methods of our previous study (15). Expression of Ccl22 was
normalized to glyceraldehyde-3-phosphate dehydrogenase
(Gapdh). The following primer pairs were used; Ccl22
(sense, 5′-GGCACCTATCCAGTGCCACA-3′ and antisense,
5′-TGGTGGACCAGCCTGAAACTC') and Gapdh (sense,
5′-TGTGTCCGTCGTGGATCTGA-3′ and antisense,
5′-TTGCTGTTGAAGTCGCAGGAG-3′). Relative expression levels were
calculated by the 2−ΔΔCq method (31) as a fold-increase or -decrease.
Primer array
Raw264.7 cells were stimulated with 10 ng/ml LPS,
100 ng/ml rmSCRG1, or 10 ng/ml LPS plus 100 ng/ml rmSCRG1 for 6 h.
Unstimulated cells were used as a control. Gene expression levels
of a range of cytokines and chemokines were measured using a
PrimerArray consisting of mouse cytokines and cytokine receptors
(PN001, Takara Bio) and PrimerArray Analysis Tool version 2.0
(Takara Bio) according to the manufacturer's instructions. Genes
whose expression levels increased more than 100-fold following LPS
stimulation and less than 100-fold following SCRG1 treatment were
identified.
Western blotting
Raw264.7 cells were serum-starved overnight and
stimulated with 100 ng/ml rmSCRG1 for various period time. Western
blotting was performed in our previously reported procedure
(15). The following primary
antibodies that has been purchased from Cell Signaling Technology
(Danvers, MA, USA) were used; anti-p44/42 mitogen-activated protein
kinase (ERK1/2), anti-phospho-ERK1/2, anti-c-Jun N-terminal kinase
(JNK), anti-phospho-JNK, anti-p38, anti-phospho-p38, anti-Akt,
anti-phospho-Akt, anti-focal adhesion kinase (FAK),
anti-phospho-FAK. β-actin level measured were detected with an
anti-β-actin antibody (Santa Cruz Biotechnology, Dallas, TX, USA)
as a loading control. The densitometry of the band measured by
ImageJ version 1.44 software was expressed as the ratio of
phosphorylation to the total molecule.
Enzyme-linked immunosorbent assay
(ELISA)
Raw264.7 cells were left unstimulated or were
stimulated with 10 ng/ml LPS, 100 ng/ml rmSCRG1, or 10 ng/ml LPS
plus 100 ng/ml rmSCRG1 for 48 h. The amount of secreted CCL22 in
the culture medium was measured using a sandwich ELISA kit for
mouse CCL22 (R&D Systems, Inc., Minneapolis, MN, USA). CCL22
levels were quantified according to the manufacturer's
instructions.
Flow cytometry
A total of 1×105 Raw264.7 cells suspended
in PBS containing 2 mM EDTA and 0.5% FBS were incubated with either
a phycoerythrin (PE)-conjugated anti-mouse BST1/CD157 (1:10),
anti-mouse β1 integrin/CD29 (1:10), or anti-mouse β2 integrin/CD18
(1:10) (all from BioLegend, Inc., San Diego, CA, USA) antibody for
1 h at 4°C. Data acquisition and analysis was performed using a
flow cytometer EPICS XL (Beckman Coulter, Brea, CA, USA).
Statistical analysis
All experiments were performed in triplicate.
Numerical data were presented as the mean ± standard deviation
(SD), and significant differences were analyzed by Student's
t-test. P<0.05 were considered statistically
significant.
Results
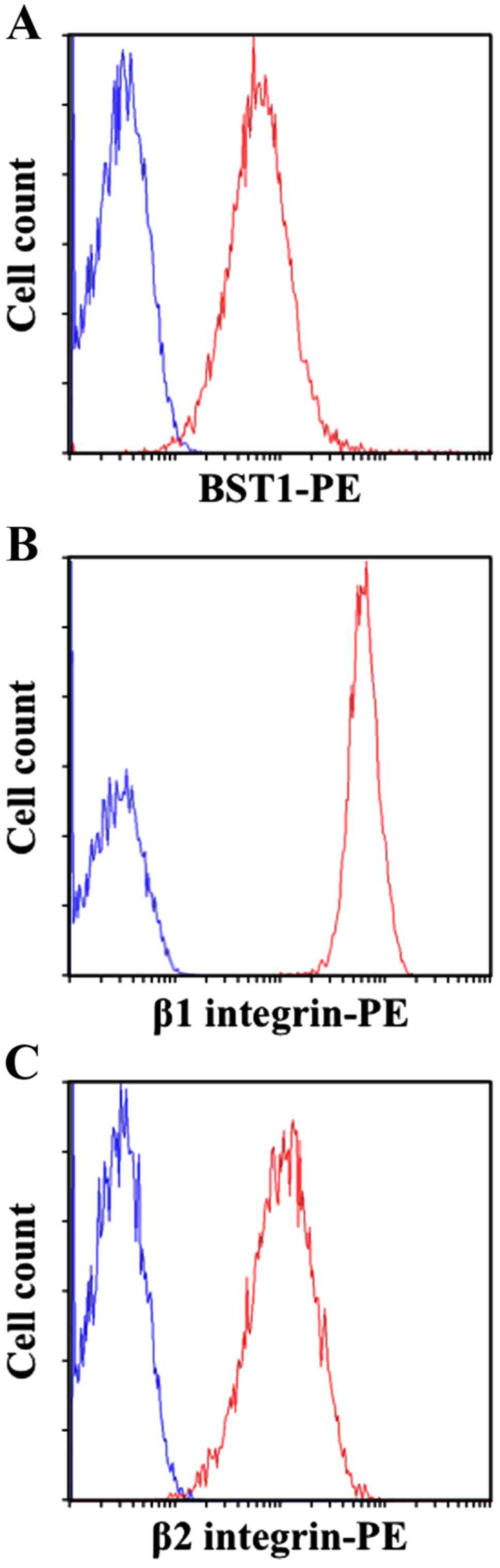
Raw264.7 cells express BST1, and β1
and β2 integrins as an SCRG1 receptor complex
Our recent study revealed that SCRG1 secreted from
MSCs forms a complex with the membrane proteins BST1 and β1
integrin, which acts as a receptor for its autocrine/paracrine
activity (15). Further evidence
for this novel receptor was provided by Lavagno et al, which
showed BST1 also interacts with β1 and β2 integrins at the
neutrophil cell surface (32).
Based on these data, we investigated the expression of BST1, β1
integrin, and β2 integrin in mouse macrophage-like Raw264.7 cells
by flow cytometry. As shown in Fig.
1, Raw264.7 cells co-expressed BST1, β1 integrin, and β2
integrin on the cell surface. These results indicate that Raw264.7
cells express the SCRG1 receptor complex.
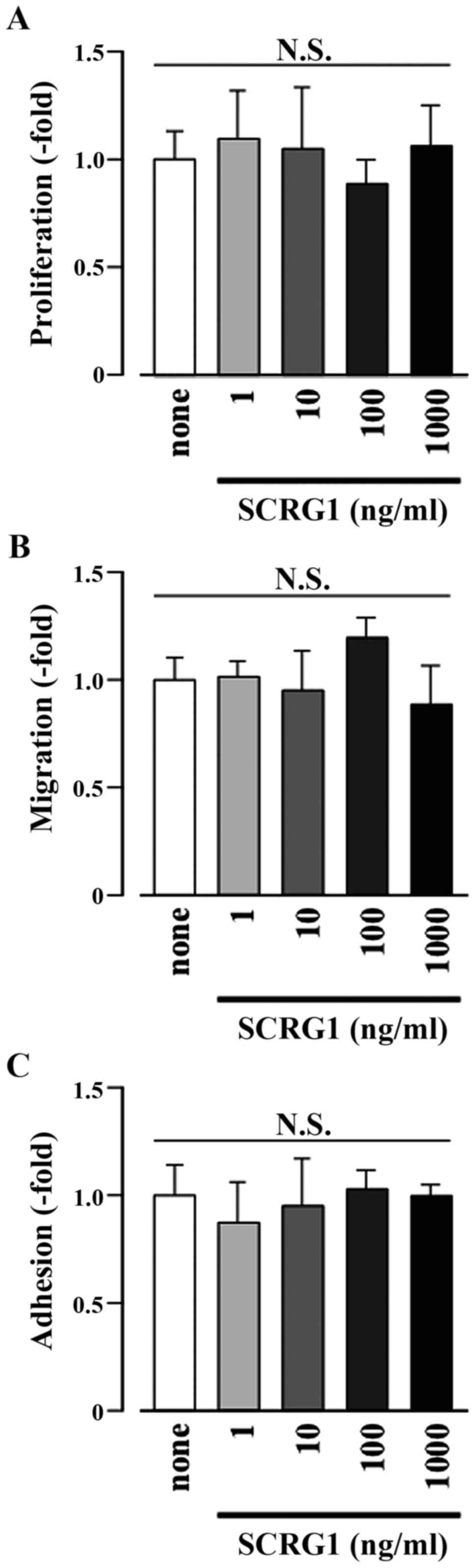
SCRG1 does not affect the statuses of
cell proliferation, migration, and adhesion of Raw264.7 cells
In MSCs, SCRG1 promotes cell migration through the
β1 integrin/focal adhesion kinase (FAK) -dependent phosphoinositide
3-kinase (PI3K)/Akt pathway (15).
Here, to understand the biological processes controlled by SCRG1 in
macrophages, the effects of SCRG1 on cell proliferation, migration,
as well as adhesion activity were investigated. As shown in
Fig. 2, rmSCRG1 did not affect
proliferative, migratory, or adhesive activities in Raw264.7 cells.
Thus, these results indicate that the bioactivity of SCRG1 in
monocyte/macrophage lineage cells is different from that in
MSCs.
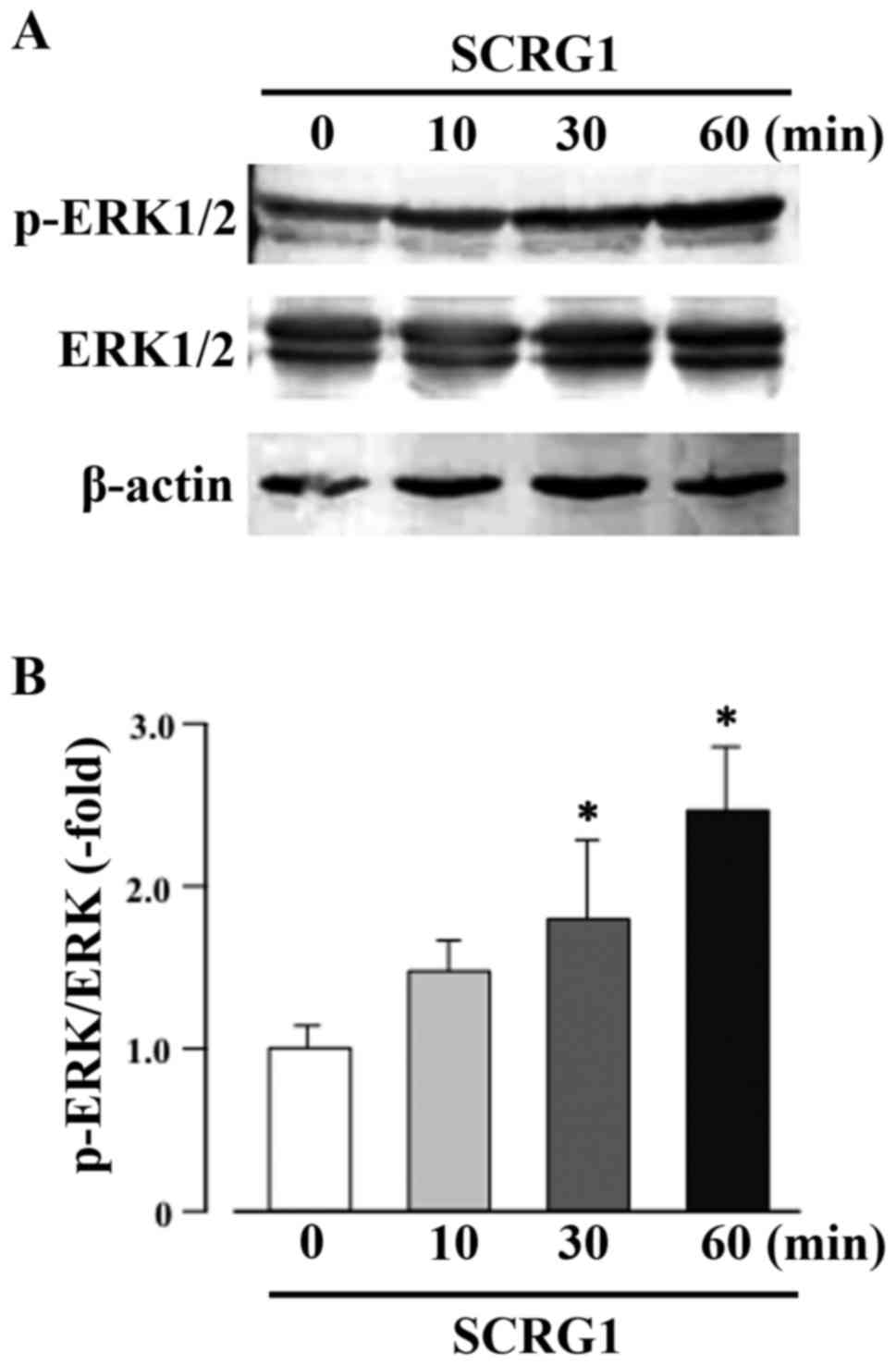
SCRG1 enhances the phosphorylation of
ERK1/2 in Raw264.7 cells
The MAPK pathway, in conjunction with the nuclear
factor-κB (NF-κB) pathway, is closely associated with the
macrophage immune response (33,34).
Accordingly, the intracellular signaling pathways induced by SCRG1
in Raw264.7 cells were investigated. Treatment of cell for 30 min
with rmSCRG1 significantly enhanced the phosphorylation of ERK1/2
(Fig. 3). In contrast,
phosphorylation of FAK, Akt, SAPK/Jun amino-terminal kinase (JNK),
or p38 MAPK by rmSCRG1 treatment in Raw264.7 cells was not observed
(data not shown). These results indicate that SCRG1 specifically
induces the activation of the ERK1/2 pathway.
SCRG1 suppresses LPS-induced CCL22
production through the activation of ERK1/2 in Raw264.7 cells
By primer array analysis, we next investigated the
effects of SCRG1 on the expression of LPS-induced chemokines and
cytokines in Raw264.7 cells. As shown in Table I, rmSCRG1 suppresses the
LPS-induced production of five chemokines and one cytokine in these
cells. In particular, LPS-induced Ccl22 expression was
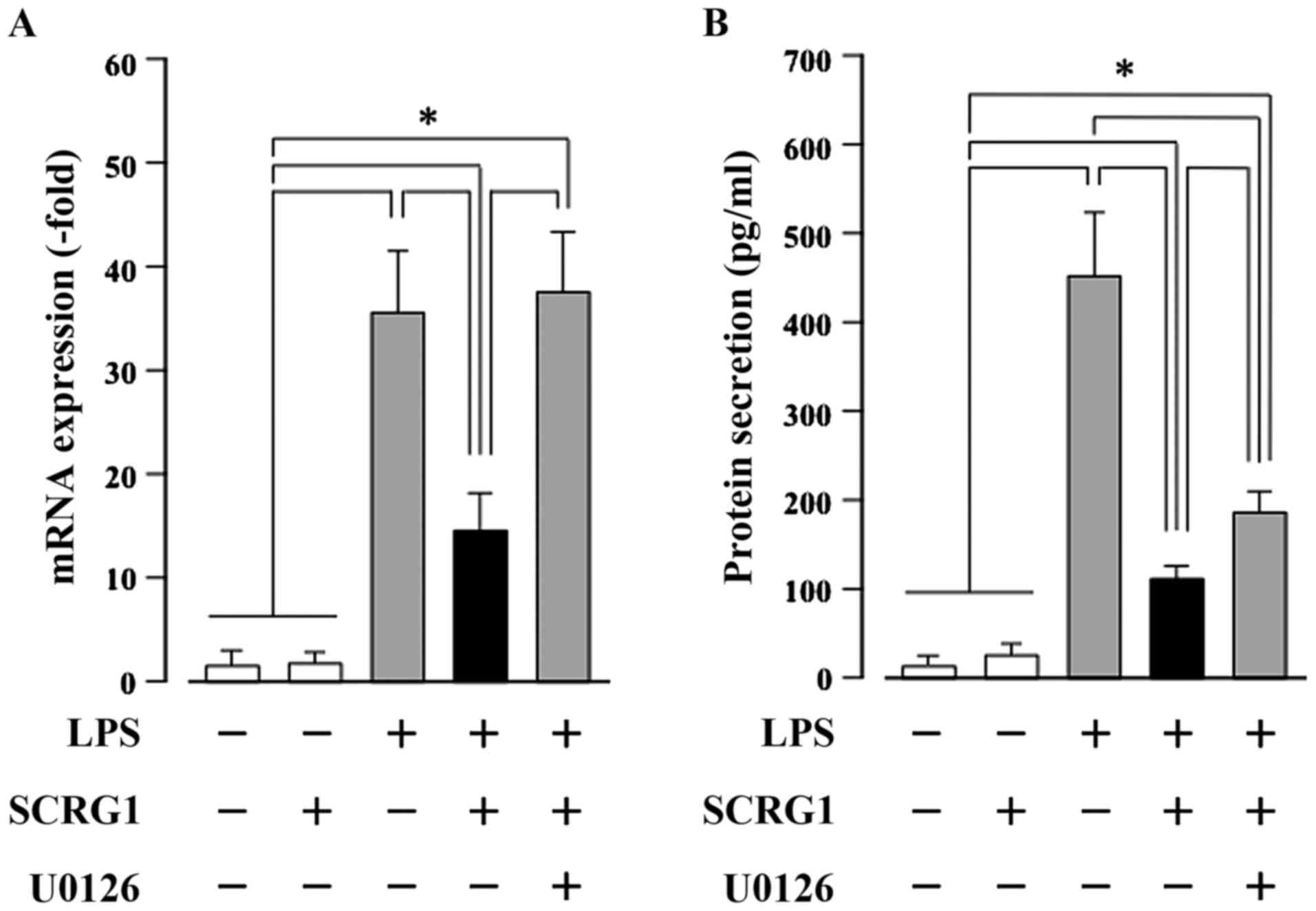
reduced to less than half by rmSCRG1 treatment. Following on from
this, we examined the association between suppression of CCL22
expression and ERK activation by rmSCRG1 in Raw264.7 cells. Both
the LPS-induced increases in CCL22 mRNA expression, and protein
secretion were significantly suppressed by rmSCRG1 treatment in
Raw264.7 cells (Fig. 4). In
addition, this suppressive effect of rmSCRG1 was completely
abolished by treatment with the MAPK/ERK kinase inhibitor, U0126.
These results indicate that LPS-induced CCL22 production in
macrophages was suppressed by treatment with SCRG1 through the
activation of an ERK1/2-mediated signal.
 | Table I.Genes whose expression increased more
than 100-fold after LPS treatment and less than 100-fold by SCRG1
treatment alone relative to unstimulated Raw264.7 cells. |
Table I.
Genes whose expression increased more
than 100-fold after LPS treatment and less than 100-fold by SCRG1
treatment alone relative to unstimulated Raw264.7 cells.
|
| Fold change
relative to unstimulated Raw 264.7 cells |
|---|
|
|
|
|---|
| Gene symbol | Gene name | LPS | LPS+SCRG1 | SCRG1 |
|---|
| Ccl22 | Chemokine (C-C
motif) ligand 22 | 4420.5 | 1520.1 | 96.335 |
| Ccl8 | Chemokine (C-C
motif) ligand 8 | 2856.4 | 1770.5 | 0.84089 |
| Cxcl11 | Chemokine (C-X-C
motif) ligand 11 | 1746.1 | 843.35 | 89.884 |
| Ccl12 | Chemokine (C-C
motif) ligand 12 | 929.29 | 749.61 | 70.521 |
| Csf1 | Colony stimulating
factor 1 | 206.50 | 125.36 | 17.876 |
| Cx3cl1 | Chemokine (C-X3-C
motif) ligand 1 | 154.34 | 115.36 | 1.3755 |
Discussion
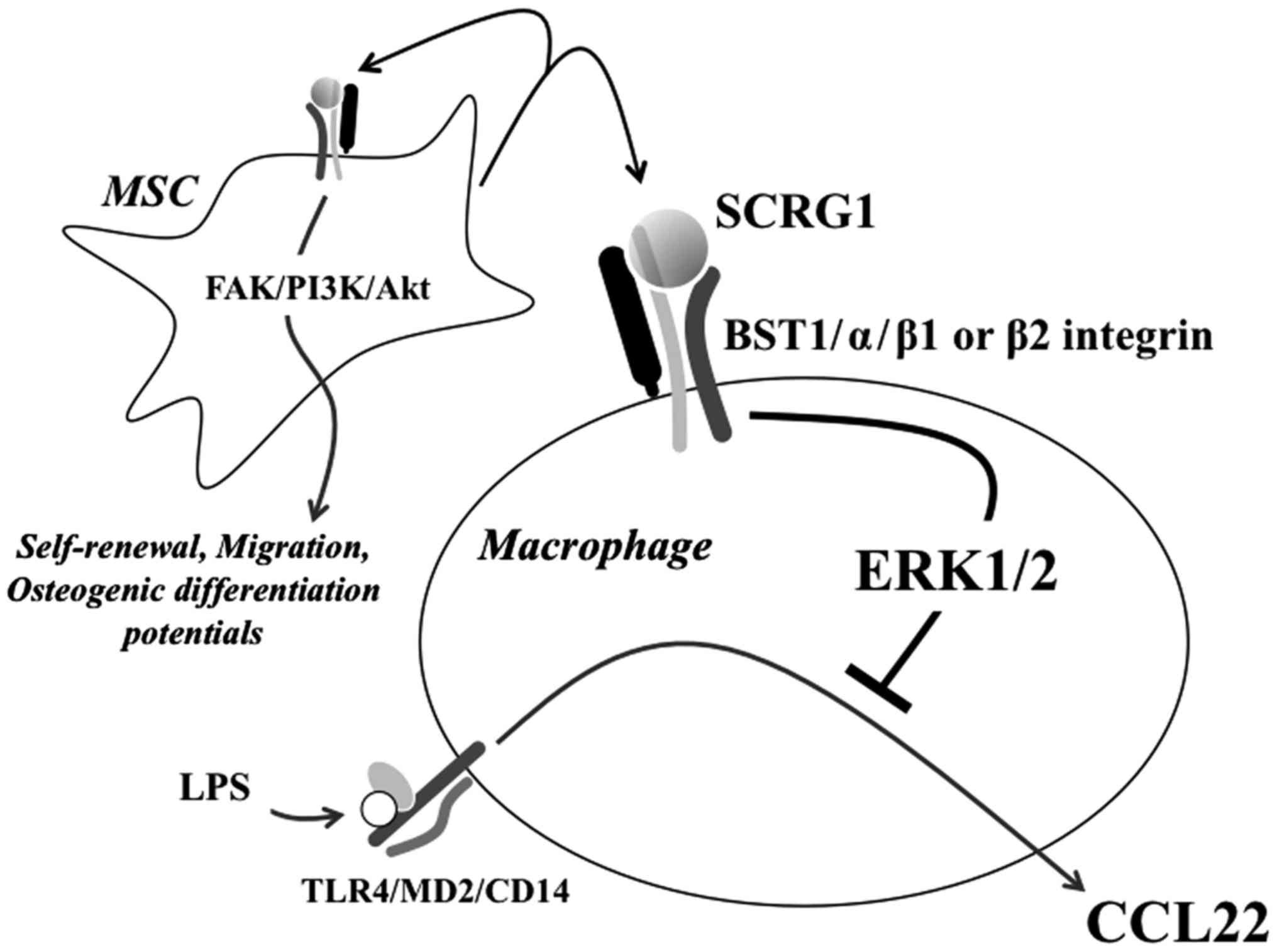
Several roles of SCRG1 suggested in this study and
our previous studies are shown in Fig.
5. Recently, we identified a novel ligand-receptor combination,
SCRG1/BST1, that maintains expressions of the stem cell markers
Oct-4 and CD271/LNGFR in MSC, as well as self-renewal, migration,
and osteogenic differentiation potential during ex vivo
expansion (15). Here, we
investigated the expression of the SCRG1 receptor components, BST1,
β1 integrin, and β2 integrin in mouse macrophage-like Raw264.7
cells. In addition to BST1, these cells also expressed β1 and β2
integrins (Fig. 1). BST1 is an
ectoenzyme with a glycosyl phosphatidylinositol anchor and a
NADase/ADP-ribosyl cyclase activity that belongs to the CD38 family
(35,36). It has been found on the cell
surface of stromal (37) and bone
marrow-derived cells (38), and it
facilitates pre-B-cell growth and induces cell migration (39). Under the condition of a complex of
BST1 with either β1 integrin or β2 integrin, BST1 has been shown to
promote the phosphorylation of FAK through the use of an agonistic
monoclonal antibody (32,40,41).
Furthermore, it has also been reported that BST-1 regulates the
adhesion and migration of leukocytes via phosphorylation of Akt and
MAPKs (42,43). In this study, we also observed that
SCRG1 enhanced the phosphorylation of ERK1/2 in Raw264.7 cells
(Fig. 3).
Notably, SCRG1 did not enhance cell proliferation,
migration, or adhesion of Raw264.7 cells (Fig. 2), indicating a different biological
function in these cells. In support of this, we demonstrated that
SCRG1 suppresses LPS-induced CCL22 production in these mouse
macrophage-like Raw264.7 cells in a MAPK/ERK-dependent manner
(Table I, Fig. 4). LPS and other microbial products
are recognized by the host's toll-like receptors (TLRs) family
members (44). LPS interacts with
a heterologous receptor involving TLR4 (45,46),
CD14 (47,48) and MD2 (49–51).
By LPS binds to the TLR4, two major signaling pathways via the
adapter molecules TIR-domain-containing adaptor inducing IFN-β
(TRIF) or myeloid differentiation factor 88 (MyD88) are activated
(45,52–55).
Significantly, both the Myd88 and TRIF pathways result in
activation of the transcription factor, NF-κB, a central regulator
of the LPS response, which induces cytokine and chemokine
production as well as stress responses in many cell types,
including macrophages (53,54).
NF-κB is involved in regulating the expression of multiple genes
involved in inflammatory and immune responses (56). Innate and adaptive immune response
are mainly regulated by MAPKs, including ERK, JNK, and p38 MAPK, in
addition to the NF-κB pathway. In macrophages, activation of the
MEK/ERK pathway by bacterial infection regulates various
inflammatory responses (57–61).
The MEK/ERK pathway is one of the most studied intracellular
signaling pathways in monocyte-derived macrophages activated by
LPS-induced pro-inflammatory responses (62–64).
In addition, findings have been previously reported that
NF-κB-dependent gene expression is controlled by the activation of
ERK1/2 without affecting DNA binding (65–69).
Therefore, our results indicate that the SCRG1-induced ERK1/2
signaling pathway controls NF-κB-dependent chemokine CCL22
expression.
CCL22, a member of the CC chemokine families, is
mainly produced by monocyte-derived macrophages, mast cells, and
inflammatory dendritic cells upon stimulation with microbial
products (70,71). CCL22 binds to its receptor CCR4
plays an important homeostatic role in leukocyte trafficking,
activation of innate immune cells, Th2 immunopathology, as well as
accumulation of regulatory T (Treg) cells in solid tumor (72–74).
CCL 22 also plays a role in recruiting Treg cells into synovial
fluid in inflammatory diseases like rheumatoid arthritis (75). CCL22 plays an important role in a
variety of other diseases, including allergic rhinitis (76), atopic dermatitis (77), and lymphoma (78). Recent studies suggested that CCL22
can be used as a biomarker for autoimmune diseases (79). On the other hand, the role of CCL
22 in regulation of immune homeostasis is unclear (80,81).
In conclusion, here we clearly demonstrate that
SCRG1, which in vivo could be derived from MSCs, suppresses
LPS-induced chemokine CCL22 production in Raw264.7 macrophages. The
mechanism appears to involve the MAPK ERK1/2 pathway, since SCRG1
induced the phosphorylation of ERK1/2 and a MAPK/ERK kinase
inhibitor U0126 ablated the suppressive effect of SCRG1 on
LPS-induced chemokine CCL22 production. These results suggest a
model whereby MSCs play their immunosuppressive role by secreting
SCRG1, which then suppresses microbial products-induced chemokine
expression in monocyte/macrophage lineage cells in a paracrine
fashion. We have additionally established fluorescently tagged
immortalized MSC lines derived from different tissues of GFP- and
tdTomato-transgenic mice (25,82).
These cell lines can be used for in vivo imaging analyses on
proliferation and differentiation of MSCs, as well as in
vivo imaging studies to test cell therapies and regenerative
medicine techniques, providing insight into diseases such as bone
and immune disorders, fibrosis, and cancer progression or
metastasis. Our findings provide new insights into the molecular
mechanisms of MSCs, as well as a novel perspective for
understanding the immune regulatory mechanisms of MSCs.
Acknowledgements
The present study was supported in part by the JSPS
KAKENHI grant nos. JP25463053 and JP16K11654 awarded to N.C.,
JP26893249 and JP16K20652 awarded to E.K. and JP26670852 and
JP16H05534 awarded to A.I.; and a grant from the Keiryokai Research
Foundation grant no. 120 awarded to N.C., 2013.
References
|
1
|
Prockop DJ: Marrow stromal cells as stem
cells for nonhematopoietic tissues. Science. 276:71–74. 1997.
View Article : Google Scholar
|
|
2
|
Uccelli A, Moretta L and Pistoia V:
Mesenchymal stem cells in health and disease. Nat Rev Immunol.
8:726–736. 2008. View
Article : Google Scholar
|
|
3
|
Shi Y, Su J, Roberts AI, Shou P, Rabson AB
and Ren G: How mesenchymal stem cells interact with tissue immune
responses. Trends Immunol. 33:136–143. 2012. View Article : Google Scholar :
|
|
4
|
Le Blanc K, Rasmusson I, Sundberg B,
Götherström C, Hassan M, Uzunel M and Ringdén O: Treatment of
severe acute graft-versus-host disease with third party
haploidentical mesenchymal stem cells. Lancet. 363:1439–1441. 2004.
View Article : Google Scholar
|
|
5
|
Stemberger S, Jamnig A, Stefanova N,
Lepperdinger G, Reindl M and Wenning GK: Mesenchymal stem cells in
a transgenic mouse model of multiple system atrophy:
Immunomodulation and neuroprotection. PLoS One. 6:e198082011.
View Article : Google Scholar :
|
|
6
|
Ortiz LA, Dutreil M, Fattman C, Pandey AC,
Torres G, Go K and Phinney DG: Interleukin 1 receptor antagonist
mediates the antiinflammatory and antifibrotic effect of
mesenchymal stem cells during lung injury. Proc Natl Acad Sci USA.
104:pp. 11002–11007. 2007; View Article : Google Scholar :
|
|
7
|
Lee RH, Pulin AA, Seo MJ, Kota DJ,
Ylostalo J, Larson BL, Semprun-Prieto L, Delafontaine P and Prockop
DJ: Intravenous hMSCs improve myocardial infarction in mice because
cells embolized in lung are activated to secrete the
anti-inflammatory protein TSG-6. Cell Stem Cell. 5:54–63. 2009.
View Article : Google Scholar :
|
|
8
|
Tögel F, Hu Z, Weiss K, Isaac J, Lange C
and Westenfelder C: Administered mesenchymal stem cells protect
against ischemic acute renal failure through
differentiation-independent mechanisms. Am J Physiol Renal Physiol.
289:F31–F42. 2005. View Article : Google Scholar
|
|
9
|
Sheikh AM, Nagai A, Wakabayashi K,
Narantuya D, Kobayashi S, Yamaguchi S and Kim SU: Mesenchymal stem
cell transplantation modulates neuroinflammation in focal cerebral
ischemia: Contribution of fractalkine and IL-5. Neurobiol Dis.
41:717–724. 2011. View Article : Google Scholar
|
|
10
|
Lee JK, Jin HK, Endo S, Schuchman EH,
Carter JE and Bae JS: Intracerebral transplantation of bone
marrow-derived mesenchymal stem cells reduces amyloid-beta
deposition and rescues memory deficits in Alzheimer's disease mice
by modulation of immune responses. Stem Cells. 28:329–343.
2010.
|
|
11
|
Lda S Meirelles, Fontes AM, Covas DT and
Caplan AI: Mechanisms involved in the therapeutic properties of
mesenchymal stem cells. Cytokine Growth Factor Rev. 20:419–427.
2009. View Article : Google Scholar
|
|
12
|
Di Nicola M, Carlo-Stella C, Magni M,
Milanesi M, Longoni PD, Matteucci P, Grisanti S and Gianni AM:
Human bone marrow stromal cells suppress T-lymphocyte proliferation
induced by cellular or nonspecific mitogenic stimuli. Blood.
99:3838–3843. 2002. View Article : Google Scholar
|
|
13
|
Ooi YY, Ramasamy R, Rahmat Z, Subramaiam
H, Tan SW, Abdullah M, Israf DA and Vidyadaran S: Bone
marrow-derived mesenchymal stem cells modulate BV2 microglia
responses to lipopolysaccharide. Int Immunopharmacol. 10:1532–1540.
2010. View Article : Google Scholar
|
|
14
|
Németh K, Leelahavanichkul A, Yuen PS,
Mayer B, Parmelee A, Doi K, Robey PG, Leelahavanichkul K, Koller
BH, Brown JM, et al: Bone marrow stromal cells attenuate sepsis via
prostaglandin E(2)-dependent reprogramming of host macrophages to
increase their interleukin-10 production. Nat Med. 15:42–49. 2009.
View Article : Google Scholar
|
|
15
|
Aomatsu E, Takahashi N, Sawada S, Okubo N,
Hasegawa T, Taira M, Miura H, Ishisaki A and Chosa N: Novel
SCRG1/BST1 axis regulates self-renewal, migration, and osteogenic
differentiation potential in mesenchymal stem cells. Sci Rep.
4:36522014. View Article : Google Scholar :
|
|
16
|
Dandoy-Dron F, Guillo F, Benboudjema L,
Deslys JP, Lasmézas C, Dormont D, Tovey MG and Dron M: Gene
expression in scrapie. Cloning of a new scrapie-responsive gene and
the identification of increased levels of seven other mRNA
transcripts. J Biol Chem. 273:7691–7697. 1998. View Article : Google Scholar
|
|
17
|
Dron M, Bailly Y, Beringue V, Haeberlé AM,
Griffond B, Risold PY, Tovey MG, Laude H and Dandoy-Dron F: Scrg1
is induced in TSE and brain injuries and associated with autophagy.
Eur J Neurosci. 22:133–146. 2005. View Article : Google Scholar
|
|
18
|
Dron M, Bailly Y, Beringue V, Haeberlé AM,
Griffond B, Risold PY, Tovey MG, Laude H and Dandoy-Dron F: SCRG1,
a potential marker of autophagy in transmissible spongiform
encephalopathies. Autophagy. 2:58–60. 2006. View Article : Google Scholar
|
|
19
|
Dron M, Dandoy-Dron F, Guillo F,
Benboudjema L, Hauw JJ, Lebon P, Dormont D and Tovey MG:
Characterization of the human analogue of a Scrapie-responsive
gene. J Biol Chem. 273:18015–18018. 1998. View Article : Google Scholar
|
|
20
|
Dron M, Tartare X, Guillo F, Haik S,
Barbin G, Maury C, Tovey M and Dandoy-Dron F: Mouse scrapie
responsive gene 1 (Scrg1): Genomic organization, physical linkage
to sap30, genetic mapping on chromosome 8 and expression in
neuronal primary cell cultures. Genomics. 70:140–149. 2000.
View Article : Google Scholar
|
|
21
|
Aggarwal S and Pittenger MF: Human
mesenchymal stem cells modulate allogeneic immune cell responses.
Blood. 105:1815–1822. 2005. View Article : Google Scholar
|
|
22
|
Spaggiari GM, Capobianco A, Abdelrazik H,
Becchetti F, Mingari MC and Moretta L: Mesenchymal stem cells
inhibit natural killer-cell proliferation, cytotoxicity and
cytokine production: Role of indoleamine 2,3-dioxygenase and
prostaglandin E2. Blood. 111:1327–1333. 2008. View Article : Google Scholar
|
|
23
|
Sato K, Ozaki K, Oh I, Meguro A, Hatanaka
K, Nagai T, Muroi K and Ozawa K: Nitric oxide plays a critical role
in suppression of T-cell proliferation by mesenchymal stem cells.
Blood. 109:228–234. 2007. View Article : Google Scholar
|
|
24
|
Lee RH, Oh JY, Choi H and Bazhanov N:
Therapeutic factors secreted by mesenchymal stromal cells and
tissue repair. J Cell Biochem. 112:3073–3078. 2011. View Article : Google Scholar
|
|
25
|
Sawada S, Chosa N, Takizawa N, Yokota J,
Igarashi Y, Tomoda K, Kondo H, Yaegashi T and Ishisaki A:
Establishment of mesenchymal stem cell lines derived from the bone
marrow of green fluorescent protein-transgenic mice exhibiting a
diversity in intracellular transforming growth factor-β and bone
morphogenetic protein signaling. Mol Med Rep. 13:2023–2031.
2016.
|
|
26
|
Igarashi Y, Chosa N, Sawada S, Kondo H,
Yaegashi T and Ishisaki A: VEGF-C and TGF-β reciprocally regulate
mesenchymal stem cell commitment to differentiation into lymphatic
endothelial or osteoblastic phenotypes. Int J Mol Med.
37:1005–1013. 2016.
|
|
27
|
Godiska R, Chantry D, Raport CJ, Sozzani
S, Allavena P, Leviten D, Mantovani A and Gray PW: Human
macrophage-derived chemokine (MDC), a novel chemoattractant for
monocytes, monocyte-derived dendritic cells and natural killer
cells. J Exp Med. 185:1595–1604. 1997. View Article : Google Scholar :
|
|
28
|
Chang Ms, McNinch J, Elias C III, Manthey
CL, Grosshans D, Meng T, Boone T and Andrew DP: Molecular cloning
and functional characterization of a novel CC chemokine, stimulated
T cell chemotactic protein (STCP-1) that specifically acts on
activated T lymphocytes. J Biol Chem. 272:25229–25237. 1997.
View Article : Google Scholar
|
|
29
|
Schaniel C, Pardali E, Sallusto F,
Speletas M, Ruedl C, Shimizu T, Seidl T, Andersson J, Melchers F,
Rolink AG and Sideras P: Activated murine B lymphocytes and
dendritic cells produce a novel CC chemokine which acts selectively
on activated T cells. J Exp Med. 188:451–463. 1998. View Article : Google Scholar :
|
|
30
|
Mantovani A, Gray PA, Van Damme J and
Sozzani S: Macrophage-derived chemokine (MDC). J Leukoc Biol.
68:400–404. 2000.
|
|
31
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
32
|
Lavagno L, Ferrero E, Ortolan E, Malavasi
F and Funaro A: CD157 is part of a supramolecular complex with
CD11b/CD18 on the human neutrophil cell surface. J Biol Regul
Homeost Agents. 21:5–11. 2007.
|
|
33
|
Arthur JS and Ley SC: Mitogen-activated
protein kinases in innate immunity. Nat Rev Immunol. 13:679–692.
2013. View
Article : Google Scholar
|
|
34
|
Huang G, Shi LZ and Chi H: Regulation of
JNK and p38 MAPK in the immune system: Signal integration,
propagation and termination. Cytokine. 48:161–169. 2009. View Article : Google Scholar :
|
|
35
|
Ishihara K and Hirano T: BST-1/CD157
regulates the humoral immune responses in vivo. Chem Immunol.
75:235–255. 2000. View Article : Google Scholar
|
|
36
|
Malavasi F, Deaglio S, Funaro A, Ferrero
E, Horenstein AL, Ortolan E, Vaisitti T and Aydin S: Evolution and
function of the ADP ribosyl cyclase/CD38 gene family in physiology
and pathology. Physiol Rev. 88:841–886. 2008. View Article : Google Scholar
|
|
37
|
Kaisho T, Ishikawa J, Oritani K, Inazawa
J, Tomizawa H, Muraoka O, Ochi T and Hirano T: BST-1, a surface
molecule of bone marrow stromal cell lines that facilitates
pre-B-cell growth. Proc Natl Acad Sci USA. 91:pp. 5325–5329. 1994;
View Article : Google Scholar :
|
|
38
|
Goldstein SC and Todd RF III: Structural
and biosynthetic features of the Mo5 human myeloid differentiation
antigen. Tissue Antigens. 41:214–218. 1993. View Article : Google Scholar
|
|
39
|
Funaro A, Ortolan E, Bovino P, Lo Buono N,
Nacci G, Parrotta R, Ferrero E and Malavasi F: Ectoenzymes and
innate immunity: The role of human CD157 in leukocyte trafficking.
Front Biosci (Landmark Ed). 14:929–943. 2009. View Article : Google Scholar
|
|
40
|
Hussain AM, Lee HC and Chang CF:
Functional expression of secreted mouse BST-1 in yeast. Protein
Expr Purif. 12:133–137. 1998. View Article : Google Scholar
|
|
41
|
Okuyama Y, Ishihara K, Kimura N, Hirata Y,
Sato K, Itoh M, Ok LB and Hirano T: Human BST-1 expressed on
myeloid cells functions as a receptor molecule. Biochem Biophys Res
Commun. 228:838–845. 1996. View Article : Google Scholar
|
|
42
|
Lo Buono N, Parrotta R, Morone S, Bovino
P, Nacci G, Ortolan E, Horenstein AL, Inzhutova A, Ferrero E and
Funaro A: The CD157-integrin partnership controls transendothelial
migration and adhesion of human monocytes. J Biol Chem.
286:18681–18691. 2011. View Article : Google Scholar :
|
|
43
|
Funaro A, Ortolan E, Ferranti B, Gargiulo
L, Notaro R, Luzzatto L and Malavasi F: CD157 is an important
mediator of neutrophil adhesion and migration. Blood.
104:4269–4278. 2004. View Article : Google Scholar
|
|
44
|
Takeda K, Kaisho T and Akira S: Toll-like
receptors. Annu Rev Immunol. 21:335–376. 2003. View Article : Google Scholar
|
|
45
|
Poltorak A, He X, Smirnova I, Liu MY, Van
Huffel C, Du X, Birdwell D, Alejos E, Silva M, Galanos C, et al:
Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: Mutations
in Tlr4 gene. Science. 282:2085–2088. 1998. View Article : Google Scholar
|
|
46
|
Qureshi ST, Larivière L, Leveque G,
Clermont S, Moore KJ, Gros P and Malo D: Endotoxin-tolerant mice
have mutations in Toll-like receptor 4 (Tlr4). J Exp Med.
189:615–625. 1999. View Article : Google Scholar :
|
|
47
|
Wright SD: CD14 and innate recognition of
bacteria. J Immunol. 155:6–8. 1995.
|
|
48
|
Ulevitch RJ and Tobias PS:
Receptor-dependent mechanisms of cell stimulation by bacterial
endotoxin. Annu Rev Immunol. 13:437–457. 1995. View Article : Google Scholar
|
|
49
|
Shimazu R, Akashi S, Ogata H, Nagai Y,
Fukudome K, Miyake K and Kimoto M: MD-2, a molecule that confers
lipopolysaccharide responsiveness on toll-like receptor 4. J Exp
Med. 189:1777–1782. 1999. View Article : Google Scholar :
|
|
50
|
Akashi S, Shimazu R, Ogata H, Nagai Y,
Takeda K, Kimoto M and Miyake K: Cutting edge: Cell surface
expression and lipopolysaccharide signaling via the toll-like
receptor 4-MD-2 complex on mouse peritoneal macrophages. J Immunol.
164:3471–3475. 2000. View Article : Google Scholar
|
|
51
|
Viriyakosol S, Tobias PS, Kitchens RL and
Kirkland TN: MD-2 binds to bacterial lipopolysaccharide. J Biol
Chem. 276:38044–38051. 2001.
|
|
52
|
O'Neill LA, Dunne A, Edjeback M, Gray P,
Jefferies C and Wietek C: Mal and MyD88: Adapter proteins involved
in signal transduction by toll-like receptors. J Endotoxin Res.
9:55–59. 2003. View Article : Google Scholar
|
|
53
|
Kawai T, Adachi O, Ogawa T, Takeda K and
Akira S: Unresponsiveness of MyD88-deficient mice to endotoxin.
Immunity. 11:115–122. 1999. View Article : Google Scholar
|
|
54
|
Kawai T, Takeuchi O, Fujita T, Inoue J,
Mühlradt PF, Sato S, Hoshino K and Akira S: Lipopolysaccharide
stimulates the MyD88-independent pathway and results in activation
of IFN-regulatory factor 3 and the expression of a subset of
lipopolysaccharide-inducible genes. J Immunol. 167:5887–5894. 2001.
View Article : Google Scholar
|
|
55
|
West AP, Koblansky AA and Ghosh S:
Recognition and signaling by toll-like receptors. Annu Rev Cell Dev
Biol. 22:409–437. 2006. View Article : Google Scholar
|
|
56
|
Ghosh S and Karin M: Missing pieces in the
NF-kappaB puzzle. Cell. 109:S81–S96. 2002. View Article : Google Scholar
|
|
57
|
Gordon S and Martinez FO: Alternative
activation of macrophages: Mechanism and functions. Immunity.
32:593–604. 2010. View Article : Google Scholar
|
|
58
|
Nathan C: Points of control in
inflammation. Nature. 420:846–852. 2002. View Article : Google Scholar
|
|
59
|
Mantovani A, Sica A, Sozzani S, Allavena
P, Vecchi A and Locati M: The chemokine system in diverse forms of
macrophage activation and polarization. Trends Immunol. 25:677–686.
2004. View Article : Google Scholar
|
|
60
|
Nathan C and Ding A: Nonresolving
inflammation. Cell. 140:871–882. 2010. View Article : Google Scholar
|
|
61
|
Martinez FO, Gordon S, Locati M and
Mantovani A: Transcriptional profiling of the human
monocyte-to-macrophage differentiation and polarization: New
molecules and patterns of gene expression. J Immunol.
177:7303–7311. 2006. View Article : Google Scholar
|
|
62
|
Rao KM: MAP kinase activation in
macrophages. J Leukoc Biol. 69:3–10. 2001.
|
|
63
|
Rao KM, Meighan T and Bowman L: Role of
mitogen-activated protein kinase activation in the production of
inflammatory mediators: Differences between primary rat alveolar
macrophages and macrophage cell lines. J Toxicol Environ Health A.
65:757–768. 2002. View Article : Google Scholar
|
|
64
|
Bain J, Plater L, Elliott M, Shapiro N,
Hastie CJ, McLauchlan H, Klevernic I, Arthur JS, Alessi DR and
Cohen P: The selectivity of protein kinase inhibitors: A further
update. Biochem J. 408:297–315. 2007. View Article : Google Scholar :
|
|
65
|
Akhtar M, Watson JL, Nazli A and McKay DM:
Bacterial DNA evokes epithelial IL-8 production by a
MAPK-dependent, NF-kappaB-independent pathway. FASEB J.
17:1319–1321. 2003.
|
|
66
|
Fiebich BL, Schleicher S, Butcher RD,
Craig A and Lieb K: The neuropeptide substance P activates p38
mitogen-activated protein kinase resulting in IL-6 expression
independently from NF-kappa B. J Immunol. 165:5606–5611. 2000.
View Article : Google Scholar
|
|
67
|
Patel DN, King CA, Bailey SR, Holt JW,
Venkatachalam K, Agrawal A, Valente AJ and Chandrasekar B:
Interleukin-17 stimulates C-reactive protein expression in
hepatocytes and smooth muscle cells via p38 MAPK and
ERK1/2-dependent NF-kappaB and C/EBPbeta activation. J Biol Chem.
282:27229–27238. 2007. View Article : Google Scholar
|
|
68
|
Tokuda M, Miyamoto R, Sakuta T, Nagaoka S
and Torii M: Substance P activates p38 mitogen-activated protein
kinase to promote IL-6 induction in human dental pulp fibroblasts.
Connect Tissue Res. 46:153–158. 2005. View Article : Google Scholar
|
|
69
|
Zampetaki A, Mitsialis SA, Pfeilschifter J
and Kourembanas S: Hypoxia induces macrophage inflammatory
protein-2 (MIP-2) gene expression in murine macrophages via
NF-kappaB: The prominent role of p42/p44 and PI3 kinase pathways.
FASEB J. 18:1090–1092. 2004.
|
|
70
|
Yamashita U and Kuroda E: Regulation of
macrophage-derived chemokine (MDC, CCL22) production. Crit Rev
Immunol. 22:105–114. 2002. View Article : Google Scholar
|
|
71
|
Layseca-Espinosa E, Korniotis S, Montandon
R, Gras C, Bouillié M, Gonzalez-Amaro R, Dy M and Zavala F:
CCL22-producing CD8α- myeloid dendritic cells mediate regulatory T
cell recruitment in response to G-CSF treatment. J Immunol.
191:2266–2272. 2013. View Article : Google Scholar
|
|
72
|
Iellem A, Mariani M, Lang R, Recalde H,
Panina-Bordignon P, Sinigaglia F and D'Ambrosio D: Unique
chemotactic response profile and specific expression of chemokine
receptors CCR4 and CCR8 by CD4(+)CD25(+) regulatory T cells. J Exp
Med. 194:847–853. 2001. View Article : Google Scholar :
|
|
73
|
Curiel TJ, Coukos G, Zou L, Alvarez X,
Cheng P, Mottram P, Evdemon-Hogan M, Conejo-Garcia JR, Zhang L,
Burow M, et al: Specific recruitment of regulatory T cells in
ovarian carcinoma fosters immune privilege and predicts reduced
survival. Nat Med. 10:942–949. 2004. View
Article : Google Scholar
|
|
74
|
Li YQ, Liu FF, Zhang XM, Guo XJ, Ren MJ
and Fu L: Tumor secretion of CCL22 activates intratumoral treg
infiltration and is independent prognostic predictor of breast
cancer. PLoS One. 8:e763792013. View Article : Google Scholar :
|
|
75
|
Flytlie HA, Hvid M, Lindgreen E,
Kofod-Olsen E, Petersen EL, Jørgensen A, Deleuran M, Vestergaard C
and Deleuran B: Expression of MDC/CCL22 and its receptor CCR4 in
rheumatoid arthritis, psoriatic arthritis and osteoarthritis.
Cytokine. 49:24–29. 2010. View Article : Google Scholar
|
|
76
|
Yanai M, Sato K, Aoki N, Takiyama Y,
Oikawa K, Kobayashi H, Kimura S, Harabuchi Y and Tateno M: The role
of CCL22/macrophage-derived chemokine in allergic rhinitis. Clin
Immunol. 125:291–298. 2007. View Article : Google Scholar
|
|
77
|
Nakazato J, Kishida M, Kuroiwa R, Fujiwara
J, Shimoda M and Shinomiya N: Serum levels of Th2 chemokines,
CCL17, CCL22 and CCL27, were the important markers of severity in
infantile atopic dermatitis. Pediatr Allergy Immunol. 19:605–613.
2008.
|
|
78
|
Niens M, Visser L, Nolte IM, van der
Steege G, Diepstra A, Cordano P, Jarrett RF, Te Meerman GJ, Poppema
S and van den Berg A: Serum chemokine levels in hodgkin lymphoma
patients: Highly increased levels of CCL17 and CCL22. Br J
Haematol. 140:527–536. 2008. View Article : Google Scholar
|
|
79
|
Jafarzadeh A, Ebrahimi HA, Bagherzadeh S,
Zarkesh F, Iranmanesh F, Najafzadeh A, Khosravimashizi A, Nemati M,
Sabahi A, Hajghani H, et al: Lower serum levels of Th2-related
chemokine CCL22 in women patients with multiple sclerosis: A
comparison between patients and healthy women. Inflammation.
37:604–610. 2014. View Article : Google Scholar
|
|
80
|
Nagata S: Apoptosis and autoimmune
diseases. Ann N Y Acad Sci. 1209:10–16. 2010. View Article : Google Scholar
|
|
81
|
Szondy Z, Garabuczi E, Joós G, Tsay GJ and
Sarang Z: Impaired clearance of apoptotic cells in chronic
inflammatory diseases: Therapeutic implications. Front Immunol.
5:3542014. View Article : Google Scholar :
|
|
82
|
Furukawa S, Kuwajima Y, Chosa N, Satoh K,
Ohtsuka M and Miura H, Kimura M, Inoko H, Ishisaki A, Fujimura A
and Miura H: Establishment of immortalized mesenchymal stem cells
derived from the submandibular glands of td to mato transgenic
mice. Exp Ther Med. 10:1380–1386. 2015.
|