Introduction
MicroRNAs (miRNAs) are short (20–22 nucleotides in
length), single-stranded, noncoding regulatory ribonucleic acids
that participate in gene expression modulation (1–5) by
complementary base paring to its target mRNA. miRNAs have been
implicated as having a role in a number of cellular events,
including proliferation, differentiation, communication and
apoptosis (6), as well as in the
regulation of multiple physiological and pathological processes,
such as DNA methylation, chromatin modification and oncogenesis
(7,8). A large majority of miRNAs exist as
members of a large family with similar nucleotide sequences,
therefore it may be hard to specifically discriminate one
individual family member from others using conventional polymerase
chain reaction (PCR).
Originally identified in Caenorhabditis
elegans in 2000, a homolog of lethal-7 (let-7)
was subsequently identified in humans and other mammalian species
that exhibited numerous functions (9). The let-7 family is the largest
miRNA family discovered to date, and it is ubiquitously expressed.
In Homo sapiens, at least nine let-7 family members
have been identified that are highly evolutionarily conserved,
including let-7a-g, let-7i and microRNA
(miR)-98 (10,11).
The let-7 family not only regulates a series of crucial
physiological functions, such as growth, development and
homeostasis, but also acts as a suppressor that impedes tumour
generation and growth (12).
Indeed, plenty of evidence revealed that let-7 was
downregulated in malignant tumours (13,14).
For example, the expression level of let-7 was previously
correlated with the degree of tumour malignancy, which suggested a
significant role for let-7 expression signatures in cancer
diagnosis and prognosis evaluation (15). However, owing to their short length
and sequence similarity, the identification of an individual
let-7 gene from the other family members, and the subsequent
accurate quantitative profiling for each mature let-7 miRNA
in neoplasms remains challenging.
Current methods that have been extensively used for
the detection and quantification of miRNAs largely depend on
poly(A) tailing and stem-loop reverse transcription-quantitative
PCR (RT-qPCR). A previous study reported a novel stem-loop RT-qPCR
in 2005 (16), in which a
specially designed stem-loop RT primer that hybridized to 6–8
nucleotides at the 3′-end of mature miRNAs and was able to reverse
transcribe them. Subsequently, the products were subjected to
TaqMan-based conventional qPCR using specific forward primers and
the aforementioned stem-loop reverse primers. Although high
sensitivity and specificity were observed using this method
compared with conventional qPCR, it may not be used extensively on
account of the expensive TaqMan probes and the low amplification
efficiency attained using TaqMan miRNA assays coupled with qPCR.
Furthermore, the linear primers used in the poly (A) tailing
approach may not distinguish between mature miRNAs and primary
miRNA precursors, which may lead to relatively poor amplification
specificity.
The present study provided a modified stem-loop
RT-based qPCR strategy for the specific and sensitive measurement
of individual let-7 miRNA family members using a SYBR
green-based miRNA qPCR assay. This approach was able to
specifically detect and quantify individual let-7 genes, of
which the expression signatures may serve as potential biomarkers
for various disorders.
Materials and methods
Cell culture and total RNA
extraction
The U87 human glioblastoma cell line was purchased
from the American Type Culture Collection (Manassas, VA, USA) and
cultured in Dulbecco's modified Eagle's medium (Invitrogen; Thermo
Fisher Scientific, Inc., Waltham, MA, USA) supplemented with 10%
(v/v) foetal bovine serum (Invitrogen; Thermo Fisher Scientific,
Inc.), 100 U/ml of penicillin and 100 µg/ml of streptomycin (both
from Invitrogen; Thermo Fisher Scientific, Inc.). Cultures were
maintained in a humidified atmosphere of 5% CO2 at 37°C.
Total RNA was isolated from 2×106 U87 cells using 1 ml
TRIzol Reagent (Invitrogen; Thermo Fisher Scientific, Inc.),
according to the manufacturer's protocol. RNA was quantified and
evaluated for purity with a spectrophotometer based on
A260 and A280 values, followed by
visualization on a 1.0% (w/v) agarose gel stained with ethidium
bromide.
Stem-loop RT-qPCR
Stem-loop RT and RT-qPCR primers were synthesized by
Genewindows Biotech. Co. Ltd. (Guangzhou, China) and are listed in
Table I. cDNA was generated by
reverse transcription using 1 µg RNA as template with ReverTra
Ace-α-Transcriptase (Toyobo Life Science, Osaka, Japan). For
amplification synthetic DNA, ~0.1 ng of each let-7 member
were mixed and used as template. Briefly, stem-loop and U6 RT
primers were combined to transcribe the total RNA. cDNA was
prepared in a series of dilutions and thus a concentration gradient
was generated prior to RT-qPCR assay. RT-qPCR was performed using
the SYBR Premix Ex Taq II kit (Tli RNaseH Plus; Takara Bio, Inc.,
Otsu, Japan), following the manufacturer's protocol. qPCR was
performed with a LightCycler 480 Real-Time PCR system (Roche
Applied Science, Rotkreuz, Switzerland) and quantitation cycle (Cq)
values were normalized to U6 small nuclear RNA (16), which was used as an internal
control. The amplification profile was: 1 cycle of denaturation at
95°C for 30 sec; followed by 45 cycles of denaturation at 95°C for
5 sec, annealing and extension at 60°C for 1 min; and fluorescence
intensity was measured at 75°C. For melting curve analysis, the
following parameters were used: Denaturation at 95°C for 5 sec,
annealing at a rate of −2.8°C/sec until 60°C, fluorescence
intensity were measured throughout the process. All experiments
were performed in triplicate. Analysis was performed using
Microsoft Excel 2013 (Microsoft Corporation, Redmond, WA, USA) and
Origin 8.0 (OriginLab, Wellesley, MA, USA) software.
 | Table I.Stem-loop RT and RT-quantitative
polymerase chain reaction primer sequences. |
Table I.
Stem-loop RT and RT-quantitative
polymerase chain reaction primer sequences.
| Gene | Sequence
(5′→3′) |
|---|
| U6 (RT) |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAAAAATATG |
| U6 (F) |
CTCGCTTCGGCAGCACA |
| U6 (R) |
AACGCTTCACGAATTTGCGT |
|
Let-7a/e/f-5p (RT) |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAACTATAC |
| Let-7b-5p
(RT) |
CTCAACTGGAGCTAGTTTCGTCGTAGGGCAGTTGAGAACCACAC |
| Let-7c
(RT) |
GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAACCAT |
| Let-7d-5p
(RT) |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAACTATGC |
| Let-7g-5p
(RT) |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAACTGTAC |
| Let-7i-5p
(RT) |
GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAACAGC |
| miR-98 (RT) |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAACAATAC |
|
Let-7a/b/c-5p (F) |
CCAGCTGGGTGAGGTAGTAGGTTGT |
| Let-7d-5p
(F) |
CCAGCTGGGAGAGGTAGTAGGTTGC |
| Let-7e-5p
(F) |
CCAGCTGGGTGAGGTAGGAGGTTGT |
| Let-7f-5p
(F) |
CCAGCTGGGTGAGGTAGTAGATTGT |
| Let-7g/i-5p
(F) |
CCAGCTGGGTGAGGTAGTAGTTTGT |
| miR-98-5p (F) |
CCAGCTGGGTGAGGTAGTAAGTTGT |
| Uni
(R1)a |
TCCAGTGCAGGGTCCGAGGTA |
| Uni
(R2)b |
CTGGTGTCGTGGAGTCGGCAATT |
| Uni
(R3)c |
CTGGAGCTAGTTTCGTCGTAGGG |
| Poly(A) (RT) |
GCGAGCACAGAATTAATACGACTCACTATAGGTTTTTTTTTTTTTTTTTTVN |
| Poly(A) (Uni
R) |
GCGAGCACAGAATTAATACGACTCA |
Poly(A) tailing and RT
Poly(A) tails were added to total RNA 3′-ends using
the miRNA First-Strand cDNA Synthesis kit (ComWin Biotech Co.,
Ltd., Beijing, China). Subsequently, RNA was reverse transcribed
with anchor primer consisting of oligo(dT) for binding poly(A), a
specific sequence for the combination with the reverse primer used
in subsequent RT-qPCR, and a binucleotide VN (V indicates any
nucleotide except for dT whereas N refers to any nucleotide). qPCR
forward primers and thermocycling profiles were similar to the
aforementioned stem-loop RT-qPCR.
Data analysis
Cq values and melting curves were obtained using the
LightCycler 480 software V1.5, and Origin 8.0 software (OriginLab)
was exploited to draw standard curves. The Cq values were plotted
against the log2 of total RNA input, and the
amplification efficiency was evaluated through correlation analysis
between these two values using Origin 8.0 software. Amplification
efficiency (E) was calculated as: E=2−1/a-1; where a is
the slope of the standard curve line. The Clustal algorithm
(Bio-Edit software V7.0.5) was applied in multiple sequence
alignment, whereas primer designation schemes were produced using
Adobe Illustrator CS4 software (Adobe Systems Incorporated, San
Jose, CA, USA).
Statistical analysis
All data were presented as the mean ± standard
deviation. Significant differences between samples were calculated
by Student's t-test. All tests were performed at least in
triplicate. All statistical analyses were carried out using the
Origin 8.0 (Additive GmbH, Friedrichsdorf, Germany) and Excel
software (Microsoft Corporation, Redmond, WA, USA). P<0.05 was
considered to indicate a statistically significant difference.
Results
Sequence alignment identified no differences in the
16 nucleotide base sequences at the 5′-ends of let-7a,
let-7b and let-7c; therefore, the same forward
primers was used for these genes. Similarly, let-7i and
let-7g shared a common forward primer. However, distinct
stem-loop structures and different reverse primers were designed
for used in SYBR Green-based RT-qPCR, which would ensure
amplification specificity. The specific stem-loop RT primers
designed for each let-7 family member is illustrated in
Fig. 1.
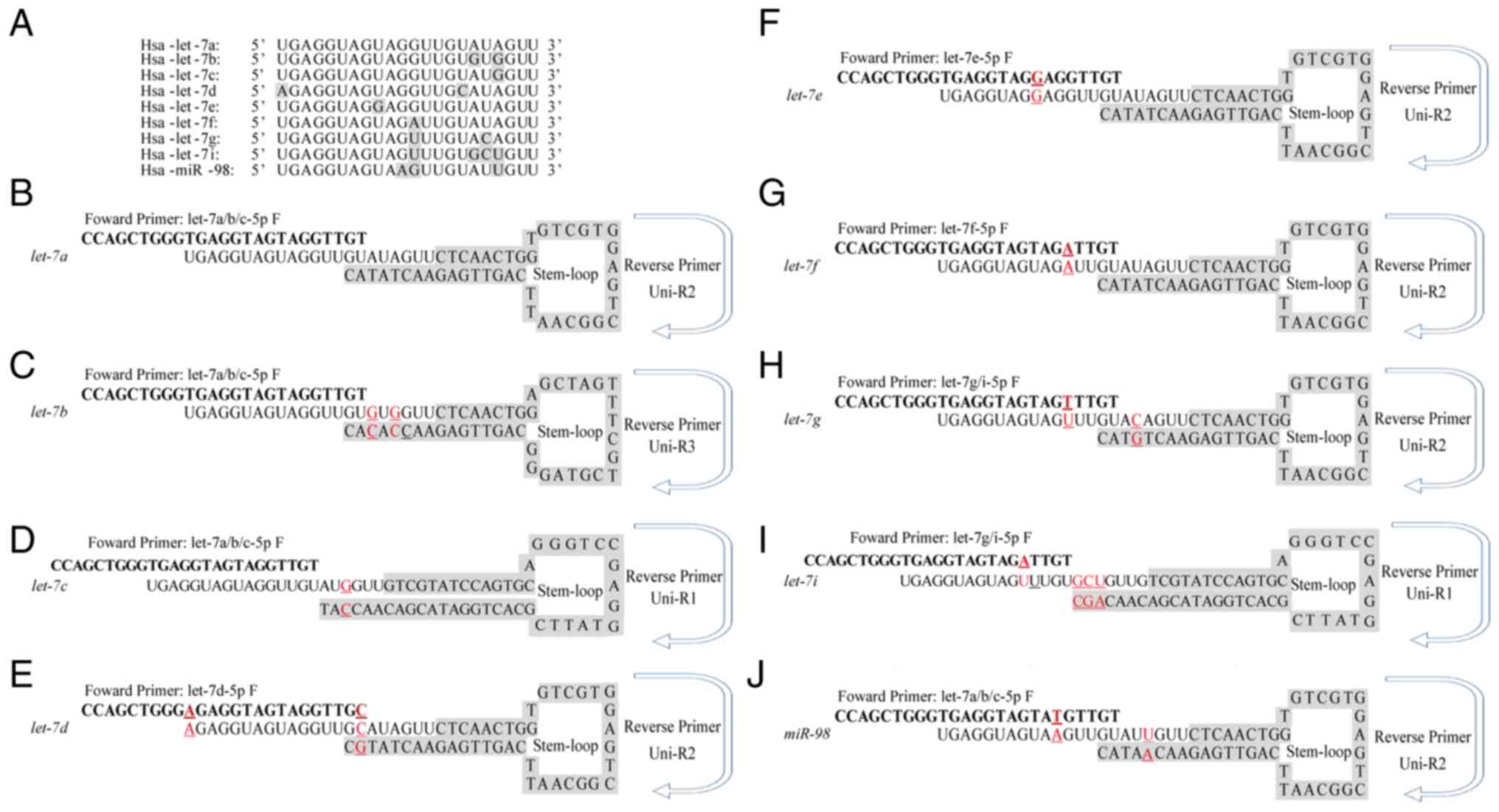 | Figure 1.Different stem-loop primers were
designed to distinguish let-7 family members. (A) Multiple
sequence alignment of nine let-7 family members was
performed to analyse the conserved nature of let-7 family.
(B-J) Sequences of the specific stem-loop primers for let-7
family members, including (B) let-7a, (C) let-7b, (D)
let-7d, (E) let-7d, (F) let-7e, (G)
let-7f, (H) let-7g, (I) let-7i and (J)
miR-98. Red underscored characters indicated variant
nucleotides that differ from let-7a. F, forward; Hsa,
Homo sapiens; R, Reverse; uni, universal. |
Robust amplification efficiency and
specificity are achieved
Total RNA was extracted from U87 cells and reverse
transcribed with a mixture of distinct stem-loop RT primers
specifically designed for let-7 family members and U6
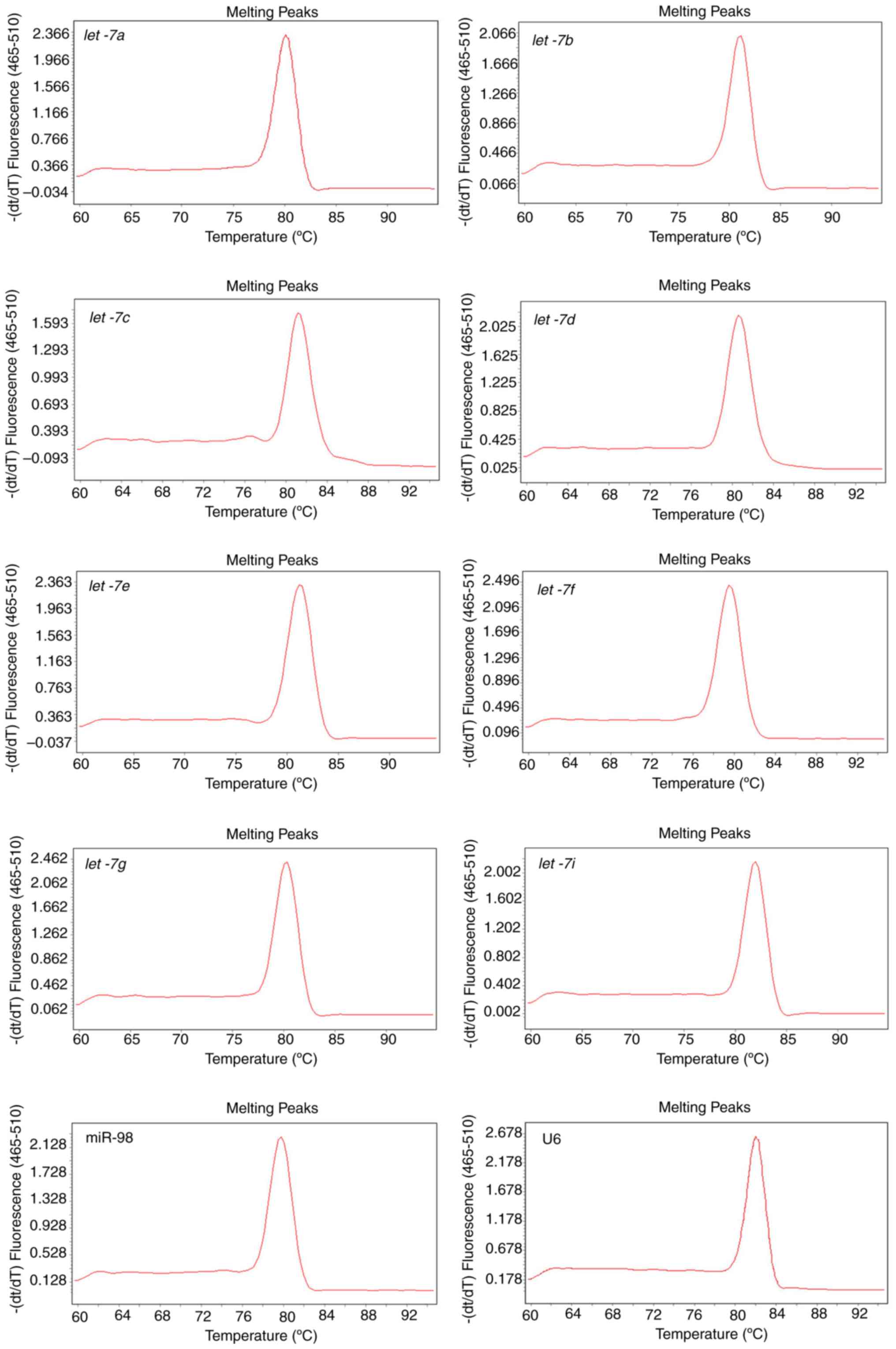
primers. Melting curves that are representative of amplification
specificity combined with amplification efficiency of let-7
and U6 were also measured. Melting curve analysis of the amplified
products demonstrated a single, sharp peak, which indicated good
specificity (Fig. 2). Except for
let-7i, which had a low concentration and therefore cannot
be detected at a high dilution ratio, other amplification
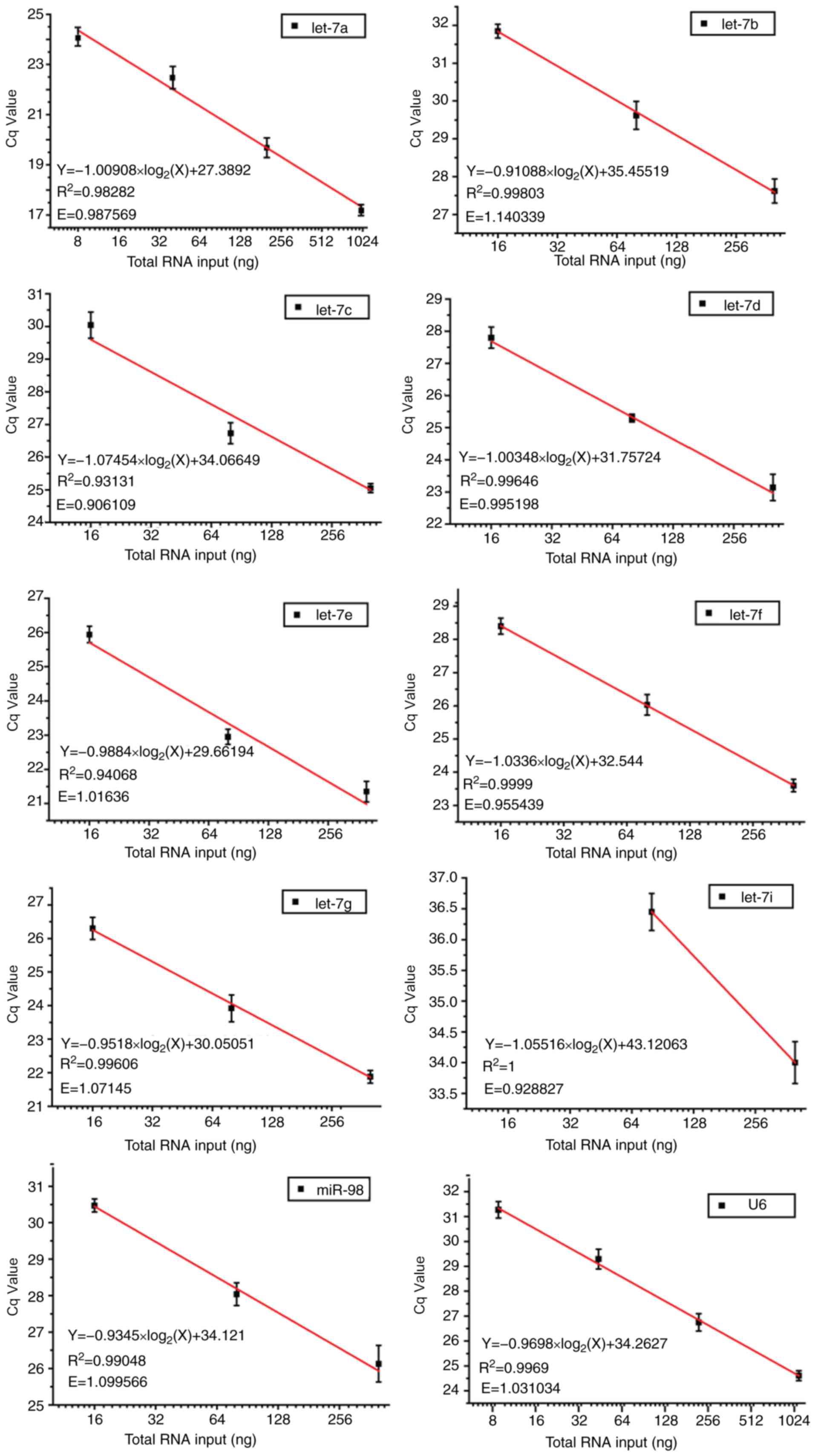
efficiencies ranged between 0.9288 and 1.1403. In addition, good
linear relationships were indicated between PCR Cq values and
log2 of RNA input (R2>0.9; Fig. 3). These data suggested that by
performing only one miRNA qPCR reaction, it was feasible to
simultaneously detect and quantify all nine let-7 family
members, and to further discriminate between let-7a,
let-7b and let-7c, as well as between let-7i
and let-7g. Notably, the mixture containing our modified RT
primers and U6 primer was still able to efficiently and
specifically quantify each of the let-7 congeners and U6
internal control, respectively, contributing to a considerable
decrease in time and cost.
Stem-loop RT-qPCR is superior to
Poly(A) tailing-based RT-qPCR in amplifying the mature let-7
family
The expression profiles of let-7 family
members in U87 cells were also assessed by the poly(A) tailing
method (Fig. 4). This method was
initially established to detect miRNAs for large-scale and
high-throughput screening, so the forward primers in the stem-loop
method and the poly(A) tailing method designed in the present study
were the same. There is only a one base difference between mature
let-7 miRNA family members, therefore, if the stem-loop
section of primers used to reverse transcribe in different
let-7 members were identical, the melting temperature values
of these distinct let-7 members were alike (Fig. 2). However, in the poly(A) method,
cross-reactions between let-7a, let-7b and
let-7c or between let-7g and let-7i cannot
result in a difference of >5°C in the melting temperature.
Therefore, multiple peaks that occurred in the melting curves in
the poly(A) tailing method were caused by other reasons.
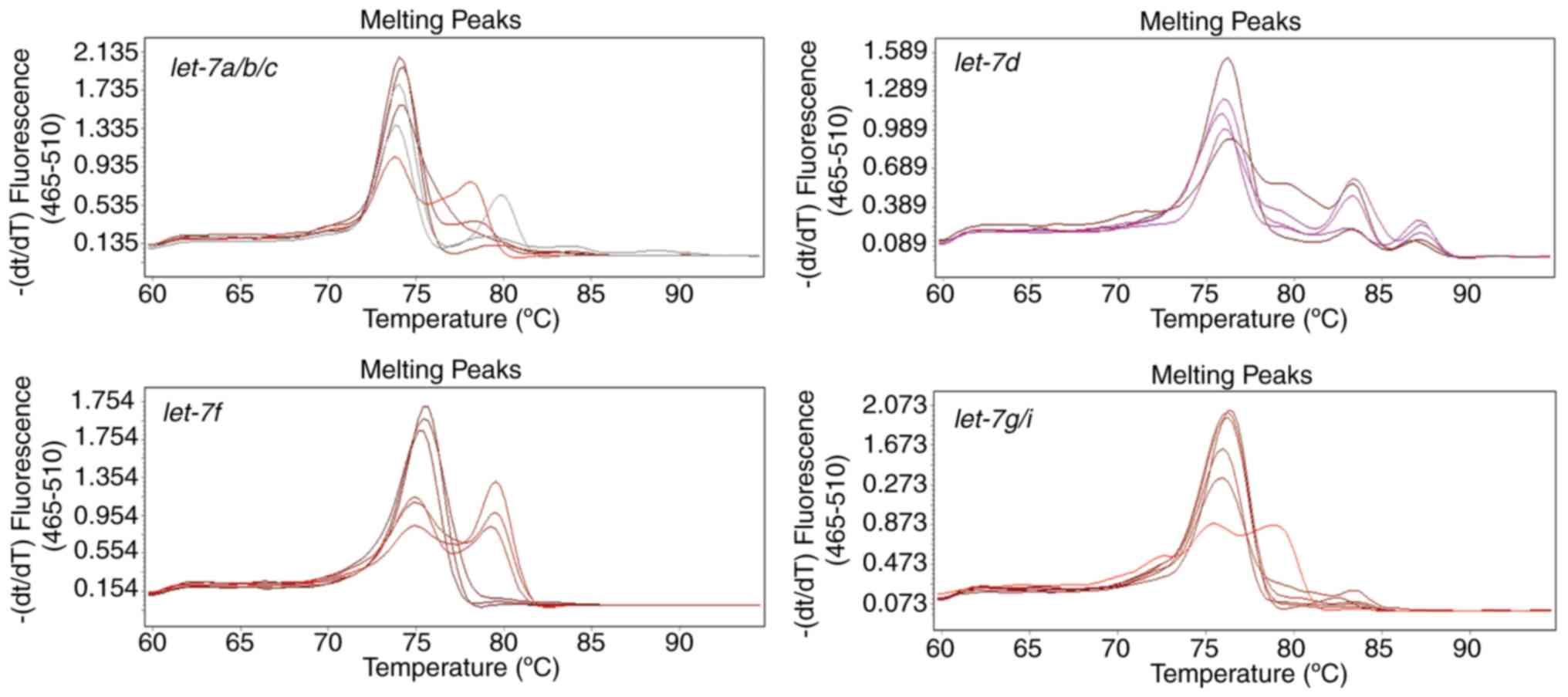
However, the melting curves of each amplification
reaction that were generated using the poly(A) method contained
multiple peaks representing different products (Fig. 4), regardless of the specific
forward primers of let-7d and let-7f. These may
because miRNA precursors were also amplified, since the linear
primers used in poly (A) tailing method lack specificity.
Modified stem-loop RT-qPCR
discriminates distinct mature let-7 family members that differ by
as little as a single nucleotide
The ability of the modified stem-loop RT-qPCR assay
to discriminate between miRNAs that differ by as little as a single
nucleotide was examined using synthetic templates that contained a
certain amount of each let-7 member and the differential Cq
values were analysed. Different concentrations of synthetic
let-7a, let-7b and let-7c were mixed and
reverse transcribed with a mixture of corresponding primers and
RT-qPCR was performed with let-7a primers alone. The results
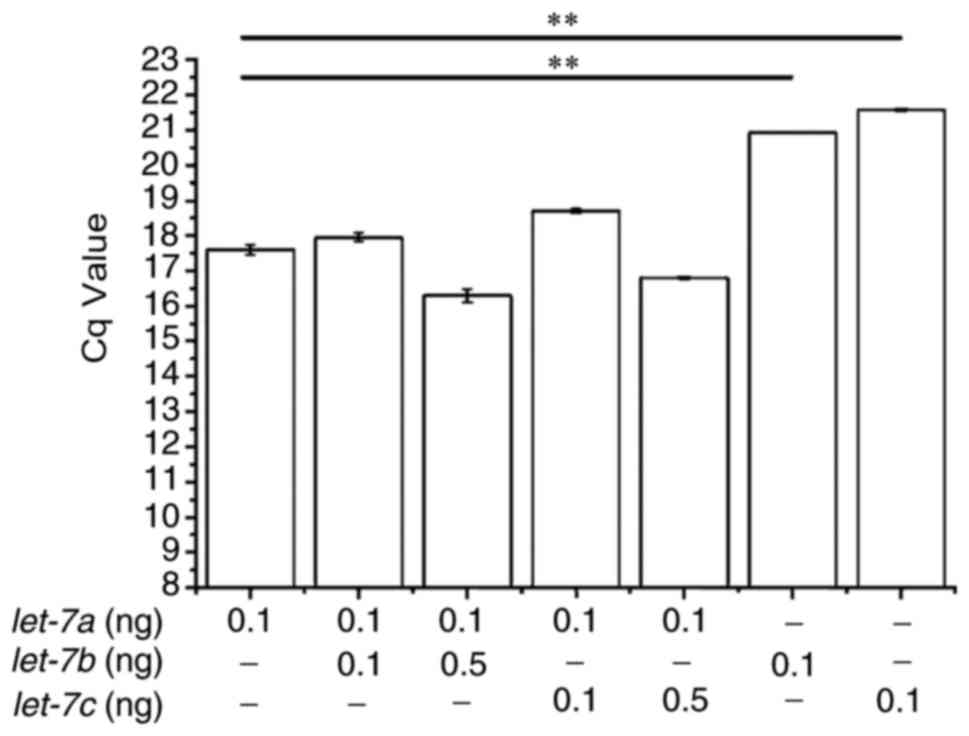
demonstrated that the addition of equal concentrations of
let-7b or let-7c did not affect the detection of
let-7a (Fig. 5). qPCR
reactions devoid of let-7a had significantly higher Cq
values compared with those that also contained synthetic
let-7a. Although 5-fold excess concentration of
let-7b or let-7c slightly interfered with
let-7a detection, this interference was not statistically
significant.
Discussion
The let-7 family of miRNAs are highly
conserved, differing by only 1–4 nucleotides among its members,
which enables these congeners to share common target mRNAs that are
downregulated by complementary sequences within their
3′-untranslated regions. Let-7 miRNA was first discovered in
C. elegans as a switch gene that controlled cell fate
transitions between larva and adult, and was later documented in
humans in repressing several cancers, including lung cancer
(17), colon cancer (18) and melanoma (19). Underexpression of let-7 in
lung cancer was reported to be associated with shorter survival
following tumorectomy (17),
whereas overexpression of let-7 suppressed cancer cell
proliferation and tumourigenicity (20–22).
Accordingly, let-7 may be regarded as a miRNA candidate for
cancer detection and monitoring.
However, small discrepancies among the let-7
family members generally escape detection by current routine
techniques, except for high-throughput sequencing (23) or stem-loop RT-qPCR combined with
TaqMan-based real-time quantification. Nevertheless, their clinical
utilization in detection and intensive research for miRNAs was
still hindered, due to the relatively high cost. Conversely,
despite the low cost of SYBR Green dyes, it cannot be used as a
substitute for TaqMan probes to simultaneously detect miRNAs with
few differences. However, the present study offered a newly
developed method based on the method previously described by Chen
et al (16). The initial
stem-loop method described by Chen et al was able to
differentiate between let-7a and let-7b, but not
between let-7a, let-7c and let-7d (16). This was likely to the fact that the
TaqMan probe used in their system restricted the amplification
efficiency. Notably, the modified method used in the present study
was able to adequately discriminate let-7a and
let-7c, which indicated that this method is able to
differentiate miRNAs that differ by as little as a single
nucleotide and therefore render it capable of being used in
numerous of clinical applications.
Using an identical upstream primer, the present
method enabled specific RT and quantification of several distinct
miRNAs without a major loss of specificity of downstream primers
that were mixed together, and in the absence of costly TaqMan
probes. More importantly, this method exhibited higher specificity
compared with conventional TaqMan. Data from the present study
indicated that the modified stem-loop RT-qPCR method was able to
differentiate between the let-7 family members that differ
by a single nucleotide, which thereby provides a method to
investigate the specific function of each family member and the
potential cooperation between these members, as well as the role
let-7 family members in various pathological conditions,
particularly in the process of tumour generation and progression.
In addition, this method is able to discriminate mature miRNAs from
their longer progenitors (16).
These characteristics and merits may facilitate the use of this
modified stem-loop RT-based qPCR into clinical applications and may
be further expanded to detect and distinguish between miRNA
congeners differing by as little as a single nucleotide.
Acknowledgements
The present study was supported in part by The
Special Fund for the Development of Shenzhen Strategic Emerging
Industries (grant no. CXZZ20130517100617030).
References
|
1
|
Tay Y, Zhang J, Thomson AM, Lim B and
Rigoutsos I: MicroRNAs to Nanog, Oct4 and Sox2 coding regions
modulate embryonic stem cell differentiation. Nature.
455:1124–1128. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Doench JG and Sharp PA: Specificity of
microRNA target selection in translational repression. Genes Dev.
18:504–511. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wightman B, Ha I and Ruvkun G:
Posttranscriptional regulation of the heterochronic gene lin-14 by
lin-4 mediates temporal pattern formation in C. elegans. Cell.
75:855–862. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Olsen PH and Ambros V: The lin-4
regulatory RNA controls developmental timing in Caenorhabditis
elegans by blocking LIN-14 protein synthesis after the initiation
of translation. Dev Biol. 216:671–680. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kim VN, Han J and Siomi MC: Biogenesis of
small RNAs in animals. Nat Rev Mol Cell Biol. 10:126–139. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bartel DP: MicroRNAs: genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bao N, Lye KW and Barton MK: MicroRNA
binding sites in Arabidopsis class III HD-ZIP mRNAs are required
for methylation of the template chromosome. Dev Cell. 7:653–662.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Michael MZ, O'Connor SM, van Holst
Pellekaan NG, Young GP and James RJ: Reduced Accumulation of
Specific MicroRNAs in Colorectal Neoplasia11Note: Susan M. O'Connor
and Nicholas G. van Holst Pellekaan contributed equally to this
work. Mol Cancer Res. 1:882–891. 2003.PubMed/NCBI
|
|
9
|
Reinhart BJ, Slack FJ, Basson M,
Pasquinelli AE, Bettinger JC, Rougvie AE, Horvitz HR and Ruvkun G:
The 21-nucleotide let-7 RNA regulates developmental timing in
Caenorhabditis elegans. Nature. 403:901–906. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Rodini CO, Suzuki DE, Saba-Silva N,
Cappellano A, de Souza JE, Cavalheiro S, Toledo SR and Okamoto OK:
Expression analysis of stem cell-related genes reveal OCT4 as a
predictor of poor clinical outcome in medulloblastoma. J
Neurooncol. 106:71–79. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Sakurai M, Miki Y, Masuda M, Hata S,
Shibahara Y, Hirakawa H, Suzuki T and Sasano H: LIN28: A regulator
of tumor-suppressing activity of let-7 microRNA in human breast
cancer. J Ster Biochem Mol Biol. 131:101–106. 2011. View Article : Google Scholar
|
|
12
|
Zhao C, Sun G, Li S, Lang MF, Yang S, Li W
and Shi Y: MicroRNA let-7b regulates neural stem cell proliferation
and differentiation by targeting nuclear receptor TLX signaling.
Proc Natl Acad Sci USA. 107:pp. 1876–1881. 2010; View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Johnson CD, Esquela-Kerscher A, Stefani G,
Byrom M, Kelnar K, Ovcharenko D, Wilson M, Wang X, Shelton J,
Shingara J, et al: The let-7 MicroRNA represses cell proliferation
pathways in human cells. Cancer Res. 67:7713–7722. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Dong Q, Meng P, Wang T, Qin W, Qin W, Wang
F, Yuan J, Chen Z, Yang A and Wang H: MicroRNA let-7a inhibits
proliferation of human prostate cancer cells in vitro and in vivo
by targeting E2F2 and CCND2. PLoS One. 5:e101472010. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Shell S, Park SM, Radjabi AR, Schickel R,
Kistner EO, Jewell DA, Feig C, Lengyel E and Peter ME: Let-7
expression defines two differentiation stages of cancer. P Natl
Acad Sci USA. 104:11400–11405. 2007. View Article : Google Scholar
|
|
16
|
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee
DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, et al:
Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic
Acids Res. 33:e1792005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Takamizawa J, Konishi H, Yanagisawa K,
Tomida S, Osada H, Endoh H, Harano T, Yatabe Y, Nagino M, Nimura Y,
et al: Reduced expression of the let-7 microRNAs in human lung
cancers in association with shortened postoperative survival.
Cancer Res. 64:3753–3756. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Akao Y, Nakagawa Y and Naoe T: let-7
microRNA functions as a potential growth suppressor in human colon
cancer cells. Biol Pharm Bull. 29:903–906. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Schultz J, Lorenz P, Gross G, Ibrahim S
and Kunz M: MicroRNA let-7b targets important cell cycle molecules
in malignant melanoma cells and interferes with
anchorage-independent growth. Cell Res. 18:549–557. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yu F, Yao H, Zhu P, Zhang X, Pan Q, Gong
C, Huang Y, Hu X, Su F, Lieberman J and Song E: let-7 regulates
self renewal and tumorigenicity of breast cancer cells. Cell.
131:1109–1123. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kumar MS, Erkeland SJ, Pester RE, Chen CY,
Ebert MS, Sharp PA and Jacks T: Suppression of non-small cell lung
tumor development by the let-7 microRNA family. Proc Natl Acad Sci
USA. 105:pp. 3903–3908. 2008; View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Lu L, Schwartz P, Scarampi L, Rutherford
T, Canuto EM, Yu H and Katsaros D: MicroRNA let-7a: a potential
marker for selection of paclitaxel in ovarian cancer management.
Gynecol Oncol. 122:366–371. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zovoilis A, Agbemenyah HY, Agis-Balboa RC,
Stilling RM, Edbauer D, Rao P, Farinelli L, Delalle I, Schmitt A,
Falkai P, et al: microRNA-34c is a novel target to treat dementias.
Embo J. 30:4299–4308. 2011. View Article : Google Scholar : PubMed/NCBI
|