Introduction
Lung cancer has one of the highest
disease-associated mortalities in the world (1). Currently, in the United States,
cancer has the second highest mortality rate, and lung cancer has
the highest rate of mortality in men and women (2). Non-small cell lung carcinoma (NSCLC)
accounts for ~85% of lung cancer cases (3); treatment of NSCLC predominantly
involves chemotherapy, radiation and surgery (4). Chemotherapy is currently the standard
treatment for lung cancer (5) and
cisplatin is widely used as an effective anti-cancer agent for the
treatment of various types of malignant cancers (4). However, resistance to cisplatin has
been observed, combined with toxicity to normal cells (6,7);
thus, clinical cisplatin resistance (CR) is a major obstacle for
the treatment of lung cancer (4).
HanAmDan-B1 (HAD-B1) is a blended herbal extract
that is composed of Panax Notoginseng Radix, Cordyceps
militaris, Panax ginseng C. A. Mey and Boswellia
carterii Birdwood. HAD-B1 was modified from the previously
developed HAD-B for use as an anti-cancer herbal medicine at the
East West Cancer Center (EWCC; Dunsan Korean Medicine Hospital of
Daejeon University, Daejeon, Korea) (8–10). A
retrospective study of HAD-B indicated it may have efficacy in
treating lung cancer (11–13) and it has been demonstrated to have
an anti-angiogenesis effect on HUVEC cells (14). Furthermore, HAD-B exhibited a safe
toxicological profile in normal cells (15).
This study was conducted to evaluate the anti-cancer
effects of HAD-B1 against the lung cancer cell strain A549CR.
Materials and methods
Preparation of HAD-B1 extract
HAD-B1 was provided by the EWCC (Table I). A voucher specimen
(#HAD-B-1-2014-10-HS) has been deposited at the Institute of
Traditional Medicine and Bioscience in Daejeon University. The
ingredients of the herb mixture (HAD-B1) were soaked for 18 h in a
bath of distilled water at 60°C and the supernatant was obtained.
The extracts were concentrated by using a rotary vacuum evaporator
at 60°C for 2 h and were dried on a flat evaporator at 60°C for 8
h, the powder produced was used for the experiments (16). The HAD-B1 was dissolved in
distilled water (DW).
 | Table I.Ingredients of HangAmDan-B1. |
Table I.
Ingredients of HangAmDan-B1.
| Scientific name | Relative amount
(g) |
|---|
| Panax Notoginseng
Radix | 25.2 |
| Cordyceps
militaris | 19.2 |
| Panax ginseng
C.A.Mey. | 19.2 |
| Boswellia
carerii Birdwood | 14.4 |
| Total amount | 78.0 |
Cell culture
A549 human lung cancer cells with CR were acquired
from the ASAN Medical Center (Seoul, Korea). The cells were
cultured in Dulbecco's modified Eagle's media (DMEM, Welgene, Inc.,
Daejeon, Korea) containing 10% fetal bovine serum (FBS, Welgene,
Inc.) and 1X antibiotics (Welgene, Inc.). A549CR cell cultures were
maintained at 37°C in a humidified atmosphere containing 5%
CO2.
A549CR cell viability assay
A549CR cells (2×103 cells/well) were
added to 96-well tissue culture plates and allowed to adhere
overnight. HAD-B1, cisplatin, afatinib, and cordycepin were
dissolved in distilled water. The cells were treated with HAD-B1 (0
and 4.8 µg/ml-20 mg/ml, 4 fold), cisplatin (0 and 0.01–30 µg/ml, 4
fold), afatinib (0 and 2.4 ng/ml-9.87 µg/ml, 4 fold), and
cordycepin (0 and 0.012–200 µg/ml, 4 fold) and incubated for 72 h
at 37°C. A total of 10 µl (5 mg/ml) MTT solution was added to each
well, and the cells were incubated for 2 h at 37°C. The
supernatants were discarded and the residual formazan crystals were
dissolved in 100 µl dimethyl sulfoxide. The absorbance was measured
at 595 nm on an Emax ELISA plate reader (Molecular Devices, LLC,
Sunnyvale, CA, USA). Measurements were made in triplicate.
Apoptosis and cell cycle analysis
A549CR cells (1×106 cells/100
mm2 dish) were treated with HAD-B1 (1 mg/ml) for 24, 48
or 72 h at 37°C. Cell viability and apoptosis were determined using
a Muse Annexin V & Dead Cell kit (EMD Millipore, Billerica, MA,
USA) in accordance with the manufacturer's protocol. Cell cycle
analysis was performed using a Muse Cell Cycle kit (EMD Millipore).
Apoptosis and cell cycle were analyzed using Muse® Cell
Analyzer (EMD Millipore).
Caspase activity assay
The enzyme activities of caspases-3 (BF3100), −8
(BF4100), and −9 (BF10100) in the cells were measured using kits
purchased from R&D Systems, Inc. (Minneapolis, MN, USA). A549CR
cells were collected using trypsin-EDTA following incubation with
Afatinib (2.96 mg/ml) or HAD-B1 (1 mg/ml) for 72 h. Collected cells
were centrifuged for 15 min at 582 × g and at room temperature, the
supernatant was discarded, and the remaining cell pellet was
incubated with Lysis-M solution (Roche Applied Science, Mannheim,
Germany) on ice for 15 min. The lysed cells were centrifuged (15
min, 15,267 × g, 4°C), and the amount of protein in the supernatant
was quantified using a Bradford assay kit (Bio-Rad Laboratories,
Inc., Hercules, CA, USA). Protein lysates at a final concentration
of 2 µg/µl) containing 0.1 M dithiothreitol were added into the
wells of a 96-wellplate, 5 µl LEHD-pNA was added to each well and
the plate was incubated at 37°C for 2 h. The absorbance was then
measured at 405 nm using a microplate reader.
Protein extraction and fluorescence
labeling of A549CR cells
A549CR cells were serum-starved by incubation in
DMEM for 4 h. The cells were subsequently treated with or without
HAD-B1 (1 mg/ml). Following incubation for 72 h, the cells were
washed twice with phosphate-buffered saline (PBS), harvested using
5 mM EDTA-PBS and centrifuged for 15 min at 582 × g, at room
temperature. The pellets were washed with PBS and centrifuged for
15 min at 582 × g, at room temperature. A549CR cells
(5×105 cells/ml) were extracted using Lysis-M mammalian
cell extraction buffer. Each protein extract (100 µg, 1 mg/ml) was
separately labeled with Cyanine3 (Cy3) and Cyanine5 (Cy5; GE
Healthcare Life Sciences, Little Chalfont, UK) according to the
manufacturer's instructions. Free dyes were removed using Sigma
Spin columns (S5059; Sigma-Aldrich; Merck KGaA, Darmstadt, Germany)
and purified samples were stored at 4°C until use.
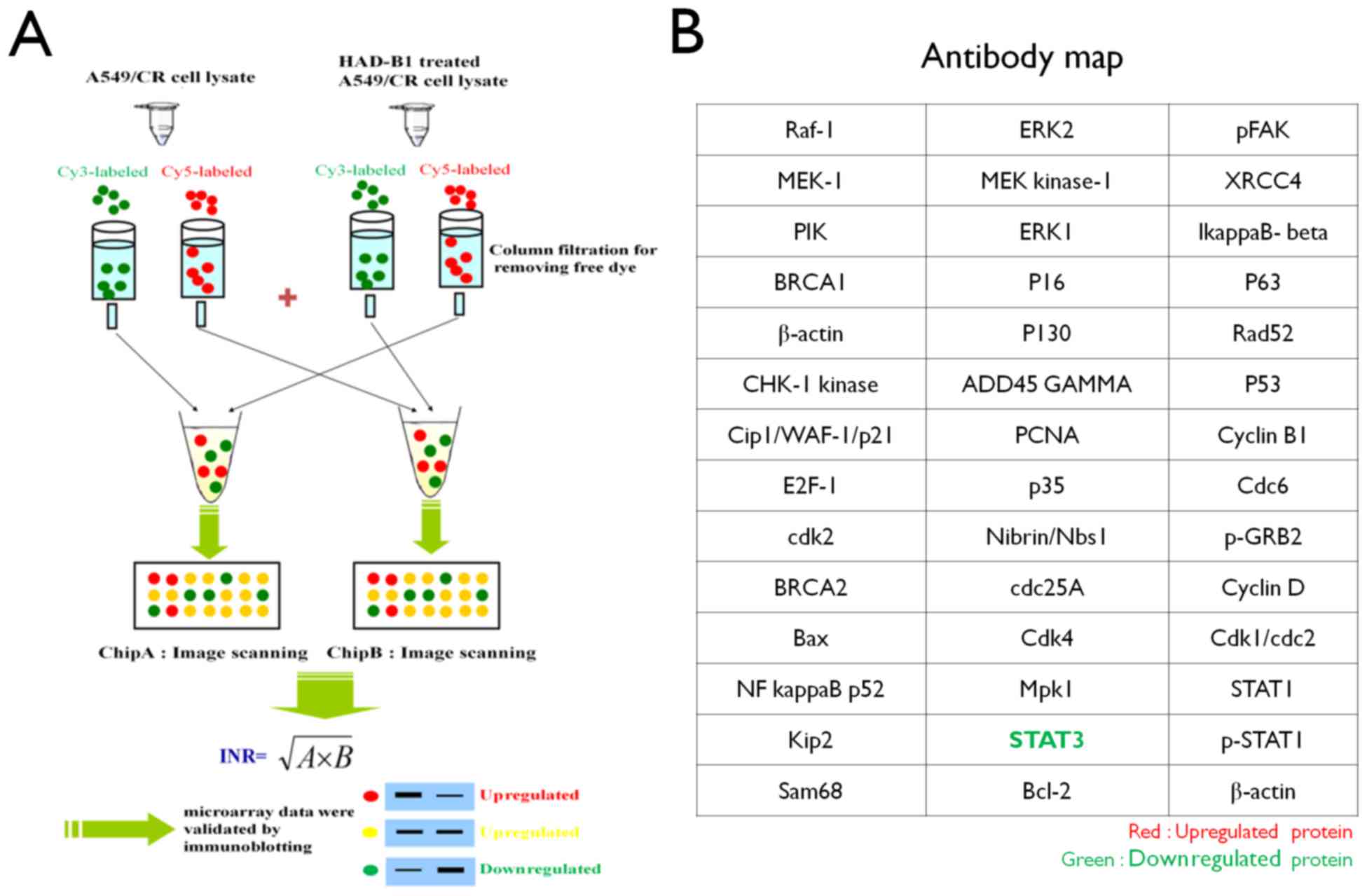
Preparation of InnoPharmaScreen (IPS)
ProteoChip-based antibody microarray
Proteochip was provided from InnoPharmaScreen Inc.
(Asan, Korea). Two chips were made with the same antibodies arrays.
A total of 42 distinct antibodies against proteins involved in cell
proliferation were spotted onto Proteochip arrays in duplicate
(Fig. 1) (17).
Profile analysis of cell cycle
proteins using IPS ProteoChip-based antibody microarray
The fluorescence-labeled cell lysates were applied
to the antibody array and incubated for 1 h at 37°C in the dark.
The slides were subsequently washed three times with PBS-0.1% Tween
(PBST), dried until completely dry by N2 gas and
analyzed using a fluorescence microarray scanner. Antibody array
slides were scanned using a GenePix 4100A microarray scanner (Axon
Instruments; Molecular Devices, LLC) with 532 and 635 nm lasers.
Image analysis was performed for each spot using the manufacturer's
software package (GenePix version 6.0; Axon Instruments; Molecular
Devices, LLC). The internally normalized ratios (INR) of all spots
were calculated using a previously described procedure (16,18).
Western blot analysis
A549CR cells were incubated with HAD-B1 (1 mg/ml)
and Afatinib (2.96 µg/ml) in DMEM (containing 2% FBS and 1X
antibiotics) for 72 h at 37°C. The cells were harvested to extract
the proteins using Lysis-M buffer containing a protease and
phosphatase inhibitor cocktail (Roche Applied Science) and
clarified by centrifugation (15 min, 15,267 × g, 4°C). Lysates
containing 40 µg protein were loaded into each well and separated
by 12% sodium dodecyl sulfate-gel electrophoresis. Gels were
subsequently soaked in transfer buffer (16 mM Tris-HCl, 30-mM
glycine, 20% methanol) and proteins were transferred to
polyvinylidene difluoride membranes. Nonspecific binding sites were
blocked by incubation with 5% non-fat dried milk in PBST (137 mM
NaCl, 27 mM KCl, 100 mM Na2HPO4, 20 mM
KH2PO4 and 0.05% Tween-20, pH 7.4). The
polyvinylidene difluoride membranes were then incubated with
primary antibodies against signal transducer and activator of
transcription 3 (STAT3; ab68153; 1:10,000; Abcam, Cambridge, UK),
and β-actin (A5441; 1:10,000; Sigma-Aldrich; Merck KGaA) in PBST
containing 5% non-fat skimmed milk (232100; BD Biosciences,
Franklin Lakes, USA) at 4°C overnight. Membranes were washed with
PBST and then incubated with secondary antibodies [Anti-Mouse IgG
(Fc Specific); A0168; 1:10,000; Sigma-Aldrich; Merck KGaA and
Anti-Rabbit IgG (whole molecule), A0545; 1:10,000; Sigma-Aldrich;
Merck KGaA] for 1 h at room temperature. Signals were developed
using an Enhanced chemiluminescence western blotting detection kit
(EzWestLumi plus, WSE-7120S; ATTO Corporation, Tokyo, Japan) and
blots were subsequently exposed to X-ray films (B2640, Agfa Gevaert
NV, Mortsel, Belgium).
STAT3 knockdown and cell viability
assay
Small interfering RNA (siRNA) of STAT3 (Human) was
purchased from Bioneer Corporation, Daejeon, Korea
(UGUUCUCUGAGACCCAUGA). Cells were transfected with STAT3 siRNA (100
nM) using Lipofectamine 2000 (11668,027; Invitrogen; Thermo Fisher
Scientific, Inc., Waltham, MA, USA) for 24 h. Transfected cells
were incubation in a 100 mm2 dish for 72 h and harvested
to extract the proteins. The extraction was performed using western
blotting. Transfection cells (2×103 cells/well) were
incubated in a 96 well plate for 72 h and cell viability assayed
using MTT assay.
Statistical analysis
All data are expressed as the mean ± standard
deviation, and statistical comparisons were performed using a
Student's t-test. Caspase activity data was compared using one-way
analysis of variance with a post hoc Dunnett's test. Statistical
analyses were performed using Microsoft Office Excel version 2007
(Microsoft Corporation, Redmond, WA, USA). P<0.05 was considered
to indicate a statistically significant difference.
Results
Inhibitory effect of HAD-B1 on the
viability of A549CR cells
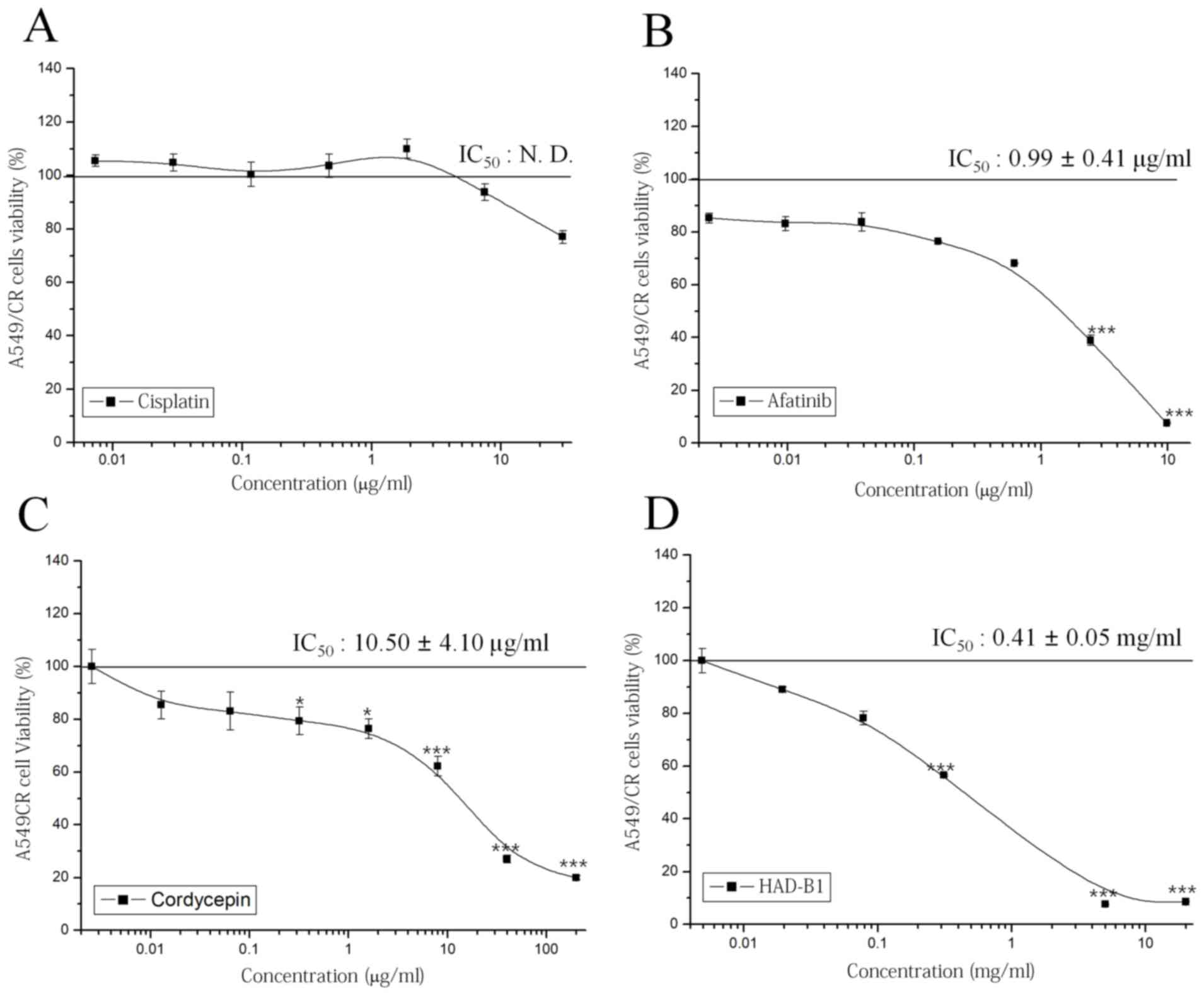
To investigate the cytotoxic effect of HAD-B1 on
A549CR cells, an MTT assay was employed. Cisplatin did not have a
significant effect on cell viability; however, afatinib, a
therapeutic agent targeted for NSCLC containing an epidermal growth
factor receptor mutation, inhibited cell viability in a
concentration-dependent manner (Fig.
2A and B). Notably, HAD-B1 significantly inhibited the growth
of A549CR cells in a concentration-dependent manner; the
half-maximal inhibition of A549CR cell growth by HAD-B1 was
observed at a concentration of 0.41±0.05 mg/ml (Fig. 2C). Cordycepin, a major component of
one of the ingredients in HAD-B1, Cordyceps militari, also
inhibited the viability of A549CR cells in a
concentration-dependent manner (Fig.
2D). This result demonstrated that HAD-B1 may have a
suppressive effect on the growth of A549CR cells.
Profiling of expression proteins in
A549CR cells treated with HAD-B1 using on antibody microarray
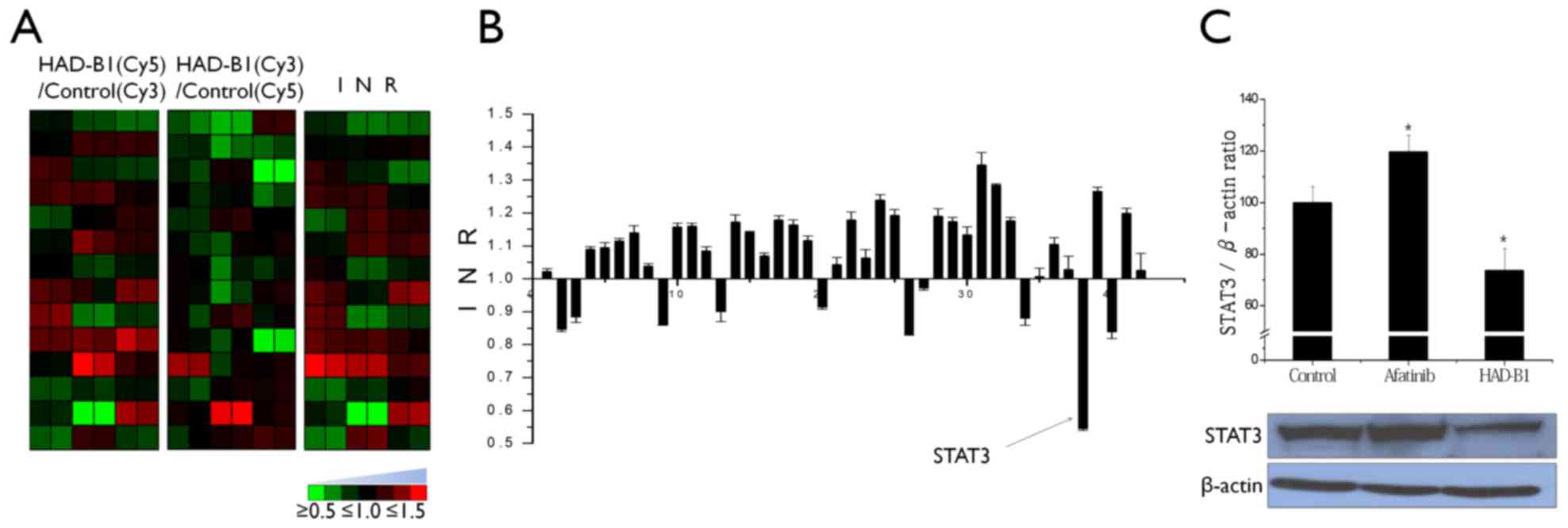
To investigate the molecular mechanism of HAD-B1 in
A549CR cells, the cellsignaling protein expression profiles of
HAD-B1-treated A549CR cells were investigated using an antibody
microarray. Lysates from cells treated with HAD-B1 (1 mg/ml) or
vehicle were labeled with a fluorescent dye (Cy5 or Cy3) and
cross-reactivity of the two samples was performed on the antibody
microarray (Fig. 1). The
fluorescence intensity of each spot was measured using a
fluorescence laser scanner and the protein expression pattern
between the samples was determined. Based on the data analysis,
HAD-B1 demonstrated the ability to decrease the expression of STAT3
in HAD-B1-treated cells (Fig. 3A and
B).
Validation of antibody
microarray-based protein profiling
To confirm the expression data from the profile
analysis of cell cycle proteins in HAD-B1-treated A549CR cells,
immunoblot analysis and an siRNA-based functional assay were
employed. Downregulation of STAT3 was confirmed by western blotting
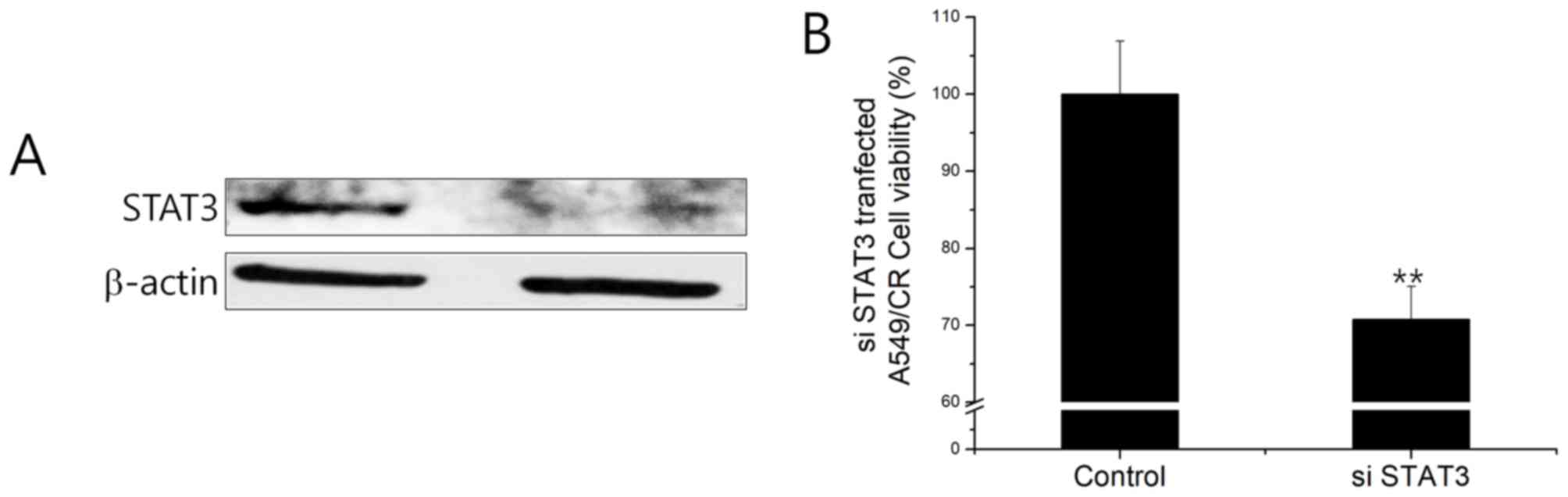
(Fig. 3C). A549CR cells
overexpressing STAT3 siRNA resulted in the downregulation of STAT3
at the protein level (Fig. 4A).
STAT3 siRNA-transfected A549CR cells demonstrated a lower cell
viability when compared with non-silencing siRNA (NS)-treated cells
(Fig. 4B). This finding indicated
that the negative effect of HAD-B1 on cell viability may be due to
the downregulation of STAT3 in A549CR cells.
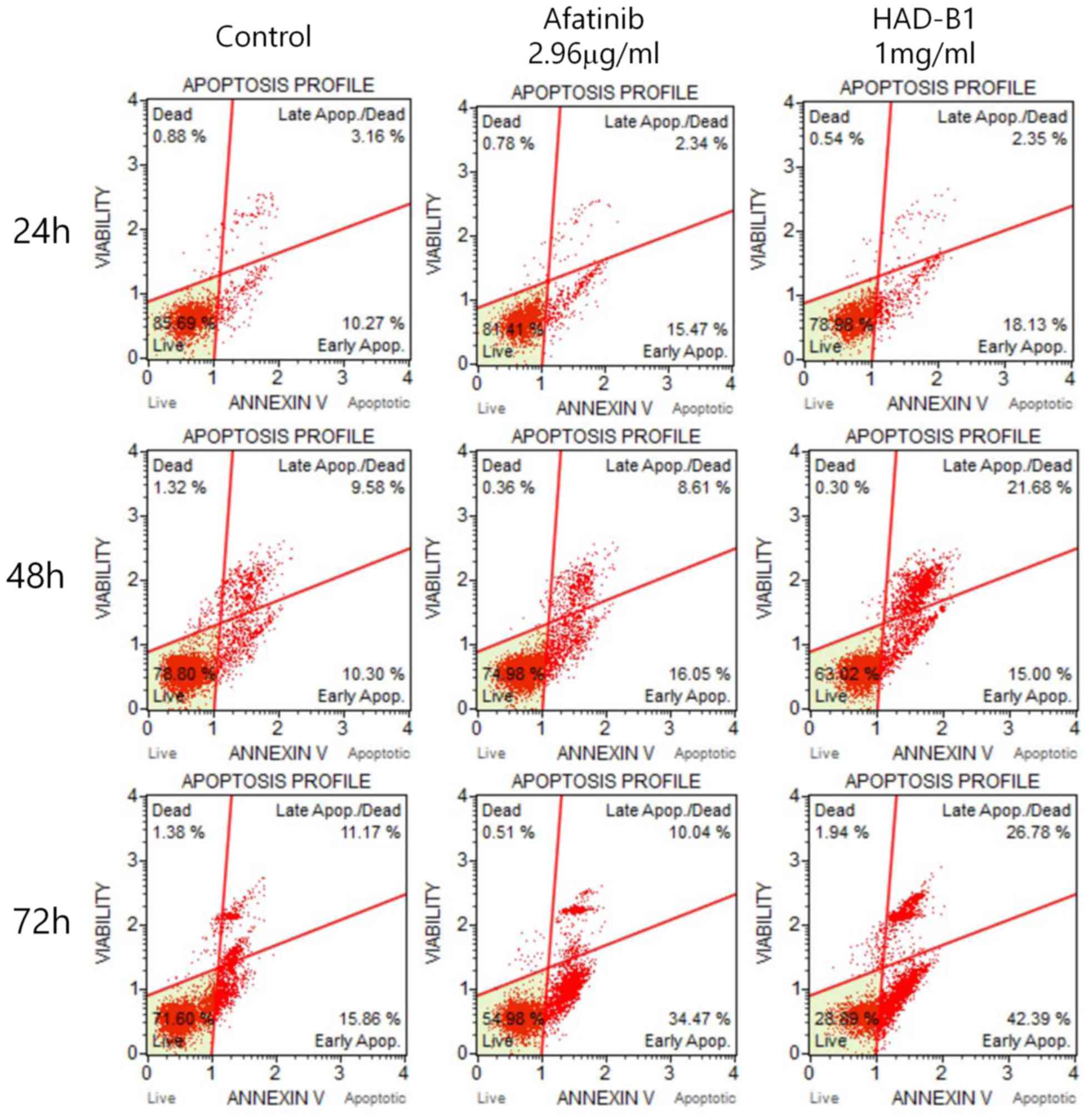
Cell cycle analysis and induction of
apoptosis in HAD-B1-treated A549CR cells
To investigate whether the toxicity of HAD-B1 on
A549CR cells was due to the induction of cell apoptosis and cell
cycle arrest, a flow cytometric assay was employed. Cells were
treated with HAD-B1 (1 mg/ml) for 24, 48 and 72 h. Induction of
cell apoptosis was analyzed by Annexin-V/propidium iodide double
staining using a Muse® Cell Analyzer. The early
apoptotic rate was markedly increased in A549CR cells treated for
72 h, compared with the vehicle- and Afatinib-treated groups
(Fig. 5). However, the early
apoptotic rate of cells treated with HADB1 for 24 and 48 h was not
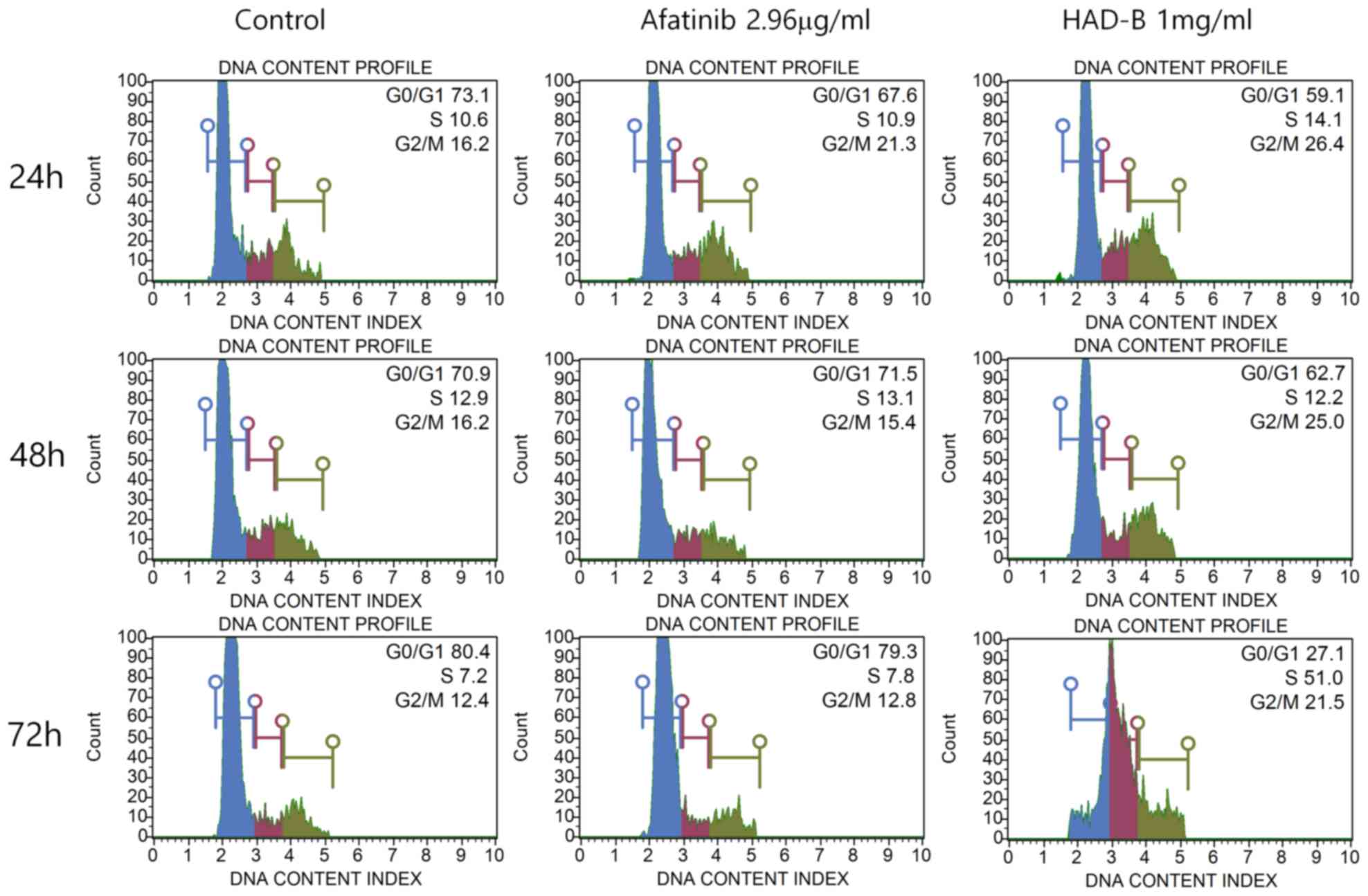
altered. Cell cycle analysis was performed using a Muse®
Cell cycle kit. The results indicated that S-phase arrest of the
cell cycle was significantly increased in cells treated with HAD-B1
for 72 h, when compared with the vehicle-treated control group and
the Afatinib-treated group (Fig.
6).
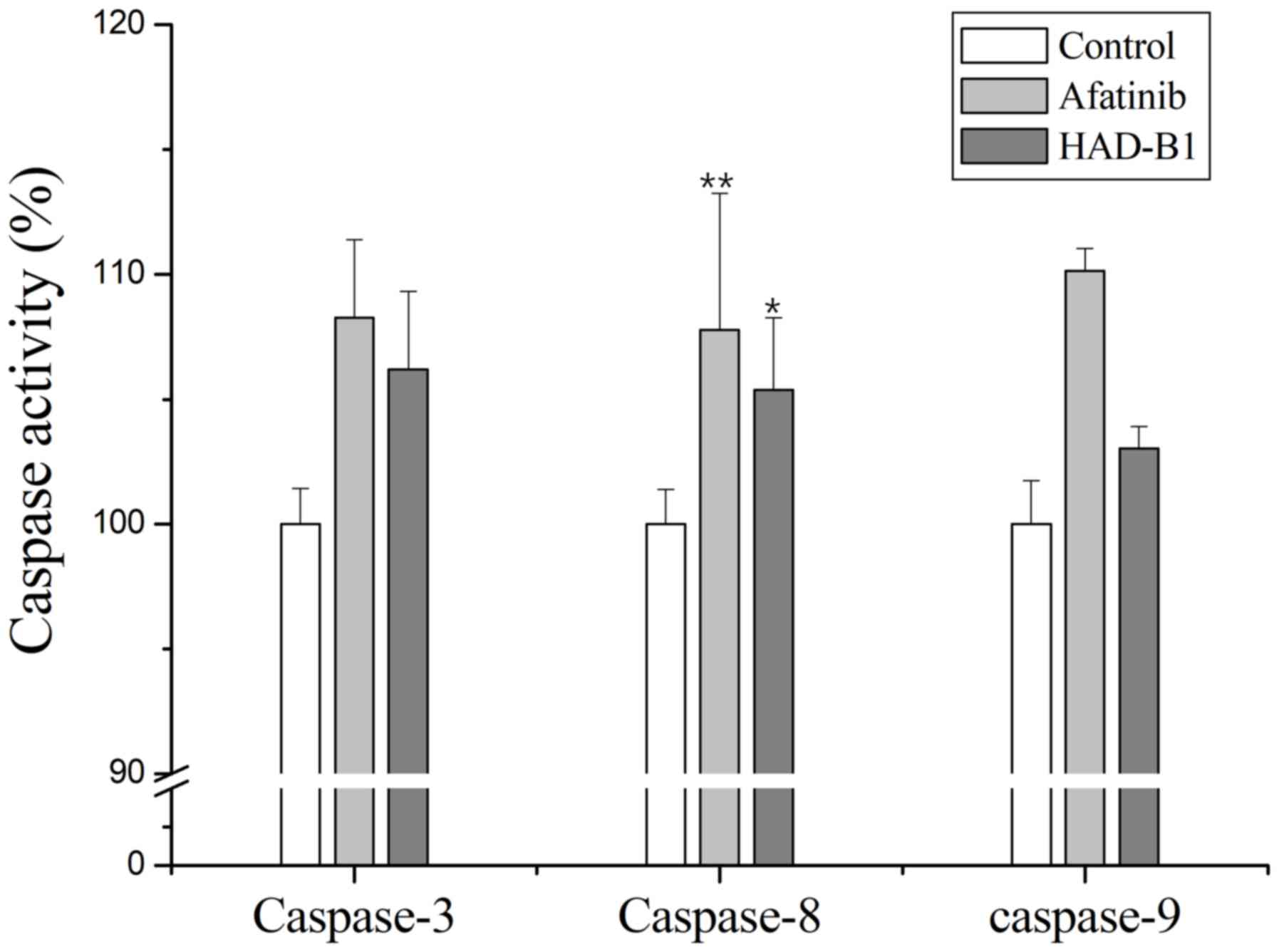
Activation of caspases in A549CR cells
treated with HAD-B1
To further investigate the induction of apoptosis in
A549CR cells treated with HAD-B1, an in vitro caspase
activity assay was performed. Caspase-3, −8 and −9 were activated
in A549CR cells treated with HAD-B1 (1 mg/ml) when compared with
the control. This result indicated that HAD-B1 may enhance caspase
activation, resulting in the induction of apoptosis (Fig. 7).
Discussion
Cisplatin is a commonly used chemotherapy agent;
however, its limitations involve drug resistance and severe side
effects (19). Due to the
difficulty in treating cancer, many patients seek other therapies
outside the field of formal medical care. These therapies are
viewed as complementary and alternative, and the interest in such
therapies is growing. According to European research, 35.9% of
patients with cancer utilize alternative medicine, and 13.9%
frequently use herbal medicines and natural products (20). In particular, interest in herbal
medicines and natural medicinal products has been revealed to
increase following the diagnosis of cancer; 13.3% of patients were
reported to have used them at diagnosis, while only 5.3% used
herbal medicines prior to diagnosis, an increase of ~3 fold
(20). Therefore, the significance
of employing natural products such as anti-cancer medications
cannot be overlooked as vinca alkaloids, paclitaxel and
epipodophyllotoxin, which, have been developed as anticancer
agents, are composed of >60% natural products (21). HAD-B1 is composed of 4 natural
products from different species of Korean medicinal plants. The
present study focused on the effect of HAD-B1 on the lung cancer
cell strain A549CR, which, is resistant to cisplatin, and the
toxicity of HAD-B1 on A549CR cells was examined. Furthermore, using
antibody microarray analysis in HAD-B1-treated A549CR cells,
alterations to STAT3 expression were detected. Western blot
analysis supported the microarray data, indicating that STAT3 was
downregulated. The STAT3 protein is ubiquitously expressed in
mammalian cells and has diverse functions during embryogenesis and
early development (22). It is
constitutively activated in a number of human cancer cells, and the
downregulation or pharmacological inhibition of STAT3 are known to
induce caspase-dependent cell death (22). The decrease of STAT3 activation
leads to downregulation of Mcl-1 gene expression in cancer cells.
When apoptotic signaling is triggered by TRAIL receptor, it
proceeds through caspase-8-mediated Bid cleavage; the decrease of
Mcl-1 protein level facilitates tBid-induced cytochrome c release
from mitochondria and leads to caspase-9 activation that synergizes
with caspase-8 to activate caspase-3 and PARP cleavage (23) Notably, Aplasia Ras homolog member I
(ARHI) mediated blockade of STAT3 signaling arrested human ovarian
cancer SKOV3 cells at S-Phase of the cell cycle, and induced
apoptosis (24). The results of
the present study indicated that the activities of caspase-3, −8
and −9 in the HAD-B1-treated group were increased. Furthermore,
inhibition of cell viability, increased apoptosis and S-phase cell
cycle arrest were also demonstrated. These results indicated that
downregulation of STAT3 in A549CR cells treated with HAD-B1
resulted in S-phase cell cycle arrest and induction of
caspase-mediated cell apoptosis.
In conclusion, HAD-B1 exhibited an anti-cancer
effect against A549CR lung-cancer cells. The present study suggests
that HAD-B1 has the potential to be a novel therapeutic agent for
treating Cisplatin Resistant NSCLC. Further studies, including
in vivo efficacy assay and mode-of-action, are required.
Acknowledgements
The present study was supported by the Daejeon
University fund (grant no. 201501160001).
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
Statistics, 2017. CA Cancer J Clin. 67:7–30. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Breathnach OS, Freidlin B, Conley B, Green
MR, Johnson DH, Gandara DR, O'Connell M, Shepherd FA and Johnson
BE: Twenty-two years of phase III trials for patients with advanced
non-small-cell lung cancer: Sobering results. J Clin Oncol.
19:1734–1742. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Meng S, Zhou Z, Chen F, Kong X, Liu H,
Jiang K, Liu W, Hu M, Zhang X, Ding C and Wu Y: Newcastle disease
virus induces apoptosis in cisplatin-resistant human lung
adenocarcinoma A549 cells in vitro and in vivo. Cancer Lett.
317:56–64. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
National Comprehensive Cancer Network
(NCCN): NCCN Clinical Practice Guidelines in Oncology (NCCN
Guidelines®)Non-Small Cell Lung Cancer. NCCN; Fort
Washington, PA: 2015
|
|
6
|
Maroun JA, Anthony LB, Blais N, Burkes R,
Dowden SD, Dranitsaris G, Samson B, Shah A, Thirlwell MP, Vincent
MD and Wong R: Prevention and management of chemotherapy-induced
diarrhea in patients with colorectal cancer: A consensus statement
by the canadian working group on chemotherapy-induced diarrhea.
Curr Oncol. 14:13–20. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
de Gramont A, Figer A, Seymour M, Homerin
M, Hmissi A, Cassidy J, Boni C, Cortes-Funes H, Cervantes A, Freyer
G, et al: Leucovorin and fluorouracil with or without oxaliplatin
as first-line treatment in advanced colorectal cancer. J Clin
Oncol. 18:2938–2947. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kim JM, Park JW, Yoo HS, Lee YW and Cho
CK: Case report of the pancreatic cancer patient after
pancreatoduodenectomy who is taking the hangam-plus to
anti-metastasis and preventing recurrence. J Korean Tradit Onco.
16:33–39. 2011.
|
|
9
|
Kim KH, Park BR, Cho CK, Lee YW, Cho EJ,
Yea SC, Yoo BC and Yoo HS: Proteome alteration in human colon
cancer cells by the treatment of HangAmDan-B. Biochip J. 5:114–122.
2011. View Article : Google Scholar
|
|
10
|
Choi YJ, Shin DY, Lee YW, Cho CK, Kim GY,
Kim WJ, Yoo HS and Choi YH: Inhibition of cell motility and
invasion by HangAmDan-B in NCI-H460 human non-small cell lung
cancer cells. Oncol Rep. 26:1601–1608. 2011.PubMed/NCBI
|
|
11
|
Zheng HM, Yoon JW, Lee YW, Cho CK, Oh DS
and Yoo HS: Case series of advanced non-small cell lung cancer
patients treated with Hang-Am Plus. Korean J Orient Int Med.
32:113–120. 2011.
|
|
12
|
Kim KS, Jung TY, Yoo HS, Lee YW and Cho
CK: Case series of advanced non-small cell lung cancer patients
treated with Hang-Am-Plus. Korean J Orient Int Med. 30:893–900.
2009.
|
|
13
|
Park BK, Yoo HS, Lee YW, Han SS, Cho JH,
Son CG and Cho CK: Retrospective cohort analysis for lung cancer
patients treated with Wheel Balance Therapy (WBT). Korean J Orient
Int Med. 29:45–56. 2008.
|
|
14
|
Bang JY, Kim EY, Yoo HS, Lee YW, Kim YS,
Cho CK, Choi Y, Jeong HJ and Kang IC: Analysis of anti-angiogenic
mechanism of HangAmDan-B (HAD-B), a Korean traditional medicine,
using antibody microarray chip. BioChip J. 4:350–355. 2010.
View Article : Google Scholar
|
|
15
|
Yoo HS, Lee HJ, Kim JS, Yoon J, Lee GH,
Lee YW and Cho CK: A toxicological study of HangAmDan-B in mice. J
Acupunct Meridian Stud. 4:54–60. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Kang HJ, Park SJ, Park YM, Yoo HS and Kang
IC: Inhibitory effects of HangAmDan-B1 (HAD-B1) on A549 lung cancer
cell proliferation and tumor growth in a xenograft model. Acad J
Sci Res. 4:187–193. 2016.
|
|
17
|
Bang JY, Kim EY, Kang DK, Chang SI, Han
MH, Baek KH and Kang IC: Pharmacoproteomic Analysis of a Novel
Cell-permeable Peptide Inhibitor of Tumor-induced Angiogenesis. Mol
Cell Proteomics. 10:M110.0052642011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Fumagalli S, Totty NF, Hsuan JJ and
Courtneidge SA: A target for Src in mitosis. Nature. 368:871–874.
1994. View
Article : Google Scholar : PubMed/NCBI
|
|
19
|
Galluzzi L, Senovilla L, Vitale I, Michels
J, Martins I, Kepp O, Castedo M and Kroemer G: Molecular mechanisms
of cisplatin resistance. Oncogene. 31:1869–1883. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Molassiotis A, Fernadez-Ortega P, Pub D,
Ozden G, Scott JA, Panteli V, Margulies A, Browall M, Magri M,
Selvekerova S, et al: Use of complementary and alternative medicine
in cancer patients: A European survey. Ann Oncol. 16:655–663. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Cragg GM and Newman DJ: Plants as a source
of anti-cancer agents. J Ethnopharma. 100:72–79. 2005. View Article : Google Scholar
|
|
22
|
Aoki Y, Feldman GM and Tosato G:
Inhibition of STAT3 signaling induces apoptosis and decreases
survivin expression in primary effusion lymphoma. Blood.
101:1535–1542. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Lirdprapamongkol K, Sakurai H, Abdelhamed
S, Yokoyama S, Athikomkulchai S, Viriyaroj A, Awale S, Ruchirawat
S, Svasti J and Saiki I: Chrysin overcomes TRAIL resistance of
cancer cells through Mcl-1 downregulation by inhibiting STAT3
phosphorylation. Int J Oncol. 43:329–337. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhu Q, Hu J, Meng H, Shen Y, Zhou J and
Zhu Z: S-Phase cell cycle arrest, apoptosis, and molecular
mechanisms of aplasia ras homolog member i-induced human ovarian
cancer SKOV3 cell lines. Int J Gynecol Cancer. 24:629–634. 2014.
View Article : Google Scholar : PubMed/NCBI
|