Introduction
Classical swine fever (CSF), which is caused by the
CSF virus (CSFV), remains one of the most important swine-related
diseases worldwide, which has yet to be eradicated in China,
despite a nationwide vaccination program (1,2).
CSFV is a single-stranded positive-sense RNA molecule ~12.3 kb that
encodes a polyprotein, which belongs to the Flaviviridae
family (3). The Shimen strain is
the standard virulent strain of CSFV in China (2), and induces severe immunopathological
symptoms associated with hemorrhagic fever, thrombocytopenia, and
lymphoid organ atrophy and severe lymphopenia, which are
accompanied by immune suppression (4). Research investigating the
immunopathogenesis of CSFV is of great importance, since the
underlying mechanisms of CSFV remain poorly understood.
Toll-like receptors (TLRs) are fundamental sensor
molecules of the host innate immune system, which detect conserved
viral structures and initiate innate immune responses (5,6).
Among the TLR family members, some viral-envelope proteins are
recognized by TLR2 and TLR4 (7);
TLR7 and TLR8 are involved in the recognition of single-stranded
RNA (ssRNA) sequences of RNA viruses (8); and TLR3 is able to detect
double-stranded RNA (dsRNA) and ssRNA viruses (9). Our previous study investigated the
gene and protein expression levels of these TLRs in response to
infection with the highly virulent CSFV Shimen strain in porcine
monocyte-derived macrophages (pMDMs) (10). Consequently, upregulation of TLR2,
4 and 7 was detected, whereas no effects on TLR3 were determined.
At present, little is currently known regarding how the tissue
expression patterns of TLRs are rapidly modulated in response to a
virulent strain of CSFV, or whether the propagation of CSFV is
influenced by specific TLR activation.
Accordingly, the present study aimed to determine
the expression patterns of TLR2, 3, 4 and 7 in tissues (heart,
liver, spleen, lung, kidney and lymph nodes) from CSFV-infected
swine compared with a control group. In addition, the relevance of
TLR2, 3, 4 and 7 activation to the CSFV life cycle in pMDMs was
examined.
Materials and methods
Infection of animals and tissue
preparation
A total of 9 Landrace pigs (castrated boars; age, 50
days; weight, 25–30 kg) provided by Professor Zhi-Zhong Jing
(Lanzhou Veterinary Research Institute, Chinese Academy of
Agricultural Sciences, Lanzhou, China) were used in the present
study. The animal procedures were approved and supervised by the
Animal Care Commission of the College of Veterinary Medicine,
Northwest A&F University (Xianyang, China). The following
housing conditions were used in the present study: Temperature,
22°C; atmosphere, −40 to −80Pa, 0.08% CO2; free access
to food and water. All pigs were free of CSFV, porcine reproductive
and respiratory syndrome virus (PRRSV) and swine influenza virus
(SIV) infection, as confirmed by the negative results from reverse
transcription (RT)-polymerase chain reaction (PCR) and antibody
(Ab) test kits used according to the manufacturer's instructions:
IDEXX CSFV Ab test, IDEXX PRRS X3 Ab test and IDEXX Influenza A Ab
test (Idexx Switzerland AG, Liebefeld, Switzerland). Subsequently,
6 pigs were injected intramuscularly with l ml CSFV Shimen strain
[105 50% tissue culture infective dose
(TCID50)/ml], which was obtained from the Control
Institute of Veterinary Bioproducts and Pharmaceuticals (Beijing,
China); the remaining pigs were used as controls. Clinical and
pathomorphological data of the infected pigs were recorded daily
postinfection. A total of 3 infected pigs were euthanized 24 h
postinfection (hpi), whereas 4 pigs, including 3 infected pigs and
1 randomly selected pig from the control group, were euthanized at
72 hpi. Subsequently, the heart, liver, spleen, lung, kidney and
lymph node tissues were collected from all euthanized pigs. The 2
remaining control pigs were used for collection of data regarding
color of conjunctiva, spirit, appetite and diarrhea data, and for
virus titer detection and pMDM preparation.
Cells, viruses and virus titer
assays
pMDMs were differentiated from the peripheral blood
mononuclear cells (PBMCs) of the 2 remaining control group pigs
(all of which were negative for CSFV, PRRSV and SIV antigens and
antibodies) using Ficoll-Paque density (1.077±0.001 g/ml) gradient
centrifugation, as described previously (10). Prior to addition of PBMCs, the
flasks were coated for 2 h with 2% sterile, endotoxin-free gelatin
in 37°C and the excess gelatin was discarded. Subsequently, flasks
were dried in the 5% CO2 incubator (37°C) overnight and
coated for 1 h with 100% FBS (Gibco; Thermo Fisher Scientific,
Inc., Waltham, MA, USA) and rinsed with 0.01 M PBS (pH 7.4). PBMCs
were seeded into gelatin and fetal FBS-coated culture plate and
were allowed to adhere for 2 h at 37°C in an atmosphere containing
5% CO2. Adherent monocytes were washed three times with
RPMI-1640 (Gibco; Thermo Fisher Scientific, Inc.) and suspended in
a mixture of RPMI-1640 containing 10% FBS and 10 mM EDTA in a 1:1
dilution. Isolated monocytes were seeded into 6-well plates at a
concentration of 1×106 cells/well in RPMI-1640
supplemented with 20% FBS and cultured at 37°C in a 5%
CO2 incubator for 8 days to obtain pMDMs.
The CSFV Shimen strain was obtained from the Control
Institute of Veterinary Bioproducts and Pharmaceuticals. The
multiplicity of infection (MOI) was determined and calculated using
the same method as described previously (11).
Stimulation of pMDMs and in vitro CSFV
inoculation
pMDMs were seeded into 12-well plates at a
concentration of 5×105 cells/well and were stimulated
with TLR-specific ligands (InvivoGen, San Diego, CA, USA):
Peptidoglycan from Staphylococcus aureus (PGN-SA; 10 µg/ml),
polyinosinic-polycytidylic acid [poly (I:C); 10 µg/ml],
lipopolysaccharide from E. coli 055:B5 (LPS-B5; 1 µg/ml) or
imiquimod (R837; 5 µg/ml), which are ligands for TLR2, 3, 4 and 7,
respectively (6). Following
stimulation for 12 h at 37°C in an atmosphere containing 5%
CO2, cells were infected with CSFV Shimen strain at an
MOI of 10. At 0 (mock), 12, 24, 48 and 72 hpi, cell lysates were
harvested and stored at −80°C until use.
RT-quantitative PCR (RT-qPCR)
Tissue samples (30 mg) and Buffer RLT (600 µl)
(Qiagen, Inc., Valencia, CA, USA) were added to sample tubes that
were prefilled with ceramic beads (MagNA Lyser Green Beads) and
were homogenized using a MagNA Lyser Instrument (both from Roche
Diagnostics, Indianapolis, IN, USA). Total RNA was prepared from
the homogenized tissue samples and cell lysates using an RNeasy
Mini kit (Qiagen, Inc.) and cDNA was synthesized with random
primers using a PrimeScript™ RT reagent kit with gDNA
Eraser, according to the manufacturer's instructions (Takara
Biotechnology Co., Ltd., Dalian, China) in 50-µl reaction mixtures
containing 2 µg total RNA. RT-qPCR was performed using a Bio-Rad
iQ5 system (Bio-Rad Laboratories, Inc., Hercules, CA, USA) and SYBR
Premix Ex Taq II (Takara Biotechnology Co., Ltd.) with
primer pairs listed in Table I
(10). qPCR was conducted
according to the following thermocycling conditions: 95°C for 30
sec, followed by 40 cycles at 95°C for 5 sec and 60°C for 30 sec.
Gene expression data were normalized by assessing the mRNA
accumulation of a housekeeping gene (β-actin) per sample. The
2−ΔΔCq method was used to calculate the relative
quantities of mRNA (12,13). For CSFV-specific detection, a
recombinant pCMV-myc plasmid containing the CSFV gene NS2 (Sangon
Biotech Co., Ltd., Shanghai, China) was used to build a
fluorescence quantitative standard curve, and quantification of
viral RNA was performed as previously described (10).
 | Table I.Primer sets used in the present
study. |
Table I.
Primer sets used in the present
study.
| Genes | Forward primer
(5′-3′) | Reverse primer
(5′-3′) | GenBank accession
number |
|---|
| CSFV |
GATCCTCATACTGCCCACTTAC |
GTATACCCCTTCACCAGCTTG | AF092448 |
| β-actin |
CAAGGACCTCTACGCCAACAC |
TGGAGGCGCGATGATCTT | DQ845171.1 |
| TLR2 |
GGAGCCTTAGAAGTAGAGTTTG |
TGTATCCACATTACCGAGGG | AB208696.2 |
| TLR3 |
TAACAACCTTCCAGGCATA |
AAGAGGAGAATCAGCGAGTG | DQ266435.1 |
| TLR4 |
TCTACATCAAGTGCCCCTAC |
ATTCTCCCAAAACCAAC | AB188301.2 |
| TLR7 |
CATGTGATCGTGGACTGC |
GTGATGCTCGCTATGTGG | EF583901.1 |
Western blot analysis
For western blotting, sample tubes were prefilled
with ceramic beads, tissue (100 mg) and radioimmunoprecipitation
assay buffer (1 ml) with Halt™ Protease Inhibitor Cocktail (cat.
no. 78410) (both from Thermo Fisher Scientific, Inc.).
Subsequently, the samples were homogenized using a MagNA Lyser
Instrument. The mixtures were shaken gently for 15 min on ice and
were then centrifuged at 14,000 × g for 15 min to pellet the tissue
debris. The supernatant was transferred to a new tube and
quantified using a bicinchoninic acid kit (Thermo Fisher
Scientific, Inc.). Subsequently, the proteins were boiled with SDS
sample loading buffer (5X), loaded 20 µl (50 µg/ml) onto gels and
separated by 12% SDS-PAGE and transferred onto polyvinylidene
difluoride membranes (EMD Millipore, Billerica, MA, USA). The
membranes were blocked with TBS-0.05% Tween containing 5% skim milk
at room temperature for 2 h and were then incubated with primary
antibodies at 4°C overnight, as previously described (10). The following primary antibodies
were used: Rabbit anti-TLR2 antibody (cat. no. LS-C116906-50;
dilution 1:200), mouse anti-TLR3 antibody (cat. no. LS-C545-100;
dilution 1:500) (both from LifeSpan Biosciences, Inc., Seattle, WA,
USA), mouse anti-TLR4 antibody (cat. no. ab8376; dilution 1:500;
Abcam, Cambridge, MA, USA), goat anti-TLR7 antibody (cat. no.
sc-13207; dilution 1:200; Santa Cruz Biotechnology, Inc., Dallas,
TX, USA) and mouse anti-β-actin (cat. no. GTX109639; dilution
1:1,000; GeneTex, Inc., Irvine, CA, USA). After several washes with
TBST the membranes were incubated with goat anti-rabbit IgG+IgA+IgM
secondary antibody horseradish peroxidase (HRP) conjugated (cat.
no. LS-C347361), or goat anti-mouse IgG secondary antibody (HRP)
(cat. no. LS-C60710), or rabbit anti-goat IgG Fc secondary
antibodies (cat. no. LS-C69762-1000) (all 1:5,000; LifeSpan
Biosciences, Inc.) for 2 h at room temperature. Unbound antibodies
were removed by washing with TBST, and the signal was detected
using an ChemiDoc™ XRS (Bio-Rad Laboratories, Inc.) with an
exposure time ranging between 30 sec and 4 min. Image Lab™ software
(5.0) (Bio-Rad Laboratories, Inc.) was used to semi-quantify the
blots.
Histological examination and
immunohistochemistry (IHC)
Tissues from the infected swine were immersion-fixed
in cold 4% paraformaldehyde fixative (pH 7.4) for 48 h at room
temperature and were then transferred to 70% ethanol prior to
processing for routine paraffin embedding. Several 5-µm sections
were prepared from each block and stained with hematoxylin and
eosin to observe pathological alterations (14). Briefly, the paraffin sections
underwent conventional dewaxing and rehydration; the cell nucleus
was then stained as follows: Cells were stained with hematoxylin
for 10 min, rinsed under running water for 5 min, incubated in 1%
hydrochloric acid ethanol for 20 sec and rinsed under running water
for 15 min, prior to rinsing with distilled water for 2 min. Cell
cytoplasm was stained as follows: Cells were stained with eosin for
3–5 min, rinsed with distilled water for 30 sec, dehydrated for
transparency and mounted with neutral gum; after drying,
microscopic structures and pathological alterations were detected
using a light microscope (Eclipse TS l00-F; Nikon, Tokyo,
Japan).
IHC staining was performed to detect the presence of
TLR antigens in the tissues as previously described, using the same
antibodies as for western blotting above (15,16).
Briefly, paraffin-embedded sections were immersed in xylene for
dewaxing and rehydrated with gradient washes in pure alcohol and
alcohol diluted with distilled water. The specimens were heated in
a modified citrate buffer (pH 6.0; Abcam) at 94°C for 45 min to
improve antigenicity. Subsequently, each section was immersed in 50
µl 3% H2O2 for 30 min at room temperature,
after which sections were washed three times in 0.01 M PBS (pH 7.4)
to remove any residual reagents. After blocking with 50 µl
nonimmunized autologous serum (Beijing Solarbio Science and
Technology Co., Ltd., Beijing, China) for 1 h at room temperature,
the sections were incubated with anti-TLR2, anti-TLR3, anti-TLR4 or
anti-TLR7 (1:200 dilution) at 4°C overnight. The sections were then
incubated with HRP-conjugated goat anti-rabbit (or anti-mouse) IgG
secondary antibody (1:5,000) for 30 min at 37°C. After incubation
with 1–3 drops of peroxidase substrate (DAB; 0.6 mg/ml of 0.3%
H2O2 in 0.01 M Tris-HCl buffer, pH 7.6) and
subsequent washing in deionized H2O, the sections were
counterstained with Gill's hematoxylin and mounted on xylene-based
crystal mounts. To identify the background staining, PBS (0.01 M,
pH 7.4) instead of primary antibody was used as a negative control.
The light microscope used to observe IHC was the same as that for
H&E.
Statistical analysis
Data are presented as the mean ± standard deviation
and were analyzed by one-way analysis of variance followed by
Tukey's multiple comparison test using SPSS 17.0 software (SPSS,
Inc., Chicago, IL, USA). P<0.05 was considered to indicate a
statistically significant difference.
Results
In vivo inoculation of pigs with
CSFV
Within 72 hpi swine from the CSFV infection group
exhibited the following clinical symptoms and pathomorphological
alterations: Redness of the conjunctiva with secretions, increased
rectal temperature (40.8–41.4°C) and anorexia (Table II).
 | Table II.Clinical and pathomorphological data
of the CSFV-infected pigs. |
Table II.
Clinical and pathomorphological data
of the CSFV-infected pigs.
|
|
|
| Clinical data | Pathomorphological
data |
|---|
|
|
|
|
|
|
|---|
| Group | Animal ID number | Rectal temperature
(°C) | Color of
conjunctiva | Spirit, appetite | Diarrhea | Splenic
infarction | Visceral
hemorrhage |
|---|
| CSFV infection 72
h | 17 | 40.8 | Red | Depressed | – | – | Renal hemorrhage |
|
| 18 | 41.4 | Red with
secretions | Depressed | √ | √ | Lymph node
hemorrhage, renal hemorrhage |
|
| 20 | 41.0 | Red with
secretions | Depressed | √ | – | Lymph node
hemorrhage, renal hemorrhage |
| Control | 22 | 39.2 | – | – | – | – | – |
|
| 24 | 39.3 | – | – | – | / | / |
|
| 25 | 39.0 | – | – | – | / | / |
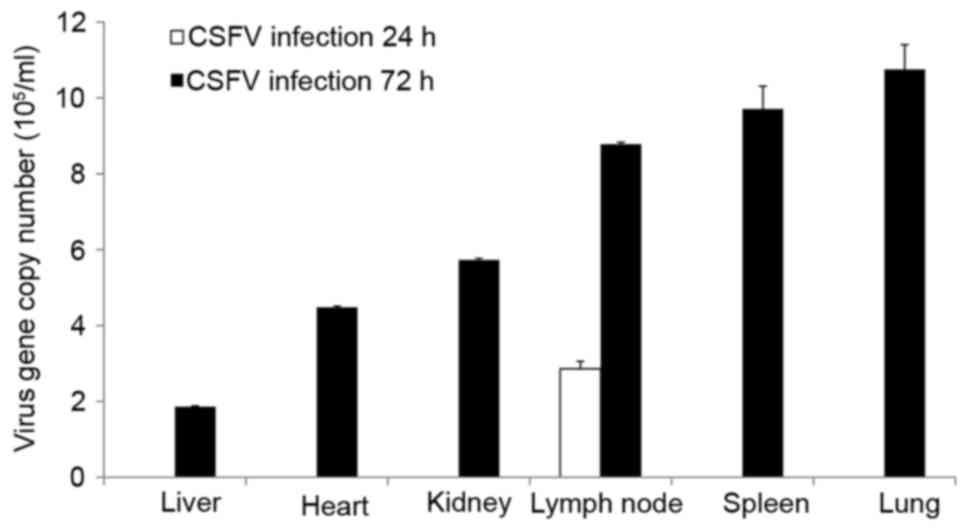
CSFV RNA was only detected in the lymph nodes at 24
hpi; however, it was detected in all organs at 72 hpi, as
determined by RT-qPCR (Fig. 1).
The tissues, ordered by the highest to the lowest amounts of viral
RNA at 72 hpi, were as follows: Lung, spleen, lymph node, kidney,
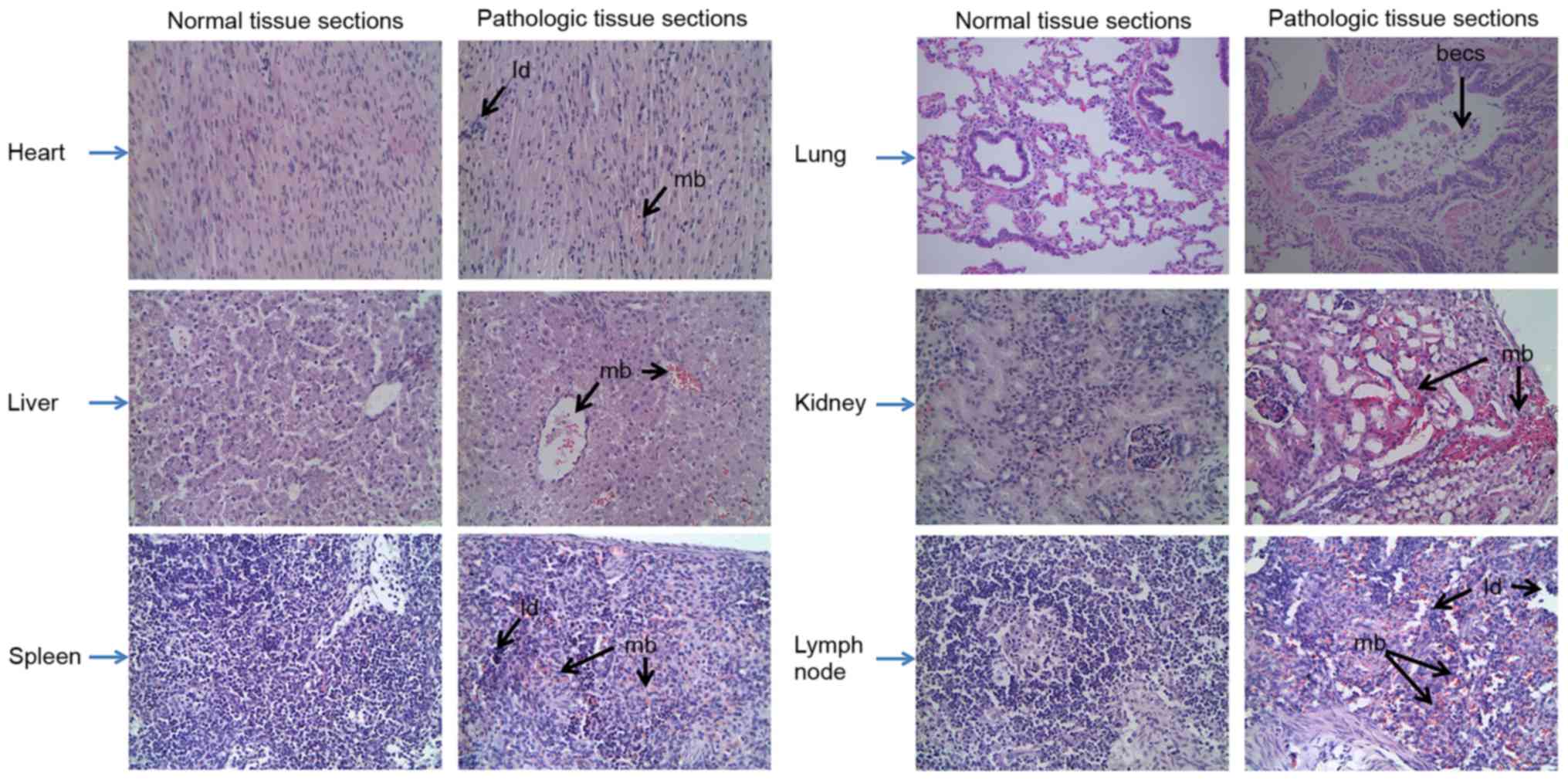
heart and liver. Furthermore, the characteristics of the various
swine organs infected with CSFV Shimen strain are presented in
Fig. 2; characteristics included
mild bleeding, the presence of lymphocyte debris and bronchial
epithelial cell shedding.
Alterations in TLR expression in
response to CSFV challenge
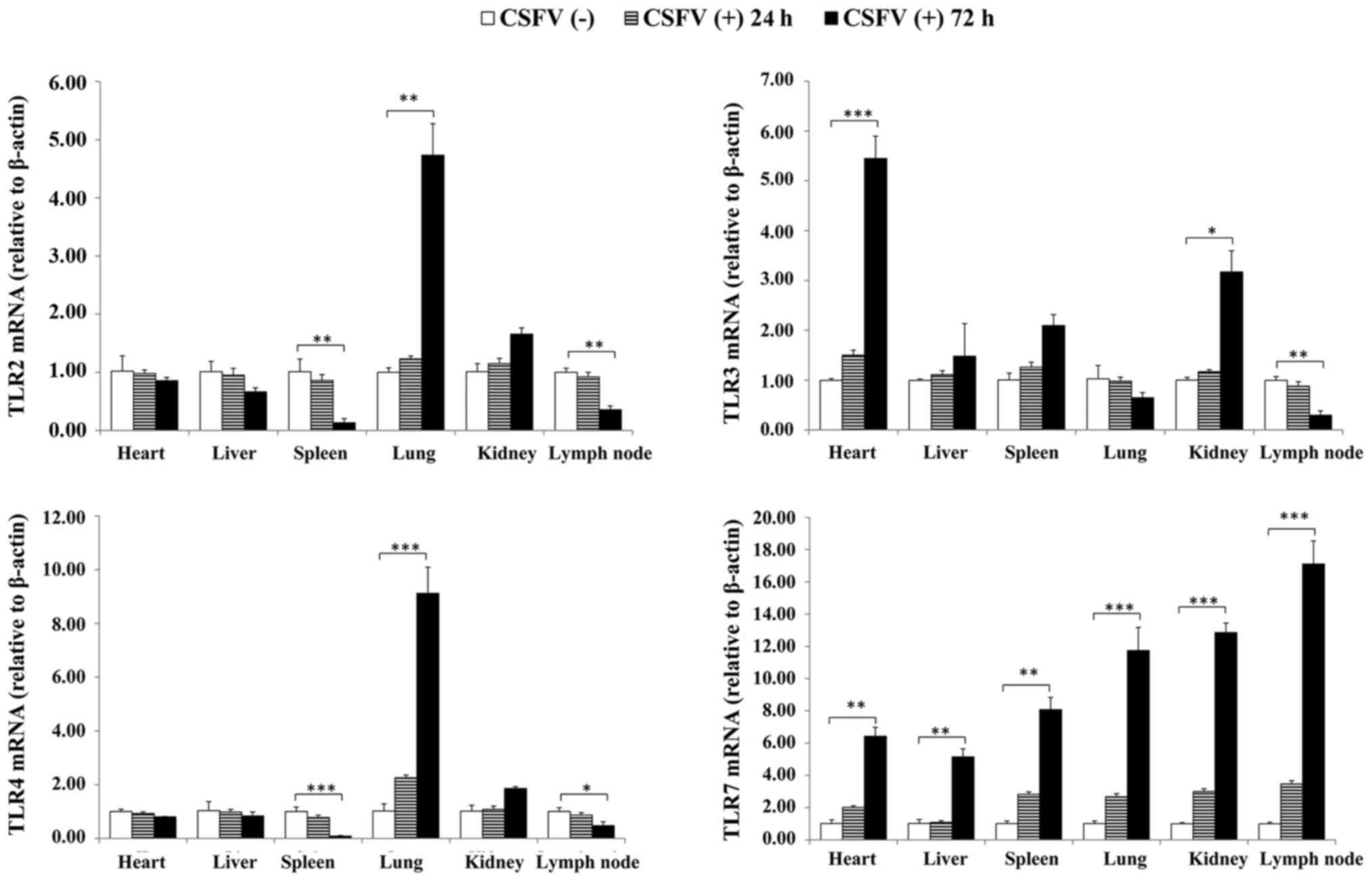
RT-qPCR was conducted to determine the expression
levels of TLR2, 3, 4 and 7 in the tissues of CSFV-infected pigs at
24 and 72 hpi. As presented in Fig.
3, the expression levels of TLR2 and TLR4 were similarly
upregulated in the lung and kidney, but were downregulated in the
spleen and lymph nodes; TLR7 was expressed at the highest levels in
all organs; and TLR3 expression was increased in the heart, spleen
and kidney, but was decreased in the lymph nodes. Notably, the
expression levels of TLR2, 3, 4, and 7 did not significantly differ
at 24 hpi. Therefore, sample collection was performed at 72 hpi for
western blotting.
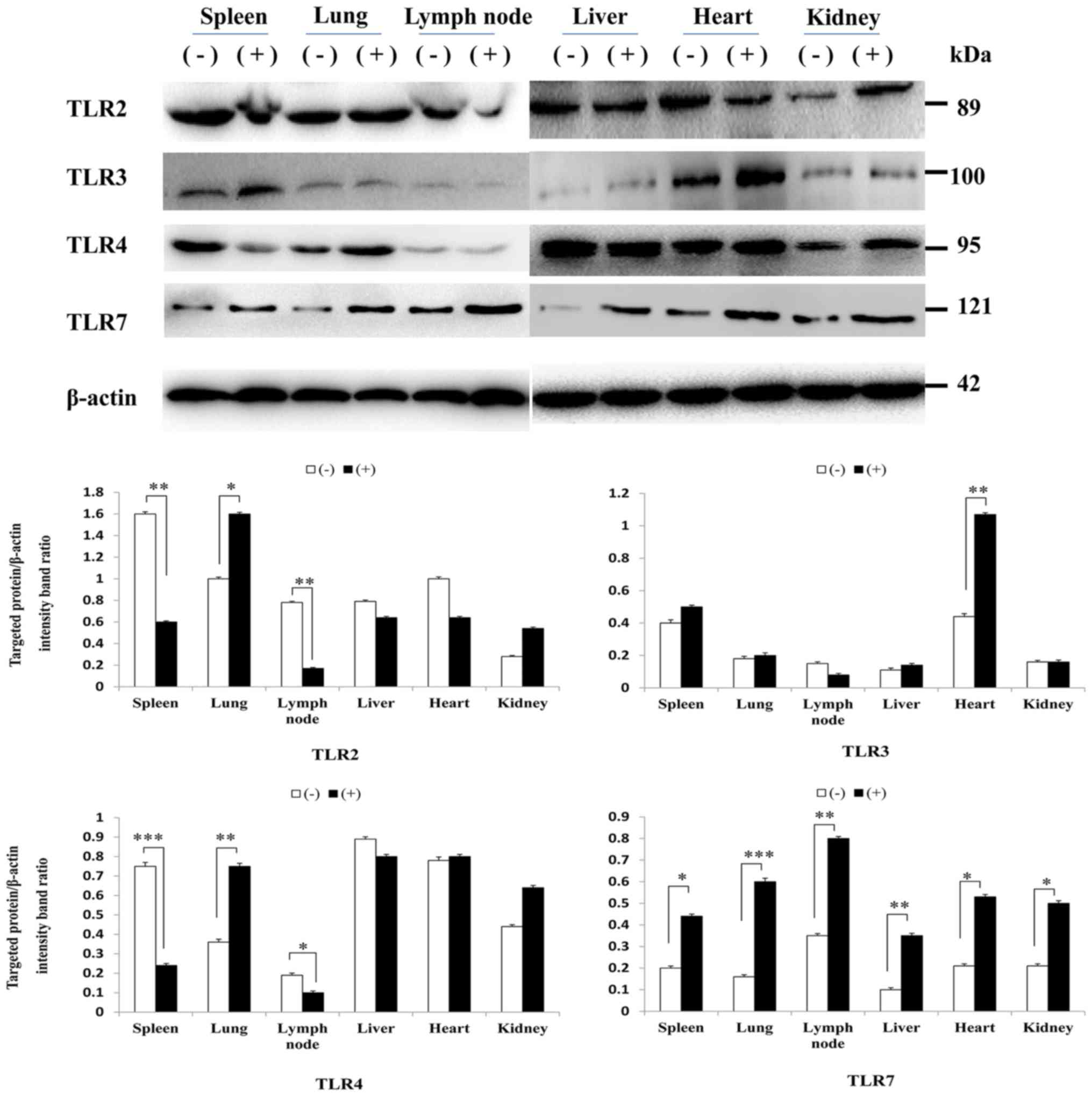
Using western blot analysis, it was indicated that
the protein expression levels of TLR2, 4 and 7 in each tissue
appeared to correlate with mRNA levels. However, the protein
expression levels of TLR3 were not consistent with mRNA levels;
there were no significant differences in TLR3 protein expression in
the kidney and lymph nodes (Fig.
4).
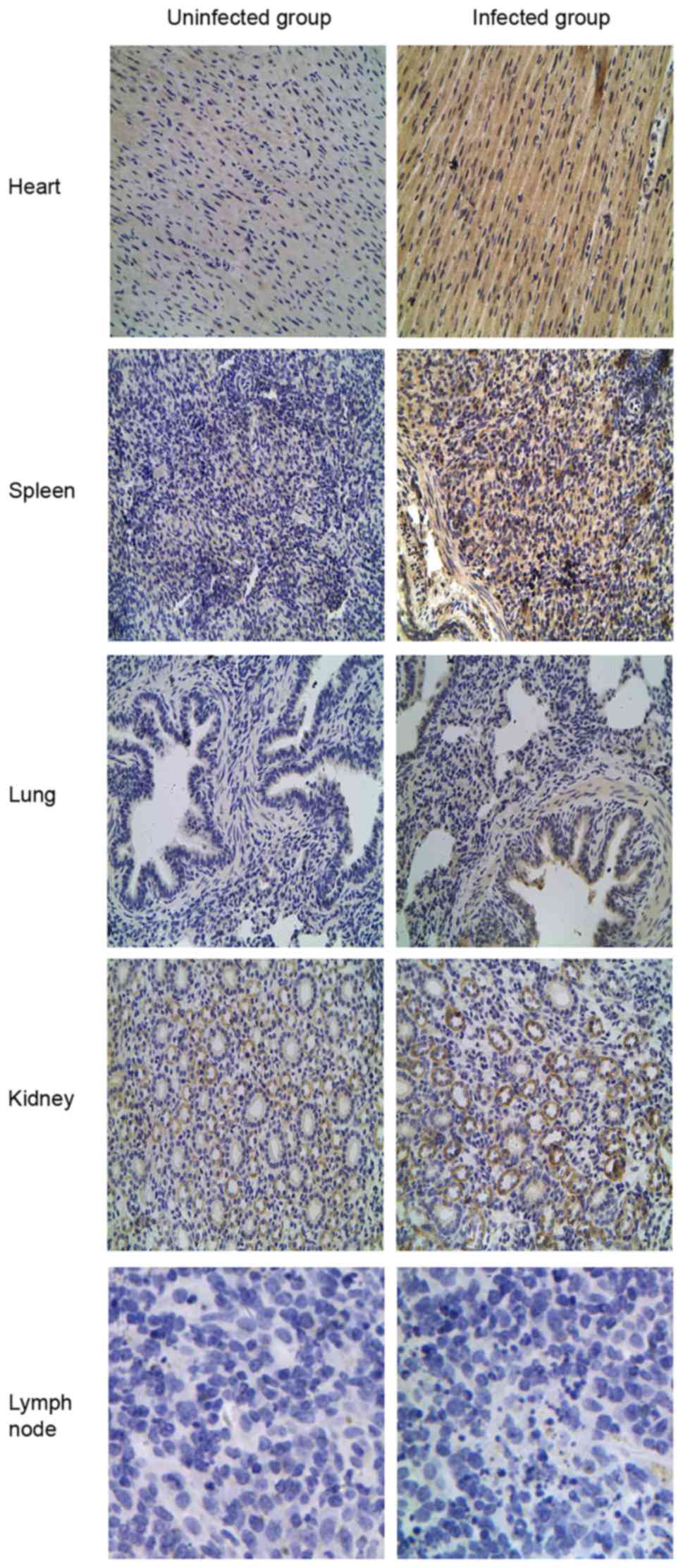
IHC staining of TLR2, 3, 4 and 7
expression in porcine tissues
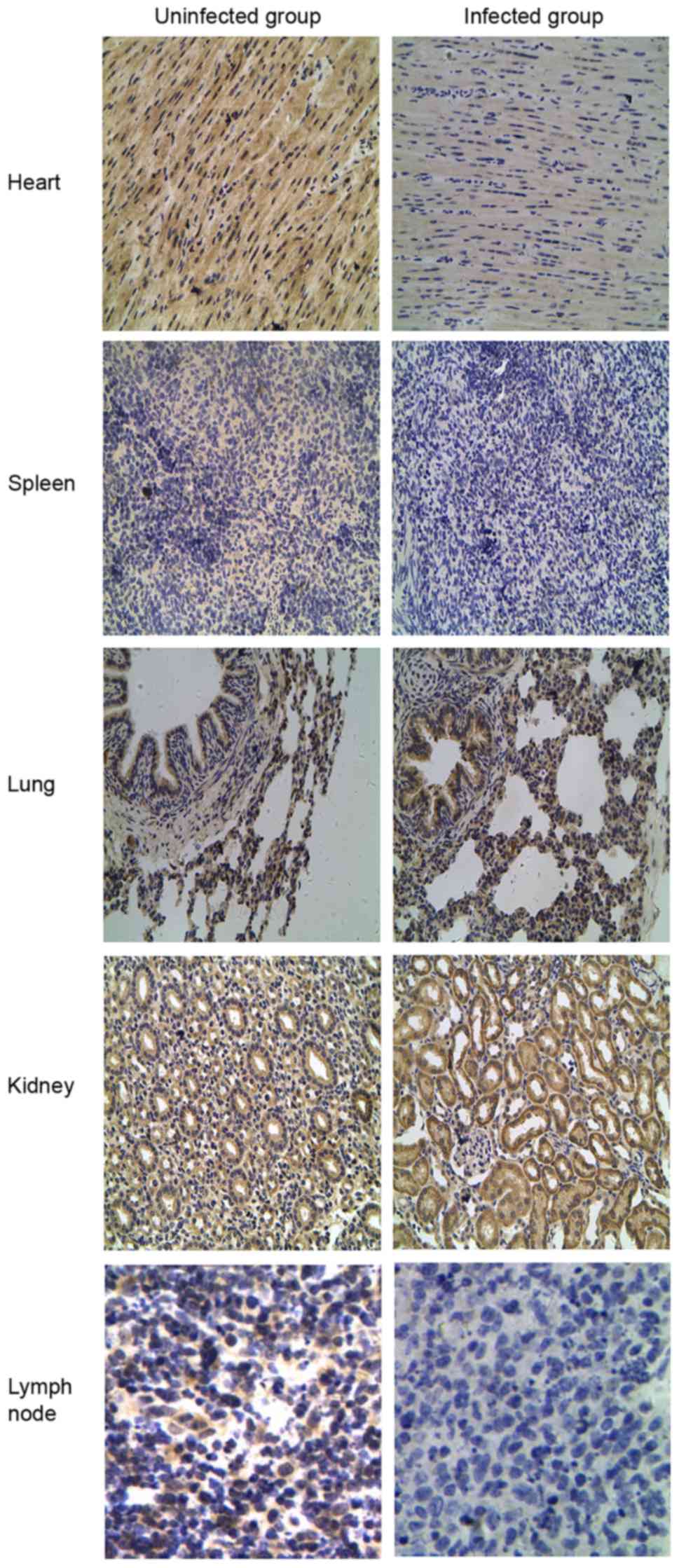
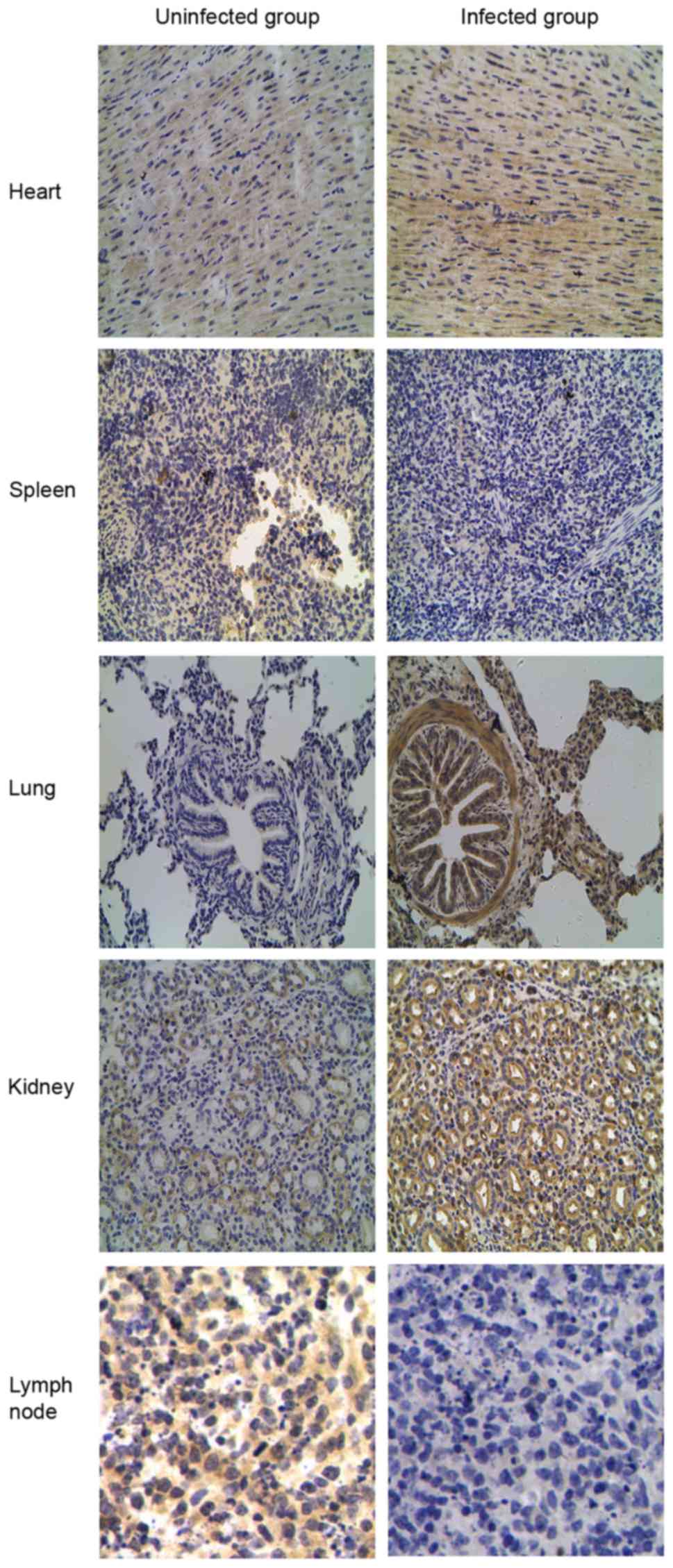
TLR tissue distributions were compared by IHC
staining between the CSFV-infected and uninfected groups to verify
the differential expression patterns of TLR2, 3, 4 and 7. Different
expression patterns for TLR2 and 4 were observed in CSFV-infected
porcine tissues (Figs. 5 and
6). Compared with the uninfected
group, TLR2 and 4 expression was most abundant in the lung and
kidney but was decreased in the spleen and lymph nodes from the
CSFV-infected group. Furthermore, analysis of TLR3-positive tissues
indicated that the heart expressed the highest levels of TLR3
compared with the other tissues (Fig.
7), and TLR3 was not detected in the lymph nodes from either
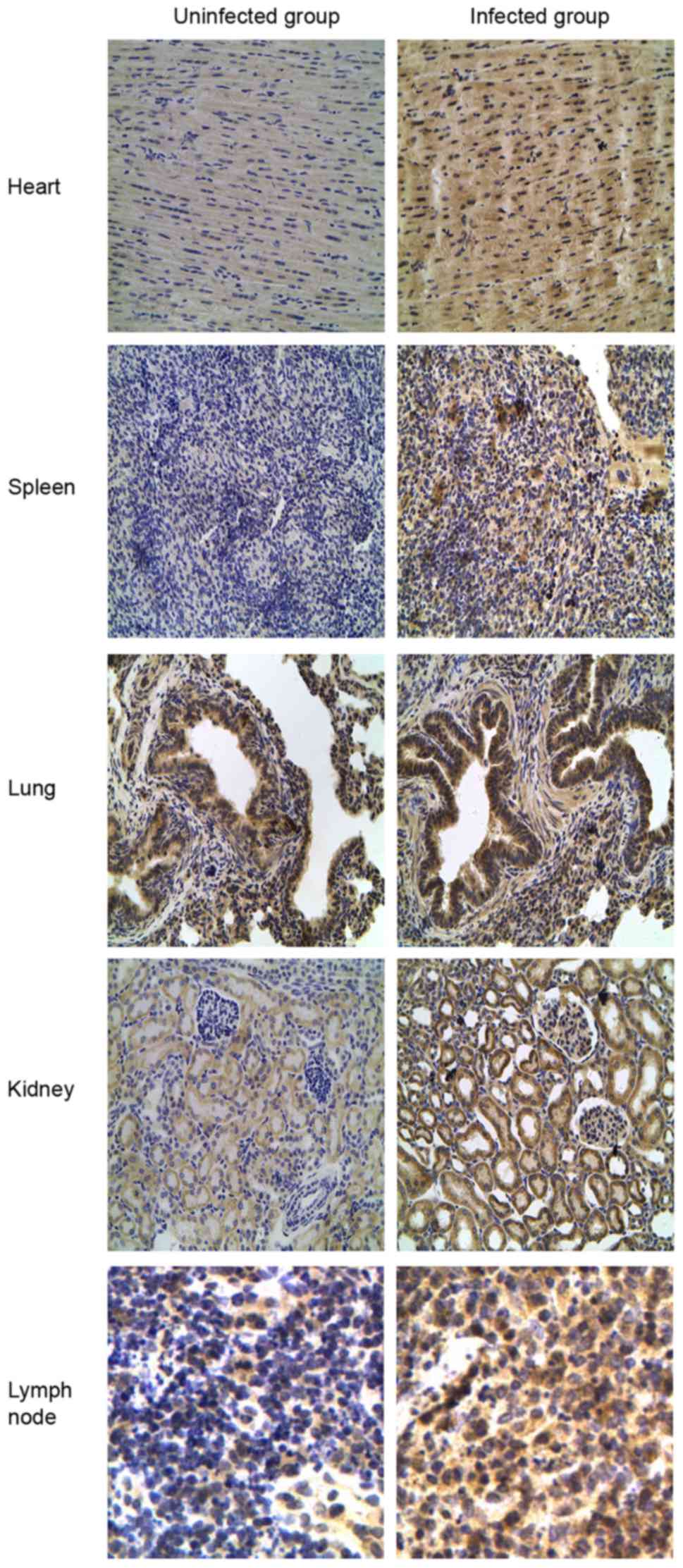
the CSFV-infected or uninfected groups. In addition, the intensity
of TLR7 staining was much stronger in all infected tissues (heart,
spleen, lung, kidney and lymph nodes) compared with in uninfected
tissues (Fig. 8).
Effects of TLR agonists on the growth
of CSFV
The present study aimed to reveal the relevance of
TLR2, 3, 4 and 7 activation to the CSFV life cycle in pMDMs by
measuring intracellular CSFV copy numbers. Therefore, following
prestimulation with PGN-SA, poly(I:C), LPS-B5 or R837, the
intracellular copy numbers of CSFV were detected by RT-qPCR. The
results indicated that R837 was able to inhibit the proliferation
of CSFV at 12, 24, 48 and 72 h.; and LPS-B5 costimulation was able
to decrease CSFV propagation at 24, 48 and 72 h. Conversely, PGN-SA
and poly(I:C) had no significant effect on CSFV proliferation at
any time-point (Fig. 9).
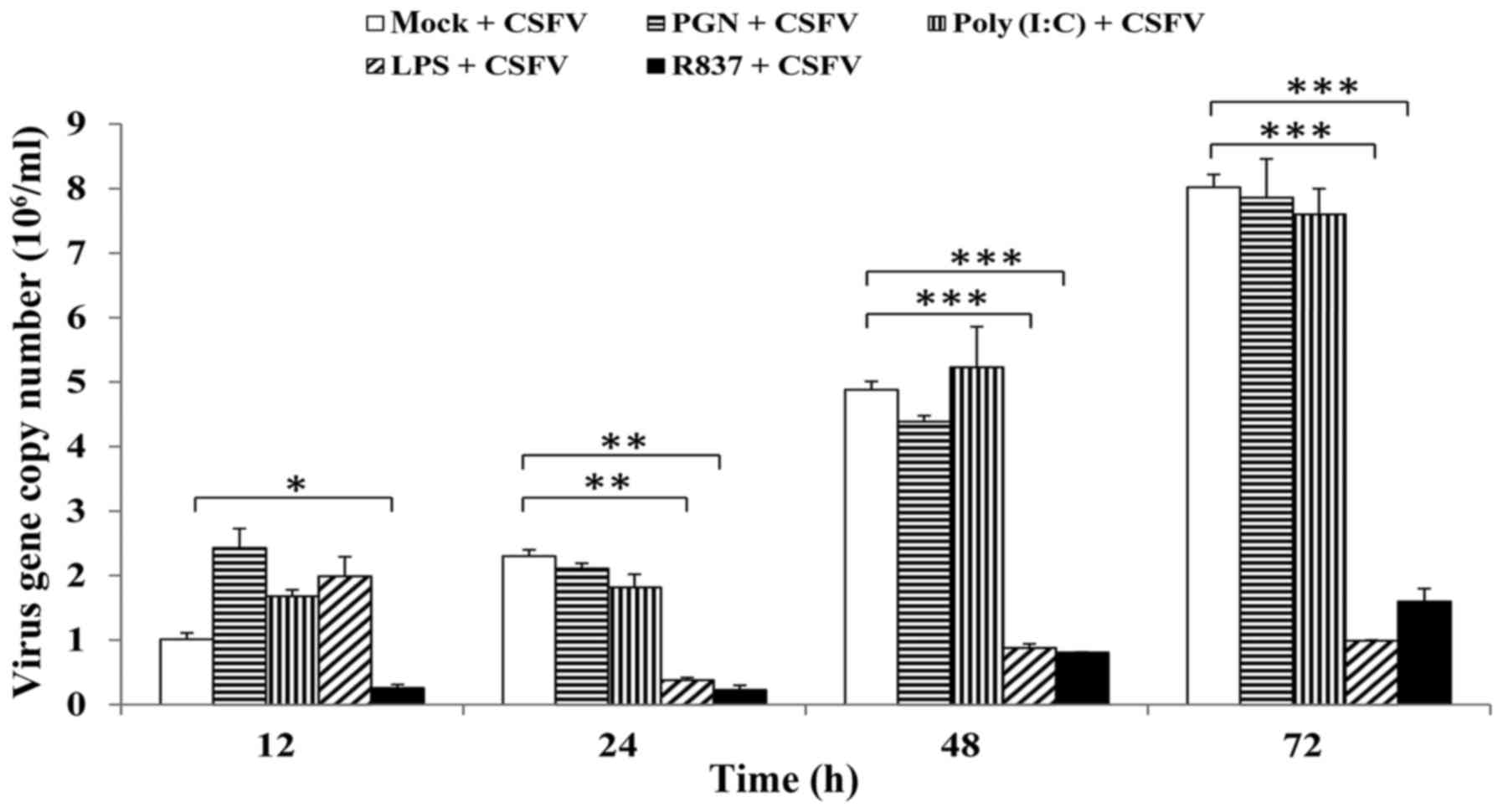 | Figure 9.Effects of TLR2, 3, 4 and 7 ligands on
CSFV proliferation. Porcine monocyte-derived macrophages were
seeded onto 12-well plates at a concentration of 5×105
cells/well and stimulated with PGN from Staphylococcus
aureus (10 µg/ml), poly (I:C) (10 µg/ml), LPS from
Escherichia coli 055:B5 (1 µg/ml) or R837 (5 µg/ml).
Following stimulation for 12 h, cells were infected with CSFV
Shimen strain at a multiplicity of infection of 10. At 0 (mock),
12, 24, 48 and 72 h postinfection, the cells were lysed, and the
copy numbers of CSFV were measured by reverse
transcription-quantitative polymerase chain reaction. pCMV-myc
plasmid encoding CSFV NS2 protein was used to build a fluorescence
quantitative standard curve for calculating the copy numbers of
CSFV in the different samples. Data are presented as the mean ±
standard deviation of three independent experiments. *P<0.05,
**P<0.01 and ***P<0.001 compared with the mock + CSFV group,
as calculated by two-way analysis of variance. CSFV, classical
swine fever virus; LPS, lipopolysaccharide; PGN, peptidoglycan;
poly (I:C), polyinosinic-polycytidylic acid; R837, imiquimod. |
Discussion
TLRs represent essential factors of the innate
immune system and provide a crucial link to the adaptive response
(17,18). It has previously been suggested
that the expression patterns of TLRs may vary in different tissues
and cell types, and in response to pathogen-associated molecular
patterns (PAMPs) (19). The aim of
the present study was to investigate TLR expression in tissues from
CSFV-infected swine at early time-points.
Due to the acidification of endosomal vesicles, TLR7
is able to detect GU-rich and AU-rich ssRNA sequences of RNA
viruses (20). Consistent with the
role of TLR7 in immune surveillance (i.e., the recognition of viral
ssRNA), the mRNA and protein expression levels of TLR7 were
increased in the examined tissues in the presence of CSFV. IHC
staining demonstrated that TLR7 was expressed in various tissues
and was most abundant in the infected group. Furthermore, CSFV
growth was significantly reduced following stimulation with R837,
which is a ligand for TLR7. Taken together, these findings
suggested that TLR7 signaling is important for detection of the
CSFV.
TLR2 and 4 are expressed on the cell surface and
primarily recognize viral glycoproteins or viral proteins released
into the extracellular space (7).
Notably, TLR2 and 4 expression was similarly associated with the
pathogenesis of CSFV in swine, thus suggesting that their target
gene expression and signaling pathways may be similar. The present
study demonstrated that CSFV infection induced distinct alterations
in TLR2 and 4 expression in tissues, including increased expression
in the lung and kidney but decreased expression in the spleen and
lymph nodes, suggesting that important roles exist for these
receptors in the establishment and resolution of infections and
inflammation. Furthermore, the proliferation of CSFV was inhibited
following treatment with LPS, which is probably due to the
activation of TLR4. However, LPS is a well-described and widely
accepted antigen, which has been reported to influence a wide
spectrum of immunological responses, not just TLR4.
TLR3 recognizes the dsRNA generated during viral
infection as a replication intermediate for ssRNA viruses (17–21).
The results of the present study indicated that TLR3 was strongly
expressed in the heart and slightly upregulated in the spleen in
response to CSFV Shimen strain infection. However, a further
experiment demonstrated that poly(I:C) had no significant effect on
the proliferation of CSFV at any time point. In a previous study,
TLR3 mediated innate responses induced by poly(I:C) were inhibited
in Shimen strain infected pMDMs (10). This contradiction between the
expression of TLR3 in pMDMs and infected tissues requires further
study.
The various expression levels of TLR2, 3, 4 and 7
detected during tissue damage in disease situations suggested that
CSFV may serve as a TLR agonist or inhibitor. In addition, it
remains to be determined as to whether the induction of these TLRs
may have therapeutic implications. In particular, the use of TLRs
themselves, PAMPs, specific cytokines and endogenous agonists, such
as proteins (e.g., high mobility group box 1), lipids (e.g.,
oxidized phospholipids) and nucleic acids, which stimulate immune
cells or nonimmune cells, may be considered targets for the
development of effective vaccine adjuvants (22–25).
In conclusion, the present study is the first, to
the best of our knowledge, to detect the distinct alterations in
TLR2, 3, 4 and 7 expression in the presence of the highly virulent
CSFV Shimen strain. Furthermore, the effects of PGN-SA, poly (I:C),
LPS-B5 and R837 on CSFV replication were observed. The results
highlighted the possible roles of TLR expression in the context of
CSFV infection and may provide a basis for the further examination
of the functions of other members of the Flaviviridae family
in the innate immune system.
Acknowledgements
The present study was supported by the National
Natural Science Foundation of China (grant no. 31472210).
References
|
1
|
Edwards S, Fukusho A, Lefèvre PC, Lipowski
A, Pejsak Z, Roehe P and Westergaard J: Classical swine fever: The
global situation. Vet Microbiol. 73:103–119. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Luo Y, Li S, Sun Y and Qiu HJ: Classical
swine fever in China: A minireview. Vet Microbiol. 172:1–6. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Paton DJ and Greiser-Wilke I: Classical
swine fever-an update. Res Vet Sci. 75:169–178. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Jamin A, Gorin S, Cariolet R, Le Potier MF
and Kuntz-Simon G: Classical swine fever virus induces activation
of plasmacytoid and conventional dendritic cells in tonsil, blood,
and spleen of infected pigs. Vet Res. 39:72008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kawai T and Akira S: The role of
pattern-recognition receptors in innate immunity: Update on
Toll-like receptors. Nat Immunol. 11:373–384. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Moresco EM, LaVine D and Beutler B:
Toll-like receptors. Curr Biol. 21:R488–R493. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lester SN and Li K: Toll-like receptors in
antiviral innate immunity. J Mol Biol. 426:1246–1264. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Heil F, Hemmi H, Hochrein H, Ampenberger
F, Kirschning C, Akira S, Lipford G, Wagner H and Bauer S:
Species-specific recognition of single-stranded RNA via toll-like
receptor 7 and 8. Science. 303:1526–1529. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Guillot L, Le Goffic R, Bloch S, Escriou
N, Akira S, Chignard M and Si-Tahar M: Involvement of toll-like
receptor 3 in the immune response of lung evithelial cells to
double-stranded RNA and influenza A virus. J Biol Chem.
280:5571–5580. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Cao Z, Guo K, Zheng M, Ning P, Li H, Kang
K, Lin Z, Zhang C, Liang W and Zhang Y: A comparison of the impact
of Shimen and C strains of classical swine fever virus on Toll-like
receptor expression. J Gen Virol. 96:1732–1745. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ning P, Zhang Y, Guo K, Chen R, Liang W,
Lin Z and Li H: Discovering up-regulated VEGF-C expression in swine
umbilical vein endothelial cells by classical swine fever virus
Shimen. Vet Res. 45:482014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Pei J, Zhao M, Ye Z, Gou H, Wang J, Yi L,
Dong X, Liu W, Luo Y, Liao M and Chen J: Autophagy enhances the
replication of classical swine fever virus in vitro. Autophagy.
10:93–110. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Fischer AH, Jacobson KA, Rose J and Zeller
R: Hematoxylin and eosin staining of tissue and cell sections. CSH
protocols. 2008:pdb prot4986. 2008.
|
|
15
|
Xiao SY, Zhang H, Guzman H and Tesh RB:
Experimental yellow fever virus infection in the golden hamster
(Mesocricetus auratus). II. pathology. J Infect Dis.
183:1437–1444. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
16
|
Benias PC, Gopal K, Bodenheimer H Jr and
Theise ND: Hepatic expression of toll-like receptors 3, 4 and 9 in
primary biliary cirrhosis and chronic hepatitis C. Clin Res Hepatol
Gastroenterol. 36:448–454. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kawai T and Akira S: Toll-like receptors
and their crosstalk with other innate receptors in infection and
immunity. Immunity. 34:637–650. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Akira S, Uematsu S and Takeuchi O:
Pathogen recognition and innate immunity. Cell. 124:783–801. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Jiang D, Liang J, Li Y and Noble P: The
role of Toll-like receptors in non-infectious lung injury. Cell
Res. 16:693–701. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lund JM, Alexopoulou L, Sato A, Karow M,
Adams NC, Gale NW, Iwasaki A and Flavell RA: Recognition of
single-stranded RNA viruses by Toll-like receptor 7. Proc Natl Acad
Sci USA. 101:pp. 5598–5603. 2004; View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Alexopoulou L, Holt AC, Medzhitov R and
Flavell RA: Recognition of double-stranded RNA and activation of
NF-kappaB by Toll-like receptor 3. Nature. 413:732–738. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Tsung A, Sahai R, Tanaka H, Nakao A, Fink
MP, Lotze MT, Yang H, Li J, Tracey KJ, Geller DA and Billiar TR:
The nuclear factor HMGB1 mediates hepatic injury after murine liver
ischemia-reperfusion. J Exp Med. 201:1135–1143. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Stewart CR, Stuart LM, Wilkinson K, van
Gils JM, Deng J, Halle A, Rayner KJ, Boyer L, Zhong R, Frazier WA,
et al: CD36 ligands promote sterile inflammation through assembly
of a Toll-like receptor 4 and 6 heterodimer. Nat Immunol.
11:155–161. 2010. View
Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kanzler H, Barrat FJ, Hessel EM and
Coffman RL: Therapeutic targeting of innate immunity with Toll-like
receptor agonists and antagonists. Nat Med. 13:552–559. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
O'Neill LA, Bryant CE and Doyle SL:
Therapeutic targeting of Toll-like receptors for infectious and
inflammatory diseases and cancer. Pharmacol Rev. 61:177–197. 2009.
View Article : Google Scholar : PubMed/NCBI
|