Introduction
Lung cancer is the most commonly diagnosed type of
cancer worldwide (1), with a
5-year survival rate of ~20% (2).
With increased air pollution exposure and smoking, the incidence of
non-small cell lung cancer (NSCLC) and its respective mortality
rate have risen, threatening the health of individuals worldwide.
Therefore, obtaining a clear understanding of the underlying
mechanisms of NSCLC is critical for improving the efficacy of
clinical treatment. Long non-coding RNAs (lncRNAs) are transcripts
that are less than 200 nucleotides in length (3). However, lncRNAs are involved in
various functions in cancer cells, including cell growth and
metastasis (4), which are
implicated in cancer progression.
The functions of a novel cytoplasmic lncRNA,
NORAD (non-coding RNA activated by DNA damage), also known
as LINC00657 or LOC647979, have been investigated (5). NORAD serves as an oncogene and is
associated with overall survival in breast cancer (6) and pancreatic cancer (7). However, its underlying mechanisms
have not been revealed. Existing evidence suggests that lncRNAs
could interact with microRNAs as competing endogenous RNAs (ceRNAs)
or ‘RNA sponges’, recruiting these molecules and reducing their
regulatory effect on target mRNAs (8,9). In
patients with pancreatic cancer, NORAD is believed to serve
as a sponge for miR-125a-3p to regulate Ras homolog family member A
(7). NORAD was reported to be
associated with epithelial-mesenchymal transition, metastasis and
poor prognosis in patients with colorectal cancer, by interacting
with miR-202-5p (10). In the
present study, NORAD was found to function as a ceRNA to inhibit
miR-136-5p. Upregulation of NORAD expression in NSCLC and tissues
was associated with increased lung cancer cell viability and
anaerobic glycolysis. This study provides novel insight on the
possible mechanism of lncRNA NORAD in regulating NSCLC.
Materials and methods
Statement of ethics
Informed consents were obtained from all of the
participating patients, and the study was approved by the Clinical
Research Ethics Committee of Suqian People's Hospital of Nanjing
Drum Tower Hospital Group (Nanjing, China).
Cell culture
The NSCLC cell lines A549, H1975, H1650, LK-2,
H1299, H460 and epithelial cell line HBE were purchased from the
American Type Culture Collection (ATCC; Manassas, VA, USA). A549
cells were cultured in F-12K medium supplemented with 10% fetal
bovine serum (FBS) (all purchased from Gibco/Thermo Fisher
Scientific, Inc., Waltham, MA, USA) at 37°C in 5% CO2.
H1975, H1650, LK-2, H1299, H460 and HBE cell lines were cultured in
RPMI-1640 medium, supplemented with 10% FBS (all purchased from
Gibco/Thermo Fisher Scientific, Inc.) at 37°C in 5%
CO2.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA was extracted from cells using Trizol
reagent (Invitrogen; Thermo Fisher Scientific, Inc.), and
synthesized into cDNA using a reverse transcription kit
(Invitrogen; Thermo Fisher Scientific, Inc.). RT-qPCR was performed
using the 7500 Fast Real-time PCR system (Applied Biosystems;
Thermo Fisher Scientific, Inc) using SYBR-Green PCR kit (Toyobo
Life Science, Osaka, Japan), according to the manufacturer's
protocols. PCR amplification conditions were: 95°C for 5 sec, 60°C
for 30 sec, 72°C for 30 sec for 40 cycles. The results were
normalized to the internal reference gene GAPDH. The primer
sequences used for RT-qPCR assays were as follows: NORAD forward,
5′-TGATAGGATACATCTTGGACATGGA-3′ and reverse,
5′-AACCTAATGAACAAGTCCTGACATACA-3′; GAPDH forward,
5′-GGAGCGAGATCCCTCCAAAAT-3′ and reverse,
5′-GGCTGTTGTCATACTTCTCATGG-3′. For the detection of miRNA
expression, reverse transcription was performed and microRNAs were
detected with stem-loop primers purchased from RiboBio (Guangzhou,
China): miR-136-5p, F: ACTCCATTTGTTTTGATGATGGA. U6 snoRNA was used
as the endogenous control: U6, F: CTCGCTTCGGCAGCACA and R:
ACGCTTCACGAATTTGCGT. Relative fold changes were calculated using
the 2−ΔΔCq method (11). All PCR assays were repeated three
times.
Plasmid construction
NORAD cDNA fragments containing either the predicted
potential microRNA binding sites, wild-type (wt) or scrambled
microRNA binding site sequences, mutation (mut) were amplified by
PCR. The plasmid was constructed by cloning NORAD cDNA into the
pcDNA3.1 vector (Invitrogen; Thermo Fisher Scientific, Inc.).
Mimics and inhibitors of miR-136-5p were purchased from Guangzhou
RiboBio Co., Ltd. (Guangzhou, China).
CCK-8 assay
Cell Counting Kit-8 (CCK-8; Dojindo Molecular
Technologies, Inc., Kumamoto, Japan) was used to detect A549 and
H460 cell proliferation. The cells (1×104 cells/well)
were seeded into 96-well plates at 37°C in 5% CO2, and
transfected with the indicated plasmid. A total of 10 µl CCK-8
solution was subsequently added and incubated was carried out for
another 4 h at 37°C. CCK-8 reagent was added at 0, 24, 48 and 72 h,
according to the manufacturer's protocol. Absorbance rate was
measured at a wavelength of 450 nm using a microplate reader.
Lactate dehydrogenase (LDH) activity,
lactate production, glucose utilization assay and intracellular ATP
level
A total of 1×106 transfected cells were
used for LDH activity and lactate production assay using the
Lactate Dehydrogenase Activity Assay kit and Lactate Assay kit
(Sigma-Aldrich; Merck KGaA, Darmstadt, Germany), according to the
manufacturer's protocols. For glucose utilization assay,
transfected cells were incubated for 24 h and replaced with
phenol-red free RPMI with 1% FBS or phenol-red free RPMI with 1%
FBS and cultured for 3 days. Glucose concentrations in media were
measured using a colorimetric glucose assay kit (BioVision, Inc.,
Milpitas, CA, USA) and normalized to the cell number. Intracellular
ATP was measured using a firefly luciferase-based ATP assay kit
(Beyotime Institute of Biotechnology, Haimen, China). The relative
ATP level is expressed as ATP value/protein value.
Dual-luciferase reporter assay
Recombinant plasmids or an empty plasmid encoding
the firefly luciferase reporter were constructed and transfected
into cells as well as the Renilla luciferase reporter
pRL-CMV plasmid (Promega Corporation, Madison, WI, USA) as an
internal control. miR-136-5p mimic or a negative control were
subsequently transfected into each group. Luciferase activities
were measured using the Dual-Luciferase Reporter Assay System
(Promega Corporation), according to the manufacturer's
protocols.
Western blot analysis
Lysis buffer RIPA was used to harvest cell protein.
Equal amounts of protein (20 µg) were added onto a 10% SDS-PAGE gel
and transferred onto polyvinylidene difluoride membranes. The
membranes were incubated with primary antibodies overnight at 4°C
following blocking with skim milk, and washed with TBST buffer 3
times. They were subsequently incubated with secondary antibody for
2 h at room temperature. Enhanced chemiluminescence reagent (Thermo
Fisher Scientific, Inc.) was added to the membranes and images were
captured using a Tanon 6100 chemiluminescent imaging system (Tanon
Science and Technology Co., Ltd., Shanghai, China). The primary
antibodies were E2F1 (dilution 1:1,000; Cell Signaling Technology,
Inc., Danvers, MA, USA; cat. no. 3742) and β-actin (dilution
1:2,000; Cell Signaling Technology, Inc., cat. no. 4970). The
secondary antibody was horseradish peroxidase-conjugated secondary
antibody (dilution 1:2,000, GTVision III Detection kit, GeneTech,
Shanghai, China; cat. no. GK500710).
RNA immunoprecipitation (RIP)
A549 and H460 cell lysates containing NORAD and
miRNAs were incubated with RIP buffer using EZ-Magna RIP
RNA-binding protein immunoprecipitation kit, (EMD Millipore,
Billerica, MA, USA), according to the manufacturer's protocols,
containing magnetic beads conjugated to human anti-argonaute2
(Ago2) antibody (EMD Millipore). Normal mouse immunoglobulin G
(IgG; EMD Millipore) was used as a negative control. NORAD and
miRNAs presented in the precipitates were assessed by RT-qPCR.
Statistical analysis
All the data are presented as means ± standard
deviation. The Student t-test, χ2 test, log-rank test
and Fisher's exact test was performed for comparisons between each
group. The correlation between NORAD, miR-136-5p and E2F1
expression was analyzed using Pearson Correlation. For comparison
between two groups, we used two-tailed Student's t-test. Multiple
group comparisons were calculated with one-way ANOVA analysis.
Least-significant difference (LSD) was used as a post hoc test.
Kaplan-Meier analysis was used to calculate overall survival.
P<0.05 was used to indicate statistically significant results.
The results are presented as the mean ± standard error of the
mean.
Results
High NORAD expression in NSCLC
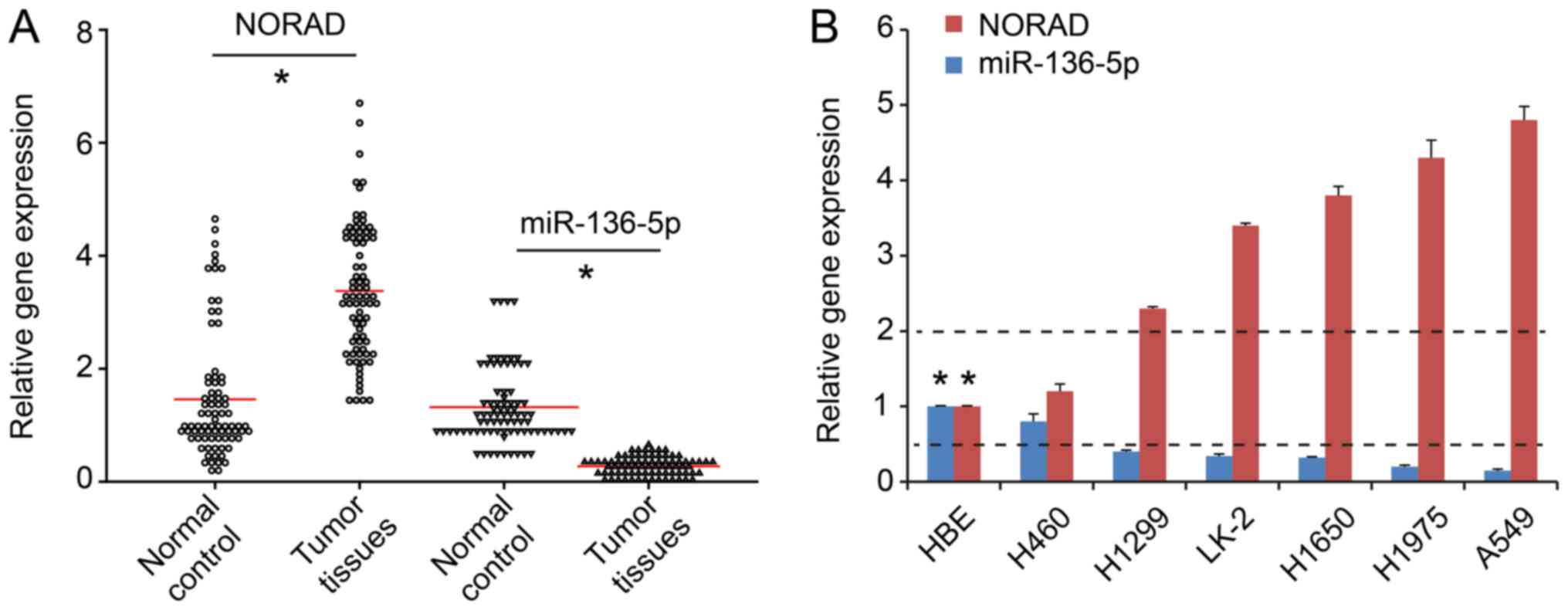
RT-qPCR demonstrated higher NORAD expression in
NSCLC tumor tissues compared with adjacent tissues (n=80;
P<0.05; Fig. 1A). Additionally,
the expression of NORAD in the NSCLC lines was significantly higher
compared with that in normal epithelial HBE cells (P<0.05;
Fig. 1B). To further elucidate the
underlying mechanism of NORAD, A549 and H460 cell lines were
selected for subsequent experiments, as they exhibited the highest
and lowest NORAD expression levels, respectively (Fig. 1B).
Effect of NORAD expression on NSCLC
cell proliferation and glycolysis
qPCR analysis indicated that the expression level of
NORAD was decreased in A549 cells transfected with the NORAD small
interfering RNAs (si1-NORAD and si2-NORAD) (Fig. 2A), and transfection with these
siRNAs inhibited A549 cell proliferation (Fig. 2B). Since aerobic glycolysis is the
primary aspect of altered cancer metabolism, a significant decrease
was indicated in LDH activity, glucose utilization, lactate
production, and an increase in intracellular ATP level in A549
cells following NORAD expression silencing (Fig. 2C).
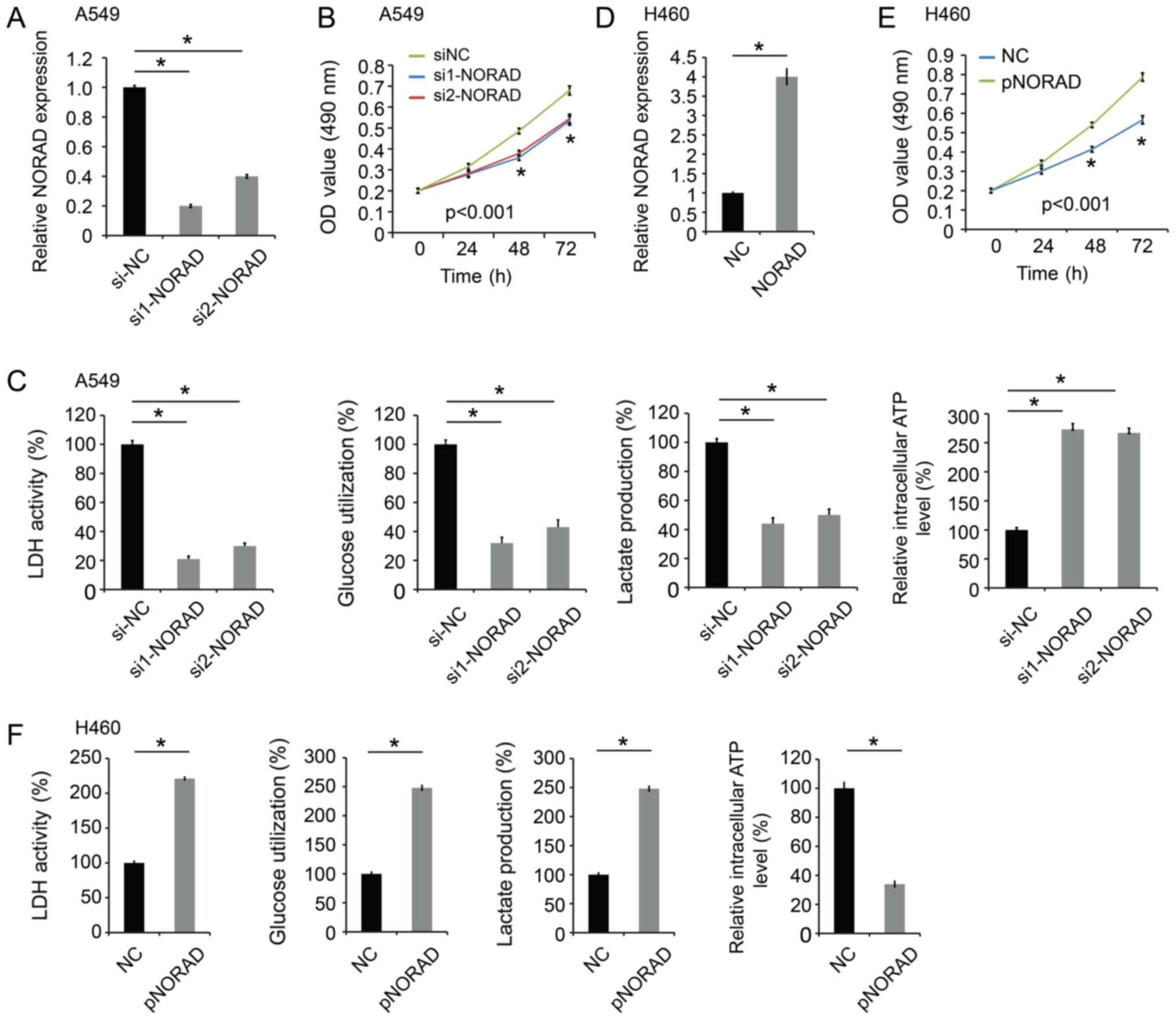 | Figure 2.NORAD promotes cell proliferation and
glycolysis. (A) The level of NORAD was decreased in A549 cells
following transfection with NORAD siRNAs (si1-NORAD and si2-NORAD).
(B) NORAD-knockdown inhibited A549 cell proliferation. (C)
NORAD-knockdown significantly decreased LDH activity, glucose
utilization, lactate production, and increased intracellular ATP
level in A549 cells. (D) The expression level of NORAD was
increased in H460 cells transfected with pcDNA3-NORAD (pNORAD). (E)
NORAD overexpression promoted cell proliferation in H460 cells. (F)
NORAD overexpression increased LDH activity, glucose utilization,
lactate production, and decreased the intracellular ATP level in
H460 cells. *P<0.05 vs. control group. NORAD, non-coding RNA
activated by DNA damage; NC, negative control; siRNA, small
interfering RNA; LDH, lactate dehydrogenase. |
In addition, the function of ectopic expression of
NORAD was examined in NSCLC. The H460 cell line was transfected
with pcDNA3-NORAD (pNORAD) (Fig.
2D). The proliferation of H460 cells with NORAD overexpression
was significantly increased compared with the control groups
(Fig. 2E). Furthermore, NORAD
overexpression increased LDH activity, glucose utilization, lactate
production, and decreased intracellular ATP level in H460 cells
(Fig. 2F). Therefore, these
results indicated that NORAD could promote NSCLC proliferation and
glycolysis.
NORAD functions as a competing
endogenous RNA and sponges miR-136-5p in NSCLC
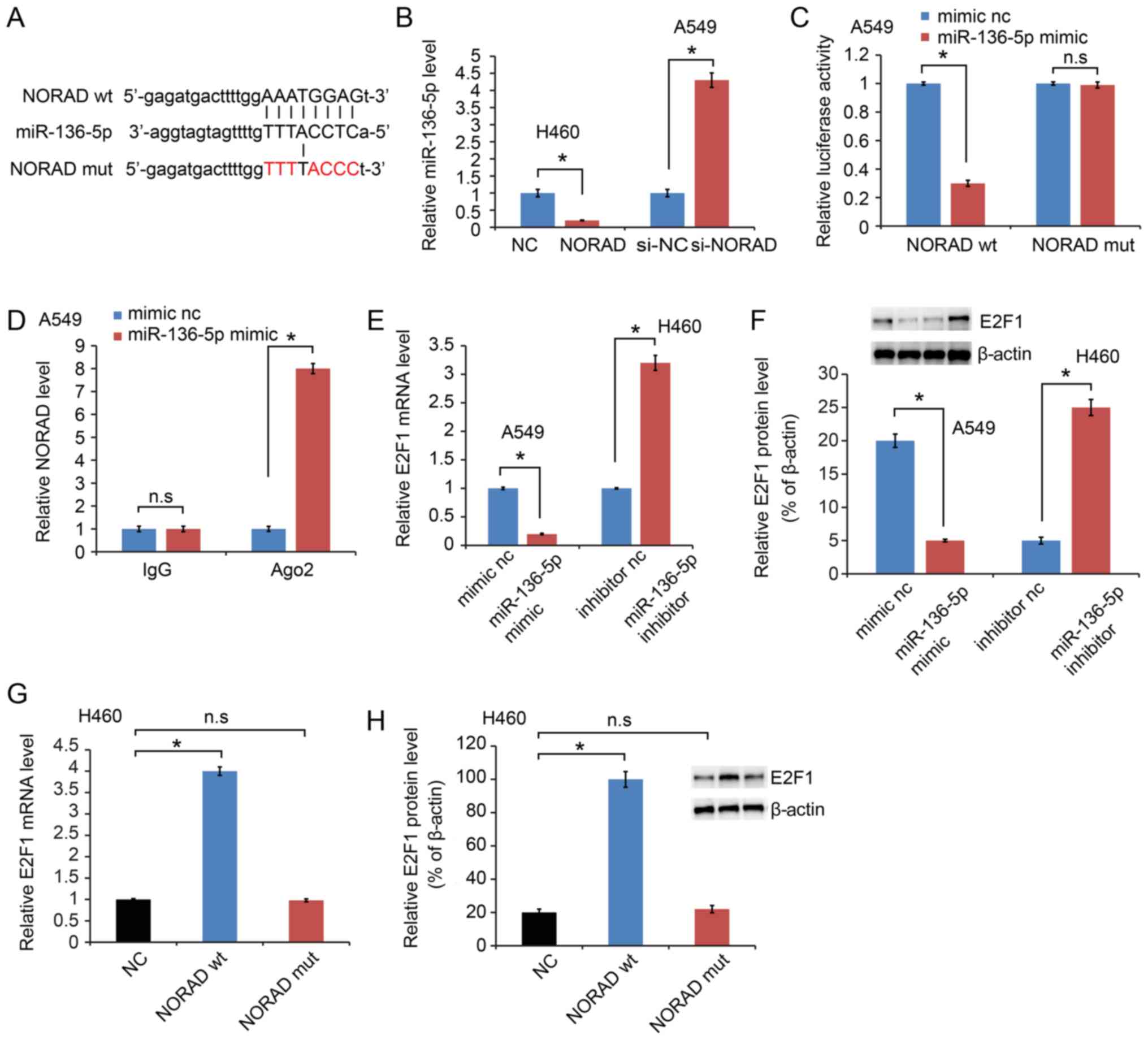
LncRNAs have been reported to sponge miRNAs and
functionally liberate other RNA transcripts (9). A Starbase 2.0 software (http://starbase.sysu.edu.cn/) was used in the present
study to predict the potential miRNA binding sites in NORAD and a
sequence complementary to miRNA-136-5p was identified (Fig. 3A). In addition, miR-136-5p was
upregulated in the si-NORAD-treated cells compared with that noted
in the negative control-treated cells, while miR-136-5p was
downregulated in phosphorylated-NORAD cells (Fig. 3B). A dual-luciferase reporter assay
was performed in A549 cells to test the binding site of miR-136-5p
in NORAD. The activity of luciferase reporters containing the
theoretical binding site of NORAD in miR-136-5p was significantly
decreased in the NORAD wt constructs, while no effect was observed
in the NORAD mut constructs (Fig.
3C). NORAD expression was further examined on magnetic beads
conjugated to the anti-Ago2 antibody and it was indicated that
NORAD was highly enriched in cells transfected with the miR-136-5p
mimic (Fig. 3D). This result
suggested that NORAD and miR-136-5p can combine with Ago2 and form
an increased RNA-induced silencing complex (RISC) and that
miR-136-5p can bind to NORAD in NSCLC cells.
E2F transcription factor 1 (E2F1) is a potential
target of miR-136-5p (12). The
results of the present study also confirmed decreased E2F1
expression following transfection with miR-136-5p mimics, while
upregulated expression was observed with the miR-136-5p inhibitor
at the mRNA (Fig. 3E) and protein
level (Fig. 3F). It was further
indicated that E2F1 was increased by NORAD wt, while NORAD mut
failed to increase E2F1 at the mRNA (Fig. 3G) and protein expression level
(Fig. 3H).
NORAD is a target of miR-136-5p
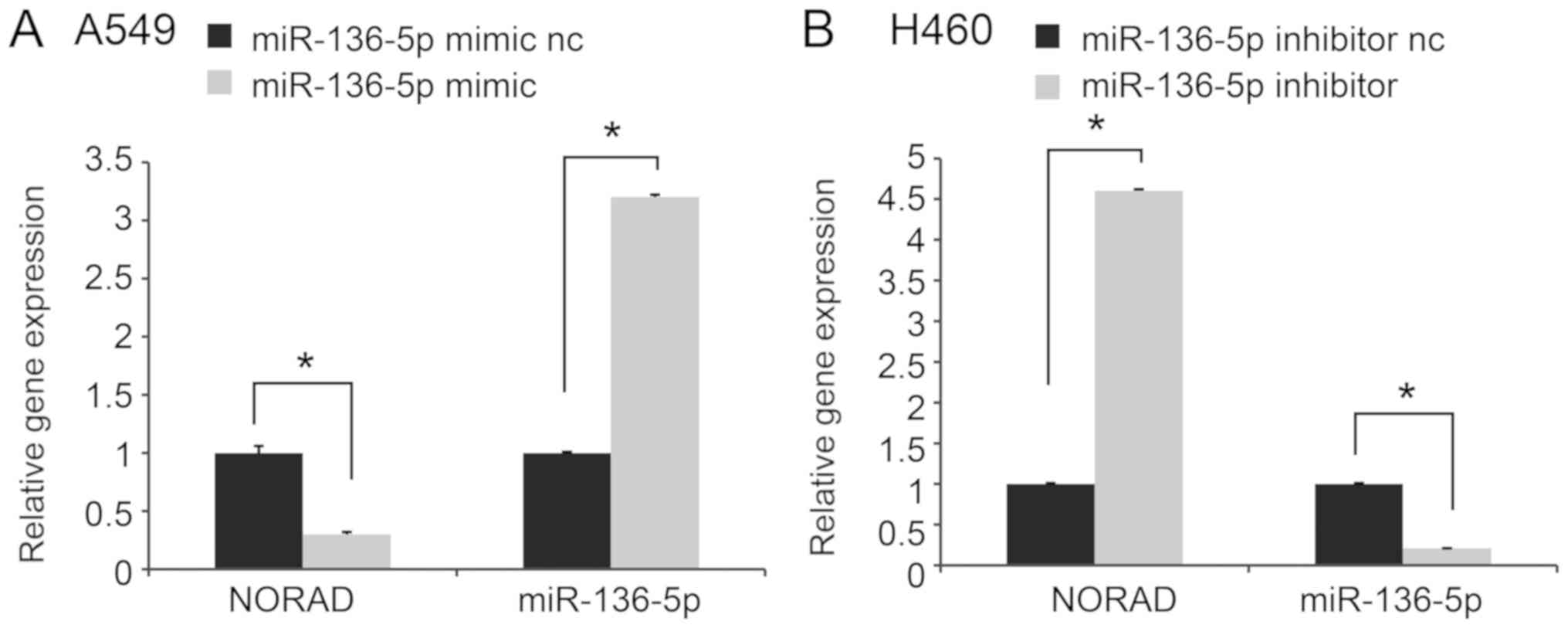
We firstly validated miR-136-5p expression after
transfection with the miR-136-5p mimic or inhibitor (Fig. 4A and B). Then NORAD was detected.
Transfection with miR-136-5p mimics inhibited NORAD expression,
while transfection with inhibitor exhibited an opposite effect
(Fig. 4A and B).
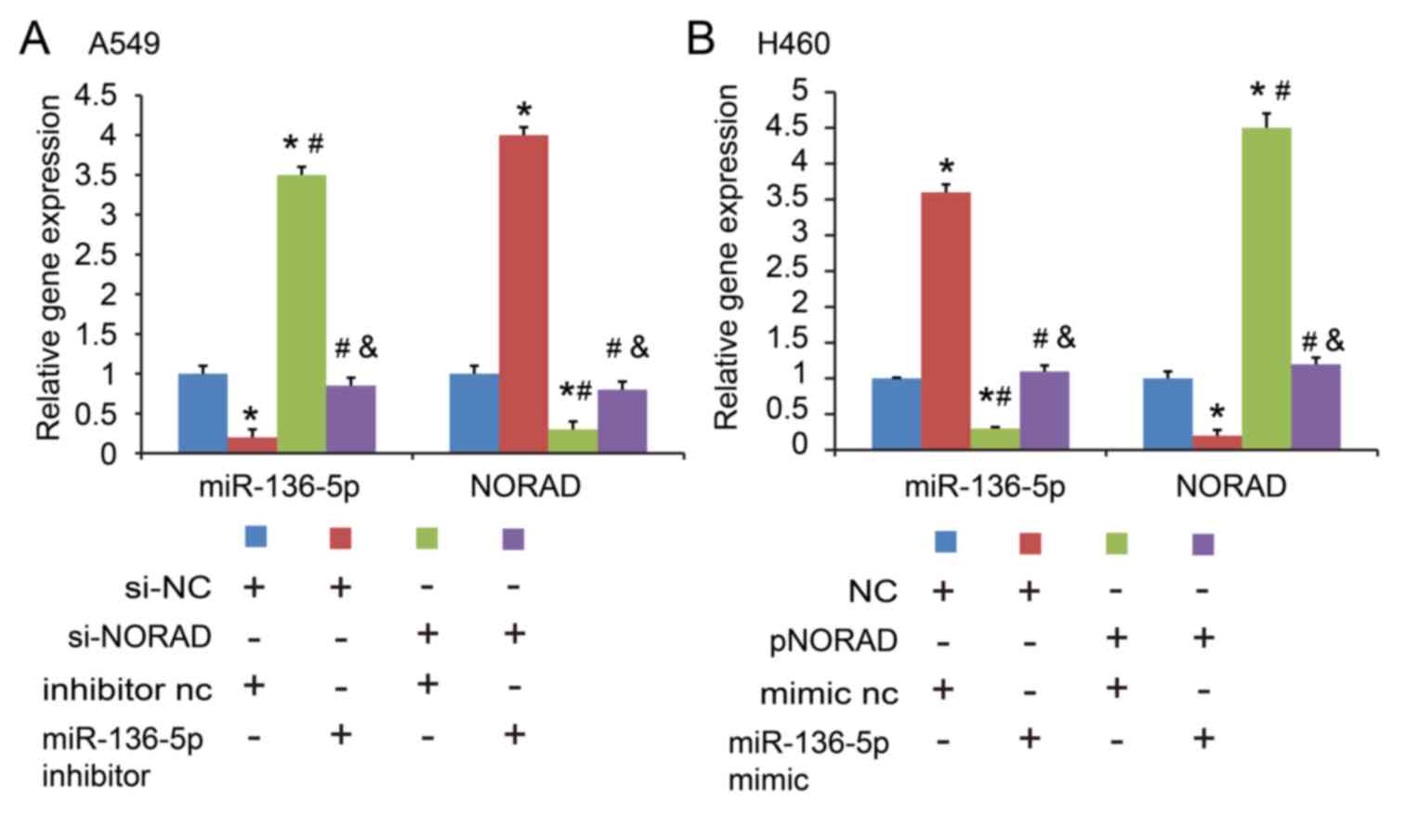
miR-136-5p reverses the promoting
effects of NORAD in NSCLC
To understand the importance of miR-136-5p binding
in NORAD promoting NSCLC progression, the present study
co-transfected A549 cells with NORAD nc/si1 or miR-136-5p
nc/inhibitor, and H460 cells with miR-136-5p mimic. The gene
expression levels are indicated in Fig. 5A and B. It was indicated that
miR-136-5p inhibitor reversed the inhibitory effect of
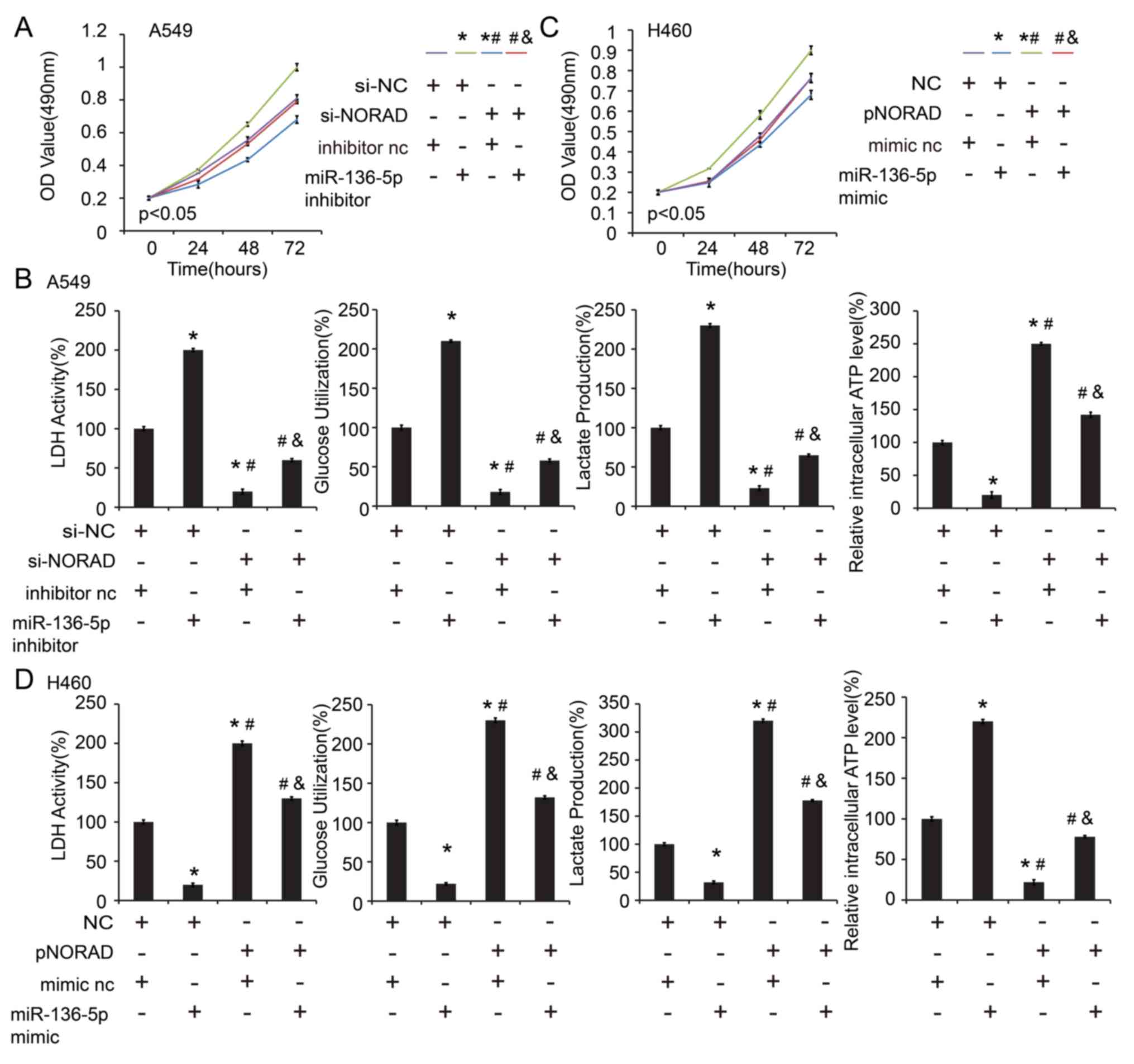
NORAD-knockdown on cell proliferation (Fig. 6A) and glycolysis (Fig. 6B) in the A549 cell line. In
addition, transfection with miR-136-5p mimic abrogated the enhanced
effect of ectopic expression of NORAD on cell proliferation
(Fig. 6C) and glycolysis (Fig. 6D) in H460 cells. E2F1 expression
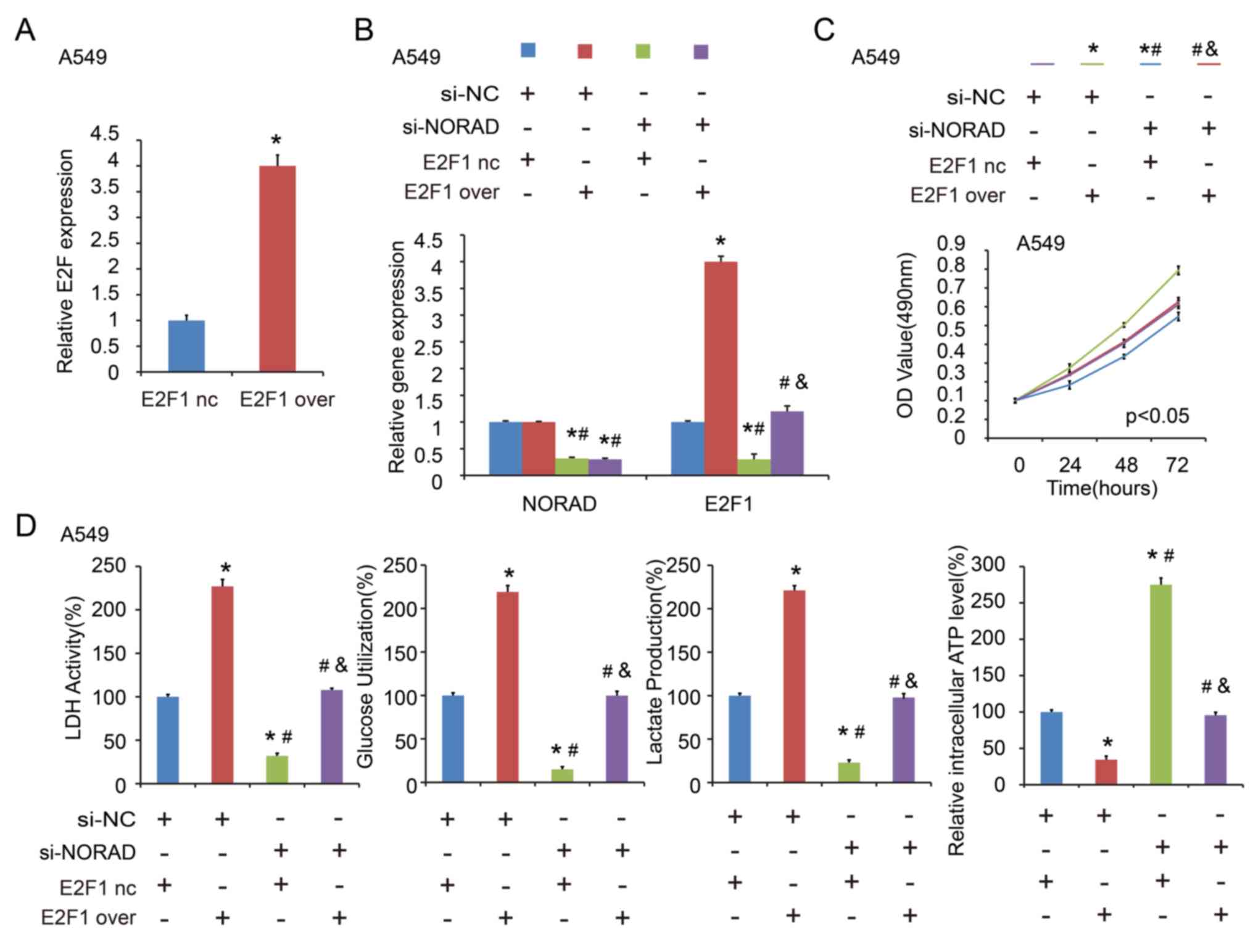
was confirmed by RT-qPCR after transfection of the E2F1
overexpression plasmid (Fig. 7A).
Similarly, in the co-transfected A549 cells with NORAD nc/si1 or
E2F1 nc/over (negative control/overexpression), it was indicated
that E2F1 overexpression abolished the si-NORAD-mediated cell
proliferation and glycolysis (Fig.
7B-D). These findings indicated that NORAD promoted NSCLC
progression in part via competitively binding with miR-136-5p.
Association of NORAD, miR-136-5p and
E2F1 in NSCLC
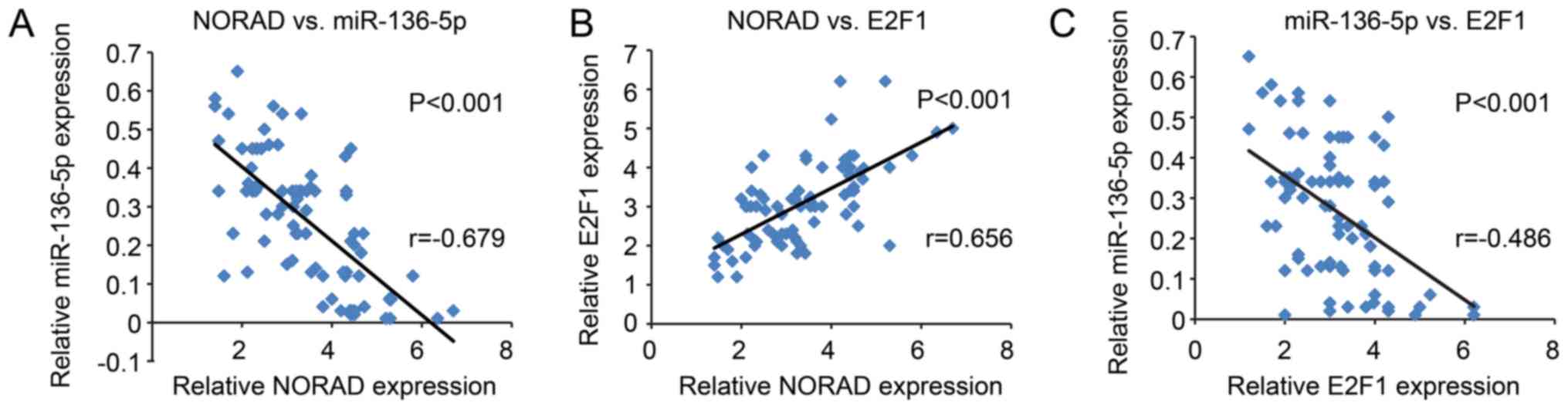
After analyzing their expression and association,
NORAD was found to be inversely correlated with miR-136-5p, while
positively correlated with E2F1 expression (Fig. 8A and B). In addition, miR-136-5p
was inversely correlated with E2F1, all of which were indicated to
be statistically significant (Fig.
8C).
Discussion
Accumulating evidence has revealed that lncRNA
deregulation promotes the growth of human malignancies (13), resulting in tumor progression and
uncontrolled tumor growth, providing insights into novel strategies
for cancer treatment. In lung cancers, numerous lncRNAs have been
investigated, in order to understand the complex underlying
mechanisms, including Hox transcript antisense RNA (HOTAIR)
(14), taurine upregulated 1
(15), tubulin α 4b (TUBA4B)
(16) and metastasis-associated
lung adenocarcinoma transcript 1 (MALAT-1) (17). However, the underlying mechanisms
of lncRNAs in lung cancer remain unclear. The present study
identified an lncRNA, named NORAD, which was increased in lung
cancer tissues, as well as in NSCLC cell lines, demonstrating that
NORAD serves oncogenic functions in NSCLC.
Cancer metabolism, characterized by aerobic
glycolysis or Warburg effect, is characterized by the unusual use
of glucose in the production of lactate even under sufficient
oxygen conditions (18). However,
the functions of lncRNAs in cancer metabolism remain unclear.
Previous research has revealed that lncRNA crystalline β-gamma
domain containing 3 (CRYBG3) interacts with lactate dehydrogenase A
to affect glycolysis (18). In
ovarian cancer, lncRNA small nucleolar RNA host gene 3 (SNHG3)
regulates mitochondrial proteomes, affecting cancer metabolism
(19). In the present study, a
significant decrease in LDH activity, glucose utilization, lactate
production, and an increase in intracellular ATP level, was
observed in lung cancer cells when the NORAD expression level was
modified. NORAD affects glycolysis, leading to a decrease in ATP.
However, we found that ATP was elevated in our data. First, we
speculate that NORAD may affect the Krebs cycle, which in turn
increases ATP. Moreover, a large amount of lactic acid is produced
during the glycolysis of malignant tumors, and is transported into
the liver through blood to provide sufficient raw materials for
gluconeogenesis, resulting in increased conversion of glucose
conversion and utilization of glucose barrier by peripheral
tissues. These procedures consume a large amount of ATP. Therefore,
when lactic acid is reduced, gluconeogenesis is reduced and ATP is
increased. The specific mechanism needs to be further elaborated.
These results indicate that NORAD could regulate metabolism in
NSCLC mainly by affecting glycolysis.
In the past decade, investigation of miRNAs has
dominated the field of non-coding RNA regulation (20,21).
miRNAs have been reported to impact, in various manners, genetic
alteration and signaling pathways (22). It is well known that miRNAs exert
their gene silencing functions through a ribonucleoprotein complex
called the RNA induced silencing complex (RISC) (23). Potential microRNA targets can be
isolated from this complex after Ago2 co-immunoprecipitation as
Ago2 is a vital component of the RISC complex necessary for siRNA
or miRNA-mediated gene silencing (24). In the present study, the data
showed that high miR-136-5p and NORAD both bind to the Ago2
complex. We proposed that miR-136-5p inhibited the level of NORAD
expression in a way similar to miRNA-mediated silencing of
protein-coding genes. Therefore, a high level of miR-136-5p leads
to RISC cleavage, resulting in decreased NORAD expression.
Conversely, low miR-136-5p leads to high NORAD expression. Our
results are consistent with a previous report of functional
interactions among microRNAs and long non-coding RNAs (25–27).
The activity of the dual-luciferase reporter assay
containing the theoretical binding site of NORAD in miR-136-5p was
significantly decreased in NORAD wt constructs, while there was no
effect in NORAD mut constructs. It was also demonstrated that
miR-136-5p mimics arrested lung cancer cell proliferation and
glycolysis, and NORAD reversed this function. These findings
demonstrated that NORAD interacts with miR-136-5p in NSCLC
proliferation. Previous research has verified that E2F1 is a target
of NORAD (28). lncRNAs act as
ceRNA to bind specific miRNAs and regulate their function. NORAD
regulates E2F1 expression directly. We hypothesized that NORAD may
inhibit miR-136-5p expression in such a manner. This suggests that
NORAD may enhance E2F1 by repressing miR-136-5p. These results
suggest a NORAD/miR-136-5p/E2F1 axis that regulates cell
proliferation and glycolysis in NSCLC. A number of limitations in
our experiments must be mentioned, including the insufficient
variety of functional experiments performed (i.e. migration and
invasion assays). Thus, further investigation is warranted.
In conclusion, the present study demonstrated that
the lncRNA NORAD promotes NSCLC cell proliferation and glycolysis
by competitively binding miR-136-5p. NORAD may be valuable as a
targeted therapy. We identified crosstalk between miR-136-5p and
NORAD, shedding novel light on the potential treatment of
NSCLC.
Acknowledgements
Not applicable.
Funding
This study was funded by the Natural Science
Foundation of Nanjing Drum Tower Hospital Group Suqian People's
Hospital (grant no. S201518).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
WG and LFW performed the in vitro studies. TW
made substantial contributions to the design of the present study,
and interpreted the data. BS and YW wrote the manuscript and
interpreted the data. WSM and XYW collected tumor tissues, acquired
data and revised the article critically for important intellectual
content. LJ and LJF participated in designing the study. All
authors read and approved the manuscript and agree to be
accountable for all aspects of the research in ensuring that the
accuracy or integrity of any part of the work are appropriately
investigated and resolved.
Ethics approval and consent to
participate
Informed consents were obtained from all of the
participating patients, and the study was approved by the Clinical
Research Ethics Committee of Suqian People's Hospital of Nanjing
Drum Tower Hospital Group (Nanjing, China).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2018. CA Cancer J Clin. 68:7–30. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Chang JY, Senan S, Paul MA, Mehran RJ,
Louie AV, Balter P, Groen HJ, McRae SE, Widder J, Feng L, et al:
Stereotactic ablative radiotherapy versus lobectomy for operable
stage I non-small-cell lung cancer: A pooled analysis of two
randomised trials. Lancet Oncol. 16:630–637. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Khandelwal A, Bacolla A, Vasquez KM and
Jain A: Long non-coding RNA: A new paradigm for lung cancer. Mol
Carcinog. 54:1235–1251. 2015. View
Article : Google Scholar : PubMed/NCBI
|
|
4
|
Huarte M: The emerging role of lncRNAs in
cancer. Nat Med. 21:1253–1261. 2015. View
Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lee S, Kopp F, Chang TC, Sataluri A, Chen
B, Sivakumar S, Yu H, Xie Y and Mendell JT: Noncoding RNA NORAD
regulates genomic stability by sequestering PUMILIO proteins. Cell.
164:69–80. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Liu H, Li J, Koirala P, Ding X, Chen B,
Wang Y, Wang Z, Wang C, Zhang X and Mo YY: Long non-coding RNAs as
prognostic markers in human breast cancer. Oncotarget.
7:20584–20596. 2016.PubMed/NCBI
|
|
7
|
Li H, Wang X, Wen C, Huo Z, Wang W, Zhan
Q, Cheng D, Chen H, Deng X, Peng C and Shen B: Long noncoding RNA
NORAD, a novel competing endogenous RNA, enhances the
hypoxia-induced epithelial-mesenchymal transition to promote
metastasis in pancreatic cancer. Mol Cancer. 16:1692017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Cesana M, Cacchiarelli D, Legnini I,
Santini T, Sthandier O, Chinappi M, Tramontano A and Bozzoni I: A
long noncoding RNA controls muscle differentiation by functioning
as a competing endogenous RNA. Cell. 147:358–369. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Tay Y, Rinn J and Pandolfi PP: The
multilayered complexity of ceRNA crosstalk and competition. Nature.
505:344–352. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhang J, Li XY, Hu P and Ding YS: LncRNA
NORAD contributes to colorectal cancer progression by inhibition of
miR-202-5p. Oncol Res. 2018. View Article : Google Scholar
|
|
11
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chen W, Yang Y, Chen B, Lu P, Zhan L, Yu
Q, Cao K and Li Q: MiR-136 targets E2F1 to reverse cisplatin
chemosensitivity in glioma cells. J Neurooncol. 120:43–53. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Lalevée S and Feil R: Long noncoding RNAs
in human disease: Emerging mechanisms and therapeutic strategies.
Epigenomics. 7:877–879. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Liu XH, Liu ZL, Sun M, Liu J, Wang ZX and
De W: The long non-coding RNA HOTAIR indicates a poor prognosis and
promotes metastasis in non-small cell lung cancer. BMC Cancer.
13:4642013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhang EB, Yin DD, Sun M, Kong R, Liu XH,
You LH, Han L, Xia R, Wang KM, Yang JS, et al: P53-regulated long
non-coding RNA TUG1 affects cell proliferation in human non-small
cell lung cancer, partly through epigenetically regulating HOXB7
expression. Cell Death Dis. 5:e12432014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chen J, Hu L, Wang J, Zhang F, Chen J, Xu
G, Wang Y and Pan Q: Low expression LncRNA TUBA4B is a poor
predictor of prognosis and regulates cell proliferation in
non-small cell lung cancer. Pathol Oncol Res. 23:265–270. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Han T, Jiao F, Hu H, Yuan C, Wang L, Jin
ZL, Song WF and Wang LW: EZH2 promotes cell migration and invasion
but not alters cell proliferation by suppressing E-cadherin, partly
through association with MALAT-1 in pancreatic cancer. Oncotarget.
7:11194–11207. 2016.PubMed/NCBI
|
|
18
|
Vaitheesvaran B, Xu J, Yee J, Q-Y L, Go
VL, Xiao GG and Lee WN: The Warburg effect: A balance of flux
analysis. Metabolomics. 11:787–796. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Li N and Zhan X and Zhan X: The lncRNA
SNHG3 regulates energy metabolism of ovarian cancer by an analysis
of mitochondrial proteomes. Gynecol Oncol. 150:343–354. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Mirzaei H, Masoudifar A, Sahebkar A, Zare
N, Sadri Nahand J, Rashidi B, Mehrabian E, Mohammadi M, Mirzaei HR
and Jaafari MR: MicroRNA: A novel target of curcumin in cancer
therapy. J Cell Physiol. 233:3004–3015. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kumar R and Xi Y: MicroRNA, epigenetic
machinery and lung cancer. Thorac Cancer. 2:35–44. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chen L and Kang C: miRNA interventions
serve as ‘magic bullets’ in the reversal of glioblastoma hallmarks.
Oncotarget. 6:38628–38642. 2015.PubMed/NCBI
|
|
23
|
Gregory RI, Chendrimada TP, Cooch N and
Shiekhattar R: Human RISC couples microRNA biogenesis and
posttranscriptional gene silencing. Cell. 123:631–640. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Karginov FV, Conaco C, Xuan Z, Schmidt BH,
Parker JS, Mandel G and Hannon GJ: A biochemical approach to
identifying microRNA targets. Proc Natl Acad Sci USA.
104:19291–19296. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Jiang H, Huang G, Zhao N, Zhang T, Jiang
M, He Y, Zhou X and Jiang X: Long non-coding RNA TPT1-AS1 promotes
cell growth and metastasis in cervical cancer via acting AS a
sponge for miR-324-5p. J Exp Clin Cancer Res. 37:1692018.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhuo M, Yuan C, Han T, Cui J, Jiao F and
Wang L: A novel feedback loop between high MALAT-1 and low
miR-200c-3p promotes cell migration and invasion in pancreatic
ductal adenocarcinoma and is predictive of poor prognosis. BMC
Cancer. 18:10322018. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yoon JH, Abdelmohsen K and Gorospe M:
Functional interactions among microRNAs and long noncoding RNAs.
Semin Cell Dev Biol. 34:9–14. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Lu HJ, Jin PY, Tang Y, Fan SH, Zhang ZF,
Wang F, Wu DM, Lu J and Zheng YL: microRNA-136 inhibits
proliferation and promotes apoptosis and radiosensitivity of
cervical carcinoma through the NF-κB pathway by targeting E2F1.
Life Sci. 199:167–178. 2018. View Article : Google Scholar : PubMed/NCBI
|