Introduction
Glioma is the most common malignant neoplasm
worldwide. Even with an increase in the risk and incidence of
glioma, it is estimated that ~60% of subjects over the age of 50 in
the USA are not screened regularly (1). In addition, a higher frequency of
astrocytoma among African-Americans (40–49 years of age) as
compared to Hispanic-Americans has been reported suggesting an
increased susceptibility to glioma risk (1,2).
Current treatment is based on surgical excision after combined
chemotherapy and radiation therapy, yet the prognosis for glioma
patients has not fundamentally improved. The development of glioma
commonly consists of simultaneous dysregulation of cell growth,
differentiation and migration functions; cell malignant
transformation; and metastasis. However, the molecular mechanisms
underlying the development of glioma are not completely
understood.
Immune responsive gene 1 (IRG1) was originally
identified as a 2.3-kb cDNA from a library synthesized from the
mRNA isolated from a macrophage cell line after LPS stimulation.
Based on sequence homology, IRG1 has been classified as a member of
the MmgE/PrpD family (3). IRG1 was
shown to play important roles in embryonic implantation and
neurodegeneration (4), and to
mediate the production of itaconic acid, which contributes to the
activity of macrophages during inflammation (5). Recent studies have found that chronic
inflammation is implicated in tumorigenesis and cancer progression
and is considered as one of the independent risk factors. IRG1
knockdown by siRNA was found to result in significantly increased
production of pro-inflammatory cytokines in LPS-tolerized
macrophages and the activation of transcription factors NF-κB and
interferon regulatory factor 3 (IRF3), which was accompanied by the
increased generation of reactive oxygen species (ROS) and
downregulation of zinc finger protein A20 (6,7). IRG1
expression and subcellular localization in murine macrophages
demonstrated a pattern typical of a TNF- and IFN-γ-co-regulated
gene. In murine ANA-1 macrophages, IRG1 expression was shown to be
induced by several pro-inflammatory cytokines and Toll-like
receptor (TLR) agonists (8). TLRs
are upregulated in many tumor cell lines and tissues, especially in
the tumors of epithelial origin, such as ovarian and breast cancers
(9). Recent studies show that the
expression of TLRs on malignant and immune cells can promote
inflammation and cell survival in the tumor microenvironment. A
variety of endogenous TLR ligands can activate TLR signaling
pathways, inducing nuclear export of tumorigenic transcription
factors and secretion of pro-inflammatory cytokines such as
interleukin-6 (IL-6), chemokine ligand 2 (CCL-2), and vascular
endothelial growth factor (VEGF), thereby promoting tumor
development, angiogenesis, invasion, metastasis and evasion of
immune surveillance (10,11).
In the present study, we investigated the role of
IRG1 in the pathogenesis of glioma and examined the association of
IRG1 expression with glioma clinicopathologic characteristics,
including survival of cancer patients. We identified IRG1-induced
effects on cell growth, invasion, and in vivo tumorigenesis,
confirmed increased IRG1 expression in primary glioma tissues and
cell lines, and found a direct correlation between IRG1 levels and
the clinical outcome of glioma patients.
Materials and methods
Cell lines and siRNA transfection
Human glioma cell lines (A172, U251, U87, SHG-44,
TJ-105, H4 and U118MG) were obtained from the the Neurosurgery
Institute of Guangdong Province (Guangzhou, China). The U-373MG
cell line was purchased from the American Type Culture Collection.
Both the U251 and U87 cell lines were kind gifts from Professor
Xin-Yuan Guan (Sun Yat-Sen University Cancer Center, Guangzhou,
China). All cell lines were cultured in Dulbecco’s modified Eagle’s
medium with 10% fetal bovine serum (FBS) and incubated in a
humidified atmosphere of 5% CO2 and 95% air at 37°C.
Transfection of siRNA was performed by RNAi-Max (Invitrogen Life
Technologies, Carlsbad, CA, USA) according to the manufacturer’s
protocol. The siRNA sequences are provided upon request. In brief,
transfection with the final concentration of 50 nM siRNA was
conducted when the cell density was ~40% in the 6-well plates.
In vivo animal models
This study was carried out in strict accordance with
the recommendations in the Guide for the Care and Use of Laboratory
Animals of the National Institutes of Health. The protocol was
approved by the Committee on the Ethics of Animal Experiments of
the Southern Medical University. All surgery was performed under
sodium pentobarbital anesthesia, and all efforts were made to
minimize suffering.
Acquisition of clinical specimens
Clinical specimens were obtained from archived
tissue samples derived from patients with glioma who underwent
surgical treatment at The First Affiliated Hospital of Zhengzhou
University, China from January, 2003 through December, 2013. Glioma
was diagnosed according to the 2007 WHO Classification of Tumors of
Glioblastoma. All tumors were staged on the basis of the pathologic
Tumor-Node-Metastasis classification of the International Union
Against Cancer (10,11). The selection criteria were as
follows: i) the subject presented with a diagnosis of glioma and no
history of other tumors; ii) the subject had complete demographic
and clinical data, such as age, gender, clinical manifestations,
tumor size, extent of resection, adjuvant therapy and date of
relapse and/or death; iii) the subject underwent evaluation by
enhanced head MRI scanning for tumor relapse or progression after
surgery at least once every six months. Written informed consent of
the patients was provided by their legal surrogates to permit
surgical procedures and use of resected tissues. This study was
approved by the Specialty Committee on Ethics of Biomedicine
Research, Zhengzhou University of China. Human tissue acquisition
and use in this study complied with the National Regulations on the
Use of Clinical Samples in China.
Collection of clinical information and
follow-up
Data were collected by review of the clinical
history. Information was recorded including the patient
characteristics, relevant symptoms or history, tumor
characteristics, extent of resection, post-surgical treatment
protocol, overall survival time and progression-free survival time.
The follow-up was conducted by telephone or direct correspondence.
Postsurgical treatment, including adjuvant radiotherapy and
chemotherapy, was fully discussed with the patient or their
relatives. The time of tumor relapse or death was verified by the
patient or their relatives, by medical recording, or by the social
security record. Overall survival (OS) was calculated in months
from the date of diagnosis to the time of death, regardless of
cause. Progression-free survival (PFS) was defined as the period
from the initial date of diagnosis to the time of tumor progression
by MRI, or to the time of death of the patient from glioma.
Immunohistochemistry
Paraffin sections (4 μm) from samples were
deparaffinized in 100% xylene and re-hydrated in descending ethanol
series and water according to standard protocols. Heat-induced
antigen retrieval was performed in 10 mM citrate buffer for 2 min
at 100°C. Endogenous peroxidase activity and non-specific antigen
were blocked with peroxidase blocking reagent containing 3%
hydrogen peroxide and serum, followed by incubation with goat
anti-human IRG1 antibody (1:100) (Santa Cruz Biotechnology, Santa
Cruz, CA, USA) overnight at 4°C. After washing, the sections were
incubated with biotin-labeled rabbit anti-goat antibody for 10 min
at room temperature, and subsequently were incubated with
streptavidin-conjugated horseradish peroxidase (HRP) (Maixin Inc.,
China). The peroxidase reaction was developed using
3,3-diaminobenzidine chromogen solution in DAB buffer substrate.
Sections were visualized with DAB and counterstained with
hematoxylin, mounted in neutral gum, and analyzed using a bright
field microscope.
Western blot analysis
Cells were lysed in RIPA buffer (50 mM Tris-HCl pH
8.0, 1 mM EDTA pH 8.0, 5 mM DTT, 2% SDS), and the protein
concentration was determined using the BCA assay (Beyotime Inc.,
China). Total protein (30 μg) was resolved using a 10% SDS-PAGE gel
and electro-transferred to polyvinylidene fluoride membranes
(Invitrogen), and blocked with 5% nonfat dry milk in Tris-buffered
saline, pH 7.5. The membranes were immunoblotted overnight at 4°C
with the following antibodies: rabbit polyclonal anti-IRG1 (1:500),
anti-E2F1 (1:500), snail (1:1,000), CDK6 (1:1,000), CDK4 (1:1,000),
vimentin (1:1,000), E-cadherin and N-cadherin (Santa Cruz
Biotechnology). An HRP-conjugated anti-rabbit IgG antibody was used
as the secondary antibody (Zhongshan Inc., China).
Colony formation assay
Approximately 100 cells were added to each well of a
6-well culture plate, and each cell group contained 2 wells. After
2 weeks of incubation, cells were washed twice with PBS and stained
with Giemsa solution. The number of colonies containing ≥50 cells
was counted under a microscope. The colony formation efficiency was
calculated as: Efficiency = (number of colonies/number of cells
inoculated) × 100%. Each experiment was performed in
triplicates.
Cell cycle distribution
To evaluate cell cycle distribution, cells were
seeded on 10 cm-diameter plates in RPMI-1640 culture medium
containing 10% NBCS. After 48 h of incubation, a total of
1×106 cells were harvested, rinsed with cold PBS, and
fixed with 70% ice-cold ethanol for 48 h at 4°C. Fixed cells were
rinsed with cold PBS followed by incubation with PBS containing 10
μg/ml propidium iodide and 0.5 mg/ml RNase A for 15 min at 37°C.
The DNA content of the labeled cells was acquired using a
FACSCaliber cytometer (BD Biosciences). Each experiment was
performed in triplicates.
In vitro cell migration assay
Cells growing in the log phase were treated with
trypsin and re-suspended as single-cell solution. A total of
1×105 cells were seeded on a fibronectin-coated
polycarbonate membrane insert in a Transwell apparatus (Corning
Inc., Corning, NY, USA). In the lower chamber, 600 μl of RPMI-1640
with 10% NBCS was added as a chemoattractant. After the cells were
incubated for 12 h, the insert was washed with PBS, and cells on
the top surface of the insert were removed by a cotton swab. Cells
adhering to the lower surface were fixed with methanol, stained
with Giemsa, and counted under a microscope in five predetermined
fields (x200). All assays were independently repeated at least
three times.
Establishment of U251 and SHG-44 cells
with stable expression of IRG1 short hairpin RNA
The preparation of the lentivirus expressing human
IRG1 short hairpin RNA was performed using the pLVTHM-GFP
lentiviral RNAi expression system (12). U251 and SHG-44 cells were infected
with lentiviral particles containing specific or negative control
vectors, and the polyclonal cells with GFP signals were selected
for further experiments using FACS flow cytometry.
Statistical analysis
All data were analyzed for statistical significance
using SPSS 13.0 software. The Mann-Whitney U test was applied to
the examination of relationship between IRG1 expression levels and
clinicopathological characteristics. Survival analysis was
performed using Kaplan-Meier method. Multivariate Cox proportional
hazards method was used for analyzing the relationship between the
variables and patient survival time. One-way ANOVA was used to
determine the differences between groups for all in vitro
analyses. A P-value of <0.05 was considered to indicate a
statistically significant result.
Results
IRG1 is upregulated in human glioma cell
lines and clinical specimens
To investigate whether IRG1 plays an important role
in gliomas, we examined IRG1 expression in several glioma cell
lines A172, U251, U87, SHG-44, TJ-105, H4, U118MG and U-373MG by
measuring mRNA and protein levels using real-time RT-PCR and
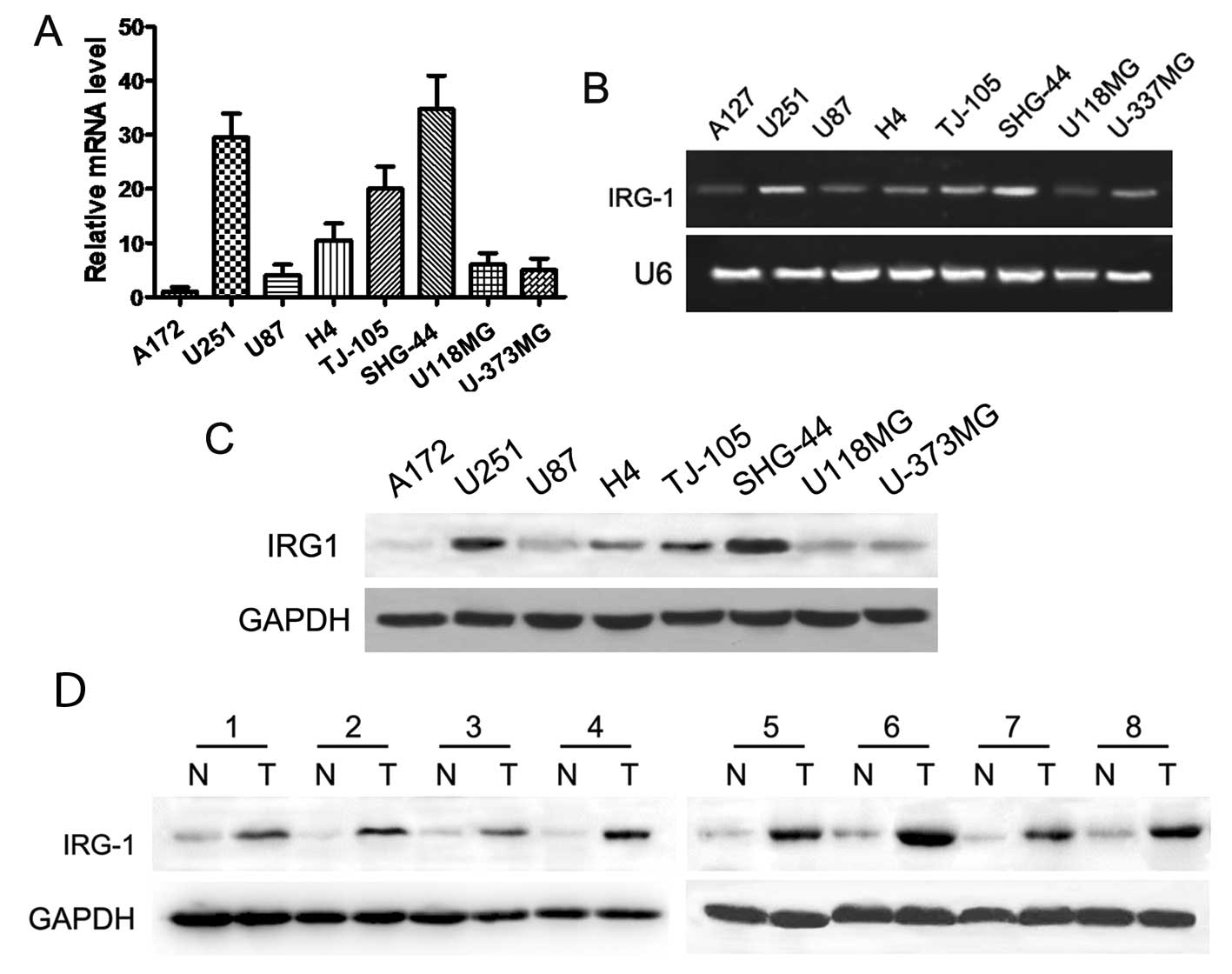
western blotting. Respectively, as shown in Fig. 1A–C the mRNA and protein levels of
IRG1 were evidently higher in the U251, SHG-44, TJ-105 and H4
glioma cell lines than the levels in the A251, U118MG and U-373MG
cell lines. More importantly, using fresh glioma specimens, we
demonstrated that the IRG1 protein level was upregulated in all 8
glioma samples compared to the matched adjacent non-tumor tissues
(Fig. 1D).
Cytoplasmic IRG1 protein is a prognostic
factor for post-surgical glioma patients
The expression levels and subcellular localization
of IRG1 protein were examined in 78 archived paraffin-embedded
glioma samples and 16 normal epithelium tissues by
immunohistochemical staining. The relationship between the
clinicopathologic characteristics and IRG1 expression in the glioma
patients who underwent surgical resection is presented in Table I. IRG1 was highly expressed in 75.6%
(59/78) of the glioma samples compared to only 37.5% (6/16) of the
normal epithelium samples, demonstrating a statistically
significant difference (P=0.006) (Table II). We also observed that IRG1
expression was positively correlated with the WHO grade I, II, III
and IV (P=0.012) (Table I).
Previous results suggest that IRG1 is highly expressed in mammalian
macrophages during inflammation, and IRG1 links cellular metabolism
with immune defense by catalyzing itaconic acid production
(10). Yet, there is no published
information as to the expression of IRG1 in relation to cancer
prognosis. We, therefore, first stained for IRG1 in glioma tissues
in 140 patients. In addition, high (defined as greater than median)
IRG1 expression in glioma cells was significantly associated with
shorter overall survival (OS) in the entire cohort (P<0.05).
 | Table ICorrelation between the
clinicopathological characteristics and IRG1 expression in 140
glioma patients. |
Table I
Correlation between the
clinicopathological characteristics and IRG1 expression in 140
glioma patients.
| | IRG1 protein | |
|---|
| |
| |
|---|
| Factor | No. | Low, n (%) | High, n (%) | P-value |
|---|
| Age (years) |
| <60 | 83 | 47 (56.6) | 36 (43.4) | |
| ≥60 | 57 | 24 (42.1) | 33 (57.9) | 0.091 |
| Gender |
| Male | 68 | 35 (51.5) | 33 (48.5) | |
| Female | 72 | 36 (50.0) | 36 (50.0) | 0.862 |
| Tumor size (cm) |
| <5 | 49 | 28 (57.1) | 21 (42.9) | |
| ≥5 | 91 | 43 (47.3) | 48 (52.7) | 0.264 |
| WHO grade |
| I | 16 | 5 (31.2) | 11 (68.8) | |
| II | 50 | 20 (40.0) | 30 (60.0) | |
| III | 34 | 28 (70.0) | 12 (30.0) | |
| IV | 40 | 18 (45.0) | 22 (55.0) | 0.012a |
 | Table IIUpregulation of IRG1 protein
expressin in glioma compared to epithelium tissues. |
Table II
Upregulation of IRG1 protein
expressin in glioma compared to epithelium tissues.
| | IRG1 protein
expression | |
|---|
| |
| |
|---|
| Group | Total | Low, n (%) | High, n (%) | P-value |
|---|
| Normal
epithelium | 16 | 10 (62.5) | 6 (37.5) | 0.006 |
| Glioma | 78 | 19 (24.4) | 59 (75.6) | |
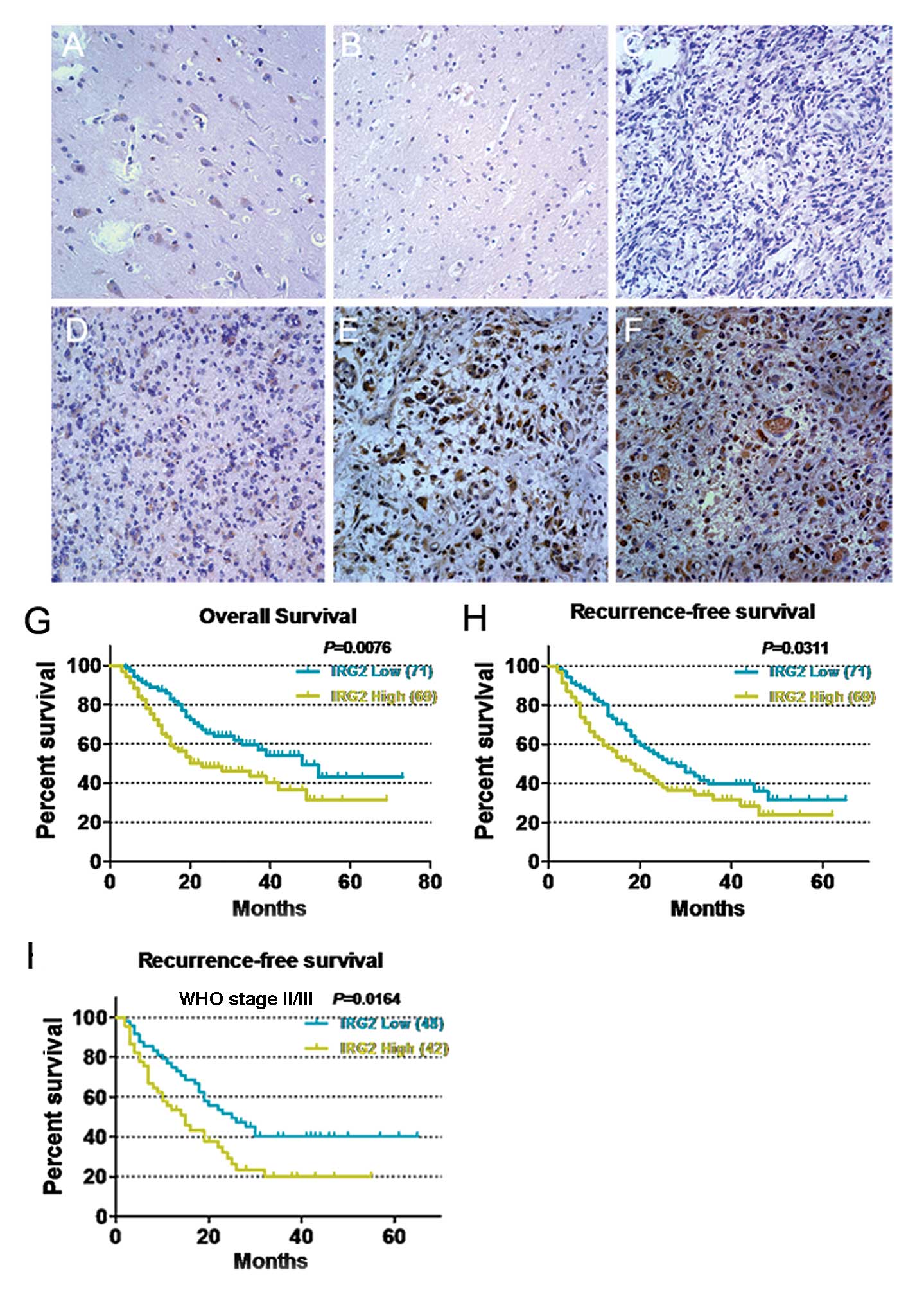
The staining signal of IRG1 was observed mainly in
the different stages of glioma tissues (Fig. 2C–F), and no signals or only weak
signals were detected in the adjacent normal brain tissues
(Fig. 2A and B). The subcellular
location of IRG1 was observed mainly in the cytoplasm of the cancer
cells.
The prognostic effect of IRG1 on glioma patient OS
was compared between patients with high and low IRG1 protein
levels. By Kaplan-Meier curve assessment, patients with a high IRG1
protein level had a significantly lower 5-year survival rate than
those with a low IRG1 protein level (P=0.0076; Fig. 2G).
Furthermore, we compared the relationship between
IRG1 expression and recurrence-free survival (RFS). The patients
with IRG1 low expression (n=69) had a long RFS compared with
patients with high IRG1 expression (n=71, P=0.0311, Fig. 2H). In the patients with WHO grade II
and III, the subgroup with high IRG1 (n=42) expression had a lower
RFS rate than the subgroup with low IRG1 expression (n=48,
P=0.0164, Fig. 2I).
IRG1-induced proliferation of glioma
cells
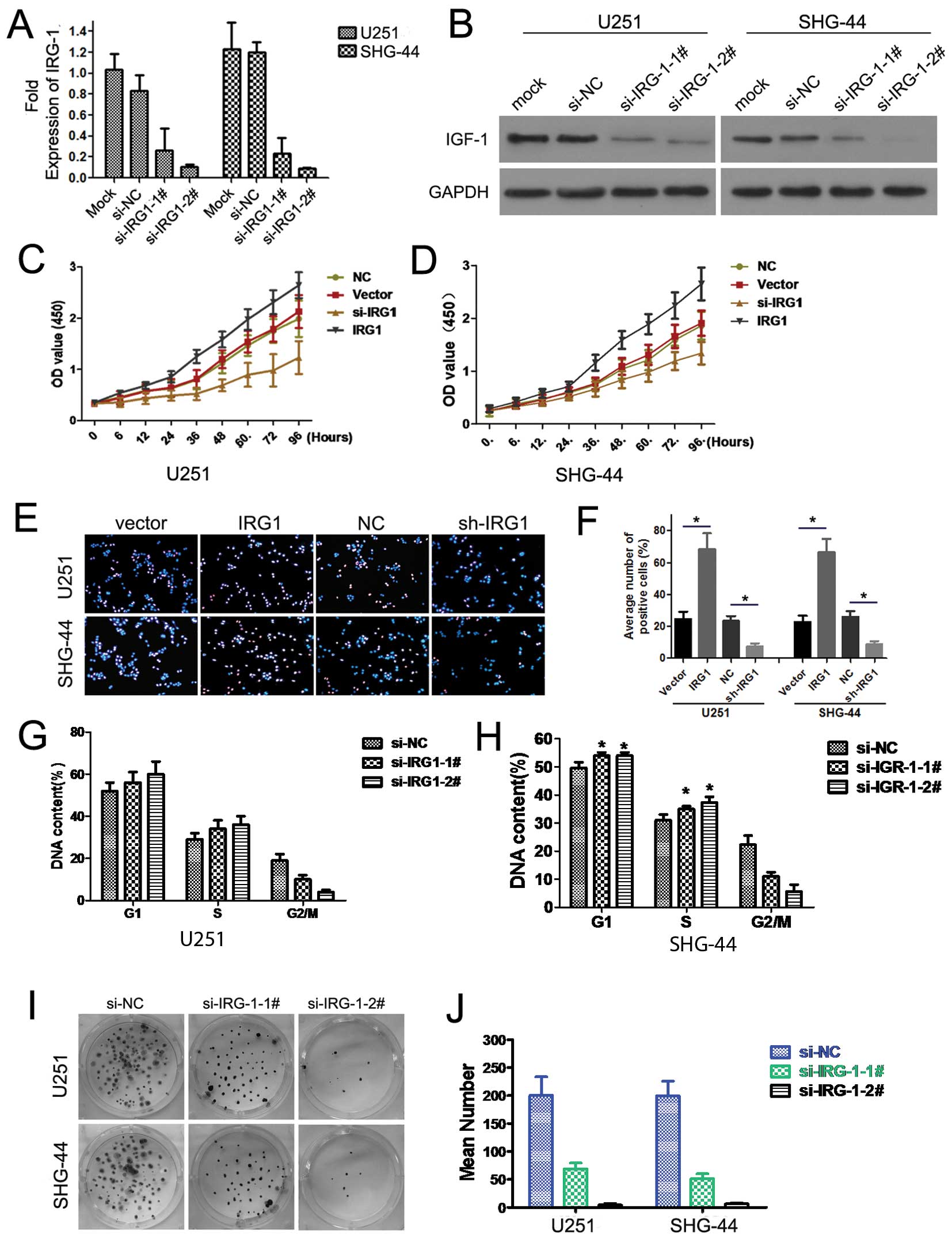
To explore the effect of IRG1 on cancer cell growth,
we used IRG1-specific siRNAs to inhibit IRG1 expression in the
cancer cells. Both si-IRG1-1 and si-IRG1-2 downregulated IRG1
expression, but the inhibitory effect of si-IRG1-2 was more
significant (Fig. 3A and B).
Therefore, we used si-IRG1-2 in the further experiments. U251 and
SHG-44 cells were transiently transfected with IRG1 and si-IRG1-2.
The results of the MTT assays (Fig. 3C
and D) revealed that the downregulation of IRG1 expression
inhibited SHG-44 cell proliferation by 43% (P<0.01), whereas
IRG1 overexpression promoted SHG-44 cell growth by 54% (P<0.01).
The EDU assay showed that overexpression of IRG1 increased the
percentage of proliferative cells from ~23 and ~25% to ~64 and ~62%
in the U251 and SHG-44 glioma cell lines; but inhibition of IRG1
using IRG1 siRNA decreased the percentage of proliferative cells in
the glioma cells (from ~23 and ~25% to ~6 and ~9% in the U251 and
SHG-44 cell lines) (Fig. 3E and F).
Next, we examined the cell cycle distribution in the population of
si-IRG1-2-transfected cells. Compared to the si-NC control, the
U251 and SHG-44 glioma cell lines transfected with si-IRG1-2
displayed an increased proportion of cells entering the G1 phase
and fewer cells in the S phase (Fig. 3G
and H; P<0.01). These results suggest that the
growth-suppressive effect of si-IRG1 was partly due to G1 phase
arrest. As shown by the colony formation assay, the
si-IRG1-transfected U251 and SHG-44 cells formed fewer and smaller
sized colonies than did the si-NC-transfected cells (Fig. 3I and J; P<0.01).
IRG1 knockdown reduces cell migration and
invasion
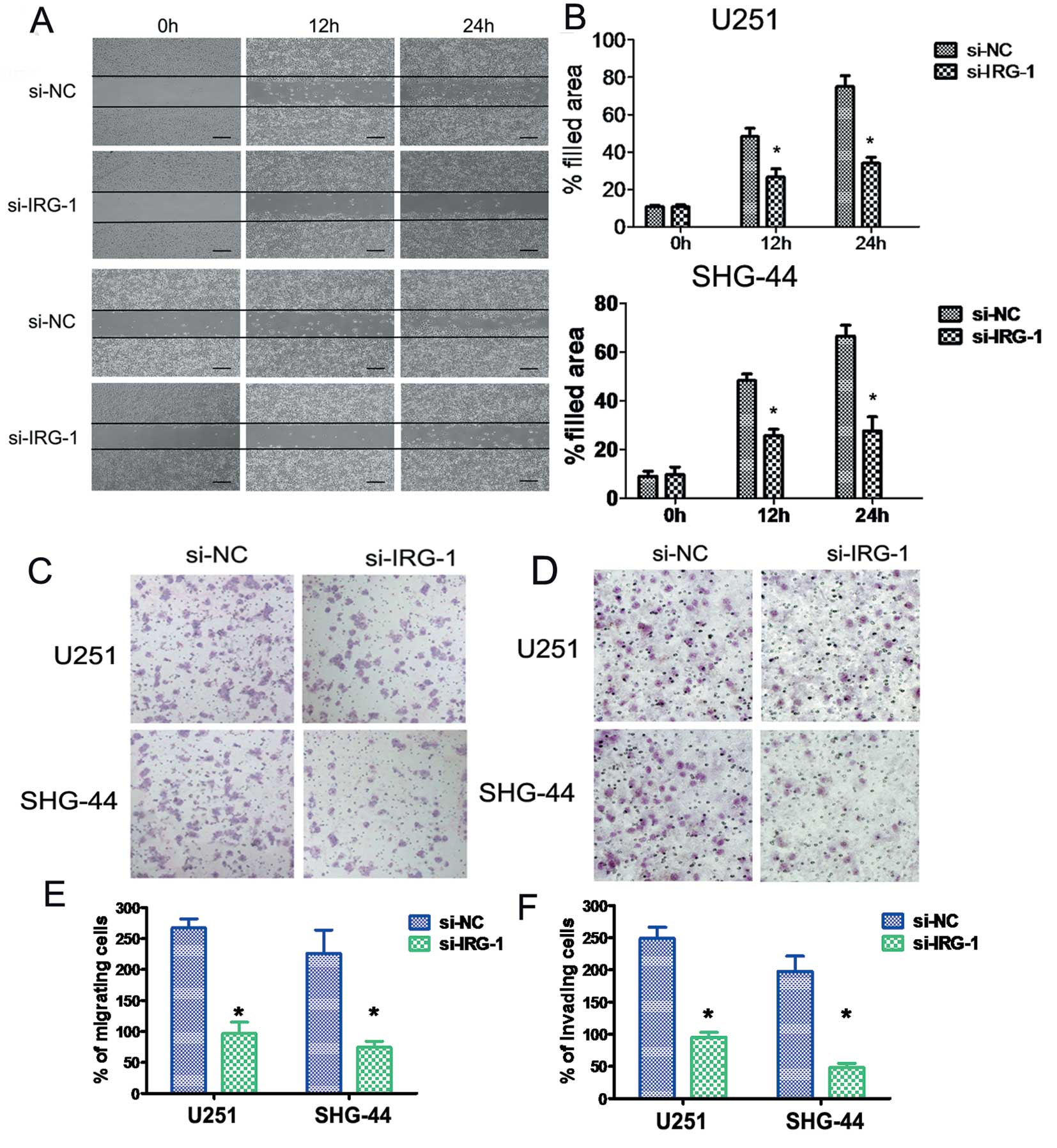
Cell migration and invasion are integral steps in
tumor development and metastasis. The wound healing assay showed
that the lateral migration of cancer cells was inhibited by IRG1
knockdown (Fig. 4A and B). When we
tested U251 and SHG-44 cell migration/invasion through an 8-μm-pore
polycarbonate membrane with or without Matrigel pre-coating, we
found that the knockdown of endogenous IRG1 significantly reduced
the ability of U251 and SHG-44 cells to migrate and invade
(P<0.05), compared to the si-NC cells (Fig. 4C–F).
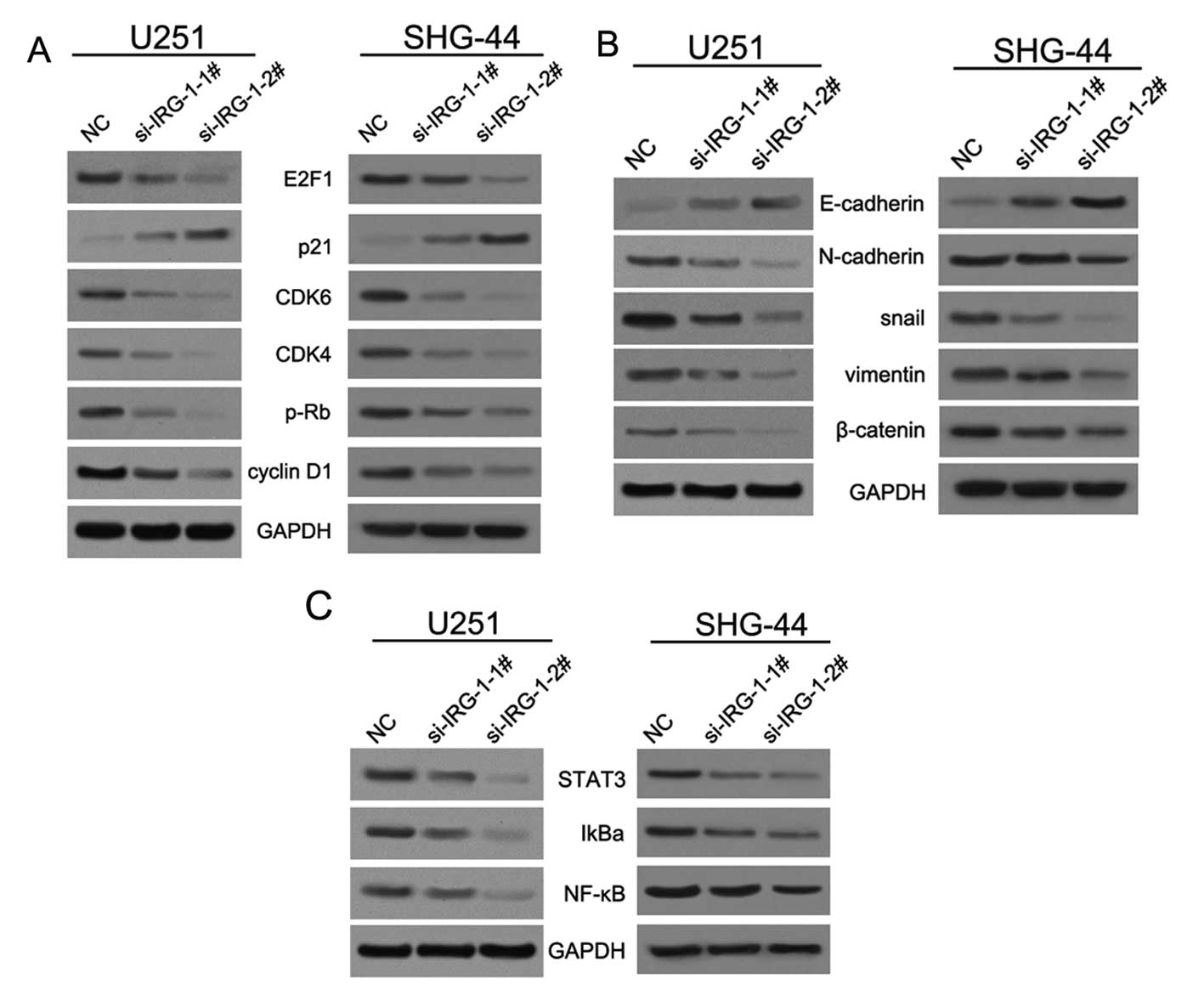
IRG1 controls the expression of genes
associated with cell cycle and epithelial-mesenchymal transition in
glioma cells
To further investigate the mechanism of IRG1
regulation of cell proliferation, migration and invasion, we
firstly examined the protein expression of the genes associated
with cell cycle and epithelial-mesenchymal transition (EMT) in
glioma U251 and SHG-44 cells with stably decreased IRG1 expression.
Knockdown of endogenous IRG1 expression induced the inhibition of
the tumor suppressor retinoblastoma protein pRB and the oncogenic
cell cycle regulator transcription factor E2F1, while upregulating
the expression of the tumor suppressor cyclin-dependent kinase
inhibitor p21. However, the levels of the cyclin-dependent kinases
CDK4 and CDK6 were depressed (Fig.
5A). In addition, we found that IRG1 inhibition decreased the
expression of the EMT marker genes snail, N-cadherin, and vimentin
(13,14), while increasing the level of
E-cadherin, an epithelial marker (Fig.
5B). IRG1 suppression did not promote the transition from
epithelial to mesenchymal morphology in glioma SHG-44 cells.
Several studies have provided evidence that
secretion of cytokines and growth factors, many of them encoded by
the NF-κB target genes, is critical for constitutive activation of
NF-κB in cancer cells (15). There
has been an increasing interest in the role of NF-κB in the
maintenance of established cancers, where it is frequently
constitutively induced and plays a major role in the
transcriptional activation of oncogenic genes (16). NF-κB signaling involves degradation
of the NF-κB inhibitor proteins IκBs through phosphorylation by the
IκB kinase (IKK). In the past study, we found that knockdown of
IRG1 expression by using si-IRG1 decreased the levels of NF-κB and
IkBα, as well as STAT3, which was shown to maintain constitutive
NF-κB activity in tumors (10,16)
(Fig. 5C). These results indicate
that the abrogation of IRG1 expression in glioma cells inhibits
NF-κB-mediated signaling.
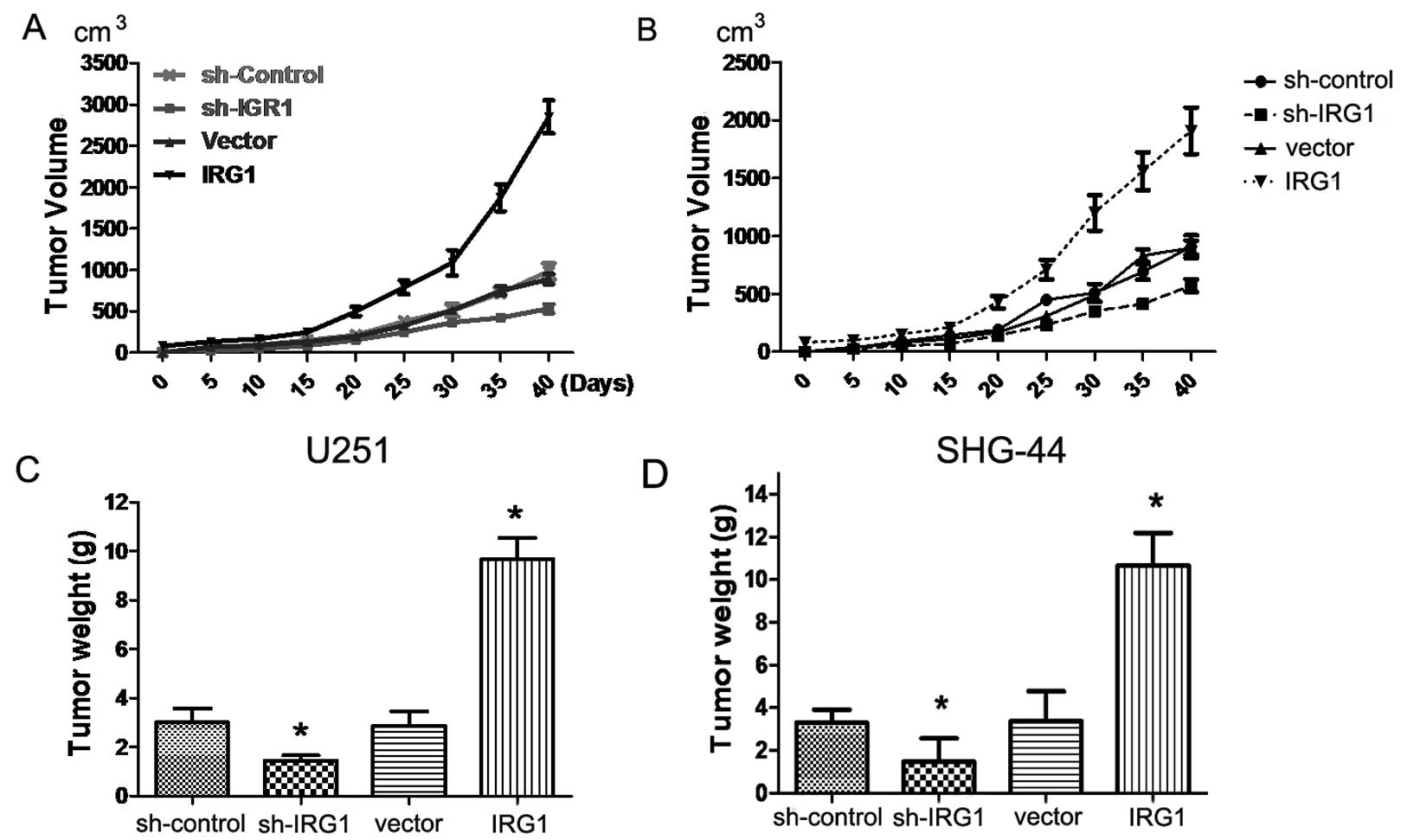
IRG1 contributes to in vivo xenograft
tumor growth
In addition to examining the biological functions of
IRG1 in vivo, we assessed the in vivo role of IRG1 by
using a xenograft transplantation model. We subcutaneously injected
sh-IRG1 and IRG1 lentiviral infection U251 and SHG-44 cells or
control cells into nude mice, and monitored tumor growth over a
40-day period. As shown in Fig. 6A and
B, the average tumor volume in mice injected with the
IRG1-depleted U251 cells was markedly (by >60%) reduced when
compared to that in the control animals (P<0.05). Tumor weight
analysis showed that sh-IRG1-transfected cells gave rise to
significantly smaller tumors than did sh-vector-transfected cells
(P<0.05) (Fig. 6C and D). These
results confirmed the previously results in vitro,
indicating that expression of IRG1 increases the cancer cell growth
of gliomas.
Discussion
Glioma is a disease characterized by uncontrolled
cell growth in brain tissues that may lead to metastases (17). It ultimately results in the death of
the patient. Tumor development from the stages of formation and
growth to those of invasion and metastasis is a complex process
that often involves changes in the tissue microenvironment, gene
mutations, angiogenesis, and other pathological processes. In this
series of events, pro-inflammatory proteins, including various
cytokines and transcription factors, play a very important role
(18). Inflammatory regulators are
associated not only with early stages of tumor growth but also with
malignant cell migration, invasion and metastasis (19).
The IRG1 gene plays a crucial role in the
orchestration of immune responses to infection (20). To clarify the role of IRG1 in glioma
pathogenesis, we analyzed IRG1 protein expression in glioma and
normal tissues by immunohistochemistry. We found that the IRG1
expression levels were higher in glioma tumors compared to those in
normal brain tissues. Increased IRG1 expression was associated with
disease progression and poor prognosis in glioma patients.
Furthermore, inhibition of IRG1 expression upregulated the tumor
suppressor p21 gene, and this may have caused inhibition of cell
proliferation and migration. These results suggest that the
molecular mechanisms underlying IRG1 activity may be associated
with pro-inflammatory factors and EMT-related proteins.
The transcription factor E2F1 was initially
identified as an adenoviral E2 promoter-binding cellular factor-1
induced by adenoviral E1A oncoprotein (21). E2F1 ectopic expression is sufficient
to promote the G1/S phase transition in the cell cycle. Our
findings indicate that IRG1 knockout can suppress E2F1 expression.
At present, cumulative evidence points to the members of the cyclin
D/CDK4/CDK6/pRB cell cycle regulatory pathway as potential targets
for cancer therapy (22). Both CDK4
and CDK6 encode cyclin-dependent serine-threonine kinases that
respond to mitogenic or pro-proliferative stimuli by complexing
with D-type cyclins and phosphorylating the tumor suppressor
protein pRB. The phosphorylated pRB is released from the complex
with E2F transcription factors, enabling E2F to initiate the
transcription of genes required for G1/S cell cycle progression,
and ultimately cellular proliferation ( 23). Here, we found that knockdown of
endogenous IRG1 expression reduced the activation of CDK4, CDK6,
and other oncogenic cell cycle regulators, including cyclin D1 and
pRd, and increased the expression of tumor suppressors, including
p21. Our results indicate that IRG1 may influence cell cycle
transition by mediating the transcription of cell cycle regulatory
proteins.
Previous studies have shown that snail, N-cadherin,
and vimentin are markers of EMT (22). In this study, we found that the
downregulation of IRG1 significantly inhibited the expression of
snail, N-cadherin and vimentin in glioma SHG-44 cells. The in
vitro Transwell cell migration assay and in vivo
xenograft model showed that the downregulation of IRG1 expression
significantly repressed the migration and pulmonary metastasis of
glioma cells. Our studies showed that IRG1 inhibition promoted the
expression of the epithelial marker E-cadherin, while reducing the
levels of the mesenchymal markers snail, N-cadherin, and vimentin
in glioma cells. These data suggest that IRG1 may increase EMT and
cancer metastasis. Several studies have provided evidence that
secretion of pro-inflammatory cytokines and growth factors is
critical for constitutive activation of NF-κB in cancer cells
(24–26). The reduced expression of IRG1 in
glioma cells may have caused inhibition of NF-κB signaling,
suggesting that IRG1 can be involved in carcinogenesis through
NF-κB signaling as well.
In summary, our study revealed that the cytoplasmic
expression of IRG1 was significantly upregulated in glioma and that
it is an important prognostic factor for glioma patients. Our in
vitro and in vivo data indicate that IRG1 is involved in
proliferation, invasion, and migration of cancer cells through
multiple molecular mechanisms, including regulation of NF-κB
signaling, the EMT pathway, and cell cycle transition, suggesting
that IRG1 plays an important role in the development of
gliomas.
Acknowledgements
The authors thank Dr Jun Wu for his support and
assistance with the preparation of the manuscript. This study was
supported by grants from National Natural Science Foundation of
China (no. 81172416).
References
|
1
|
Hamza MA and Gilbert M: Targeted therapy
in gliomas. Curr Oncol Rep. 16:3792014. View Article : Google Scholar
|
|
2
|
Kaufmann JK and Chiocca EA: Glioma virus
therapies between bench and bedside. Neuro Oncol. 16:334–351. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lohkamp B, Bauerle B, Rieger PG and
Schneider G: Three-dimensional structure of iminodisuccinate
epimerase defines the fold of the MmgE/PrpD protein family. J Mol
Biol. 362:555–566. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lee CG, Demarquoy J, Jackson MJ and
O’Brien WE: Molecular cloning and characterization of a murine
LPS-inducible cDNA. J Immunol. 152:5758–5767. 1994.PubMed/NCBI
|
|
5
|
Gautam A, Dixit S, Philipp MT, et al:
Interleukin-10 alters effector functions of multiple genes induced
by Borrelia burgdorferi in macrophages to regulate Lyme
disease inflammation. Infect Immun. 79:4876–4892. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Li Y, Zhang P, Wang C, et al: Immune
responsive gene 1 (IRG1) promotes endotoxin tolerance by increasing
A20 expression in macrophages through reactive oxygen species. J
Biol Chem. 288:16225–16234. 2013. View Article : Google Scholar
|
|
7
|
Lee CG, Jenkins NA, Gilbert DJ, Copeland
NG and O’Brien WE: Cloning and analysis of gene regulation of a
novel LPS-inducible cDNA. Immunogenetics. 41:263–270.
1995.PubMed/NCBI
|
|
8
|
Shi S, Blumenthal A, Hickey CM, Gandotra
S, Levy D and Ehrt S: Expression of many immunologically important
genes in Mycobacterium tuberculosis-infected macrophages is
independent of both TLR2 and TLR4 but dependent on IFN-alphabeta
receptor and STAT1. J Immunol. 175:3318–3328. 2005.PubMed/NCBI
|
|
9
|
Muccioli M, Sprague L, Nandigam H, Pate M
and Benencia F: Toll-like receptors as novel therapeutic targets
for ovarian cancer. ISRN Oncol. 2012:6421412012.PubMed/NCBI
|
|
10
|
Hall CJ, Boyle RH, Astin JW, et al:
Immunoresponsive gene 1 augments bactericidal activity of
macrophage-lineage cells by regulating β-oxidation-dependent
mitochondrial ROS production. Cell Metab. 18:265–278.
2013.PubMed/NCBI
|
|
11
|
Chiba Y, Mizoguchi I, Mitobe K, et al:
IL-27 enhances the expression of TRAIL and TLR3 inhuman melanomas
and inhibits their tumor growth in cooperation with a TLR3 agonist
poly(I:C) partly in a TRAIL-dependent manner. PLoS One.
8:e761592013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Song Q, Li Y, Zheng X, et al: MTA1
contributes to actin cytoskeleton reorganization and metastasis of
nasopharyngeal carcinoma by modulating Rho GTPases and Hedgehog
signaling. Int J Biochem Cell Biol. 45:1439–1446. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Lathia JD, Li M, Hall PE, et al: Laminin
alpha 2 enables glioblastoma stem cell growth. Ann Neurol.
72:766–778. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Cheng L, Huang Z, Zhou W, et al:
Glioblastoma stem cells generate vascular pericytes to support
vessel function and tumor growth. Cell. 153:139–152. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Islam KN, Bae JW, Gao E and Koch WJ:
Regulation of nuclear factor κB (NF-κB) in the nucleus of
cardiomyocytes by G protein-coupled receptor kinase 5 (GRK5). J
Biol Chem. 288:35683–35689. 2013.
|
|
16
|
Cheah SC, Lai SL, Lee ST, Hadi AH and
Mustafa MR: Panduratin A, a possible inhibitor in metastasized A549
cells through inhibition of NF-kappa B translocation and
chemo-invasion. Molecules. 18:8764–8778. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhao L, Wang H, Sun X and Ding Y:
Comparative proteomic analysis identifies proteins associated with
the development and progression of colorectal carcinoma. FEBS J.
277:4195–4204. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wang F, Qiao Y, Yu J, et al: FBX8 acts as
an invasion and metastasis suppressor and correlates with poor
survival in hepatocellular carcinoma. PLoS One. 8:e654952013.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Huang J, Ye X, Guan J, et al: Tiam1 is
associated with hepatocellular carcinoma metastasis. Int J Cancer.
132:90–100. 2013. View Article : Google Scholar
|
|
20
|
Michelucci A, Cordes T, Ghelfi J, et al:
Immune-responsive gene 1 protein links metabolism to immunity by
catalyzing itaconic acid production. Proc Natl Acad Sci USA.
110:7820–7825. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kovesdi I, Reichel R and Nevins JR: E1A
transcription induction: enhanced binding of a factor to upstream
promoter sequences. Science. 231:719–722. 1986. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang WJ, Wu SP, Liu JB, et al: MYC
regulation of CHK1 and CHK2 promotes radioresistance in a stem
cell-like population of nasopharyngeal carcinoma cells. Cancer Res.
73:1219–1231. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Lee HH, Lee SR and Leem SH:
Tristetraprolin regulates prostate cancer cell growth through
suppression of E2F1. J Microbiol Biotechnol. 24:287–294. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kim MK, Kang YJ, Kim DH, et al: A novel
hydroxamic acid derivative, MHY218, induces apoptosis and cell
cycle arrest through downregulation of NF-κB in HCT116 human colon
cancer cells. Int J Oncol. 44:256–264. 2014.PubMed/NCBI
|
|
25
|
Kim A, Yim NH, Im M, Jung YP, Kim T and Ma
JY: Suppression of the invasive potential of highly malignant tumor
cells by KIOM-C, a novel herbal medicine, via inhibition of NF-κB
activation and MMP-9 expression. Oncol Rep. 31:287–297.
2014.PubMed/NCBI
|
|
26
|
Zhang W and Grivennikov SI: Top Notch
cancer stem cells by paracrine NF-κB signaling in breast cancer.
Breast Cancer Res. 15:3162013.PubMed/NCBI
|