Introduction
The number of patients suffering from cancers
worldwide is increasing, and one of the most challenging issues in
oncology continues to be the problem of developing active drugs
economically and in a timely manner. In fact, the likelihood of
approval from pre-clinical discovery to phase I clinical trial is
lowest for oncology drugs (7%) compared with drugs for other
indications (1). Considering the
high cost and time-consuming nature of the clinical development of
oncology drugs, better pre-clinical platforms for drug screening
are urgently required.
Traditionally, the activity of anticancer drugs has
been evaluated in two-dimensionally (2D)-cultured cancer cell
lines. However, it is now being recognized that 2D-cultured cells
are unable to simulate the microenvironment of the original tumors,
which grow three-dimensionally (3D) (2–7). This
is speculated to be relevant to the fact that many drugs proving to
be clinically futile were pre-clinically evaluated to be ‘active’
using 2D-cultured cell line-based models. 3D-culture systems have
received attention as a means to avoid certain drawbacks of
2D-culture models, by recapitulating the tumor microenvironment, at
least in part (2–7).
In the present study, to investigate the utility of
3D-culture models in testing the activity of chemotherapeutic
drugs, we compared 2D- and 3D-culture of breast cancer cell lines
and primary cells obtained from a breast cancer patient and grown
as a patient-derived xenograft (PDX).
Materials and methods
Breast cancer cell lines and PDX
Two estrogen receptor
(ER)-positive/HER2-non-amplified (luminal-type; T-47D and
MCF-7), two ER-negative/HER2-amplified (HER2-type; BT-474
and HCC-1954) (8), and two
ER-negative/HER2-non-amplified (triple-negative type; BT-549
and MDA-MB-231) (8) breast cancer
cell lines were purchased from the American Type Culture Collection
(ATCC, Manassas, VA, USA). The cells were maintained in
RPMI-1640® (Sigma-Aldrich, St. Louis, MO, USA)
supplemented with 10% fetal bovine serum (FBS; Gemini-Bio-Products,
Inc., Woodland, CA, USA), 100 U/ml penicillin, 100 U/ml
streptomycin, and 2 mM glutamine. All cells were cultured at 37°C
in a humidified atmosphere with 5% CO2 and were in
logarithmic growth phase upon initiation of the experiments. The
cells were passaged for ≤3 months before fresh cells were obtained
from frozen early passage stocks received from the supplier.
The breast cancer tissue sample was collected from a
surgically resected primary tumor from a patient who underwent
surgery at Kobe University Hospital. The patient gave informed
consent for the research use of the tumor samples, as approved by
the Research Ethics Board at Kobe University Graduate School of
Medicine. The generation of PDX was as previously described
(9). In brief, the breast cancer
tissues were cut into fragments (~1 mm3 in size) using a
razor blade, and 4×106 cells were transplanted
orthotopically into inguinal mammary fat pad regions of female
NOD-SCID mice (Clea, Tokyo, Japan). When the size of the tumors
reached ~1–2 cm in diameter, the mice were sacrificed, and the
xenograft tumor was excised. In this study, a piece of fresh PDX
tissue originating from a luminal-type breast cancer tumor was
disag-gregated by automated mechanical method
(Medimachine®; Azone, Osaka, Japan). The primary cells
were cultured in Dulbecco’s modified Eagle’s medium/nutrient
mixture F12® (Life Technologies, Grand Island, NY, USA)
supplemented with 2% FBS, 100 U/ml penicillin and 100 U/ml
streptomycin, and cultured at 37°C in a humidified atmosphere with
5% CO2. All animal experiments were carried out under
the approval of the Kobe University Medical School Animal Care and
Use Committee.
Drugs
Paclitaxel (PTX), doxorubicin (ADR), and
5-fluorouracil (5FU) were purchased from Wako (Osaka, Japan). Stock
solutions were prepared in dimethyl sulfoxide (DMSO) and stored at
20°C. The drugs were diluted in fresh media before each experiment,
with final DMSO concentrations <0.1%.
Drug sensitivity test
Cells (100 μl/well, n=6) were plated (day 0) in 2D
plates [Falcon 96-well Tissue Culture Plate® (353072);
Corning Inc., NY, USA] or 3D plates [NanoCluture 96-well
Plate® (NCP-LH-96); SCIVAX, Kanagawa, Japan]. The
numbers of cells seeded per well were as follows (determined in a
preparatory experiment): BT-474, 5,000 (2D) and 15,000 cells/well
(3D); T-47D, MCF-7 and BT-549, 2,500 (2D) and 10,000 cells/well
(3D); HCC-1954, 1,000 (2D) and 5,000 cells/well (3D); MDA-MB-231,
500 (2D) and 2,500 cells/well (3D). Drugs were added on day 3. The
concentration of each drug was adjusted to achieve 0.1, 1 and 10×
the areas under the curve (AUC) obtained in clinical
pharmacokinetic studies (Table I)
(10). The number of viable cells
was evaluated using a CellTiter-Glo® luminescent assay
(Promega, Madison, WI, USA) on day 6. A series of bright field
images were recorded on days 0–6 using a BZ-X710®
inverted microscope (Keyence, Osaka, Japan).
 | Table IDrug concentrations applied. |
Table I
Drug concentrations applied.
| Package insert
data | | | |
|---|
|
| | | |
|---|
| Drug name | Dose
(mg/m2) | Maximum drug
concentration (μg/ml) | AUC (μg·h/ml) | MW | Clinical dose for
breast cancer (mg/m2) | Estimated AUC
(μM·h) | Concentration for
cell exposure (1× of AUC, μM) |
|---|
| PTX | 180 | 4.47±1.29 | 16.46±3.76 | 853.91 | 175 | 13.66 | 0.19 |
| ADR | 50 | 8.70±2.73 | 62.40±40.7 | 579.98 | 60 | 43.35 | 0.60 |
| 5FU | - | - | 20–24 | 130.08 | - | 2.86 | 0.04 |
Protein extraction and western
blotting
Cells treated with PTX (1× the AUC, Table I) were washed once with ice-cold PBS
and scraped immediately after adding lysis buffer [20 mM Tris (pH
7.5), 150 mM NaCl, 2 mM EDTA, 10% glycerol, 1% NP40] containing
protease and phosphatase inhibitors (100 mM NaF, 1 mM
phenylmethylsulfonyl fluoride, 1 mM Na3VO4, 2
mg/ml aprotinin, 5 mg/ml leupeptin). Cell lysates were centrifuged
at 14,000 × g for 10 min at 4°C to pellet insoluble material, and
the supernatant protein extracts were collected. Aliquots of
protein extracts were separated by electrophoresis on precast 7.5%
polyacrylamide gels, followed by transfer to polyvinylidene
difluoride membranes. Membranes were probed with an antibody to
detect cleaved poly(ADP-ribose) polymerase (PARP) (Asp214)(D64E10)
(1:1,000; #5625; Cell Signaling Technology Beverly, MA, USA), and
separately with a β-actin antibody (1:3,000; A1978; Sigma-Aldrich).
Each primary antibody was detected using Amersham ECL Plus Western
Blotting Detection Reagents (GE Healthcare, Buckinghamshire,
UK).
Hypoxia assay
Cells in maintenance media were plated (day 0) in 2D
(Corning Inc.) or 3D 96-well plates (SCIVAX), as described above.
The hypoxia probe LOX-1® (SCIVAX) was added on day 3.
LOX-1 is a phosphorescent light-emitting iridium complex; its
phosphorescence, which is quenched by oxygen, and increases in
response to low levels of oxygen, can be monitored using a
fluorescence microscope (11). A
BZ-X710 microscope was used to record the fluorescence images on
day 4.
Histologic and immunohistochemical
examinations
Cells were harvested by scraping into 7.5% formalin.
On the following day, the sediment containing the cell button was
scooped out and processed into paraffin wax. The paraffin-embedded
cell button (cell block) was sectioned at a 4-μm thickness and
these were assessed immunohistochemically using the following
primary antibodies: Ki-67 (1:5; #IR626; Dako, Glostrup, Denmark),
caspase-3 (1:200; #NCL-CPP32; Leica-Novocastra, Newcastle upon
Tyne, UK), and caspase-8 (1:150; #NCL-CASP-8; Leica-Novocastra).
For antigen retrieval, a citrate buffer (pH 6.0) was used for 10
min at 100°C and 30 min cooling to room temperature. A MACH 2
Double Stain system (Biocare Medical, Concord, CA, USA) was used to
detect antigen-antibody reactions, followed by brown coloring by
DAB staining. Appropriate positive controls were employed for all
conditions.
Results
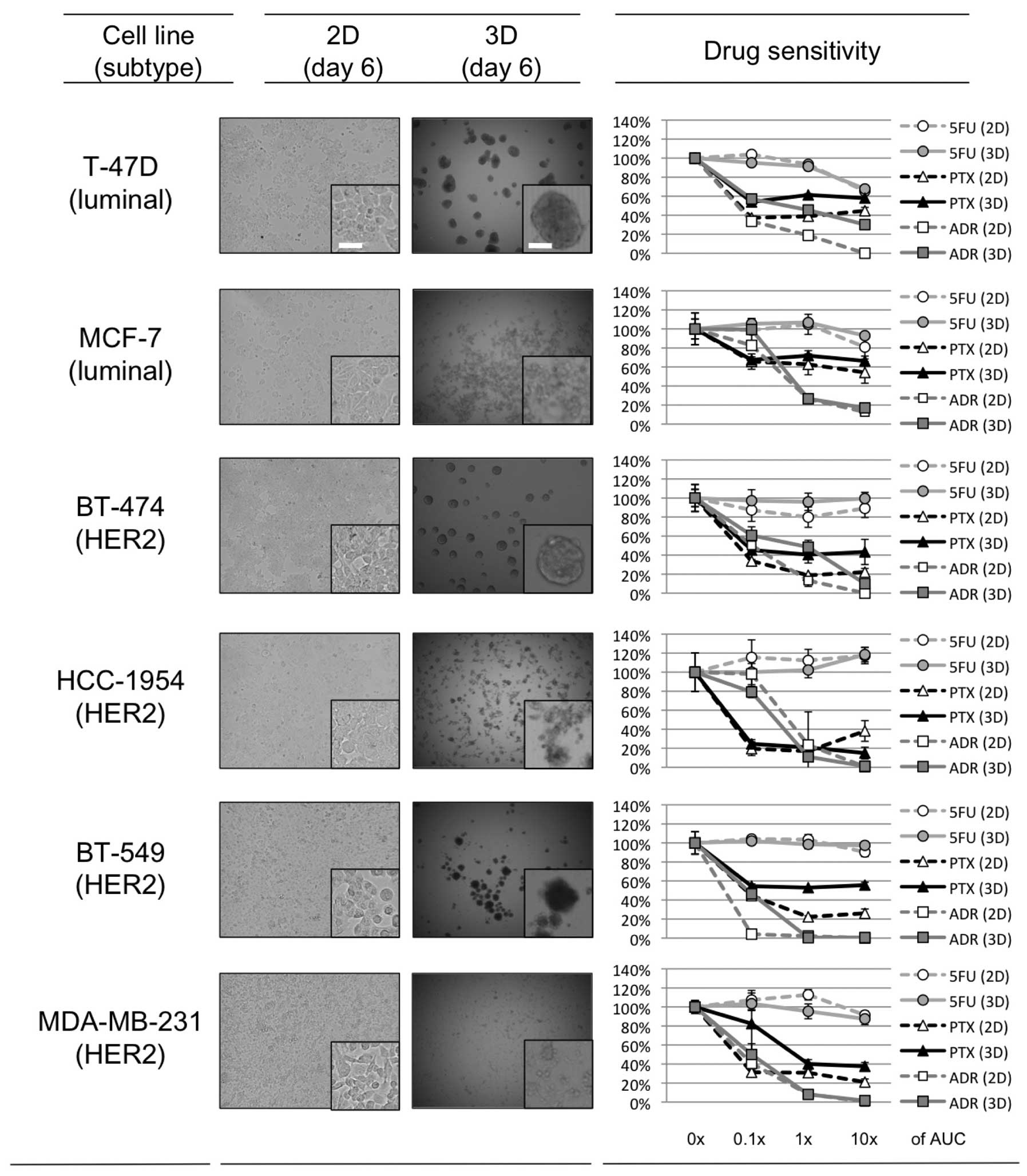
Spheroid formation in 3D-culture
Approximately one day after seeding on 3D-culture
plates, 3 of the 6 breast cancer cell lines, T47-D, BT-474 and
BT-549, started to form dense multicellular spheroids (MCSs). The
size of these MCSs plateaued ~3 days after seeding, and the maximal
size of the spheroids was ~200–300 μm for each of these cell lines.
For MCF-7, HCC-1954 and MDA-MB-231, the cells accumulated in the
3D-culture plates somewhat more than in the 2D-culture plates, and
although spheroids formed, they were looser than those produced by
the other 3 cell lines (Fig.
1).
Drug sensitivity in 2D- and
3D-culture
The relative growth rate of the 6 breast cancer cell
lines in the presence or absence of the 3 chemotherapeutic agents
(PTX, ADR, and 5FU) in 2D- or 3D-cell culture is shown in Fig. 1. The range of drug concentrations
used was set based on clinical pharmacokinetic data (0.1, 1, and
10× the AUC; Table I) in order to
explore the clinical relevance of the results. For 5FU, there was
no clear difference in sensitivity between the 2D- and 3D-culture
in any of the cell lines tested. The 3 cell lines that developed
dense 3D-MCSs (T-47D, BT-474 and BT-549) tended to show relative
resistance to PTX and ADR in the 3D-culture as compared to this
resistance in 2D. In contrast, the other 3 that developed loose
3D-spheroids (MCF-7, HCC-1954 and MDA-MB-231) tended to have
similar sensitivity to PTX and ADR in the 2D- and 3D-culture. These
findings confirm that the formation of dense MCSs in 3D-culture
plays a role in determining the sensitivity of the cell lines to
PTX and ADR.
Apoptosis induced by PTX
To explore the mechanism of differential sensitivity
to PTX in the 2D- and 3D-cultures, 3 cell lines were subjected to
further study; BT-549 and BT-474 as representatives of cell lines
forming dense 3D-MCSs, and MCF-7 as an example of the loose 3D
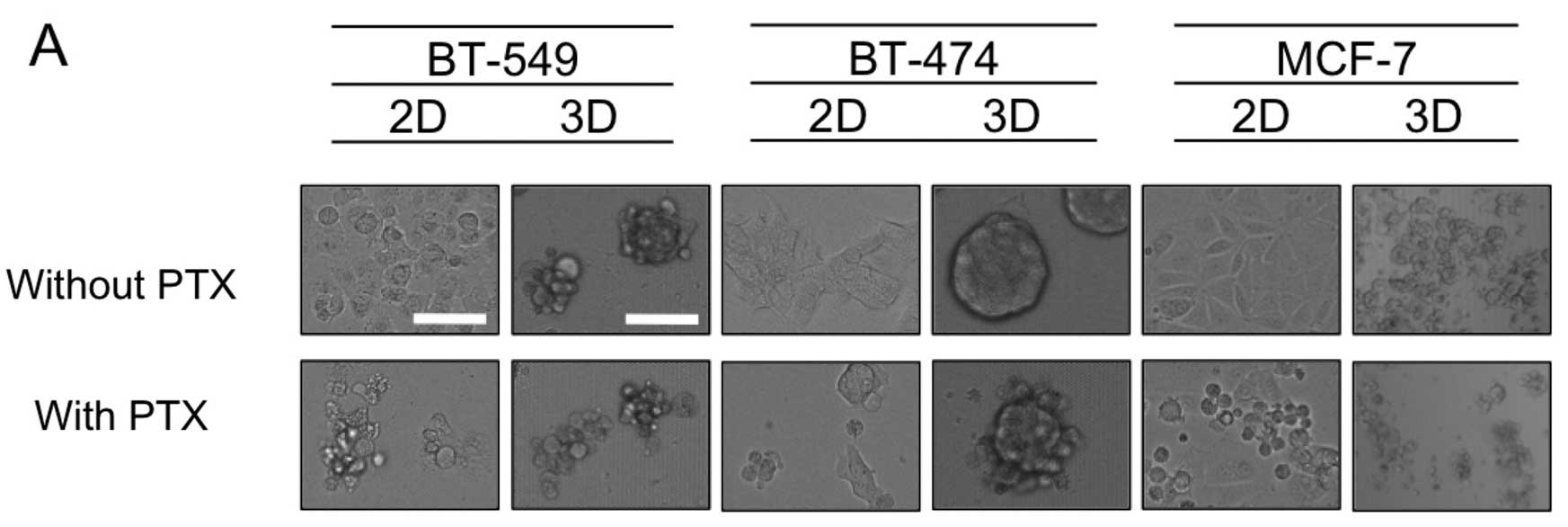
phenotype. Fig. 2A shows that for
BT-549 and BT-474 cells, exposure to PTX (1× the AUC) resulted in
cell shrinkage, indicative of apoptosis when grown in 2D-culture,
yet dense 3D-MCSs remained in the 3D-cultures. The same PTX
treatment of MCF-7 cells yielded significant cell shrinkage in both
the 2D- and 3D-cultures (Fig. 2A).
Consistent with these findings, treatment of BT-549 or BT-474 cells
with PTX resulted in much smaller increases in cleaved PARP in the
3D-culture than in the 2D-culture, whereas in MCF-7 (with loose
MCSs) the same PTX treatment resulted in a similar degree of
increase in cleaved PARP (Fig. 2B).
These results confirmed that relative resistance to PTX in the cell
lines with dense MCSs in 3D-culture than in 2D-culture was in part
due to reduced apoptosis.
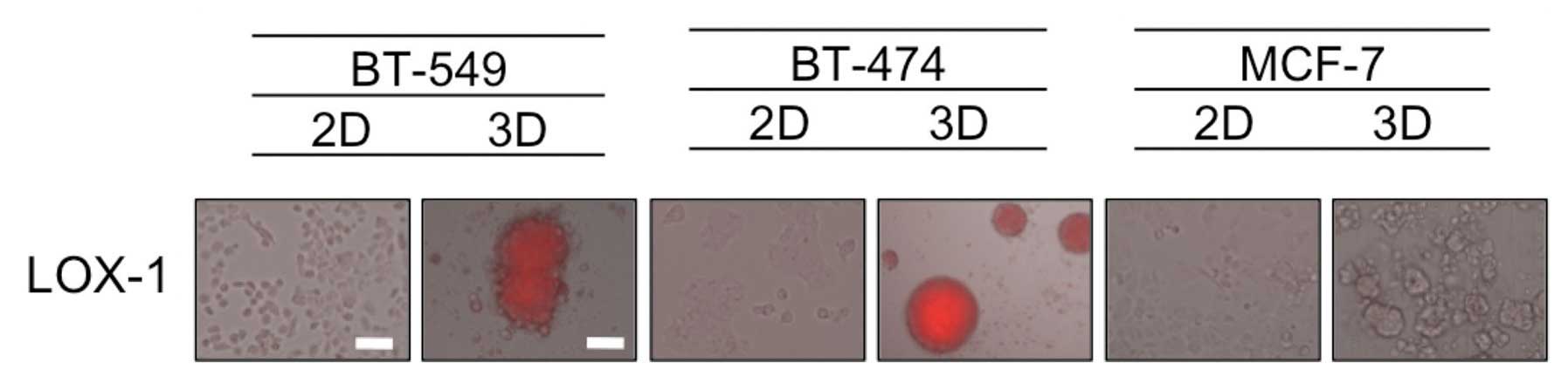
Hypoxia in dense MCSs
It has been speculated that the formation of dense
MCSs leads to hypoxia inside the spheroids mimicking in vivo
tumors, and hypoxia is known to be one of the factors associated
with resistance to chemotherapeutic drugs (12). Therefore, we evaluated the oxygen
status in 2D- and 3D-cultures by utilizing the hypoxia probe LOX-1,
that generates signals that are visible using a fluorescence
microscopy. As shown in Fig. 3,
hypoxic areas were observed inside the dense 3D-MCSs from the
BT-549 and BT-474 cell lines, but not in the loose spheroids formed
by MCF-7 cells or cells grown in 2D. These findings suggest that
the hypoxia status in dense 3D-MCSs may be one of the causes of
their drug resistance.
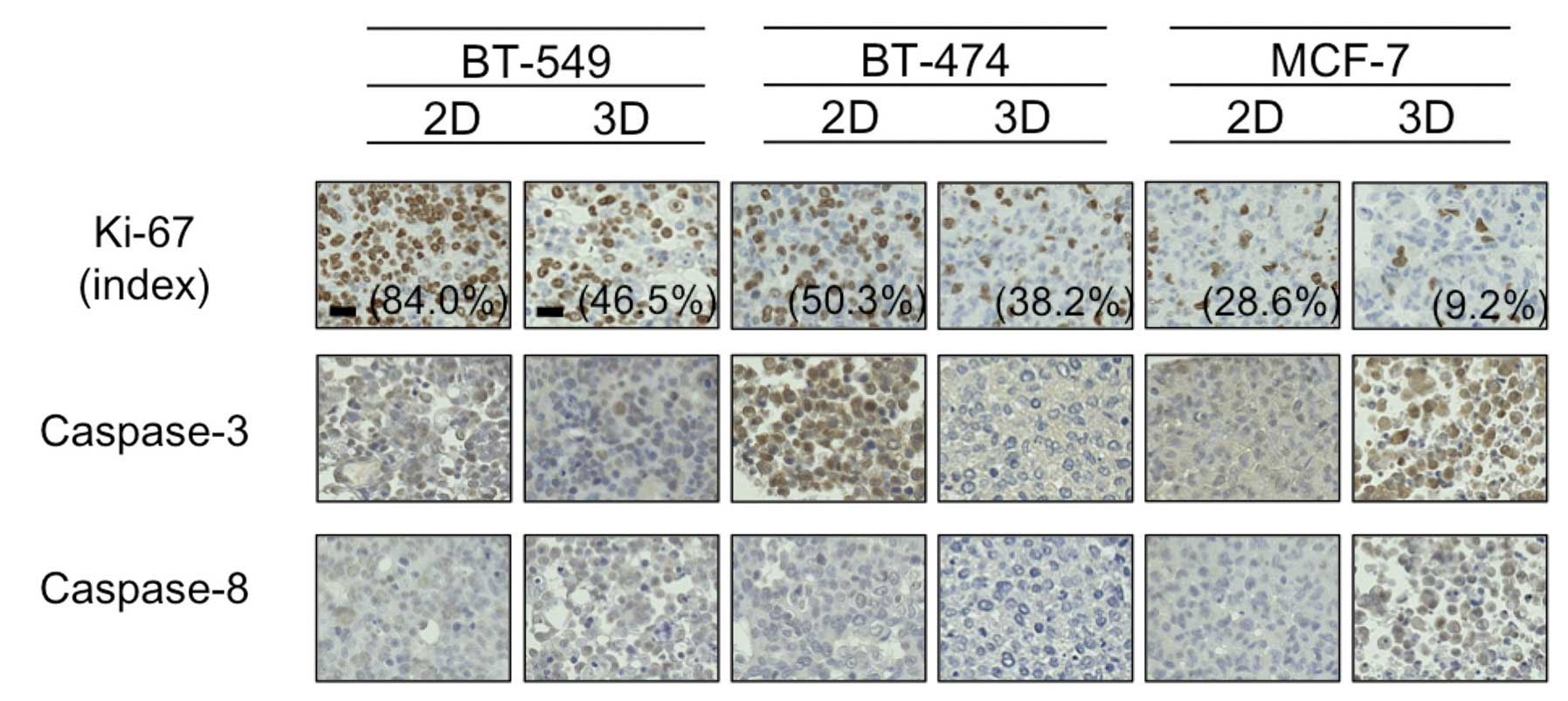
Ki-67 and caspase expression
Hypoxia has been reported to cause cancer cell
dormancy in the G0 phase (13),
thus we next evaluated staining using the antibody Ki-67, which is
reported to be positive in cells except for those in the G0 phase.
As shown in Fig. 4, the Ki-67 index
was higher in the 2D-culture than that in the 3D-culture for all 3
cell lines tested. The difference was particularly marked for the
BT-549 cells, which formed dense 3D-MCSs (84.0% Ki-67-positive
cells in 2D vs. 46.5% in 3D).
Following this, due to a study suggesting that
downregulation of caspase-3 and -8 in hypoxic conditions may
mediate resistance to PTX in breast cancer cells (14), we immunohistochemically evaluated
the expression of these proteins in 2D- and 3D-cultures. As shown
in Fig. 4, the expression levels of
caspase-3 and -8 were almost identical between the 2D- and
3D-cultures for all 3 cell lines tested, with the exception of
BT-474 cells, in which caspase-3 expression was much higher in the
2D-culture than this level in the 3D-culture. In addition,
caspase-3 was detected mainly in the nuclei of 2D-cultured BT-474
cells, indicative of active caspase-3. These findings suggested to
us that hypoxia in 3D-MCSs may lead to cell dormancy and/or
downregulation of caspase-3, and result in resistance to PTX.
The potential of the 3D-primary culture
to simulate tumor growth in vivo
To explore the relevance of a 3D-culture to tumor
growth in vivo, we compared an excised tumor and its 2D- and
3D-primary cultures in terms of expression of Ki-67, caspase-3, and
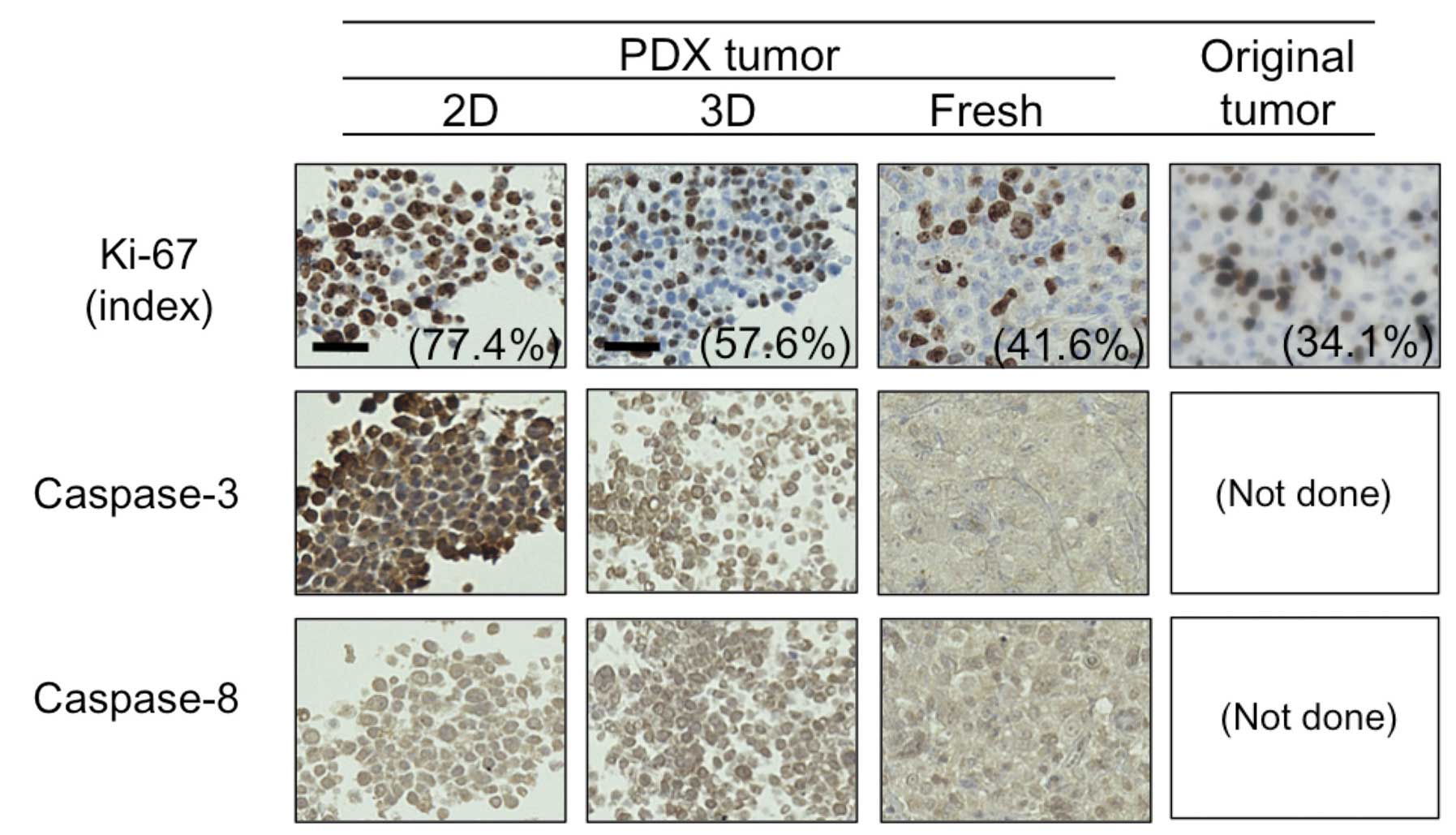
caspase-8. We utilized a PDX tumor for this purpose. As shown in
Fig. 5, the proportion of cells
positive for Ki-67 in the 2D- and 3D-primary cultured cells
originating from PDX, fresh PDX tumor, and the patient’s original
tumor were 77.4, 57.4, 41.6 and 34.1%, respectively. Moreover,
consistent with the BT-474 cells (Fig.
4), caspase-3 expression was much higher in the PDX tumor and
its 2D-primary culture than in its 3D-primary culture, whereas no
difference was observed in caspase-8 expression between the 2D- and
3D-culture. These findings confirm that a 3D-primary culture rather
than a 2D-primary culture may better indicate in vivo tumor
dormancy and anti-apoptotic features.
Discussion
In the present study, we demonstrated that certain
breast cancer cell lines formed dense 3D-MCSs, and the formation of
MCSs was associated with decreased sensitivity to chemotherapeutic
drugs (Fig. 1). Our data also
revealed that the drug resistance may be caused by hypoxia in MCSs,
which was associated with an increased cell population in the G0
phase and/or downregulation of proapoptotic molecules such as
caspase-3 (Figs. 2–4). Moreover, tumor dormancy observed in
the in vivo PDX tumor was better represented by its
3D-primary culture than its 2D-primary culture (Fig. 5).
2D-cultured cancer cell lines grown on plastic
surfaces are considered unable to precisely simulate tumor
conditions in vivo (2–7). It
may therefore, be predicted that the development of anticancer
drugs based on screening using 2D-cultured cell lines is not
efficient. The fact that numerous anticancer drugs are eliminated
during clinical development indicates that anticancer activity
tends to be overestimated on a 2D-culture-based screening platform.
In contrast, 3D-culture systems have been shown to better simulate
the in vivo tumor microenvironment than a 2D-culture
(3–7); several previous studies have shown
that 2D-cultured cells tended to overestimate the efficacies of
chemotherapeutic drugs compared with 3D-cultured cells (15–17).
Consistent with these studies, our present study showed that the
cell lines producing dense 3D-MCSs are more resistant to PTX and
ADR in a 3D- than in 2D-culture (Fig.
2A). When measuring PTX-induced apoptosis, this tendency was
even more apparent (Fig. 2B),
suggesting that the formation of 3D-MCSs may protect cells from
cell death. These findings suggest that a 3D-culture potentially
avoids the overestimation of antitumor efficacy observed in a
2D-culture. It is noteworthy that the phenomena were observed with
the clinically achievable drug concentrations calculated based on
the AUC in cancer patients, which were less than the maximum drug
concentrations (Cmax, Table I).
While drug concentrations around Cmax have conventionally been
employed for testing in in vitro experiments, these
concentrations may be inappropriate since Cmax does not persist for
hours in vivo (18).
Therefore, we believe that drugs which kill 3D-growing cells at
concentrations less than Cmax should be evaluated to be active.
Potential advantages of 3D-culture systems over
2D-culture systems are that only the former may feature oxygen
gradients which exist in in vivo tumors. Hypoxia is known to
cause drug resistance through several mechanisms. One conventional
mechanism is that of hypoxia-induced cell cycle arrest. Sullivan
et al demonstrated that hypoxia induced an increased G0
non-cycling population associated with etoposide resistance in
breast cancer cell lines (13).
Consistent with this result, one of the cell lines forming dense
MCSs in 3D, BT-549, had a much lower Ki-67 labeling index
(representing a greater G0 population) in the 3D than that in the
2D-cultured cells. However, another cell line that formed dense
3D-MCSs, BT-474, had only a slightly lower Ki-67 in 3D than in 2D,
with a difference of 12.1% (Fig.
4), which was actually smaller than that in MCF-7 cells which
formed only loose MCSs. This indicated that drug resistance
associated with dense 3D-MCSs could not always be attributable to
an increased G0-dormant cell population. Therefore, we evaluated
the expression of caspase-3 and -8, as the downregulation of these
two pro-apoptotic molecules, along with change in expression of
another six molecules, were identified to be potentially involved
in hypoxia-induced protection against PTX-induced apoptosis by a
previous study (14). Partially
consistent with this study, BT-474 expressed lower levels of
caspase-3 in the 3D- than in 2D-culture. Furthermore, we found that
tumor dormancy and downregulation of caspase-3 observed in the
original patient tumor and/or PDX tumor was better maintained in
the 3D-primary cultured cells than in the 2D-primary cultured cells
(Fig. 5). These findings suggest
that 3D-culture may provide a better drug screening platform; one
that produces more clinically relevant results.
Several limitations of our study warrant mention.
Firstly, this study was derived from the nature of an in
vitro model. At present, there are various 3D-culture systems
besides the one we used, yet none of them are considered to be a
standard method, and it is unclear which system is the most
clinically relevant (3–7). One clear observation is that
3D-culture of cell lines will never accurately, fully represent the
tumor microenvironment in vivo, because the latter have
interactions with stromal tissues or blood perfusions. Co-culture
with stromal cells or primary culture in 3D-conditions may
partially solve these issues and are under investigation in our
laboratory. Animal models such as PDXs are being evaluated as a
potential drug screening platform for the next generation (19–22),
however, difficulty in the development and expansion of the PDX
model, and their high-cost will preclude their use in
high-throughput screening. Therefore, refining 3D-culture systems
as drug screening platforms is worth driving forward. Secondly,
although our results support that hypoxia in 3D-culture may play a
role in drug resistance, we did not prove the underlying molecular
mechanisms of this in the present study. In addition, we did not
explore many other potential mechanisms of drug resistance induced
by hypoxia, such as enhanced drug efflux or autophagy, or
inhibition of senescence or DNA damage (12,23).
These mechanisms of hypoxia-induced drug resistance are reported to
be mediated mainly by hypoxia-inducible factor-1α (HIF-1α)
(12,24,25).
In our study, however, we were unable to detect HIF-1α with
immunohistochemistry or western blotting, even in dense MCSs (data
not shown).
In conclusion, 3D-cultured breast cancer cells show
relative drug resistance as compared to 2D-cultured cells when
forming dense 3D-MCSs, and the resistance is associated with
hypoxia. 3D-MCSs could be utilized as an in vitro cell-based
drug-testing platform.
Acknowledgements
We thank Ms. Megumi Izumi and staffs of Kobe
University Hospital Advanced Tissue Staining Center (KATS) for
their excellent technical support. This study was supported by the
Global Centers of Excellence Program (H.M.), Grant-in-Aid for
Scientific Research (C) (T.M.), and by a Research Grant from the
Takeda Science Foundation (T.M).
References
|
1
|
Hay M, Thomas DW, Rosenthal J, et al:
Clinical development success rates for investigational drugs. Nat
Biotechnol. 32:40–51. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
2
|
Yamada KM and Cukierman E: Modeling tissue
morphogenesis and cancer in 3D. Cell. 130:601–610. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hirschhaeuser F, Menne H, Kunz-Schughart
LA, et al: Multicellular tumor spheroids: an underestimated tool is
catching up again. J Biotechnol. 148:3–15. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Rimann M and Graf-Hausner U: Synthetic 3D
multicellular systems for drug development. Curr Opin Biotechnol.
23:803–809. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Breslin S and O’Driscoll L:
Three-dimensional cell culture: the missing link in drug discovery.
Drug Discov Today. 18:240–249. 2013. View Article : Google Scholar
|
|
6
|
Lovitt CJ, Shelper TB and Avery VM:
Advanced cell culture techniques for cancer drug discovery.
Biology. 3:345–367. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Weigelt B, Ghajar CM and Bissell MJ: The
need for complex 3D culture models to unravel novel pathways and
identify accurate biomarkers in breast cancer. Adv Drug Deliv Rev.
69–70:42–51. 2014. View Article : Google Scholar
|
|
8
|
Lacroix M and Leclercq G: Relevance of
breast cancer cell lines as models for breast tumours: an update.
Breast Cancer Res Treat. 83:249–289. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Nobutani K, Shimono Y, Takai Y, et al:
Absence of primary cilia in cell cycle-arrested human breast cancer
cells. Genes Cells. 19:141–152. 2014. View Article : Google Scholar
|
|
10
|
Gamelin E, Delva R, Jacob J, et al:
Individual fluorouracil dose adjustment based on pharmacokinetic
follow-up compared with conventional dosage: results of a
multicenter randomized trial of patients with metastatic colorectal
cancer. J Clin Oncol. 26:2099–2105. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang S, Hosaka M, Yoshihara T, et al:
Phosphorescent light-emitting iridium complexes serve as a
hypoxia-sensing probe for tumor imaging in living animals. Cancer
Res. 70:4490–4498. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Rohwer N and Cramer T: Hypoxia-mediated
drug resistance: novel insights on the functional interaction of
HIFs and cell death pathways. Drug Resist Updat. 14:191–201. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Sullivan R and Graham CH: Hypoxia prevents
etoposide-induced DNA damage in cancer cells through a mechanism
involving hypoxia-inducible factor 1. Mol Cancer Ther. 8:1702–1713.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Flamant L, Notte A, Michiels C, et al:
Anti-apoptotic role of HIF-1 and AP-1 in paclitaxel exposed breast
cancer cells under hypoxia. Mol Cancer. 9:1912010. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Karlsson H, Fryknas M, Nygren P, et al:
Loss of cancer drug activity in colon cancer HCT-116 cells during
spheroid formation in a new 3-D spheroid cell culture system. Exp
Cell Res. 318:1577–1585. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Vinci M, Gowan S, Boxall F, et al:
Advances in establishment and analysis of three-dimensional tumor
spheroid-based functional assays for target validation and drug
evaluation. BMC Biol. 10:292012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lee JM, Mhawech-Fauceglia P, Lee N, et al:
A three-dimensional microenvironment alters protein expression and
chemosensitivity of epithelial ovarian cancer cells in vitro. Lab
Invest. 93:528–542. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
No authors listed. Dishing out cancer
treatment. Nat Biotechnol. 31:852013. View
Article : Google Scholar : PubMed/NCBI
|
|
19
|
Marangoni E, Vincent-Salomon A, Auger N,
et al: A new model of patient tumor-derived breast cancer
xenografts for preclinical assays. Clin Cancer Res. 13:3989–3998.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Cottu P, Marangoni E, Assayag F, et al:
Modeling of response to endocrine therapy in a panel of human
luminal breast cancer xenografts. Breast Cancer Res Treat.
133:595–606. 2012. View Article : Google Scholar
|
|
21
|
Zhang X, Claerhout S, Prat A, et al: A
renewable tissue resource of phenotypically stable, biologically
and ethnically diverse, patient-derived human breast cancer
xenograft models. Cancer Res. 73:4885–4897. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Fong EL, Martinez M, Yang J, et al:
Hydrogel-based 3D model of patient-derived prostate xenograft
tumors suitable for drug screening. Mol Pharm. 11:2040–2050. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Housman G, Byler S, Sarkar S, et al: Drug
resistance in cancer: an overview. Cancers. 6:1769–1792. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Li J, Shi M, Cao Y, et al: Knockdown of
hypoxia-inducible factor-1alpha in breast carcinoma MCF-7 cells
results in reduced tumor growth and increased sensitivity to
methotrexate. Biochem Biophys Res Commun. 342:1341–1351. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Doublier S, Belisario DC, Sapino A, et al:
HIF-1 activation induces doxorubicin resistance in MCF7 3-D
spheroids via P-glycoprotein expression: a potential model of the
chemoresistance of invasive micropapillary carcinoma of the breast.
BMC Cancer. 12:4–18. 2012. View Article : Google Scholar
|