Introduction
Breast cancer is the most common invasive cancer and
the second cause of cancer-related death in women (1,2). Each
year more than a half a million new cases of breast cancer are
diagnosed in the US and Europe (3),
and in China. Breast cancer is a very heterogeneous disease. Some
patients are cured by the surgical removal of the primary tumor
while other patients suffer from metastasis and progression of the
disease, despite adjuvant therapy. Therefore, it is essential to
develop effective and safer therapeutic modalities against breast
cancer.
Glycosylation is one of the important steps of
protein post-translational modifications, and ~50% of proteins are
glycosylated (4). Protein
glycosylation plays a role in a variety of cellular biological
functions, such as cell-cell and cell-substrate adhesion, membrane
organization, cell immunogenicity and protein targeting (5). Specific changes in the glycosylation
patterns of cell surface glycoprotein have been shown to enhance
the metastatic potential of tumor cells. Aberrant expression of
fucosylated glycans has also been detected in various types of
tumors.
The fucosyltransferase (FUT) family is a group of
fucosylation synthases that catalyze the transfer of L-fucose (Fuc)
from an activated GDP-β-L-Fuc to various acceptor molecules such as
N-acetyllactosamine. The transfer of Fuc residue from the
donor substrate, GDP-Fuc, is catalyzed to the oligosaccharide
acceptor in a1,2-(FUT1 and FUT2), a1,3/4-(FUT3, FUT4, FUT5, FUT6,
FUT7, FUT9, FUT10 and FUT11) and a1,6-linkage (FUT8). Frequent
fucosylation, the final step within the glycosylation machinery,
results in glycans that are involved in various cellular processes
such as cell-cell recognition, adhesion and inflammation or tumor
metastasis. Fucosylation is suggested to have paramount importance
in the invasion and metastatic process of cancer stem cells (CSCs)
(6), for example increasing FUT4
and FUT7 expression promoted neoplastic cell proliferation and
hepatocellular carcinoma cell growth in vitro, respectively,
and reducing FUT3/6 expression suppressed colon carcinoma cell
proliferation (7,8).
MicroRNAs (miRNAs) comprise a class of small
non-coding RNAs implicated in post-transcriptional RNA regulation.
These RNA molecules are ~22 nt in length (9). By binding to complementary sequences
in the 3′UTRs of targeted mRNAs, miRNAs degrade or inhibit their
translation and regulate a range of cellular functions such as
differentiation, proliferation, apoptosis and migration of tumor
cells (10). Accumulating evidence
demonstrates that various types of miRNAs participate in the
regulation of tumorigenesis and metastasis (11,12).
Recent studies support the assumption that modulating the levels of
miR-146a or miR-146b could have therapeutic potential to suppress
breast cancer metastasis (13). It
was reported for the first time that anti-miR-17 molecules reduced
breast cancer cell migration in vitro and metastasis in
vivo (14). miR-184 was
identified as a putative breast tumor suppressor in pubertal mouse
mammary gland (15).
In the present study, we evaluated the expression
level of the FUT4 gene in the MCF-7 and MDA-MB-231 cell lines and
clinical breast cancer samples. We also investigated whether FUT4
participates in the regulation of tumor invasion and
tumorigenicity. In addition, the present study aimed to determine
the association between miR-493-5p and FUT4 in human breast cancer,
in order to provide a better understanding of the mechanisms
underlying breast cancer invasion and tumorigenicity.
Materials and methods
Cell culture
Cell lines MDA-MB-231 and MCF-7 were obtained from
KeyGen Co. (Nanjing, China) and cultured in Dulbecco's modified
Eagle's medium (DMEM), 10% fetal bovine serum (FBS) and 1%
penicillin-streptomycin (both from Gibco, Grand Island, NY, USA) at
37°C in a humidified atmosphere under 5% CO2.
Patient tissue collection and RNA
extraction
Twenty-nine breast cancer and matched adjacent
tissue samples were resected from patients at the Second Affiliated
Hospital of Dalian Medical University (Dalian, China) from July
2011 to June 2014. Informed consent forms and the entire protocol
were approved by the Ethics Committee of the Second Affiliated
Hospital of Dalian Medical University. The samples were stored at
−80°C, and total RNA was extracted with TRIzol reagent (Invitrogen,
Carlsbad, CA, USA).
Quantitative real-time PCR analysis
The concentration and quality of each total RNA
sample was determined using A260/A280 spectrophotometric reading.
cDNA was synthesized with TaqMan reverse transcription reagents
(Applied Biosystems, Branchbury, NJ, USA), following the
manufacturer's recommendations. Real-time PCR was carried out using
7500 Fast Real-time PCR system (Applied Biosystems). Reactions were
run in 3 independent experiments. The relative expression level of
FUT4 was normalized to GAPDH. The primer sequences were:
5′-TCCTACGGAGAGGCTCAG-3′ and 5′-TCCTCGTAGTCCAACACG-3′. RT-PCR for
miR-493-5p was performed using Real-Time PCR Universal Reagent
(GenePharma, Shanghai, China). U6 was used as an internal
control.
Western blot analysis
Protein was extracted from the cells using 1X
radioimmunoprecipitation assay lysis buffer (Santa Cruz
Biotechnology, Santa Cruz, CA, USA), subjected to SDS-PAGE, and
then transferred to polyvinylidene difluoride membranes. The
membranes were blocked in 5% skimmed milk for 2 h, probed with the
antibody against human FUT4 (1:1,000 dilution; Abcam, Cambridge,
UK) or GAPDH (1:1,000 dilution) at 4°C overnight, and with
peroxidase-conjugated secondary antibody (1:1,000 dilution) (both
from Santa Cruz Biotechnology), and then visualized by
chemiluminescence (GE Healthcare, Fairfield, CT, USA).
In vivo tumorigenesis
All animal experiments were performed according to
the protocol of the Dalian Committee on Animal Care using 5- to
6-week-old male athymic nude mice. Cells (1×107) were
subcutaneously injected into the right flank of each nude mouse.
The length (L) and width (W) of each tumor were measured every 7
days with calipers, and the volume was calculated.
Lentivirus production and infection
The FUT4 coding sequence (CDS) was obtained and
inserted into the NotI and BamHI sites of the
pGLV5/H1/GFP+Puro lentiviral plasmid, respectively. The FUT4 shRNA
sequences were inserted into the BamHI and EcoRI
sites of the pGLV3/H1/GFP+Puro lentiviral plasmid. Lentiviral
plasmids were co-transfected with PG-P1-VSVG, PG-P2-REV and
PG-P3-RRE plasmids into 293T cells (Invitrogen), and
virus-containing supernatants were prepared according to the
manufacturer's instructions. For the lentiviral infection, the
cells cultured in 6-well tissue culture plates were infected with
the lentiviral vectors at a multiplicity of infection of 40 for 24
h. The medium was replaced with fresh complete medium. After 2
days, the cells were observed by fluorescence microscopy to confirm
that >90% of the cells were GFP-positive. Subsequently, the
GFP-positive cells were screened by addition of 5 µg/ml
puromycin.
In vitro ECM invasion assay
Cell invasion was assessed using the Matrigel
invasion chamber (Corning, Corning, NY, USA) in triplicate. The
cells (1.0×105) were harvested in serum-free medium
containing 0.1% BSA and plated to the upper chamber precoated with
Matrigel. Medium containing 10% FBS in the lower chamber served as
the chemoattractant. After the cells were incubated at 37°C in a
humidified incubator with 5% CO2 for 24 h, the
non-invading cells were removed with cotton swabs. The invasive
cells that had attached to the lower surface of the inserted
membrane were fixed in 100% methanol at room temperature and
stained with Wright-Giemsa. The number of invasive cells on the
lower surface of the membrane was then counted under a
microscope.
Luciferase reporter assay
A pmirGLO Dual-Luciferase miRNA target expression
vector was used for 3′UTR luciferase assays (Promega, Madison, WI,
USA). The target genes of miRNA-493-5p were selected based on
target scan algorithms (microRNA.org
(http://www.microrna.org/microrna/home.do), MicroCosm
(http://www.ebi.ac.uk/enright-srv/microcosm/htdocs/targets/)
and TargetScan (http://www.targetscan.org/). For 3′UTR luciferase
assay, the cells were co-transfected with hsa-miR-493-5p mimics or
pmirGLO Dual-Luciferase miRNA target expression vectors and the
wild-type or mutant target sequence using Lipofectamine 2000.
Luciferase assay was performed using the
Dual-Luciferase® reporter assay system (Promega) after
transfection at 48 h. Data are presented as the mean value ± SD for
triplicate experiments.
Statistical analysis
Statistical analyses were carried out using the SPSS
17.0 program. An independent sample t-test was run to analyze the
significance of the differences found. P-value <0.05 was
considered to indicate a statistically significant result. All data
are presented as the mean ± SD.
Results
Expression of FUT4 in human breast cancer
cell lines and tissues
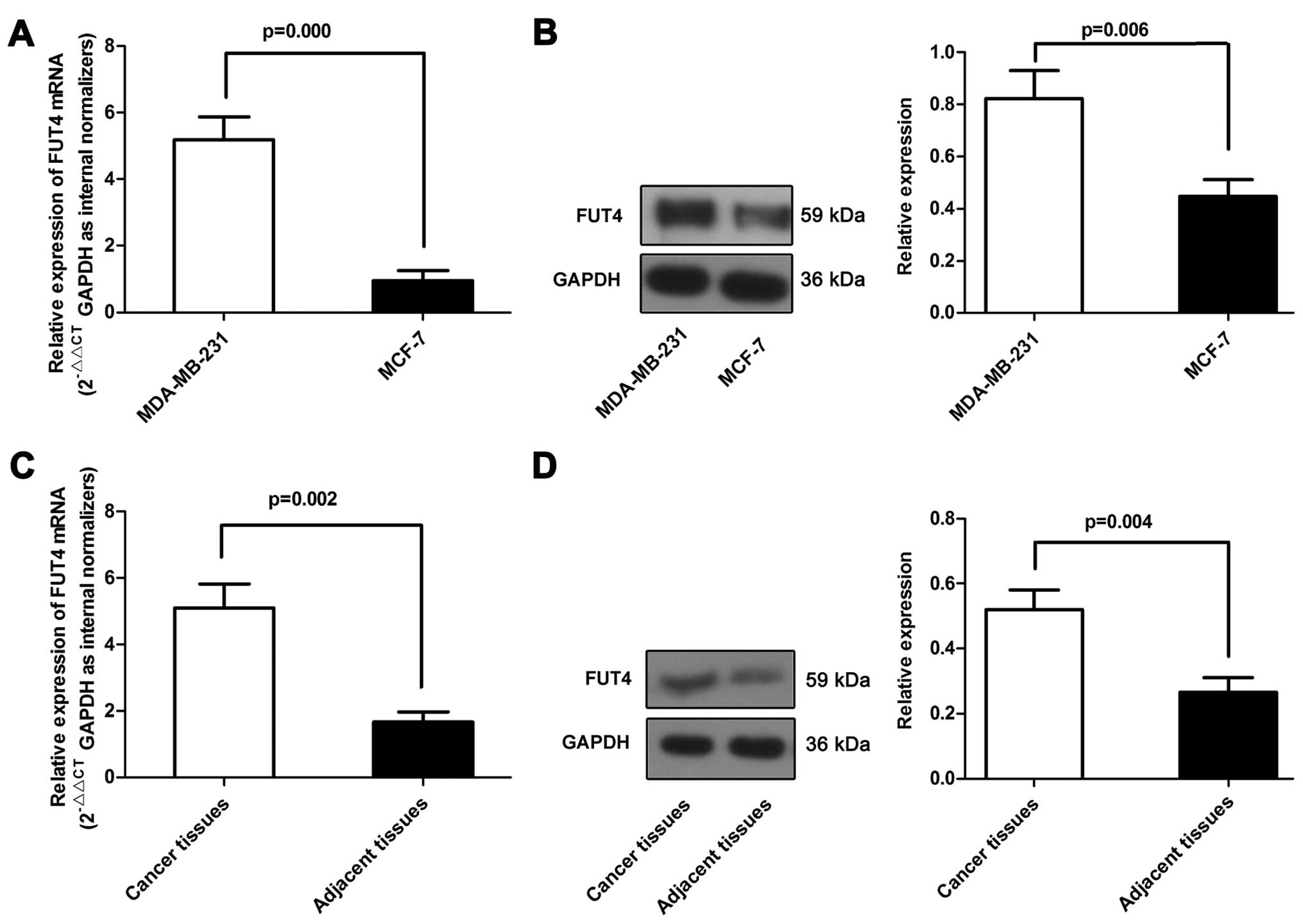
We first determined the expression of FUT4 in breast
cancer cell lines, primary breast cancer and the matched adjacent
tissue samples. As shown in Fig. 1A and
B, the FUT4 (5.76-fold) expression level was significantly
higher in the MDA-MB-231 cell line than that in the MCF-7 cell
line. Furthermore, analysis of FUT4 expression in pairs of the
primary breast cancer and matched adjacent tissue samples revealed
that FUT4 was upregulated in the primary breast cancer tissue
samples (Fig. 1C and D). These data
support the assumption that the differential expression of FUT4 may
be associated with breast cancer.
Effect of FUT4 overexpression or
silencing on the invasive ability and tumorigenicity of breast
cancer cells
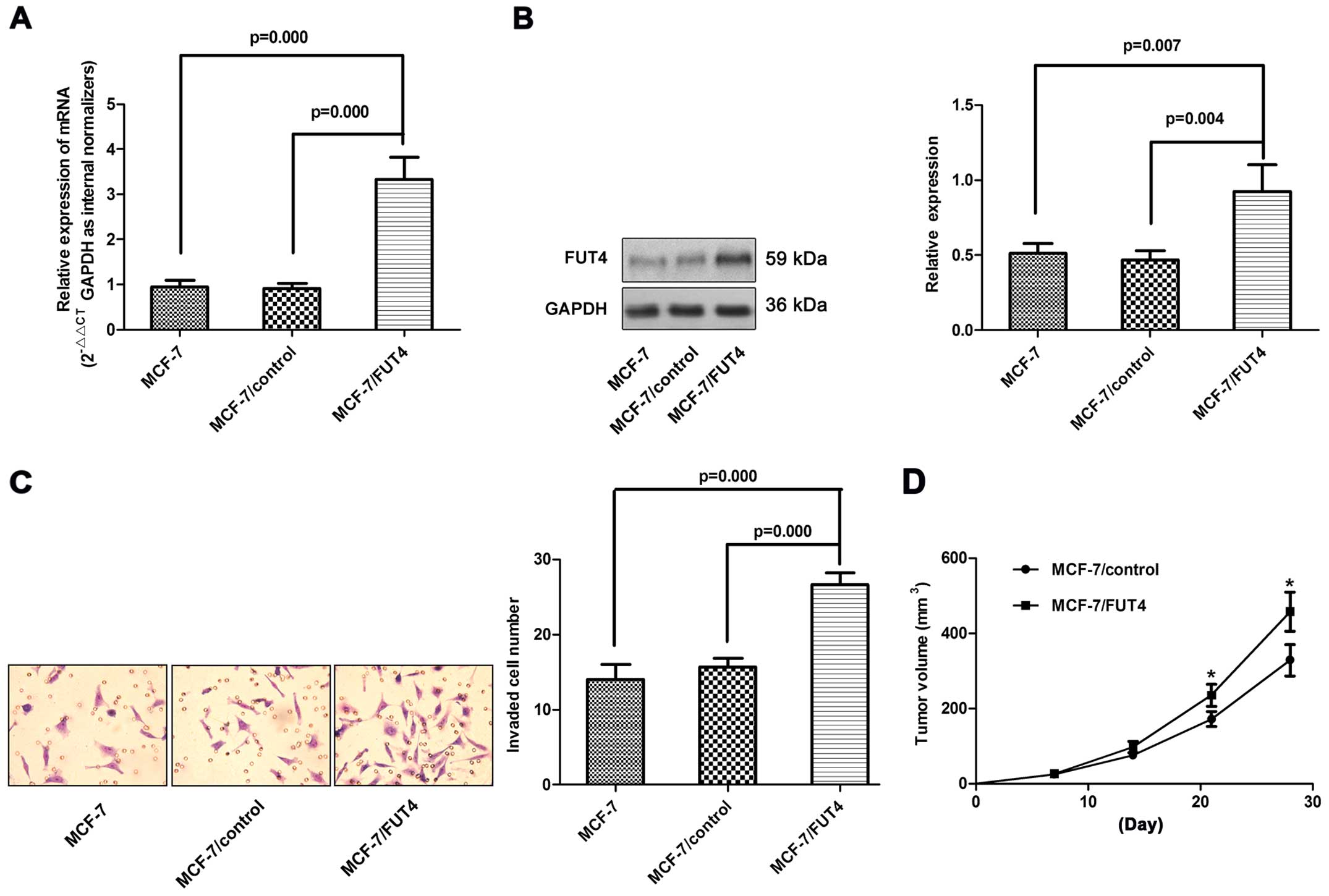
To test whether FUT4 plays a role in regulating
invasion, we infected MCF-7 cells with lentiviral vectors
containing FUT4. After infection, the MCF-7/FUT4 cells expressed
high exogenous FUT4 (Fig. 2A and B)
in comparison with the uninfected controls or GFP-expressing
cells.
Overexpression of FUT4 increased the invasive
ability of the MCF-7 cells in vitro (Fig. 2C). In order to further confirm that
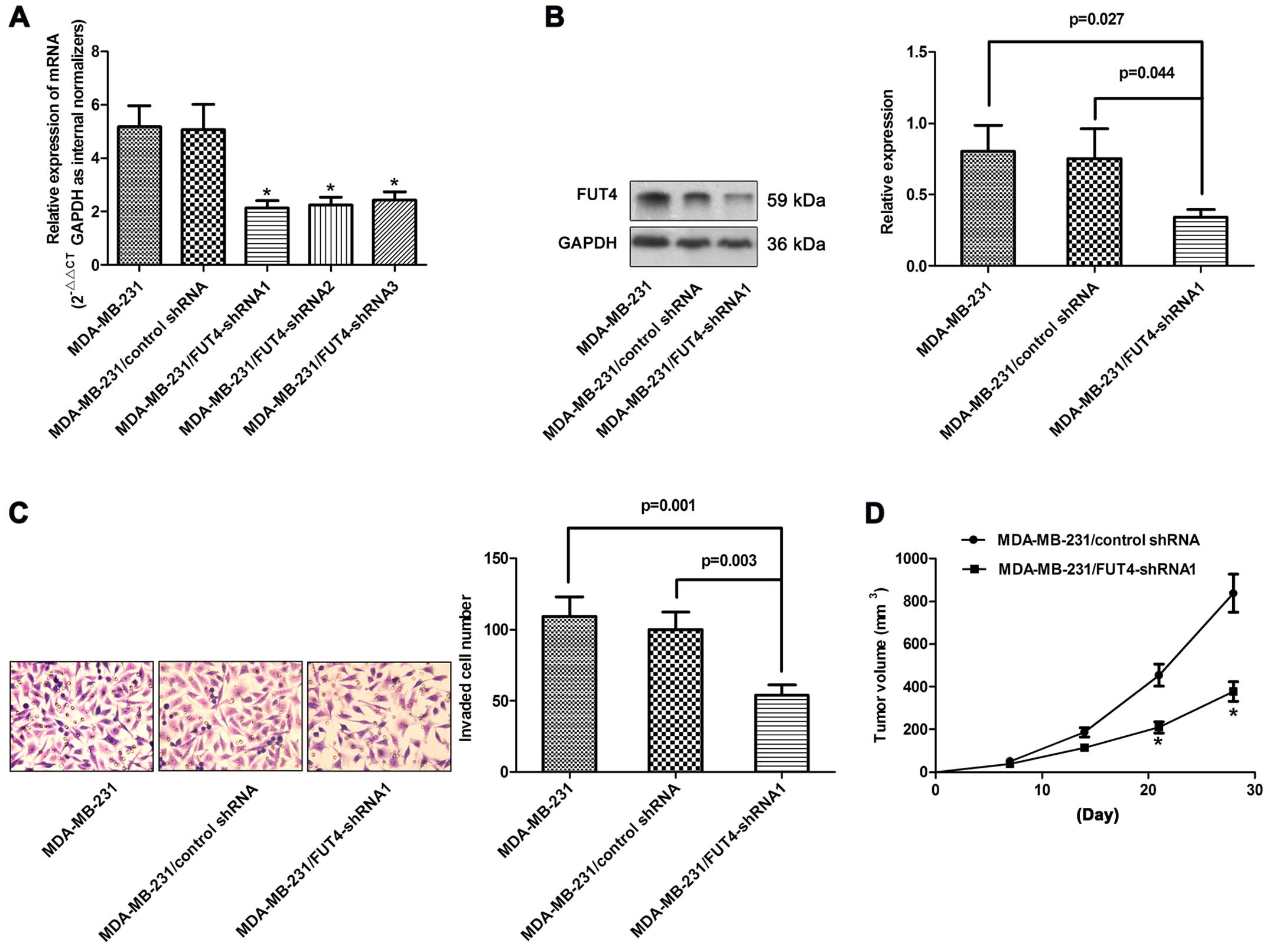
FUT4 affects invasive ability, we used shRNA delivered in a
lentiviral vector tagged with GFP to regulate FUT4 expression in
the MDA-MB-231 cells. Three different shRNA lentiviruses
significantly reduced the FUT4 expression >60% at the mRNA level
and >50% at the protein level (Fig.
3A and B). The invasive ability was significantly decreased to
55% in the MDA-MB-231/FUT4-shRNA cells compared with the invasive
ability noted in the control cells (Fig. 3C).
In order to identify whether FUT4 further affects
the tumorigenicity in vivo, MCF-7/control, MCF-7/FUT4,
MDA-MB-231/control shRNA and MDA-MB-231/FUT4-shRNA1 were
subcutaneously injected into nude mice and tumor formation was
monitored. On day 28, the mice were sacrificed under anesthesia and
tumor weights were measured. Tumors grew faster in the MCF-7/FUT4
and MDA-MB-231/control shRNA groups when compared with the rates in
the MCF-7/control and MDA-MB-231/FUT4-shRNA1 groups, respectively
(Figs. 2D and 3D). These data suggest that FUT4 enhanced
the cell growth in vivo.
Differential expression of miR-493-5p in
breast cancer cell lines and tissues
To understand the mechanisms by which miRNAs execute
their function by FUT4, we adopted 3 bioinformatic algorithms
(TargetScan, MicroCosm and miRanda) to identify a large number of
potential miRNAs. Among these candidates, miR-493-5p was selected
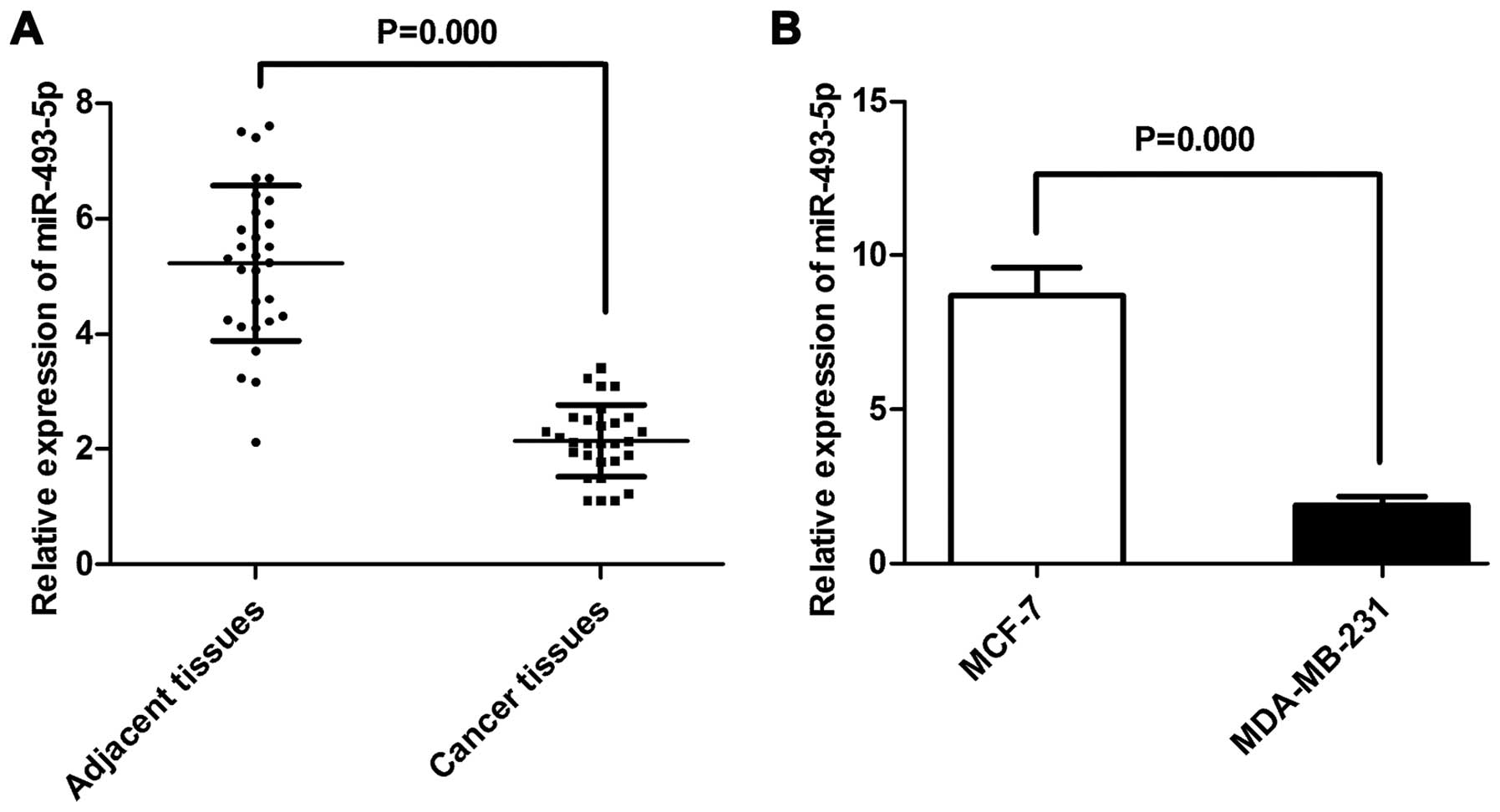
for further study. Total RNA was isolated from the cell lines, the
breast cancer tissue and the matched adjacent tissue samples, and
the miR-493-5p levels were determined using TaqMan real-time PCR.
The miR-493-5p expression level was higher in the MCF-7 cells than
that in the MDA-MB-231 cells, while the tissue samples showed the
same tendency (Fig. 4A and B).
miR-493-5p directly targets and inhibits
FUT4 expression
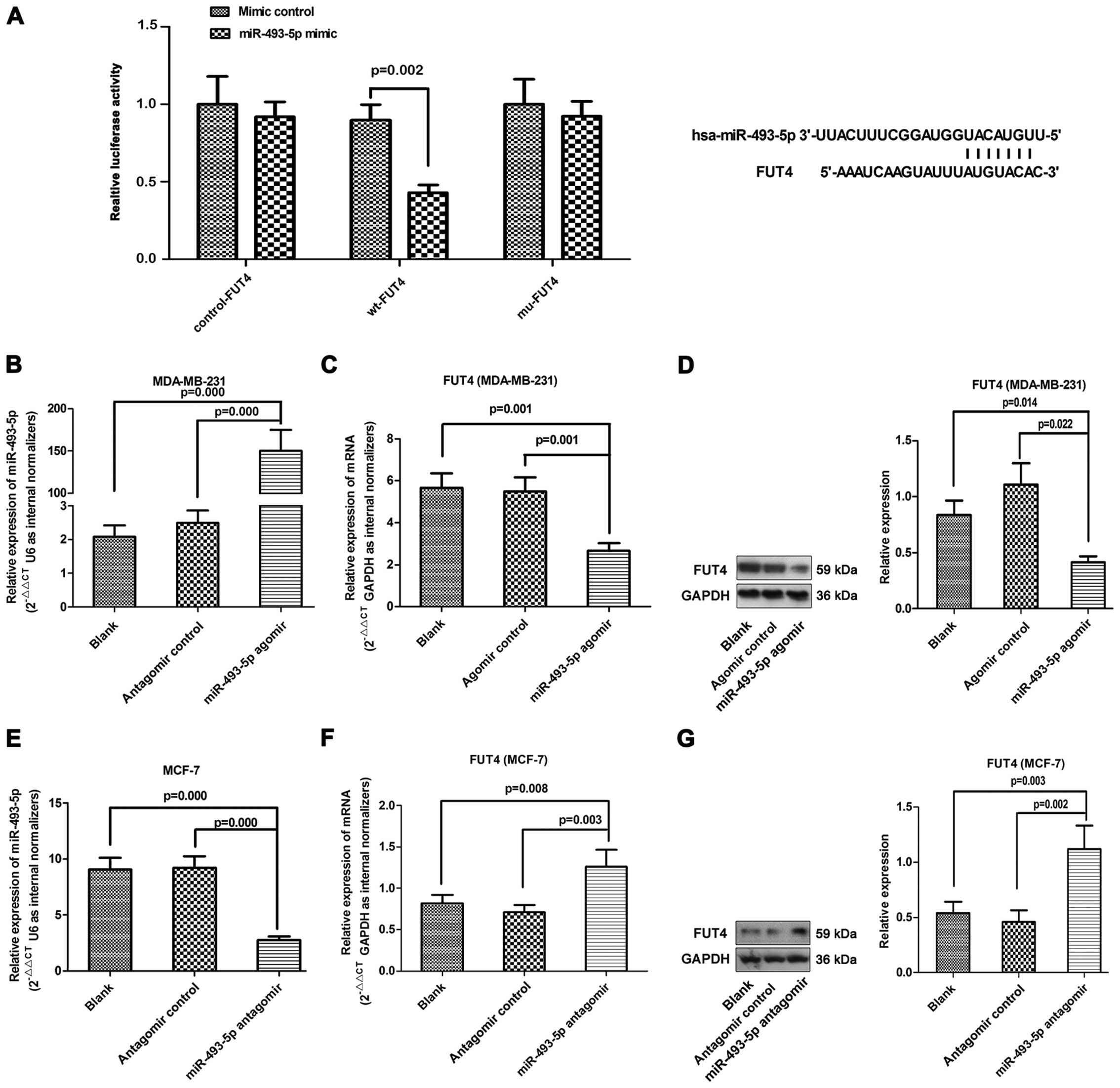
To test whether FUT4 is a direct target of
miR-493-5p, luciferase activity assays were performed using
luciferase reporters. The 3′-untranslated region (3′-UTR) of the
FUT4-mRNA, which contains a predicted target site for miR-493-5p,
was cloned into the GP-miRGLO plasmid. In addition, 3 nucleotides
within the predicted miR-493-5p target site were mutated in the
GP-miRGLO-FUT4-3′-UTR mutant plasmid. When the 3′-UTR of the FUT4
mRNA was involved in the luciferase transcript, forced miR-493-5p
expression decreased luciferase activity by 35% (Fig. 5A), whereas, miR-493-5p mimics had no
effect on the luciferase activity of the reporters containing a
mutant FUT4 3′-UTR. The results indicated that miR-493-5p targeted
the 3′-UTR of the FUT4 mRNA and repressed its expression.
To determine whether FUT4 expression is indeed
regulated by miR-493-5p in vitro, the MDA-MB-231 cells were
transfected with miR-493-5p agomir, while the MCF-7 cells were
transfected with the miR-493-5p antagomir. As expected, miR-493-5p
agomir-treated cells showed higher expression of miR-493-5p, and
miR-493-5p antagomir inhibited miR-493-5p expression (Fig. 5B and E). The MDA-MB-231 cell line
with miR-493-5p expression showed a significant attenuation of FUT4
expression at both the mRNA and protein levels in vitro.
Moreover, inhibition of miR-493-5p in MCF-7 cells led to the
opposite changes in FUT4 expression (Fig. 5C, D, F and G).
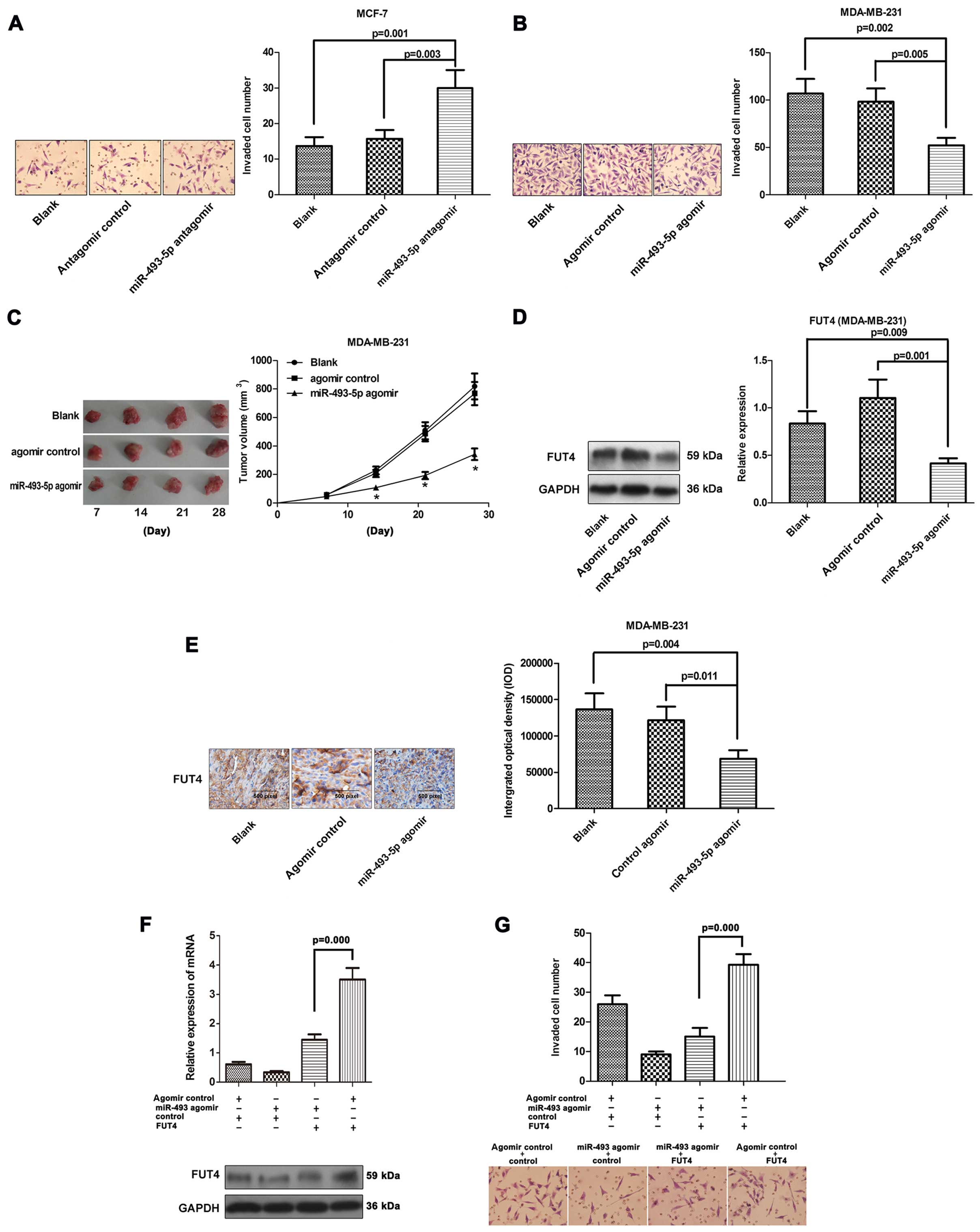
miR-493-5p attenuates the invasiveness in
vitro and the tumorigenicity in human breast cancer by targeting
FUT4 in vivo
Since the expression levels of miR-493-5p were lower
in the MDA-MB-231 cells and breast cancer tissues when compared
with the levels in MCF-7 cells and matched adjacent tissues, we
examined how alterations in miR-493-5p influence metastatic
behavior. MCF-7 and MDA-MB-231 cells were transfected with
antagomir or agomir of miR-493-5p, respectively. The effects of the
antagomir and the agomir on the invasive rate of MDA-MB-231 and
MCF-7 cells were studied with Transwell assay (Fig. 6A and B). Forced miRNA-493-5p
overexpression resulted in 41% reduction in the invasive potential
of the MDA-MB-231 cells compared to the control cells. The invasive
potential was increased in the MCF-7 cells transfected with the
antagomir. The enhanced expression of FUT4 significantly increased
or reversed the invasive capabilities of the MCF-7 cells (Fig. 6F and G).
To ascertain whether miR-493-5p could further affect
the tumorigenicity in vivo, MDA-MB-231, MDA-MB-231/NC and
MDA-MB-231/agomir cells were injected into nude mice and tumor
formation was monitored. The tumor volume in the MDA-MB-231/agomir
group was decreased compared to that noted in the MDA-MB-231 and
MDA-MB-231/NC groups (Fig. 6C).
Downregulation of FUT4 was also observed in the MDA-MB-231/agomir
group (Fig. 6D and E).
Discussion
Changes in fucosylation can be observed in nearly
all malignancies, such as breast (16), HCC (17), colon (18) and lung cancer (19). High incidence of sLex in ER-negative
breast tumors coincided with high expression of
glycosyltransferases FUT3, FUT4 and ST3GAL6 (20). Previous studies have demonstrated a
significant high expression of FUT4 in breast cancer tissues and
sera in comparison to normal tissues and sera (21). The present study revealed that the
expression profile of the FUT4 gene varied between MDA-MB-231 and
MCF-7 cell lines with different metastatic potential. Compared to
the MCF-7 cells, MDA-MB-231 cells showed upregulated expression of
FUT4 (5.76-fold). Comparison of FUT4 expression levels in 29 pairs
of clinical primary breast cancer tissues and their matched
adjacent tissues revealed that FUT4 was upregulated in the primary
breast cancer tissues. FUT4 with different expression patterns in
the two cell lines and breast cancer tissues imply its role as an
indicator and functional regulator of breast cancer metastasis.
A previous study found that stable transfection of
antisense sequences directed at the human Lewis α (1,3/1,4) FUT
gene, FUT3 led to the inhibition of E-selectin-mediated
adenocarcinoma cell adhesion (22).
Elevated expression of FUT3/6 conferred increased sLex production.
sLex+ cells (with high FUT expression levels) were
significantly more invasive than sLex− cells (with low
FUTs). This suggests an association between FUT expression and
invasive activity. Inactivation of the expression of FUT1 and FUT2
reduced cell adhesion efficiency and inhibited cell growth
(23). FUT6, a member of the a1,3/4
FUT subfamily, was found to regulate the P13K/AKT signaling pathway
in HCC cells (24). Whether the
FUT4 and its encoded protein affect tumor invasion, proliferation
and tumorigenicity in breast cancer remains a question. The present
study aimed to explore FUT4. The altered level of FUT4 led to
mediated invasiveness and tumorigenicity in MCF-7 and MDA-MB-231
cell lines both in vitro and in vivo. Taken together,
these data suggest that fucosylation is closely related to the
activities of invasion and proliferation in breast cancer
cells.
MicroRNAs (miRNAs) are attractive candidates as
upstream regulators of metastatic progression since miRNAs can
post-transcriptionally regulate entire sets of genes (25,26).
Recent research has revealed the vital roles of miRNAs in
tumorigenesis (27,28). miRNAs often act as oncogenes or
tumor suppressors, regulating many cellular processes. miR-335 was
found to suppress metastasis and migration through targeting of the
progenitor cell transcription factor SOX4 and extracellular matrix
component tenascin C (29). A few
miRNAs have been identified to target glycosylation enzymes to
date. miR-378 targeted UDP-N-acetyl-D-galactosamine:
polypeptide N-acetylgalactosaminyltransferase 7 and enhanced the
glycosylation of nephronectin, a ligand for integrin α8β1 (30). miR-148 targeted the enzyme core 1
β1, 3-galactosyltrans-ferase 1 and modulated IgA1
O-glycosylation and the level of secreted
galactose-deficient IgA1 (31).
miR-199a targeted the glycosylation enzyme ST6GAL1, reduced the
sialylation of Necl-2, indirectly reduced the protein level of
Necl-2, and enhanced ErbB2/ErbB3 signaling (32). It was reported that miR-493
inhibited the formation of metastatic foci and thus prevented liver
metastasis of colon cancer cells (33). miR-493, was found to be reduced in
lung cancer cells and tissues and to suppress cell invasion and
proliferation by blocking E2F1 (34). miR-493 inhibited cell invasion and
migration by blocking FZD4 and RhoC in bladder cancer (35).
miR-493-5p is prominently upregulated in male breast
cancer (36). However, miR-493, as
analyzed by NanoString's nCounter human miRNA expression profiling,
was down-regulated in TNBCs vs. normal breast tissues (37). The role of miR-493-5p remains
unclear in breast cancer cell lines and in breast cancer at
present. In the present study, we showed that the level of
miR-493-5p, a potentially candidate tumor-suppressor miRNA, was low
in MDA-MB-231 cells and breast cancer tissues. Introduction of
miR-493-5p into MDA-MB-231 cells resulted in a reduction in tumor
cell invasion and tumorigenicity. Furthermore, miR-493-5p was found
to target the 3′-UTR of FUT4, causing decreased FUT4 expression at
the mRNA and protein levels, thus controlling the invasiveness of
breast cancer cells. This result indicated that miR-493-5p may
serve as a specific modulator of human breast cancer invasion and
tumorigenicity through targeting FUT4.
In summary, the FUT4 gene was demonstrated to
regulate the invasiveness and tumorigenicity of human breast
cancer. The aberrant expression of miR-493-5p was associated with
breast cancer invasiveness and tumorigenicity. Moreover, we found
that miR-493-5p targeted the 3′-UTR of FUT4 mRNA and reduced the
protein expression level of FUT4. Our results provide new insight
into the mechanisms underlying the invasion and tumorigenicity of
breast cancer, and the application of miR-493-5p may be a potential
strategy for breast cancer therapeutics.
Acknowledgments
The present study was supported by grants from the
National Natural Science Foundation of China (81271910, 81472014),
and from the Natural Science Foundation of Liaoning Province, China
(2014023043).
References
|
1
|
Jemal A, Siegel R, Ward E, Murray T, Xu J
and Thun MJ: Cancer statistics, 2007. CA Cancer J Clin. 57:43–66.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
He D, Wang C, Cao L, Zhou J, Huang J and
Fang H: Epidemic trend of female breast cancer incidence in Min
Hang District, Shanghai. China Cancer. 13:108–110. 2010.
|
|
3
|
Stellrecht CM, Vangapandu HV, Le XF, Mao W
and Shentu S: ATP directed agent, 8-chloro-adenosine, induces AMP
activated protein kinase activity, leading to autophagic cell death
in breast cancer cells. J Hematol Oncol. 7:232014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Apweiler R, Hermjakob H and Sharon N: On
the frequency of protein glycosylation, as deduced from analysis of
the SWISS-PROT database. Biochim Biophys Acta. 1473:4–8. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Spiro RG: Protein glycosylation: Nature,
distribution, enzymatic formation, and disease implications of
glycopeptide bonds. Glycobiology. 12:43R–56R. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Desiderio V, Papagerakis P, Tirino V,
Zheng L, Matossian M, Prince ME, Paino F, Mele L, Papaccio F,
Montella R, et al: Increased fucosylation has a pivotal role in
invasive and metastatic properties of head and neck cancer stem
cells. Oncotarget. 6:71–84. 2015.
|
|
7
|
Wang QY, Guo P, Duan LL, Shen ZH and Chen
HL: α-1,3-Fucosyltransferase-VII stimulates the growth of
hepatocarcinoma cells via the cyclin-dependent kinase inhibitor
p27Kip1. Cell Mol Life Sci. 62:171–178. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hiller KM, Mayben JP, Bendt KM, Manousos
GA, Senger K, Cameron HS and Weston BW: Transfection of alpha(1,3)
fucosyltransferase antisense sequences impairs the proliferative
and tumorigenic ability of human colon carcinoma cells. Mol
Carcinog. 27:280–288. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sanchez-Mejias A and Tay Y: Competing
endogenous RNA networks: Tying the essential knots for cancer
biology and therapeutics. J Hematol Oncol. 8:302015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Thomson DW, Bracken CP and Goodall GJ:
Experimental strategies for microRNA target identification. Nucleic
Acids Res. 39:6845–6853. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Calin GA and Croce CM: MicroRNA-cancer
connection: The beginning of a new tale. Cancer Res. 66:7390–7394.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Croce CM: miRNAs in the spotlight:
Understanding cancer gene dependency. Nat Med. 17:935–936. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Hurst DR, Edmonds MD, Scott GK, Benz CC,
Vaidya KS and Welch DR: Breast cancer metastasis suppressor 1
up-regulates miR-146, which suppresses breast cancer metastasis.
Cancer Res. 69:1279–1283. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Liu S, Goldstein RH, Scepansky EM and
Rosenblatt M: Inhibition of rho-associated kinase signaling
prevents breast cancer metastasis to human bone. Cancer Res.
69:8742–8751. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Phua YW, Nguyen A, Roden DL, Elsworth B,
Deng N, Nikolic I, Yang J, Mcfarland A, Russell R, Kaplan W, et al:
MicroRNA profiling of the pubertal mouse mammary gland identifies
miR-184 as a candidate breast tumour suppressor gene. Breast Cancer
Res. 17:832015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lattová E, Tomanek B, Bartusik D and
Perreault H: N-glycomic changes in human breast carcinoma MCF-7 and
T-lymphoblastoid cells after treatment with herceptin and
herceptin/Lipoplex. J Proteome Res. 9:1533–1540. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Shu H, Zhang S, Kang X, Li S, Qin X, Sun
C, Lu H and Liu Y: Protein expression and fucosylated glycans of
the serum haptoglobin-β subunit in hepatitis B virus-based liver
diseases. Acta Biochim Biophys Sin. 43:528–534. 2011. View Article : Google Scholar
|
|
18
|
Mejías-Luque R, López-Ferrer A, Garrido M,
Fabra A and de Bolós C: Changes in the invasive and metastatic
capacities of HT-29/M3 cells induced by the expression of
fucosyltransferase 1. Cancer Sci. 98:1000–1005. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Vasseur JA, Goetz JA, Alley WR Jr and
Novotny MV: Smoking and lung cancer-induced changes in
N-glycosylation of blood serum proteins. Glycobiology.
22:1684–1708. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Julien S, Ivetic A, Grigoriadis A, QiZe D,
Burford B, Sproviero D, Picco G, Gillett C, Papp SL, Schaffer L, et
al: Selectin ligand sialyl-Lewis x antigen drives metastasis of
hormone-dependent breast cancers. Cancer Res. 71:7683–7693. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yan X, Lin Y, Liu S, Aziz F, Yan Q and
Fucosyltransferase IV: (FUT4) as an effective biomarker for the
diagnosis of breast cancer. Biomed Pharmacother. 70:299–304. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Weston BW, Hiller KM, Mayben JP, Manousos
GA, Bendt KM, Liu R and Cusack JC Jr: Expression of human
alpha(1,3)fucosyltransferase antisense sequences inhibits
selectin-mediated adhesion and liver metastasis of colon carcinoma
cells. Cancer Res. 59:2127–2135. 1999.PubMed/NCBI
|
|
23
|
Palumberi D, Aldi S, Ermini L, Ziche M,
Finetti F, Donnini S and Rosati F: RNA-mediated gene silencing of
FUT1 and FUT2 influences expression and activities of bovine and
human fucosylated nucleolin and inhibits cell adhesion and
proliferation. J Cell Biochem. 111:229–238. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Guo Q, Guo B, Wang Y, Wu J, Jiang W, Zhao
S, Qiao S and Wu Y: Functional analysis of
α1,3/4-fucosyltransferase VI in human hepatocellular carcinoma
cells. Biochem Biophys Res Commun. 417:311–317. 2012. View Article : Google Scholar
|
|
25
|
Lim LP, Lau NC, Garrett-Engele P, Grimson
A, Schelter JM, Castle J, Bartel DP, Linsley PS and Johnson JM:
Microarray analysis shows that some microRNAs downregulate large
numbers of target mRNAs. Nature. 433:769–773. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Naidu S, Magee P and Garofalo M:
MiRNA-based therapeutic intervention of cancer. J Hematol Oncol.
8:682015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Voorhoeve PM, le Sage C, Schrier M, Gillis
AJ, Stoop H, Nagel R, Liu YP, van Duijse J, Drost J, Griekspoor A,
et al: A genetic screen implicates miRNA-372 and miRNA-373 as
oncogenes in testicular germ cell tumors. Cell. 124:1169–1181.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ma L, Teruya-Feldstein J and Weinberg RA:
Tumour invasion and metastasis initiated by microRNA-10b in breast
cancer. Nature. 449:682–688. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Tavazoie SF, Alarcón C, Oskarsson T, Padua
D, Wang Q, Bos PD, Gerald WL and Massagué J: Endogenous human
microRNAs that suppress breast cancer metastasis. Nature.
451:147–152. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kahai S, Lee SC, Lee DY, Yang J, Li M,
Wang CH, Jiang Z, Zhang Y, Peng C and Yang BB: MicroRNA miR-378
regulates nephronectin expression modulating osteoblast
differentiation by targeting GalNT-7. PLoS One. 4:e75352009.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Serino G, Sallustio F, Cox SN, Pesce F and
Schena FP: Abnormal miR-148b expression promotes aberrant
glycosylation of IgA1 in IgA nephropathy. J Am Soc Nephrol.
23:814–824. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Minami A, Shimono Y, Mizutani K, Nobutani
K, Momose K, Azuma T and Takai Y: Reduction of the ST6
β-galactosamide α-2,6-sialyltransferase 1 (ST6GAL1)-catalyzed
sialylation of nectin-like molecule 2/cell adhesion molecule 1 and
enhancement of ErbB2/ErbB3 signaling by microRNA-199a. J Biol Chem.
288:11845–11853. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Okamoto K, Ishiguro T, Midorikawa Y, Ohata
H, Izumiya M, Tsuchiya N, Sato A, Sakai H and Nakagama H: miR-493
induction during carcinogenesis blocks metastatic settlement of
colon cancer cells in liver. EMBO J. 31:1752–1763. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Gu Y, Cheng Y, Song Y, Zhang Z, Deng M,
Wang C, Zheng G and He Z: MicroRNA-493 suppresses tumor growth,
invasion and metastasis of lung cancer by regulating E2F1. PLoS
One. 9:e1026022014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Ueno K, Hirata H, Majid S, Yamamura S,
Shahryari V, Tabatabai ZL, Hinoda Y and Dahiya R: Tumor suppressor
microRNA-493 decreases cell motility and migration ability in human
bladder cancer cells by downregulating RhoC and FZD4. Mol Cancer
Ther. 11:244–253. 2012. View Article : Google Scholar
|
|
36
|
Lehmann U, Streichert T, Otto B, Albat C,
Hasemeier B, Christgen H, Schipper E, Hille U, Kreipe HH and Länger
F: Identification of differentially expressed microRNAs in human
male breast cancer. BMC Cancer. 10:1092010. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Cascione L, Gasparini P, Lovat F, Carasi
S, Pulvirenti A, Ferro A, Alder H, He G, Vecchione A, Croce CM, et
al: Integrated microRNA and mRNA signatures associated with
survival in triple negative breast cancer. PLoS One. 8:e559102013.
View Article : Google Scholar : PubMed/NCBI
|