Introduction
Colorectal cancer (CRC) is one of the most common
malignancies and the incidence has increased significantly in
recent years (1), particularly in
developing countries such as China, where the annual increase of
4.71% exceeds the global increase rate of 2% (2). To date, the etiology and pathogenesis
of CRC have remained to be fully elucidated. Previous studies have
suggested that genetic factors are involved in 5–15% of all cases
of CRC (3). The majority of the
remaining cases, which are classified as sporadic CRC, are affected
by environmental factors (4),
including age, gender, diet, obesity and levels of physical
inactivity. One of the common characteristics of these potential
risk factors is that they can cause alterations in the structure of
the intestinal flora (5–8). These alterations (also called
dysbiosis) were hypothesized to be closely correlated with the
pathogenesis of CRC (9–11).
The human intestine harbors ~1014
microbes, whose genetic content is 100 times higher than that of
the human genome (12). The
intestine and its microbe population are collectively referred to
as a super-organism, which constitutes a large and complex
ecosystem, and is directly involved in functions including
digestion and nutrient absorption, energy supply, fat metabolism,
immune regulation and disease resistance (13,14).
The microflora in the human gut contains beneficial commensal
bacteria and certain potential pathogenic bacteria. These two types
of bacteria interact with their human host, leading to
physiological inflammation, which can prevent diseases including
inflammatory bowel disease (IBD) and CRC. However, when these
potentially pathogenic bacteria increase in number or their
virulence is enhanced, this physiological inflammation is likely to
become a pathological inflammation and possibly cancer-causing
(15,16). Accordingly, the developments of
methods to increase the number of beneficial bacteria or suppress
harmful bacteria would be advantageous (17).
The term probiotics refers to preparations
containing a class of active microorganisms in the human intestine
that can bring health benefits to the host, such as improvement of
the host micro-ecological balance (18,19).
The majority of the probiotics available at present include lactic
acid bacteria (20), in addition
to the genera Lactobacillus, Bifidobacterium and
Enterococcus (21). Studies
in animal models and humans have demonstrated that the consumption
of probiotics is effective in various medical conditions, can delay
senility (22), and improve acute
infectious diarrhea (23,24), antibiotic-associated diarrhea
(21,25), constipation (26–28),
ulcerative colitis (29,30) and Crohn's disease (31,32).
However, the majority of these previous studies focused on the
interaction between probiotics and the mammalian host. It has
remained elusive whether probiotics serve a beneficial role by
adjusting intestinal microbes or inhibiting certain detrimental
pathogens. In the light of these previous studies, it was suggested
that it may be feasible to reduce the incidence of CRC by the
application of probiotics and optimization of the gut microflora
(17). The aim of the present
study was to assess whether perioperative oral probiotics were able
to alter the microbial composition and improve gut microecology in
patients with CRC.
Materials and methods
Patients
A total of 60 patients with CRC who were due to
undergo radical colorectomy at Shanghai Jiao Tong University
Affiliated Shanghai Sixth People's Hospital (Shanghai, China), were
selected between September 2011 and December 2013. The inclusion
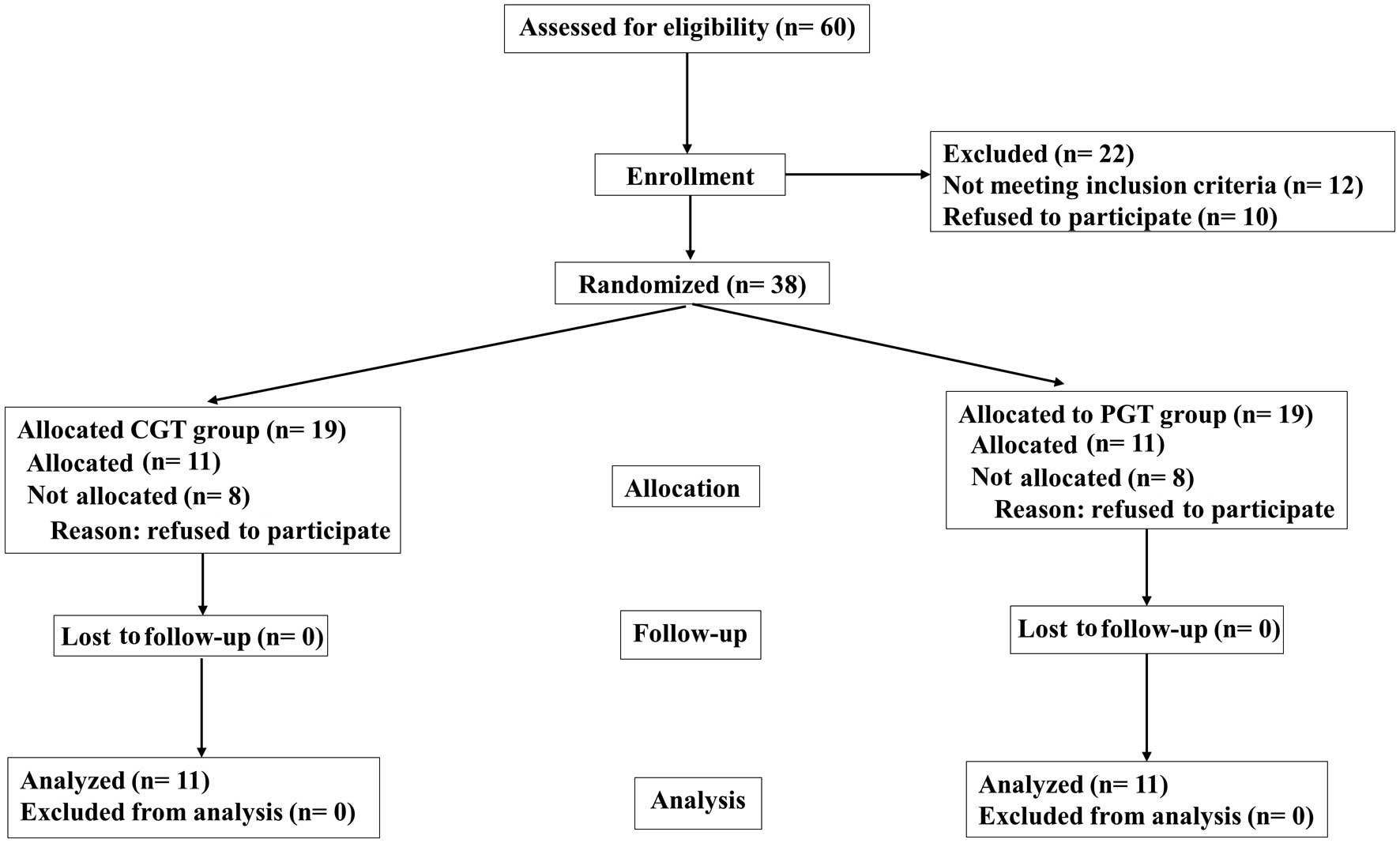
and exclusion criteria are listed in (Table I). A total of 60 patients were
assessed for eligibility; however, only 38 were finally enrolled
due to the fact that 22 failed to meet the inclusion criteria or
refused to participate in the study. A total of 22 patients
completed the trial (Fig. 1). All
eligible patients (n=22) were randomized and divided into two
treatment groups: The CGT group (n=11) received perioperative
placebos and the PGT group (n=11) received probiotics. There were
no significant differences in gender, age, body mass index (BMI),
cancer stage and time between onset of symptoms and hospital
admission between the two groups. No significant differences in the
levels of albumin, hemoglobin or creatinine were observed between
the two groups (Table II).
 | Table IInclusion and exclusion criteria for
enrolment of individuals in the present study. |
Table I
Inclusion and exclusion criteria for
enrolment of individuals in the present study.
| Colorectal cancer
patients | Healthy
individuals |
|---|
| Inclusion
criteria | Inclusion
criteria |
| Age 40–75
years | Age 40–75
years |
| Diagnosis
confirmed by biopsy and histological analysis | BMI 18.5–30
kg/m2 |
| Previous radical
resection and no distant metastasis (including liver) | |
| Exclusion
criteria | Exclusion
criteria |
| Age >75
years | BMI >30
kg/m2 |
| Pregnancy | Pregnancy |
| Known lactose
intolerance | Known lactose
intolerance |
| Clinically
significant immunodeficiency | Clinically
significant immunodeficiency |
| Usage of
antibiotics and additional gastrointestinal disorders (e.g. Crohn's
disease or ulcerative colitis) | Usage of
antibiotics and additional gastrointestinal disorders (e.g. Crohn's
disease or ulcerative colitis) |
| Received
antibiotics for the past 3 months prior to surgery | Received
antibiotics for the past 3 months prior to surgery |
| Evidence of
infection | Evidence of
infection |
| Probiotics or
prebiotics and excessive fiber intake within 2 weeks | Probiotics or
prebiotics and excessive fiber intake within 2 weeks |
| Patient subjected
to emergency operation | Patient subjected
to emergency operation |
| Bowel preparation
for colonoscopy within 6 days prior to surgery | Bowel preparation
for colonoscopy within 6 days prior to surgery |
| Patient subjected
to proctectomy with low rectal anastomosis or surgery for polypoid
lesion | Patient subjected
to proctectomy with low rectal anastomosis or surgery for polypoid
lesion |
| Laparoscopic
surgery | Laparoscopic
surgery |
| Patients received
preoperative chemotherapy or radiation therapy | History or
presence of other tumors |
 | Table IIStatistical data of subjects of the
present study. |
Table II
Statistical data of subjects of the
present study.
| Parameter | HGT | CGT | PGT | P-value |
|---|
| Sample type | Tissue | Tissue | Tissue | |
| Number of
individuals (n) | 11 | 11 | 11 | >0.05 |
| Male/female | 6/5 | 6/5 | 6/5 | >0.05 |
| Age (years) | 71±5.4 | 68±7.3 | 65±9.6 | >0.05 |
| BMI
(kg/m2) | 22.2±2.2 | 24.5±4.3 | 24.9±4.2 | >0.05 |
| Stage (A/B/C) | – | 4/4/3 | 3/5/3 | >0.05 |
| Location | | | | |
| Ascending colon
(n) | 3 | 2 | 2 | >0.05 |
| Descending colon
(n) | 1 | 1 | 2 | >0.05 |
| Sigmoid colon
(n) | 3 | 3 | 1 | >0.05 |
| Rectum (n) | 4 | 5 | 6 | >0.05 |
| Pre-operative
albumin (g/dl) | 42.2±2.6 | 36.5±3.4 | 40.1±3.3 | >0.05 |
| Pre-operative Hb
(g/l) | 126±12.4 | 123.2±19.6 | 125.3±17.7 | >0.05 |
| Creatinine
(mg/dl) | 1.1±0.2 | 1.2±0.13 | 1.1±0.16 | >0.05 |
In order to analyze further differences in the
microbiota between patients with CRC and the healthy population, an
additional 11 healthy volunteers (HGT group) were enrolled, who
were part of a larger population-based case-control study at the
Shanghai Jiao Tong University (Table
II). The inclusion and exclusion criteria for the controls are
listed in Table I.
Study design, probiotics treatment and
patient care
A single-center prospective randomized control study
design was used. Equal randomization was accomplished using a
computer-generated random allocation schedule. The probiotic and
placebo were sealed with aluminum foil and appeared identical in
the two groups, with an identical smell and taste. Only one nurse
not directly involved in the trial was able to solve the treatment
codes in the event of an emergency. An acid-resistant coating was
used to prepare the capsules containing the probiotics and placebo.
Patients in the PGT group received an encapsulated probiotics
preparation (Shanghai Xinyi Pharmaceutical Co., Ltd., Shanghai,
China) containing live combined Bifidobacterium longum,
Lactobacillus acidophilus and Enterococcus faecalis
(1:1:1) with no less than 1.0×107 CFU/g viable cells,
three times/day, with a total daily dose of 6.0×107 CFU
for five days. Patients in the CGT group received encapsulated
maltodextrin (Haiyan Liuhe Pharmaceutical Industry Co., Ltd.,
Jiangsu, China) three times a day. During the study period, no
parenteral or enteral nutritional supplementation was administered.
All patients received a regular diet preoperatively and
conventional bowel preparation was performed on pre-operative day
1, including the administration of a low-residue diet and
mechanical bowel preparation. All patients were given Soffodex
(Jiangxi Hygecon Pharmaceutical Co., Shangrao, China), containing
2.4 g monobasic sodium phosphate and 0.9 g dibasic sodium
phosphate. Parenteral hydration was administered on the morning of
the surgery supplied via a central venous catheter. A 12F catheter
was placed through a jejunal limb during surgery for gastric
aspiration to reduce colon anastomotic fluxion. For prophylaxis,
500 mg metronidazole (Sine Wanxiang Pharmaceutical Co., Shanghai,
China) and 1 g ceftriaxone (Shanghai Roche Pharmaceutical Co.,
Shanghai, China) were administered 1 h prior to induction and
continued for 48 h following the operation. All subjects were
interviewed by the study nurse, and reactions to the product,
medications taken and adverse events that occurred during the
five-day period were recorded.
Healthy subjects underwent a standard bowel
cleansing preparation with oral polyethylene glycol electrolyte
(139.12 g/2,000 ml; Jiangxi Hygecon Pharmaceutical Co.) the evening
prior to colonoscopy. Endoscopically, the colonic mucosa appeared
normal in all subjects. Colonic mucosal tissue biopsies (~1×2 mm
each) were collected from the ascending colon (n=3), descending
colon (n=1), sigmoid colon (n=3) and rectum (n=4) during
colonoscopy.
Ethics statement
The present study was approved by the Ethics
Committee of the Sixth People's Hospital Affiliated to Shanghai
Jiaotong University on Human Subjects in Medical Research [protocol
number 2011 (L)-11] and all individuals provided written informed
consent prior to participating in the study.
Sample collection and DNA extraction
Colorectal mucosa tissue samples were collected from
the tumor sites during surgery. Normal colorectal mucosa tissue
samples were collected from the cecum, ascending colon, transverse
colon, descending colon, sigmoid colon, and rectum during
colonoscopy. Fisher's exact test was used for comparison of
different anatomic sites. All samples from the study participants,
including healthy individuals, were placed in liquid nitrogen
immediately and transported to the laboratory within 30 min of
collection. DNA was extracted from all intestinal tissue samples
using MoBio Powersoil DNA extraction kits (MoBio Laboratories,
Inc., Carlsbad, CA, USA), according to the manufacturer's
instructions. Intestinal tissue samples were completely lysed by
overnight incubation at 56°C in buffer ATL (Qiagen, Düsseldorf,
Germany) and proteinase K (Shanghai SAIBAISHENG Gene Technology
Co., Ltd., Shanghai, China). The standard tissue protocol was
modified to extend the 70°C incubation step from 10 to 30 min.
DNA was eluted from a column in a final volume of
200 µl elution buffer and stored at −20°C prior to the
amplification steps.
Bacterial 16S rRNA amplification and
pyrosequencing
The V3 region of the bacterial 16S rRNA gene was
amplified from extracted DNA using primers (Sangon Biotech Co.,
Ltd., Shanghai, China) 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 533R
(5′-TTACCGCGGCTGCTGGCAC-3′) (33),
with a sample barcode sequence and the FLX Titanium adapters (Roche
Diagnostics, Basel, Switzerland) were used to amplify the V3 region
of each tissue sample by polymerase chain reaction (PCR). The PCR
mixture was composed as follows: 10 ng template, 0.4 ml FastPfu
Polymerase (Beijing TransGen Biotech Co., Ltd., Beijing, China), 4
ml 56 FastPfu buffer (Transgene, Illkirch-Graffenstaden, France), 2
ml deoxynucleotide diphosphate (2.5 mM each; Takara Bio, Inc.,
Otsu, Japan), 0.4 ml forward primers (5 mM) and 0.4 ml reverse
primers (5 mM). The reaction was performed in an ABI GeneAmp H 9700
Cycler (Applied Biosystems Life Technologies, Foster City, CA, USA)
and the cycling parameters were as follows: 5 min denaturation at
95°C followed by 25 cycles of 30 sec at 95°C (denaturation), 30 sec
for annealing at 55°C and 30 sec at 72°C (elongation), with a final
extension at 72°C for 5 min. Triplicate PCR reactions were
performed on each sample. Amplified products from stool samples
were verified by gel electrophoresis using 5 ml PCR reaction
mixture in a 2.0% agarose gel. The PCR products were purified using
the AxyPrepDNA Gel extraction kit (Corning Life Sciences, Corning,
NY, USA) and quantified on a QuantiFluor™-ST Fluorometer (Promega
Corporation, Madison, WI, USA). The products from different samples
were mixed at equal ratios for pyro-sequencing using the Roche GS
FLX 454 Sequencer (Roche Diagnostics) according to the
manufacturer's instructions. All pyrosequencing reads were then
removed of their primers, barcodes and adaptor sequences, and were
further screened and filtered according to the standards for
quality control as follows: The elimination of sequences that did
not perfectly match the proximal PCR primer (over two mismatches to
the primers), those with short sequencing length (<200
nucleotides) sequences that contained mononucleotide repeats of six
nucleotides, sequences with ambiguous characters, or sequences with
a read quality score <25. Finally, a total of 416,599
high-quality sequences from 22 samples were produced, which
accounted for 72.9% of valid sequences according to barcode- and
primer-sequence filtering.
Bioinformatics analysis of sequencing
data
In order to gain high-quality and more precise
bioinformation, effective sequences that contained certain point
mutations and macromolecular homopolymers were narrowed down using
QIIME, version 1.17 (http://qiime.org/) based on the following methods and
parameters: Number of mismatching of primers, <2; maximum number
of 3′ primers, 3; no ambiguous bases or >6 homologous
mono-bases; and sequence length, <200 base pairs (34). The optimized sequences were then
clustered into operational taxonomic units (OTUs) using Usearch,
version 7.1 (http://drive5.com/uparse/) with a criterion of minimum
similarity of 97%. Chimera sequences arising from the PCR
amplification were detected and excluded from the OTUs using UCHIME
version 4.2.40 (http://drive5.com/usearch/manual/uchime_algo.html)
(35). Using the usearch_global
command, representative OTUs were aligned to the optimized
sequences and the abundance of OTUs per sample was obtained for
performing the following further analysis.
Applying Bayesian algorithms of ribosomal database
project (RDP) classifier (http://rdp.cme.msu.edu/), representative OTUs were
identifed in the following databases: 16S bacteria and archaeal
ribosomes [SILVA (36); release
115; http://www.arb-silva.de]; RDP (21) (release 11.1; http://rdp.cme.msu.edu/); Greengene (37) (release 13.5; http://greengenes.secondgenome.com/); ITS fungus
[Unite (33); release 5.0;
http://unite.ut.ee/index.php]; and
functional genes (FGR; release 7.3; http://fungene.cme.msu.edu/). Subsequently, analysis
with Good's coverage estimator, diversity estimators (Shannon and
Simpson), richness estimators [Chao and abundance-based coverage
estimators (ACE)] were performed and the rarefaction curve was
generated using the Mothur software package (http://www.mothur.org/wiki/Main_Page) at an 80%
confidence level. In addition, the Bray-Curtis dissimilarities were
used to construct a cluster dendrogram. A metagenomic biomarker
discovery approach was employed with linear discriminant analysis
(LDA) coupled with effect size measurement (LEfSe), which was used
to perform a non-parametric Wilcoxon sum-rank test followed by LDA
analysis using online software (http://huttenhower.sph.harvard.edu/galaxy/) to assess
the effect size of each differentially abundant taxon.
Statistical analysis
Results were analysed using SPSS, version 18.0 for
Windows (SPSS, Inc., Chicago, IL, USA). Quantitative values are
expressed as the mean ± standard deviation. Comparison of
categorical data between groups was performed using the Pearson
χ2 test or, where indicated, Fisher's exact test.
Analysis of variance or the Kruskal-Wallis test was used for
continuous variables, as appropriate. P<0.05 was considered to
indicate a statistically significant difference.
Sequence data
The 16S sequence data generated in the present study
were submitted to the GenBank Sequence Read Archive with the
accession number SRP043381.
Results
Probiotics simultaneously increase
richness and diversity of mucosal microbes
A total of 416,599 high-quality, classifiable reads
were obtained, with an average of 12,624±2,423 (n=33) reads/sample.
UPARSE, version 7.1 (http://drive5.com/uparse/) was used to cluster the
above reads. At a 3% dissimilarity level, 45,453 OTUs in all
samples and an average of 1,377 OTUs (n=33) were identified per
sample.
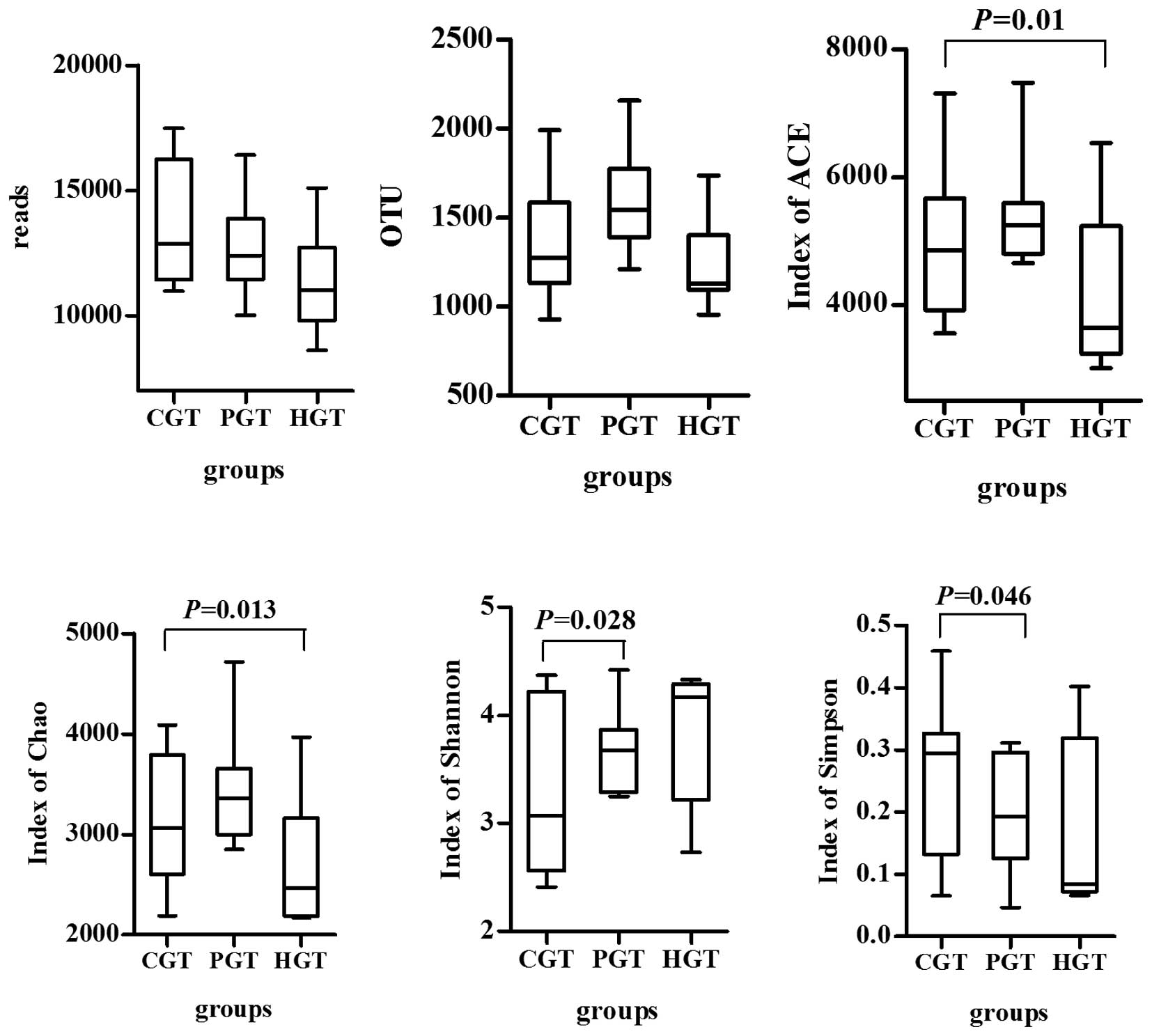
Chao and ACE indices
The PGT group exhibited significantly higher Chao
(P=0.013) and ACE (P=0.01) indices in colorectal mucosa compared
with those in the HGT group. There were no significant differences
between the HGT and CGT groups or CGT and PGT groups (Fig. 2).
Shannon and Simpson indices
Compared with the HGT group, the Shannon (P=0.028)
and Simpson (P=0.046) indices were significantly reduced in the CGT
group. This demonstrated that the mucosal microflora diversity was
reduced in patients with CRC, while the diversity of the intestinal
flora increased following oral administration of probiotics.
Probiotics modulate gut microflora
structure
The gut micro-flora structure was compared at
various bacterial levels. At the phylum level, there were no
significant differences in classification among the three groups
(27 in the CGT group, 29 in the PGT group and 25 in the HGT group).
A micro-flora whose abundance was >0.1% contributed to 99.8% of
the total category, including Firmicutes, Bacteroidetes,
Proteobacteria, Actinobacteria and Fusobacteria. At the genus
level, 188 were present in the CGT group, 198 in the PGT group and
201 in the HGT group. Bacteria whose abundance was >0.01%
contributed to 96.86% of the total category in the HGT group,
compared with 99.76% in the CGT group and 99.13% in the PGT group.
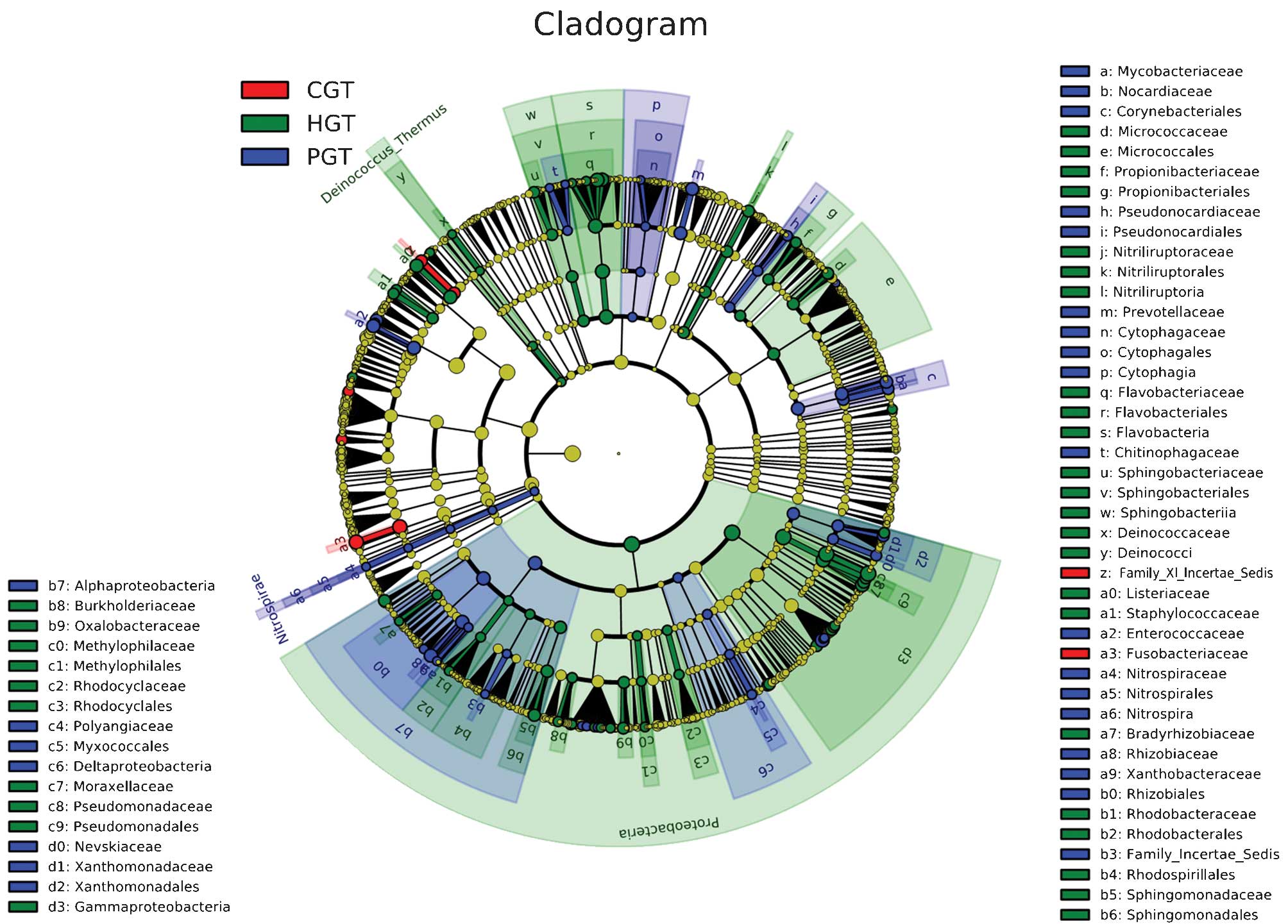
A cladogram representation of the structure of the mucosal
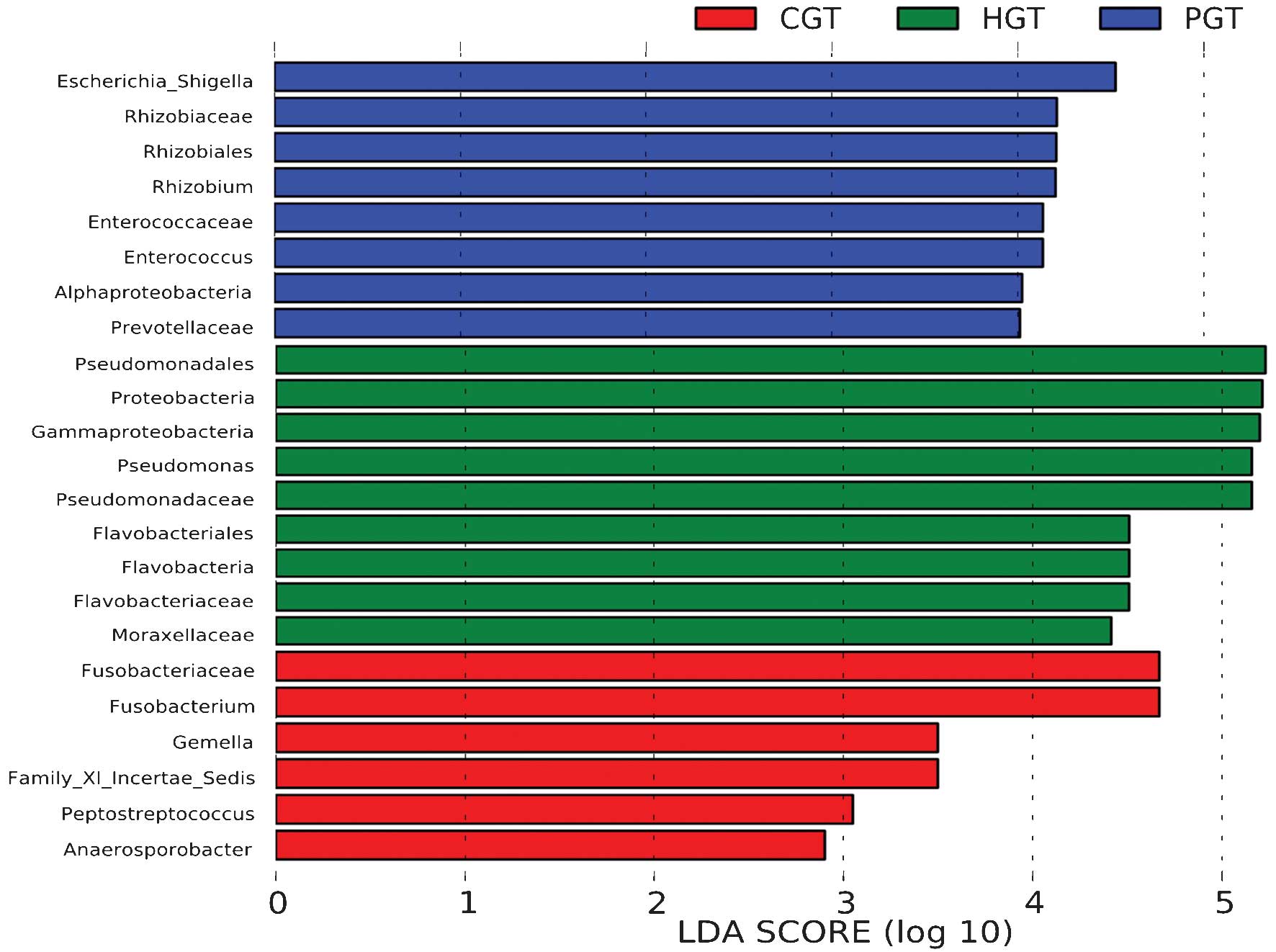
microbiota (Fig. 3) and a bar
graph showing the predominant bacteria (Fig. 4) were generated by LEfSe. The
greatest differences in taxa between the three communities are
displayed. Pyrosequencing data demonstrated that probiotics-treated
patients exhibited a significant reduction in
Peptostreptococcus, Comamonas, Fusobacterium
and expansion of Enterococcus and Proteobacteria in the
mucosa-adherent microbiota.
Reduction of a fusobacterium bacterial
group is associated with probiotics administration
To determine whether microbial alterations in the
mucosa-associated microbiota mediate the effects of probiotics in
CRC, a taxonomy-based comparison was performed to determine the
differences between the microbiota of CRC and healthy individuals
(Table III).
Fusobacterium, which constitutes less than 0.1% of total
bacteria in healthy mucosal tissue, was the most prevalent in the
mucosa of patients with CRC (10.08 vs. 0.01%; P=0.03). In addition,
probiotics treatment was observed to be associated with a
significant (~6-fold) reduction in the relative abundance of
Fusobacterium (1.91%; P=0.03). Accordingly, it was concluded
that a reduction of the mucosa-associated Fusobacterium
group may be a potential mediator of the effects of probiotics on
CRC.
 | Table IIIComparison of gut microflora
structure at phylum and genus levels. |
Table III
Comparison of gut microflora
structure at phylum and genus levels.
|
Phylum/genus/species | Relative abundance
(%)
| P-value |
|---|
| HGT | CGT | PGT |
|---|
| Firmicutes | 40.21a | 60.97 | 66.44a | 0.019 |
|
Bacillus | 35.81a | 53.83 | 63.51a | 0.017 |
|
Brochothrix | 3.05 | 0.12 | 0.19 | >0.001 |
|
Carnobacterium | 0.47a | 0.70 | 0.88a | 0.018 |
|
Enterococcus | 0.03b | 0.33a | 2.83a,b | 0.033 |
|
Coprococcus | 0.14 | 0.61a | 0.01a | 0.020 |
|
Peptostreptococcus | 0.00a | 0.23a | 0.09 | 0.036 |
| Bacteroidetes | 11.06 | 10.12 | 8.49 | >0.050 |
| Flavobacteria | 8.32a | 1.83a | 2.18a | >0.001 |
|
Epilithonimonas | 0.14a,b | 0.03a | 0.03b | 0.04 |
|
Flavobacterium | 4.67a,b | 1.03a | 1.33b | >0.001 |
|
Sphingobacteria | 0.50a | 0.08a | 0.20a | 0.001 |
|
Pedobacter | 0.10a | 0.01a | 0.01a | 0.001 |
|
Sphingobacterium | 0.36a | 0.04a | 0.05a | >0.001 |
| Proteobacteria | 46.05b | 11.55a | 19.65a,b | >0.001 |
|
Alphaproteobacteria |
3.94b | 1.26a,b | 3.74a | 0.032 |
|
Caulobacter | 1.36a | 0.02a | 0.03a | >0.001 |
|
Brevundimonas | 0.68a | 0.04a | 0.04a | >0.001 |
|
Rhizobium | 0.19a | 0.98b | 3.29a,b | 0.010 |
|
Sphingomonas | 0.98a | 0.06a | 0.04a | >0.001 |
|
Betaproteobacteria |
2.05a | 1.56 | 0.93a | 0.031 |
|
Acidovorax | 0.33a | 0.02a | 0.03a | 0.001 |
|
Comamonas | 0.39 | 1.04a | 0.08a | 0.007 |
|
Janthinobacterium | 0.54a | 0.04a | 0.05a | >0.001 |
|
Gammaproteobacteria | 39.56a | 6.66a | 14.46a | 0.001 |
|
Buttiauxella | 0.37a | 0.03a | 0.05a | >0.001 |
|
Escherichia-Shigella | 0.22a | 2.92 | 6.39a | 0.004 |
|
Rahnella | 0.66a | 0.12a | 0.24a | >0.001 |
|
Acinetobacter | 4.84a | 0.63a | 1.09a | >0.001 |
|
Enhydrobacter | 0.71a | 0.11a | 0.23a | >0.001 |
|
Psychrobacter | 0.56a | 0.05a | 0.07a | >0.001 |
|
Pseudomonas | 30.25a | 1.79a | 3.99a | >0.001 |
|
Nevskia | 0.04a | 0.03a | 0.18a | >0.001 |
|
Stenotrophomonas | 0.13b | 0.04a,b | 0.13a | 0.013 |
| Fusobacteria | 0.02a | 14.75a | 1.94a | 0.030 |
|
Fusobacterium | 0.01a | 10.08a | 1.91 | 0.032 |
| Actinobacteria | 1.91 | 1.46 | 2.58 | >0.050 |
|
Rhodococcus | 0.09a | 0.93 | 1.70a | 0.018 |
|
Nesterenkonia | 0.49a | 0.04a | 0.05a | 0.040 |
|
Propionibacterium | 0.16a,b | 0.02a | 0.07b | 0.003 |
Discussion
Numerous studies have demonstrated that consumption
of certain probiotics or prebiotics, which have been applied to
various medical conditions, is able to favorably alter the
pre-disposing factors of CRC, and is a promising approach for
prevention of CRC (38). In
vitro studies have demonstrated that lactic acid bacteria
significantly reduce the growth and viability of the human colon
cancer cell line HT-29 (39). An
in vivo study in laboratory animals have additionally
demonstrated that L. acidophilus and B. longum are
able to reduce DNA damage and protect against
1,2-dimethylhy-drazine-induced genotoxicity (40). However, unlike in vitro
behavior, commensal bacteria that inhabit the human intestine are
characteristic of relative stability and colonization resistance.
This symbiotic flora can exclude foreign microbes from habitat.
Accordingly, probiotics may serve a beneficial role by adjusting
the microfloral structure rather than directly interacting with
intestinal epithelial cells. To date, there have been few human
studies supporting the benefits of oral administration of
probiotics in high-risk populations with CRC (41).
It is of importance for a healthy intestinal
microecosystem to maintain bacterial diversity (42). In the present study, the results
demonstrated an enhanced abundance of the gut microflora following
intake of prebiotics. A lower diversity was present in the gut
microflora in patients with CRC, which was associated with the
increased abundance of certain pathogens. The results were
consistent with those of previous studies (38). Probiotic treatment of patients with
CRC can improve the abundance of gut microflora by enhancing the
diversity of the microbiota to approach a normal level (43). A possible explanation for this
association is that probiotics (B. longum, L.
acidophilus and E. faecalis) are able to quantitatively
and/or qualitatively alter the intestinal microflora and normalize
dysbiosis.
Linear discriminant analysis has indicated that
certain potential pathogens, including Fusobacterium
(44) and
Peptostreptococcus (45)
sequences, are significantly enriched in the CRC mucosal
microflora. However, other bacteria, mostly belonging to the phylum
of Proteobacteria, are reduced in CRC (46). It is reported that
Fusobacterium may be associated with IBD, which includes
ulcerative colitis and Crohn's disease (47), while IBD is one of the three
highest risk factors of CRC. It is suggested that these potential
pathogens or beneficial bacteria can be seen as the 'core
microflora' and are directly involved in the development of CRC
(48). The analysis of the present
study indicated that administration of probiotics is able to
effectively reduce pathogenic Fusobacterium and
Peptostreptococcus populations in patients with CRC. This
result agrees with a previous study, which used the Simulator of
the Human Intestinal Microbial Ecosystem model, in which L.
acidophilus or Lactobacillus casei were demonstrated to
increase lactic acid bacteria, with a concomitant reduction in
fecal coliforms and clostridia (49).
Taken together, the observations of the present
study demonstrated that probiotic supplements are able to
effectively alter the composition, richness and diversity of the
gut microflora, inhibit certain potential pathogens including
Fusobacterium and Peptostreptococcus, and increase
the number of certain beneficial microorganisms. Although
short-term administration of probiotics cannot achieve a
significant clinical effect, the results of the present study
provided a basis for microbiota-associated prevention, diagnosis,
prognosis and the development of treatment strategies for CRC.
Acknowledgments
The present study was supported by grants from the
National Natural Science Foundation of China (grant no. 81230057)
and Shen Kang Hospital Development Center of Shanghai (grant no.
SHDC12012106).
References
|
1
|
Siegel R, Naishadham D and Jemal A: Cancer
statistics, 2013. CA Cancer J Clin. 63:11–30. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Li D, Wu C, Zheng Y, Zhong W and Wu F:
Incidence and mortality of colorectal cancer in Shanghai from 2003
to 2007. Zhongguo Zhongliu. 20:413–418. 2011.
|
|
3
|
Woolf CM: A genetic study of carcinoma of
the large intestine. Am J Hum Genet. 10:42–47. 1958.PubMed/NCBI
|
|
4
|
Winawer SJ, Zauber AG, Gerdes H, O'Brien
MJ, Gottlieb LS, Sternberg SS, Bond JH, Waye JD, Schapiro M and
Panish JF: Risk of colorectal cancer in the families of patients
with adenomatous polyps National Polyp Study Workgroup. N Engl J
Med. 334:82–87. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Magrone T and Jirillo E: The interaction
between gut microbiota and age-related changes in immune function
and inflammation. Immun Ageing. 10:312013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
O'Toole PW: Changes in the intestinal
microbiota from adulthood through to old age. Clin Microbiol
Infect. 18:44–46. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
DiBaise JK, Frank DN and Mathur R: Impact
of the gut microbiota on the development of obesity: Current
concepts. Am J Gastroenterol. 1(Suppl): 22–27. 2012. View Article : Google Scholar
|
|
8
|
Kallus SJ and Brandt LJ: The intestinal
microbiota and obesity. J Clin Gastroenterol. 46:16–24. 2012.
View Article : Google Scholar
|
|
9
|
Rubinstein MR, Wang X, Liu W, Hao Y, Cai
GF and Han YW: Fusobacterium nucleatum promotes colorectal
carcinogenesis by modulating e-cadherin/β-catenin signaling via its
FadA adhesin. Cell Host Microbe. 14:195–206. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Sobhani I, Amiot A, Le Baleur Y, Levy M,
Auriault M, Van Nhieu JT and Delchier JC: Microbial dysbiosis and
colon carcinogenesis: could colon cancer be considered a
bacteria-related disease? Therap Adv Gastroenterol. 6:215–229.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Tjalsma H, Boleij A, Marchesi JR and
Dutilh BE: A bacterial driver-passenger model for colorectal
cancer: Beyond the usual suspects. Nat Rev Microbiol. 10:575–582.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ley RE, Peterson DA and Gordon JI:
Ecological and evolutionary forces shaping microbial diversity in
the human intestine. Cell. 124:837–848. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Purchiaroni F, Tortora A, Gabrielli M,
Bertucci F, Gigante G, Ianiro G, Ojetti V, Scarpellini E and
Gasbarrini A: The role of intestinal microbiota and the immune
system. Eur Rev Med Pharmacol Sci. 17:323–333. 2013.PubMed/NCBI
|
|
14
|
Xu X, Xu P, Ma C, Tang J and Zhang X: Gut
microbiota, host health and polysaccharides. Biotechnol Adv.
31:318–337. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Compare D and Nardone G: The
bacteria-hypothesis of colorectal cancer: Pathogenetic and
therapeutic implications. Transl Gastrointest Cancer. 3:44–53.
2014.
|
|
16
|
Rowland I: Probiotics and colorectal
cancer risk. Br J Nutr. 91:805–807. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Uccello M, Malaguarnera G, Basile F,
D'agata V, Malaguarnera M, Bertino G, Vacante M, Drago F and Biondi
A: Potential role of probiotics on colorectal cancer prevention.
BMC Surg. 12:S352012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Delia P, Sansotta G, Donato V, Frosina P,
Messina G, De Renzis C and Famularo G: Use of probiotics for
prevention of radiation-induced diarrhea. World J Gastroenterol.
13:912–915. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Famularo G, Mosca L, Minisola G,
Trinchieri V and De Simone C: Probiotic lactobacilli: A new
perspective for the treatment of inflammatory bowel disease. Curr
Pharm Des. 9:1973–1980. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ljungh A and Wadström T: Lactic acid
bacteria as probiotics. Curr Issues Intest Microbiol. 7:73–89.
2006.PubMed/NCBI
|
|
21
|
Allen SJ, Wareham K, Wang D, Bradley C,
Hutchings H, Harris W, Dhar A, Brown H, Foden A, Gravenor MB, et
al: Lactobacilli and bifidobacteria in the prevention of
antibiotic-associated diarrhoea and Clostridium difficile diarrhoea
in older inpatients (PLACIDE): A randomised, double-blind,
placebo-controlled, multicentre trial. Lancet. 382:1249–1257. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Mackowiak PA: Recycling metchnikoff:
Probiotics, the intestinal microbiome and the quest for long life.
Front Public Health. 1:522013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Allen SJ, Martinez EG, Gregorio GV and
Dans LF: Probiotics for treating acute infectious diarrhoea.
Cochrane Database Syst Rev. 10:CD0030482010.
|
|
24
|
Chen CC, Kong MS, Lai MW, Chao HC, Chang
KW, Chen SY, Huang YC, Chiu CH, Li WC, Lin PY, et al: Probiotics
have clinical, microbiologic and immunologic efficacy in acute
infectious diarrhea. Pediatr Infect Dis J. 29:135–138. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hempel S, Newberry SJ, Maher AR, Wang Z,
Miles JN, Shanman R, Johnsen B and Shekelle PG: Probiotics for the
prevention and treatment of antibiotic-associated diarrhea a
systematic review and meta-analysis. JAMA. 307:1959–1969. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Koebnick C, Wagner I, Leitzmann P, Stern U
and Zunft HJ: Probiotic beverage containing Lactobacillus casei
Shirota improves gastrointestinal symptoms in patients with chronic
constipation. Can J Gastroenterol. 17:655–659. 2003.PubMed/NCBI
|
|
27
|
Ouwehand AC, Lagström H, Suomalainen T and
Salminen S: Effect of probiotics on constipation, fecal
azoreductase activity and fecal mucin content in the elderly. Ann
Nutr Metab. 46:159–162. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
de Milliano I, Tabbers MM, van der Post JA
and Benninga MA: Is a multispecies probiotic mixture effective in
constipation during pregnancy? 'A pilot study'. Nutr J. 11:802012.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Persborn M and Söderholm JD: Commentary:
the effects of probiotics on barrier function and mucosal pouch
microbiota during maintenance treatment for severe pouchitis in
patients with ulcerative colitis-authors' reply. Aliment Pharmacol
Ther. 38:1406–1407. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Tursi A, Brandimarte G, Papa A, Giglio A,
Elisei W, Giorgetti GM, Forti G, Morini S, Hassan C, Pistoia MA, et
al: Treatment of relapsing mild-to-moderate ulcerative colitis with
the probiotic Vs.L#3 as adjunctive to a standard pharmaceutical
treatment: A double-blind, randomized, placebo-controlled study. Am
J Gastroenterol. 105:2218–2227. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Müllner K, Miheller P, Herszényi L and
Tulassay Z: Probiotics in the management of Crohn's disease and
ulcerative colitis. Curr Pharm Des. 20:4556–4560. 2014. View Article : Google Scholar
|
|
32
|
Huebner C, Ding Y, Petermann I, Knapp C
and Ferguson LR: The probiotic Escherichia coli Nissle 1917 reduces
pathogen invasion and modulates cytokine expression in Caco-2 cells
infected with Crohn's disease-associated E. coli LF82. Appl Environ
Microbiol. 77:2541–2544. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Kõljalg U, Nilsson RH, Abarenkov K,
Tedersoo L, Taylor AF, Bahram M, Bates ST, Bruns TD,
Bengtsson-Palme J, Callaghan TM, et al: Towards a unified paradigm
for sequence-based identification of fungi. Mol Ecol. 22:5271–5277.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Hamady M, Walker JJ, Harris JK, Gold NJ
and Knight R: Error-correcting barcoded primers for pyrosequencing
hundreds of samples in multiplex. Nat Methods. 5:235–237. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Edgar RC: UPARSE: Highly accurate OTU
sequences from microbial amplicon reads. Nat Methods. 10:996–998.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Quast C, Pruesse E, Yilmaz P, Gerken J,
Schweer T, Yarza P, Peplies J and Glöckner FO: The SILVA ribosomal
RNA gene database project: Improved data processing and web-based
tools. Nucleic Acids Res. 41:D590–D596. 2013. View Article : Google Scholar :
|
|
37
|
DeSantis TZ, Hugenholtz P, Larsen N, Rojas
M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P and Andersen GL:
Greengenes, a chimera-checked 16S rRNA gene database and workbench
compatible with ARB. Appl Environ Microbiol. 72:5069–5072. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Appleyard CB, Cruz ML, Isidro AA, Arthur
JC, Jobin C and De Simone C: Pretreatment with the probiotic VSL#3
delays transition from inflammation to dysplasia in a rat model of
colitis-associated cancer. Am J Physiol Gastrointest Liver Physiol.
301:G1004–G1013. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Fotiadis CI, Stoidis CN, Spyropoulos BG
and Zografos ED: Role of probiotics, prebiotics and synbiotics in
chemoprevention for colorectal cancer. World J Gastroenterol.
14:6453–6457. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Wollowski I, Rechkemmer G and Pool-Zobel
BL: Protective role of probiotics and prebiotics in colon cancer.
Am J Clin Nutr. 73:451S–455S. 2001.PubMed/NCBI
|
|
41
|
Bertkova I, Hijova E, Chmelarova A,
Mojzisova G, Petrasova D, Strojny L, Bomba A and Zitnan R: The
effect of probiotic microorganisms and bioactive compounds on
chemically induced carcinogenesis in rats. Neoplasma. 57:422–428.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Vipperla K and O'Keefe SJ: The microbiota
and its metabolites in colonic mucosal health and cancer risk. Nutr
Clin Pract. 27:624–635. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Wang J, Tang H, Zhang C, Zhao Y, Derrien
M, Rocher E, van-Hylckama Vlieg JE, Strissel K, Zhao L, Obin M and
Shen J: Modulation of gut microbiota during probiotic-mediated
attenuation of metabolic syndrome in high fat diet-fed mice. ISME.
9:1–15. 2015. View Article : Google Scholar
|
|
44
|
Kostic AD, Gevers D, Pedamallu CS, Michaud
M, Duke F, Earl AM, Ojesina AI, Jung J, Bass AJ, Tabernero J, et
al: Genomic analysis identifies association of Fusobacterium with
colorectal carcinoma. Genome Res. 22:292–298. 2012. View Article : Google Scholar :
|
|
45
|
Wang T, Cai G, Qiu Y, Fei N, Zhang M, Pang
X, Jia W, Cai S and Zhao L: Structural segregation of gut
microbiota between colorectal cancer patients and healthy
volunteers. ISME J. 6:320–329. 2012. View Article : Google Scholar :
|
|
46
|
Allen-Vercoe E, Strauss J and Chadee K:
Fusobacterium nucleatum: An emerging gut pathogen? Gut Microbes.
2:294–298. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Strauss J, Kaplan GG, Beck PL, Rioux K,
Panaccione R, Devinney R, Lynch T and Allen-Vercoe E: Invasive
potential of gut mucosa-derived Fusobacterium nucleatum positively
correlates with IBD status of the host. Inflamm Bowel Dis.
17:1971–1978. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Arthur JC and Jobin C: The struggle
within: Microbial influences on colorectal cancer. Inflamm Bowel
Dis. 17:396–409. 2011. View Article : Google Scholar
|
|
49
|
Chaikham P and Apichartsrangkoon A:
Effects of encapsulated Lactobacillus acidophilus along with
pasteurized longan juice on the colon microbiota residing in a
dynamic simulator of the human intestinal microbial ecosystem. Appl
Microbiol Biotechnol. 98:485–495. 2014. View Article : Google Scholar
|