Introduction
Globally, liver cancer (LC) is the commonest type of
hepatic malignancy and the third leading cause of cancer-related
death (1). Transarterial
chemoembolization (TACE) is mainly used to treat unresectable
advanced LC and can also be used for complementary treatment
without radical resection. As most patients with LC are diagnosed
at advanced stages, which causes them to miss the best opportunity
for curative therapy via resection, transplantation or ablation,
TACE serves an important role in the treatment of LC. The main
mechanism of the treatment of LC relies on the difference in the
blood supply between LC and normal liver tissue. A total of 95–99%
of the LC blood supply is derived from the hepatic artery and
70–75% of the liver tissue blood supply is derived from the portal
vein (2). TACE can effectively
block the blood supply from the hepatic artery while continuously
releasing high-concentration chemotherapy drugs (cisplatin and
fluorouracil) to directly kill tumor cells and has little effect on
normal liver tissue (3). Tumor
stage, portal hypertension and preoperative levels of α
fetoprotein, alkaline phosphatase, γ glutamyl transpeptidase and
monocytes are important clinical indicators to predict the efficacy
of TACE (4). Serum IL-6 and IL-8
levels may help to identify patients who benefit from TACE in terms
of objective response to treatment and overall survival before
intervention (5). Circulating MMP-2
is also associated with the prognosis of LC following TACE
(6). However, molecular markers of
TACE-related prognostic indicators are lacking.
Exosomes are microcapsule structures with a lipid
bilayer having a diameter of 30–150 nm that are secreted by various
normal cells and tumor cells and are naturally present in various
body fluids (7). They contain
proteins, lipids, messenger RNA (mRNA), microRNA (miRNA), long
noncoding RNA (lncRNA), circular RNA (circRNA) and transcription
factors (8,9). Exosomes serve a crucial role in the
carcinogenesis of LC (10,11).
In the process of carcinogenesis, exosomes are
associated with viral spread and affect host immunity, which causes
DNA damage and LC. For example, exosomal miR-122 promotes viral RNA
replication and transmission between hepatitis virus C-infected
hepatocytes and normal hepatocytes via the AGO2-miR-122-HSP90
complex (12). lncRNA FAL1/miR-1236
can promote tumor metastasis (13).
Noncoding (nc)RNAs can be transferred to immune cells through
exosomes to induce the expansion or differentiation of
immunosuppressive cells in LC. For example, exosomal miR-23a-3p can
upregulate the expression of programmed death-ligand 1 through the
PTEN-PI3K/AKT pathway (14).
An increasing number of studies have demonstrated
that exosomal circRNAs serve a key role in the prediction and
diagnosis of LC. For example, exosomal circRNA-100338 can promote
LC metastasis (15). The circRNA
Cdr1, as a competitive endogenous RNA, is associated with the
progression of LC (16). Exosomal
circPTGR1 promotes LC metastasis via the miR449a-hepatocyte growth
factor receptor (MET) pathway (17). In addition, exosomal circRNAs
secreted from adipocytes can promote the growth of LC by targeting
the deubiquitination-related protein USP7 (18).
The present study first compared the expression
patterns of exosomal circRNAs during TACE. Hepatic artery blood was
collected from 5 patients with LC at three time points: Prior to
TACE treatment (prior TACE), following the first TACE operation
(TACE1) and following the second TACE operation (TACE2). Then, the
exosomes were extracted from the blood and the alterations of the
circRNAs in exosomes were further analyzed. A novel circRNA was
screened (circ-G004213). The present study demonstrated that
circ-G004213 was associated with the prognosis of patients with LC.
By constructing circRNA-miRNA-mRNA interaction, it was found that
circ-G004213 interacted with miR-513b-5p and downstream target gene
PRPF39. Previous studies found that miR-513b-5p can suppress cell
promotion in experimental electromagnetic cancer cells and suppress
cell promotion and migration in ovarian cancer (19,20).
In addition, PRPF39 was associated with cisplatin sensitivity
(21). The present study was
performed in order to further confirm the role and mechanism of
circ-G004213/miR-513b-5p/PRPF39 in the cisplatin sensitivity of
LC.
Materials and methods
Human specimens
Hepatic artery blood was collected from five male
patients between May 2017 and May 2019 with LC (age range between
26 and 66 years old) at three time points: Prior TACE, TACE1 and
TACE2. A total of 10 ml of hepatic artery blood was drawn from each
patient at each time point, EDTAK2 was used to anticoagulant and
the plasma was separated (2,000 × g, 10 min at 4°C) and immediately
frozen at −80°C prior to the extraction of exosomes. All five
patients were enrolled from the Department of Gastroenterology at
the First Affiliated Hospital of Chengdu Medical College (Chengdu,
China) from 2018–2019. The other 50 patients with LC' blood samples
were obtained from the tissue specimen bank of the First Affiliated
Hospital of Chengdu Medical College (2017–2020; Table I). LC was diagnosed according to the
2017 specifications for the diagnosis and treatment of primary
liver cancer using TNM and Barcelona Clinic Liver Cancer staging
(BCLC) tumor staging assessment (22). Child-Pugh score was used to evaluate
the changes in liver function of the patients following TACE
treatment (23). None of the
patients with LC received chemotherapy or radiotherapy before TACE.
The clinical, imaging and pathologic records of the patients were
reviewed retrospectively. Informed consent was signed in writing by
all patients and the study protocol was approved by the ethics
committee of the First Affiliated Hospital of Chengdu Medical
College (approval no. 2017009).
 | Table I.Association between
clinicopathological variables and circ-G004213 expression in
patients with LC following TACE. |
Table I.
Association between
clinicopathological variables and circ-G004213 expression in
patients with LC following TACE.
|
|
| circ-G004213 |
|
|---|
|
|
|
|
|
|---|
|
Characteristics | Number | High | Low | P-value |
|---|
| Sex |
|
|
| 0.496 |
|
Male | 39 | 18 | 21 |
|
|
Female | 11 | 7 | 4 |
|
| Age (years) |
|
|
| 0.999 |
|
≤50 | 8 | 3 | 5 |
|
|
>50 | 42 | 19 | 23 |
|
| HBV |
|
|
| 0.716 |
|
Positive | 41 | 23 | 18 |
|
|
Negative | 9 | 6 | 3 |
|
| AFP (ng/ml) |
|
|
| 0.033a |
|
≤200 | 7 | 6 | 1 |
|
|
>200 | 43 | 15 | 28 |
|
| TNM stage |
|
|
| 0.009a |
| I and
II | 21 | 15 | 6 |
|
| III and
IV | 29 | 9 | 20 |
|
| BCLC stage |
|
|
| 0.022a |
| A and
B | 23 | 16 | 7 |
|
| C and
D | 27 | 9 | 18 |
|
| Child-Pugh
score |
|
|
| 0.484 |
|
5–6 | 29 | 16 | 13 |
|
|
7–9 | 18 | 7 | 11 |
|
| ≥10 | 3 | 1 | 2 |
|
Isolation of exosomes
Exosomes in plasma were separated by an exoEasy Maxi
kit (Qiagen China Co., Ltd.). XBP buffer (3 ml) was added to 3 ml
of plasma. The centrifuge tube was immediately inverted gently to
mix the solution and kept at room temperature. The plasma/XBP
buffer mixture was then added to the exoEasy rotary column and
centrifuged at 500 × g for 1 min at 4°C. The flow through was
discarded and the rotating column was placed in the same collecting
tube. XWP buffer (10 ml) was added and centrifuged at 5,000 × g for
5 min at 4°C to remove the residual volume from the rotating
column. All the flow through in the collection tube was discarded.
The rotating column was moved into a new collecting tube. Next, 400
µl-1 ml of Xe buffer was added to the membrane and incubated for 1
min at 4°C. The test tube was centrifuged at 500 × g for 5 min at
4°C and the eluate was collected. Finally, the eluate was added to
the exoEasy rotary column and incubated for 1 min and the tube was
centrifuged at 5,000 × g for 5 min at 4°C; then, the eluate was
collected and transferred to a new collection tube. Finally, the
exons were resuspended in PBS and stored at −80°C.
Detection of exosomes
Exosomes were detected by three methods. First,
exosomes were observed under a transmission electron microscope
(TEM; HT7700; Hitachi, Ltd.). For an enhanced view of the plasma
membrane, the exosomes were embedded in polymer at 65°C for 48 h.
Then, they were stained with uranium acetate for 10 min and lead
acetate for 10 min (room temperature).
Western blotting was used to detect the typical
exosomal markers, as below. Equal amounts of protein were subjected
to SDS-PAGE. The protein extracted from blood was used as the
control. The sources and dilutions of the antibodies that were used
were as follows: Mouse anti-CD9 (Abcam; 1:400), mouse anti-CD63
(Abcam; 1:400), rabbit anti-tumor susceptibility gene (TSG)101
(Abcam; 1:1,000) and mouse anti-β-actin (Abcam; 1:1,000), which was
used as an internal control. Finally, the exosomes were tested with
a Malvern Nano-Nano Analyzer (MNNA; Nanosight NS300; Malvern
Instruments, Ltd.) to measure their size.
CircRNA sequencing analysis
High-throughput full transcriptome sequencing and
subsequent bioinformatics analysis were performed. According to the
manufacturer's instructions, the RNA library was constructed with
rRNA-depleted RNA using the TruSeq Stranded Total RNA library
preparation tool (Illumina, Inc.). The Bioanalyzer 2100 system
(Agilent Technologies, Inc.) was used to test and quantify the
quality of the library. A total of 10 picomolar libraries were
denatured to generate single-stranded DNA molecules. These
molecules were captured on Illumina flow cells and amplified in
situ in clusters. Finally, 150 cycles of sequencing were
performed on an Illumina HiSeq 4000 sequencer (Illumina, Inc.)
according to the manufacturer's instructions. Paired end readings
were taken from the Illumina HiSeq 4000 sequencer (Illumina, Inc.).
After trimming of the 3′ adapter, low-quality readings were removed
by Cutadapt software (v1.9.3; http://pypi.python.org/pypi/cutadapt#downloads). High
quality trim reading analysis was then performed. Star software
(http://www.starsoftwarein.com/) was used
to compare high-quality reading data with the reference
genome/transcriptome, DCC software (https://www.hpc.dtu.dk/?page_id=3270) was used to
detect and identify circRNAs and the circRNA database (v0.1;
http://www.circbase.org/.) was used for
annotation. The data were normalized by edgeR software (v3.12;
http://bioconductor.org/biocLiteR) and
differential expression was analyzed. The relevant data of the
present study have been uploaded to GEO depository with the number
GSE 165183. Gene Ontology (GO) was used to study the differentially
expressed molecules (http://www.http://geneontology.org). The Kyoto
Encyclopedia of Genes and Genomes (KEGG; http://www.genome.jp/kegg) database was also used.
Based on the GO and KEGG pathway analyses, the differentially
expressed circRNAs were annotated and their functions
determined.
Cell culture
The human 293T cells and LC cell lines HepG2 were
derived from Wuhan tumor cell lines in China (Procell Life Science
& Technology Co., Ltd.). HepG2 and 293T cells were identified
by STR analysis. Mycoplasma conjugation test was performed in all
cells and these cells have been used in our previous study
(24). The acisplatin-resistant
subline (HepG2/CIS) of HepG2 cells was established by repeated
subculture in cisplatin (25).
Cisplatin was purchased from Sigma-Aldrich (Merck KGaA) and
dissolved in dimethyl sulfoxide to obtain a 10 mM stock solution.
All cells were cultured in DMEM (HyClone; Cytiva) containing 10%
fetal bovine serum (FBS, HyClone; Cytiva) and preserved at 37°C and
5% CO2. For comparison of the IC50 values of
CIS among the groups of HepG2 cells, cell viability was evaluated
through MTT assays as described previously (24). HepG2 cells (1×104) were
plated in 96-well plates. Once the cells reached 70% confluence,
CIS (0, 5, 10, 20, 40, 100 and 200 µM) was added. After the cells
were incubated for an additional 72 h, the MTT reagent was added
and allowed to incorporate for 4 h. The optical density at 570 nm
was determined using an ELISA plate reader (Model 550; Bio-Rad
Laboratories, Inc.).
Plasmid construction and
transfection
For overexpression of circ-G004213, the full-length
339 bp cDNA of circ-G004213 was cloned into the vector pLCDH. The
circ-G004213 and 2581-bp to 2600-bp fragments of PRPF39 were
amplified from the cDNA of HepG2 cells and cloned into pmiRGLO
(Promega Corporation) for luciferase analysis. All constructed
plasmids were confirmed by sequencing. In addition, HepG2 cell
lines with stable overexpression or knockout of PRPF39 were
constructed with lentiviral vectors (24,25).
Short interfering (si)RNA, miRNA mimics or inhibitors were
synthesized by Guangzhou RiboBio Co., Ltd. siRNA-circ-G004213,
5′-AGAUUCUUAGACUCCAGAU-3′; siRNA-NC (negative control),
5′-UUCUCCGAACGUGUCACGU-3′. HepG2 cells were seeded in a six-well
plate, and 100 pM of siRNA-circ-G004213 was transfected into HepG2
cells using TurboFect transfection reagent (Thermo Fisher
Scientific, Inc.). miRNA mimics or inhibitors (Beckman Coulter,
Inc.) were diluted to a final concentration of 20 nM in serum-free
DMEM (HyClone; Cytiva). For transient transfection, HepG2 and 293T
cells were transfected with Lipofectamine® 2000
(Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions. There was an interval of 24 h between
transfection and subsequent experiments.
Luciferase reporter assay
Double fluorescein reporter gene detection was
carried out according to the manufacturer's instructions (24,25).
The Wild-type (WT) and mutant (MUT) circ-G004213 UTRs and PRPF39
UTRs were amplified and cloned into pmirGLO (Promega Corporation).
For dual luciferase assays, 293T cells were cotransfected with the
pmirGLO plasmid and miR-523b-5p mimic and pRLTK reporter gene
carrier, respectively, with assistance of Lipofectamine®
2000 (Invitrogen; Thermo Fisher Scientific, Inc.). The firefly and
Renilla luciferase activities were measured by applying dual
luciferase reporter assay system (Promega Corporation) 48 h
following transfection according to the manufacturer's
instructions. The luciferase activity was measured by a Multi-Mode
Microplate Reader (Synergy 2; BioTek Instruments, Inc.). All the
assays were repeated at least 3 times.
In vivo experiments
A total of 120 6-week-old BALB/c nude mice from
Chengdu Dossy. Experimental Animal Co., Ltd. were subcutaneously
inoculated in the flank region with HepG2 cells (5×106;
circ-G004213-OE, vector-NC, si-circ-G004213 or si-NC). Then 10 mice
in each group were subcutaneously injected with the cell mixture.
The mice were fed in SPF animal center of Chengdu Medical College
with the humidity at 50%, the temperature at 25°C and 14-h
light/10-h dark cycle. The tumor volume was calculated using the
following formula: Tumor volume (mm3)=π/6 × length ×
width2. When the tumor volume reached 100
mm3, CIS (10 µM) was intraperitoneally injected
(25). The endpoint was tumors up
to 1,000 mm3. When they reached the endpoint, the mice
were euthanized with 100% CO2 with a flow rate of 40%
and the endpoint was recorded. The animal experiments were approved
by the research ethics committee of Chengdu Medical College
(approval no. 2017CYA-021).
CircRNA-miRNA-mRNA interactions
To further study the relationship between circRNAs
and miRNAs, the miRanda and TargetScan databases (http://www.microrna.org/microrna/home.do) were used to
identify targeted miRNAs. From these predictions it was
hypothesized that the circRNAs might act as miRNA sponges. Based on
the binding sites of differentially expressed circRNAs and miRNAs,
a network between circRNAs and miRNAs was constructed. According to
the reverse splicing site coordinates of circRNAs, the coordinates
of mRNAs (defined as circRNA-related genes) were found through the
RefSeq database (v34; http://www.ncbi.nlm.nih.gov/refseq/).
Reverse transcription-quantitative
(RT-q) PCR
To detect the expression of circRNAs and mRNA,
RT-qPCR was performed according to the manufacturer's protocols.
Total RNA from blood samples lysates was isolated using
TRIzol® reagent (Thermo Fisher Scientific, Inc.). cDNA
was synthesized with the PrimeScript RT Master Mix (Takara
Biotechnology Co., Ltd.) from 500 ng RNA. The PCR analyses were
performed with SYBR Premix Ex TaqII (Takara Biotechnology Co.,
Ltd.) with Light Cycler 480 II Real-Time PCR System (Roche
Diagnostics). PCR protocol consisted of one cycle at 95°C for 10
sec followed by 40 cycles at 95°C for 5 sec and at 60°C for 45 sec.
The relative gene expression data was analyzed by the
2−ΔΔCq method (26).
GAPDH was used as an endogenous control gene for circRNAs and mRNA.
To determine the abundance of circRNA, the divergent primers were
designed for the circular transcripts. Primers for circRNAs and
related mRNAs are shown in Table
SI.
miR-4677-3p (Assay ID ap04309), miR-513b-5p (Assay
ID ap002658), miR-580-3p (Assay ID ap01950), miR-219a-1-3p (Assay
ID ap01258) and miR-6888-3p (Assay ID ap08601) were detected
according to the RT-qPCR. Mir-X miRNA First-Strand Synthesis Kit
(Takara Biotechnology Co., Ltd.) was used to reverse transcribe
total RNA into mature miRNA. U6 was used as an endogenous control
for miRNA expression analysis. Magnetic Resonance Elastography
(MRE) was used for analysis. The threshold cycle (Cq) was defined
as the fractional cycle number at which the fluorescence passed the
fixed threshold. All assays were performed in triplicates.
Western blot analysis
Proteins from HepG2 transplanted tumor were
extracted with common protein extraction reagent (Beyotime
Institute of Biotechnology) and 1% phenylmethylsulfonyl fluoride.
Protein concentrations were determined by BCA using an assay kit
(Bio-Rad Laboratories, Inc.). Protein lysates (50 µg) were loaded
by 4% sodium dodecyl sulfate-polyacrylamide gel and separated by
10% gel electrophoresis and transferred onto a polyvinylidene
difluoride membrane (EMD Millipore). Membranes were incubated in
blocking buffer (Tris-buffered saline containing 5% skimmed milk)
for 1 h at 37°C. The antibodies used were as follows: β-actin (cat.
no. 8H10D10; Cell Signaling Technology, Inc.) PRPF39 (cat. no.
PA5-21627; Invitrogen; Thermo Fisher Scientific, Inc.). The
dilution factor of all antibodies was 1:1,000. The first antibody
were incubation at 4°C overnight. After washing, the membranes
underwent hybridization with horseradish peroxidase-conjugated
secondary antibody horseradish peroxidase conjugated anti-rabbit
IgG (1:3,000; cat. no. 14708; Cell Signaling Technology, Inc.) and
anti-mouse IgG (1:3,000; cat. no. 7076; Cell Signaling Technology,
Inc.) for 1 h at room temperature. Following washing signals, were
detected by chemiluminescence using western blotting luminol
reagent (Santa Cruz Biotechnology, Inc.). Protein levels were
quantified by scanning blots on a Gel Doc EZ imager (Bio-Rad
Laboratories, Inc.) and analysis with Quantity One 1D image
analysis software 4.4.0 (Bio-Rad Laboratories, Inc.).
Statistical analysis
All statistical analyses were performed using SPSS
20.0 (IBM Corp.). Medcalc v18.9 (Medcalc Software Bvba) software
was used to calculate the sample size to ensure that it was
sufficient to detect the pre-specified effect amount. To determine
the significant difference, Student's t-test was used for
comparison between the two groups and one-way analysis of variance
was used for comparison of multiple groups. Bonferroni's was used
as a post hoc test. The Fisher's exact test was used to test the
association between the two categorical variables. Each experiment
was repeated at least three times. In Kaplan-Meier curves survival
analysis, log-rank test was used. P<0.05 was considered to
indicate a statistically significant difference.
Results
Profiling and characteristics of
exosomal circRNAs in patients with LC following TACE
patients with LC were enrolled from the affiliated
hospital of Chengdu Medical College between 2018 and 2019. All
patients were male, with ages ranging between 26 and 66 years old.
Of these patients, four had hepatitis virus B (HBV) infection
(Table SII). During the TACE
process, cisplatin (50 mg) and fluorouracil (250 mg) were injected
into the tumor tissue through the hepatic artery for local
chemotherapy and the main blood supply of the tumor was embolized
with lipiodol. The tumor and iodized oil deposits observed during
treatment are shown in Fig. S1.
Through TNM and Barcelona Clinic Liver Cancer staging (BCLC) tumor
staging assessment, it was confirmed that following performing TACE
twice, case 1, 2 and 3 achieved a partial response and case 4 and 5
achieved a stable disease state (Table
SII). In addition, the Child-Pugh score was used to evaluate
the changes in liver function of the patients following TACE
treatment. The results demonstrated that TACE treatment did not
damage the liver function of the patients (Table SII).
Under TEM, the diameter of the exosomes ranged from
30 to 150 nm and the membrane was clear and relatively complete
(Fig. 1A). The exosomes expressed
typical exosome markers, such as CD9, TSG101 and CD63. However,
there was no expression of CD63 and TSG101 and low expression of
CD9 in the blood. There was no expression of β-actin in the
exosomes (Fig. 1A). The MNNA
demonstrated that the peak of the isolated exosomes occurred at
approximately 150 nm (Fig. 1A).
CircRNA-seq identified 390 differentially expressed circRNAs
between the prior TACE and TACE1 groups (fold change ≥2.0;
P<0.05). Heat maps of 390 differentially expressed circRNAs were
generated to illustrate the distinguishable circRNA expression
profile of the samples (Fig. 1B).
The volcano plot analysis demonstrated 305 upregulated and 85
downregulated circRNAs (Fig. 1C).
Hierarchical cluster analysis clearly identified 489 differentially
expressed circRNAs between the TACE and TACE2 groups (Fig. 1D). The volcano plot analysis
demonstrated 422 upregulated and 67 downregulated circRNAs
(Fig. 1E).
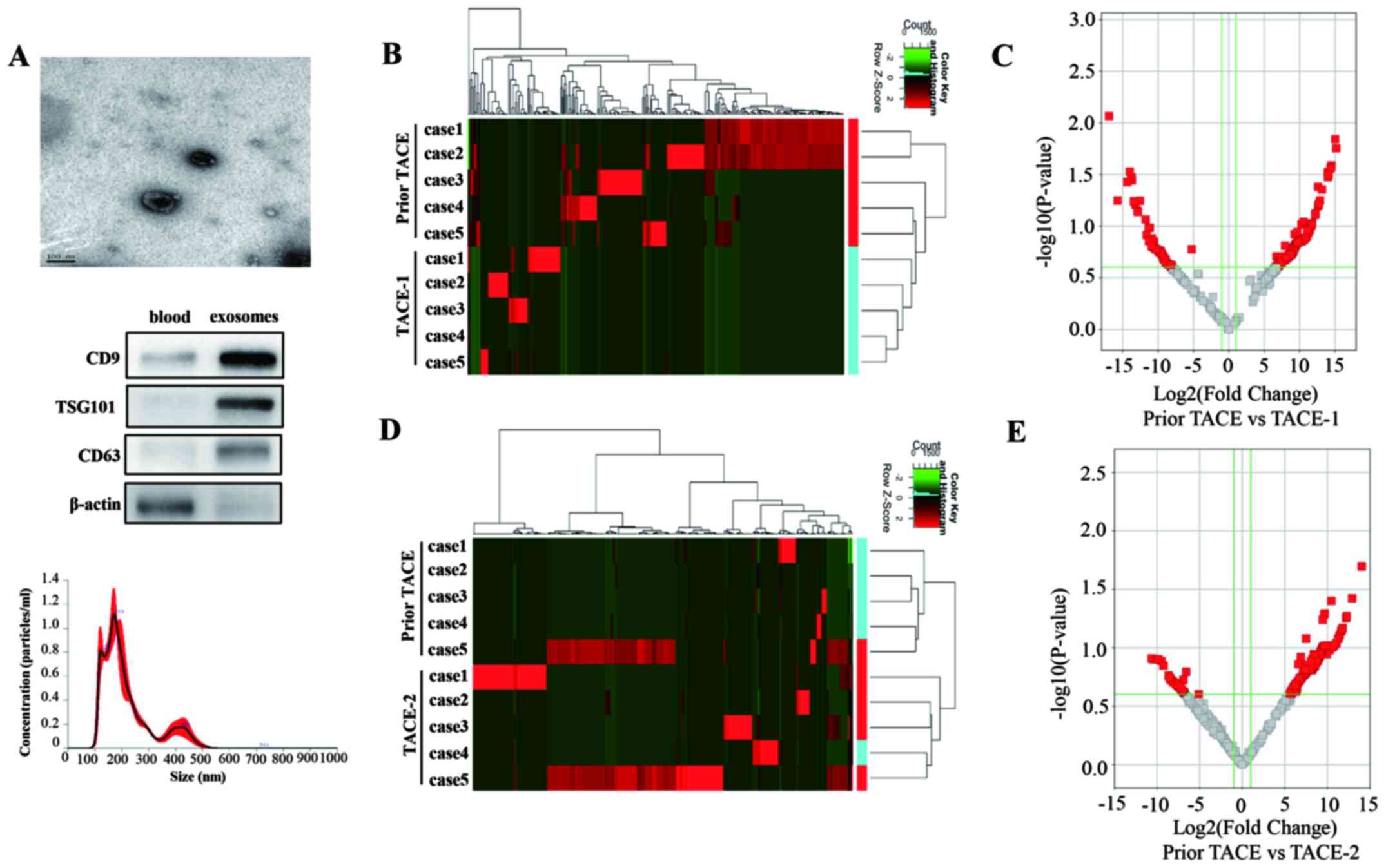 | Figure 1.Profiling and characteristics of
exosomal circRNAs in patients with LC following TACE. (A) TEM image
of exosomes with negative staining to enhance the view of membrane
structures (scale bar=100 nm). Western blotting detected CD9, CD63,
TSG101 and β-actin in exosomes (blood used as control). Malvern
Nano-Nano Analyzer demonstrated the diameter distribution of
isolated exosomes. (B) Heat map of 390 differentially expressed
circRNAs between prior TACE and TACE1. Each column represents one
sample; each row represents one probe set. Red color (upregulated
expression) or green (downregulated expression). The dendrogram on
the right reveals the sample clustering; the dendrogram on the top
reveals the gene clustering. (C) Volcano plot using fold change and
P-value. The red rectangle represents differentially expressed
circRNAs between prior TACE and TACE1, P<0.05, fold change ≥2.0.
(D) Heat map of 489 differentially expressed circRNAs between prior
TACE and TACE2. Each column represents one sample; each row
represents one probe set. Red color (upregulated expression) or
green (downregulated expression). The dendrogram on the right
reveals the sample clustering; the dendrogram on the top reveals
the gene clustering. (E) Volcano plot using fold change and
P-value. The red rectangle represents differentially expressed
circRNAs between prior TACE and TACE2, P<0.05, fold change ≥2.0.
circRNA, circular RNA; LC, liver cancer; TACE, transarterial
chemoembolization; TEM, transmission electron microscope; TACE1,
after the first TACE operation; TACE2, after the second TACE
operation. |
GO analysis of differentially
expressed circRNA-derived genes
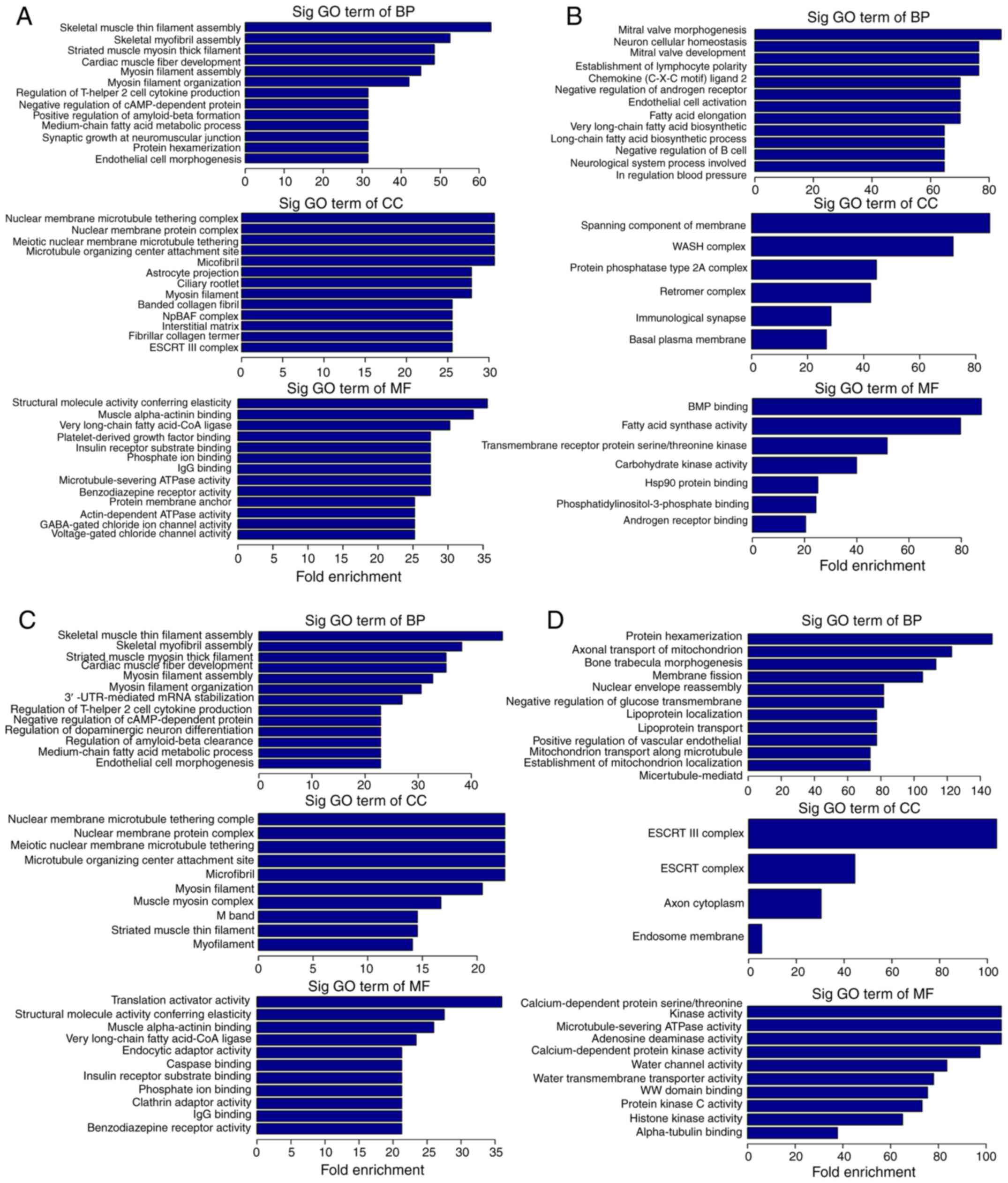
The GO analysis included three categories:
Biological process (BP), cell component (CC) and molecular function
(MF). The BP analysis demonstrated that between the prior TACE and
TACE1 groups, the differentially expressed circRNAs were
significantly associated with myosin filament assembly, negative
regulation of androgen receptor and negative regulation of B cells
(Fig. 2A and B). Significant CC
terms associated with the differentially expressed circRNAs between
the prior TACE and TACE1 groups were associated with the myosin
filament, nuclear membrane protein complex, immunological synapse
and basal plasma membrane (Fig. 2A and
B). For MF, the differentially expressed circRNAs between the
prior TACE and TACE1 groups were associated with IgG binding, very
long-chain fatty acid-CoA ligase and insulin receptor substrate
binding (Fig. 2A and B). Between
the prior TACE and TACE2 groups, the differentially expressed
circRNAs associated with BP were significantly associated with
myosin filament assembly, the medium-chain fatty acid metabolic
process, membrane fission and lipoprotein transport (Fig. 2C and D). Significant CC terms
associated with the differentially expressed circRNAs were
associated with the myosin filament, nuclear membrane protein
complex, ESCRT complex and endosome membrane (Fig. 2C and D). For MF, the differentially
expressed circRNAs were associated with IgG binding, very
long-chain fatty acid-CoA ligase, insulin receptor substrate
binding, kinase activity and water channel activity (Fig. 2C and D).
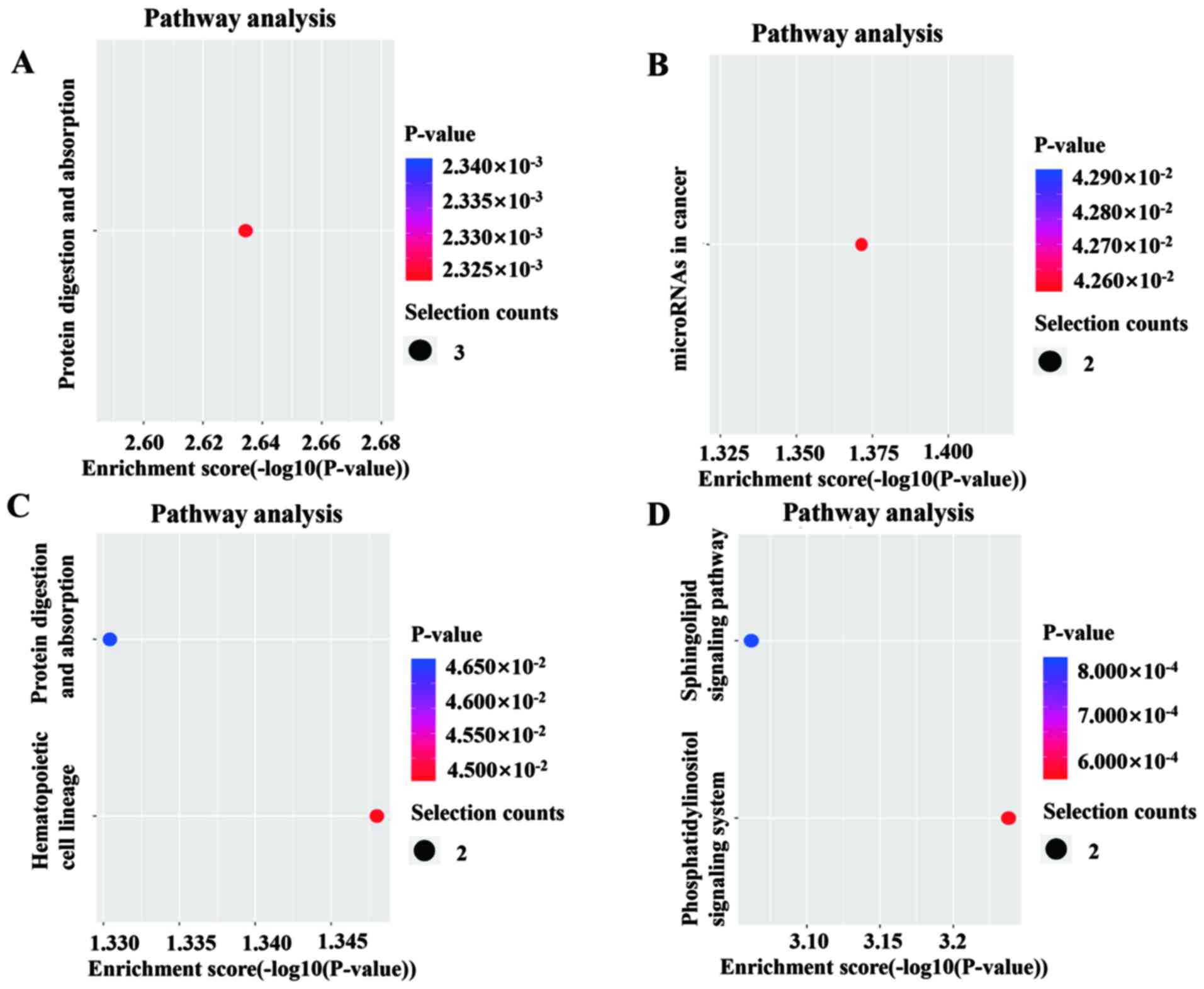
KEGG pathway analysis of
differentially expressed circRNA-derived genes
The KEGG pathway dot plot shows the significantly
enriched pathways with the enrichment score [-log10 (P-value)]
values. KEGG pathway analysis predicted the pathways affected by
the variations in circRNAs in exosomes from patients with LC
following TACE (Fig. 3). Between
the TACE and TACE1 groups, the protein digestion and absorption
pathway was shown to be activated and the miRNAs involved in the
cancer pathway were inhibited (Fig. 3A
and B). Pathway analysis demonstrated that between the prior
TACE and TACE2 groups, the hematopoietic cell lineage and protein
digestion and absorption pathways were both upregulated and the
sphingolipid signaling and phosphatidylinositol signaling system
pathways were downregulated (Fig. 3C
and D).
Circ-G004213 promotes the cisplatin
sensitivity of HepG2 cells and improves survival in vivo
To identify the circRNAs that demonstrated
significant and consistent changes during TACE treatment, the
intersection of the circRNAs with significant changes (fold change
>10, P<0.01) were selected for prior TACE vs. TACE2 and TACE
vs. TACE2 (Fig. 4A). It was found
that circ-G004213 and circ_GABRG3 were upregulated during TACE
treatment (Fig. S2A and B).
Because the sensitivity of LC cells to cisplatin is significantly
associated with the effect of TACE treatment, the expression of
circ-G004213 and circ_GABRG3 was detected in HepG2 and
cisplatin-resistant HepG2/CIS cells. Notably, as shown in Fig. 4B, circ-G004213 was weakly expressed
in HepG2/CIS cells. In addition, circ-G004213 was downregulated by
CIS induction (Fig. 4C).
Overexpressing and knockout of circ-G004213 was constructed in
HepG2 cells (Fig. S3A). To further
determine the exact role of circ-G004213 in the sensitivity to CIS,
we measured the IC50 of CIS in HepG2 cells. As shown in
Fig. 4D, overexpression of
circ-G004213 enhanced the sensitivity of HepG2 cells to CIS. By
contrast, knockout of circ-G004213 increased the tolerance of HepG2
cells to CIS (Fig. 4D). CIS (10 µM)
was injected intraperitoneally; circ-G004213 overexpression reduced
tumor growth and circ-G004213 knockout increased tumor growth in
vivo (Fig. 4E). In addition,
the survival rate of each group was recorded. As shown in Fig. 4F, treatment with CIS prolonged the
survival rate of the circ-G004213-overexpressing cells, but a
shorter survival rate was observed in the circ-G004213 knockdown
cells compared with the control cells.
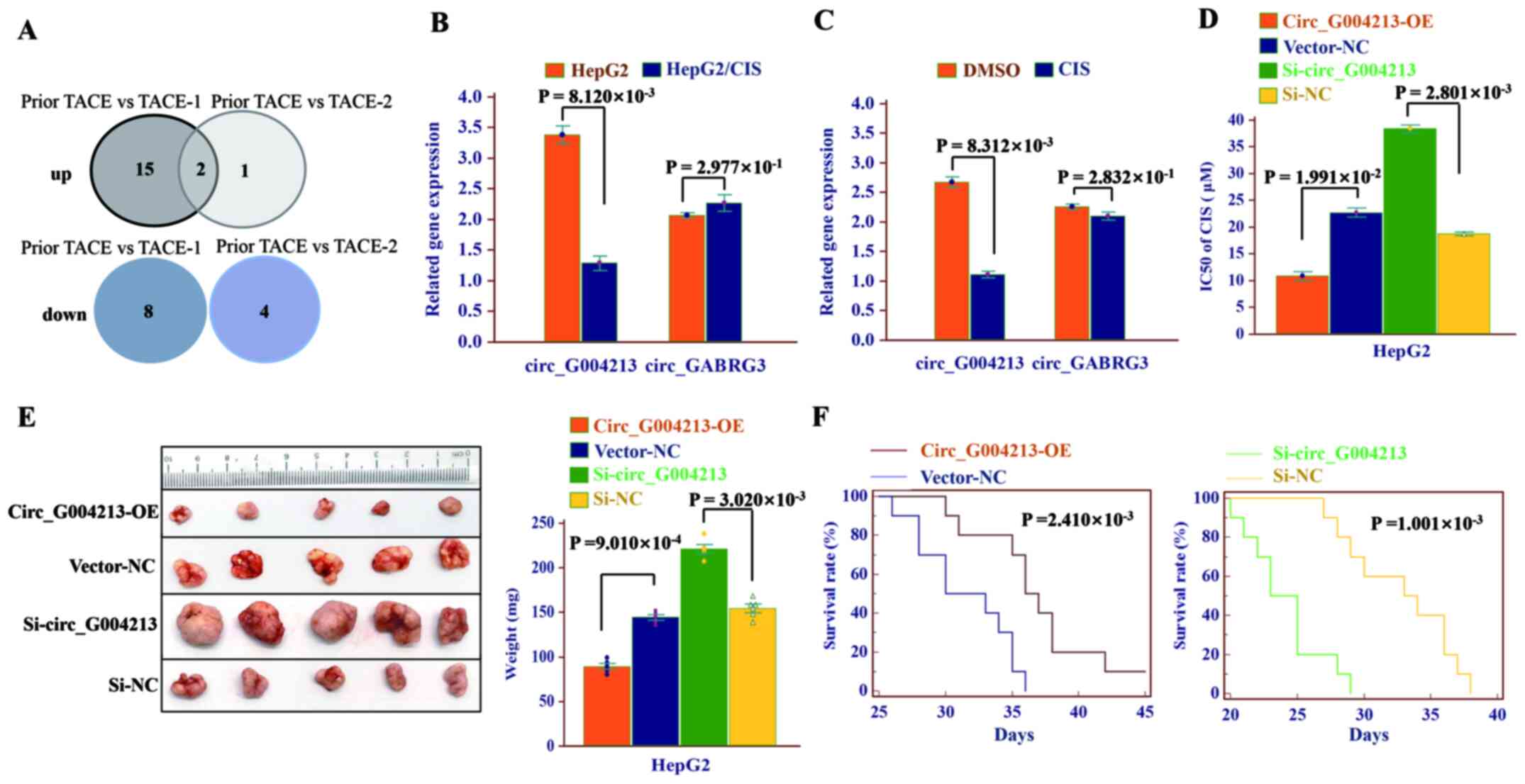 | Figure 4.Circ-G004213 promotes the cisplatin
sensitivity of HepG2 cells and improves survival in vivo. (A) A
Venn diagram demonstrated the intersection of the circRNAs with
significant changes (fold change >10; P<0.01) between prior
TACE vs. TACE2 and prior TACE vs. TACE2. (B) RT-qPCR detection of
circ-G004213 and circ_GABRG3 expression in the cisplatin-resistant
cell line HepG2/CIS compared with HepG2. (C) RT-qPCR detection of
circ-G004213 and circ_GABRG3 expression, when HepG2 cells were
treated with CIS (10 µM) for 3 days. (D) circ-G004213
overexpressing (circ-G004213-OE), negative control (Vector-NC),
circ-G004213 knockout (si-circ-G004213) and control (si-NC) HepG2
cells were treated with gradually increasing concentrations of CIS
for 3 days and the IC50 values of CIS were compared
among the groups. (E) Volume and weight of the tumor. (F) When mice
succumbed, the survival time of each group was recorded and the
Kaplan-Meier survival curves for each group were analyzed (n=10 per
group). circRNA, circular RNA; TACE, transarterial
chemoembolization; TACE1, after the first TACE operation; TACE2,
after the second TACE operation; CIS, cisplatin; RT-qPCR, reverse
transcription-quantitative PCR; si, short interfering. |
Circ-G004213 regulates CIS sensitivity
via regulation of miR-513b-5p/PRPR39
CircRNAs can act as miRNA sponges to regulate the
expression levels of other related RNAs via miRNA response
elements. miRNAs bind to mRNAs to inhibit post-transcriptional gene
expression. Based on the first five predicted miRNA targets of
circ-G004213 and the first five predicted mRNA targets of each
miRNA, the circRNA-miRNA-mRNA network was constructed (Fig. 5A). miR-513b-5p was significant and
continued to decrease following TACE, so it was selected for
further analysis (Fig. 5B).
Luciferase reporter analysis was used to determine whether
miR-513b-5p was directly targeted by circ-G004213. A double
luciferase reporter vector was constructed containing full-length
WT circ-G004213 or a version where the miR-513b-5p-binding site was
mutated (Fig. 5C). A significant
decrease in luciferase reporter activity was detected in the 293T
cells cotransfected with miR-513b-5p mimic and WT but not the
mutant vector (Fig. 5D).
Overexpressing and knockout of miR-513b-5p was constructed in HepG2
cells (Fig. S3B). In summary,
these experiments demonstrated that circ-G004213 can be used as a
sponge for miR-513b-5p and the miR-513b-5p inhibitor can improve
the sensitivity of HepG2 cells to CIS (Fig. 5E). By contrast, miR-513b-5p mimics
increased HepG2 tolerance to CIS (Fig.
5E).
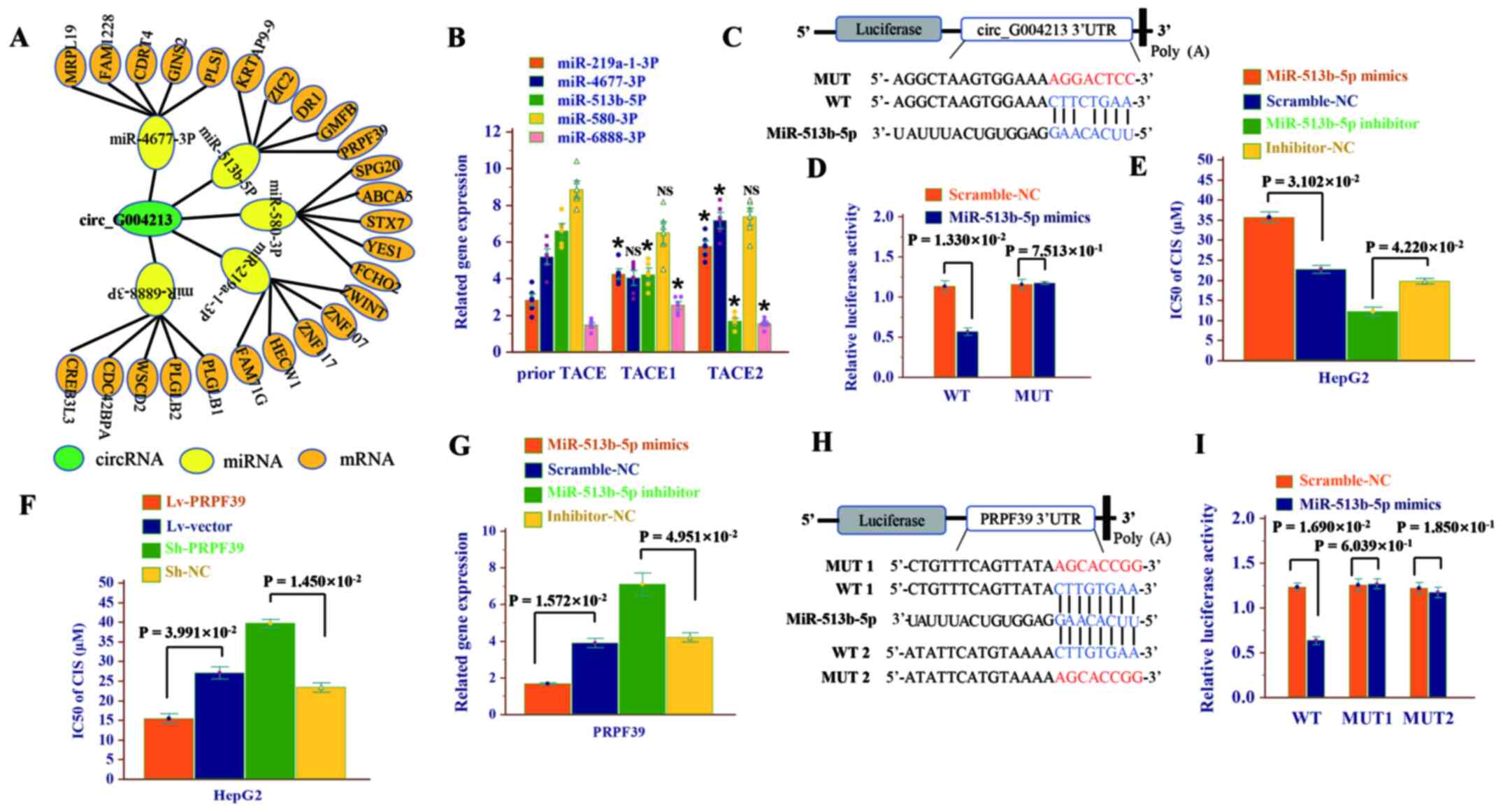 | Figure 5.Circ-G004213 regulates CIS
sensitivity via regulation miR-513b-5p/PRPR39. (A)
CircRNA-miRNA-mRNA network analysis. (B) The five miRNAs most
highly ranked as the candidate miRNAs during TACE treatment.
Triangles or circles represent samples in different groups,
*P<0.05 and NS vs. the corresponding samples of the prior TACE
group. (C) A schematic of WT and MUT circ-G004213 luciferase
reporter vectors. (D) The luciferase activity of WT circ-G004213
3′UTR or mutant circ_0025202 3′ UTR following transfection with
miR-513b-5P mimics in 293T cells. (E) HepG2 cells transfection with
miR-513b-5P mimics, mimics-NC, miR-513b-5P inhibitor and
inhibitor-NC were treated with gradually increasing concentrations
of CIS for three days and the IC50 values of CIS were
compared among the groups. (F) PRPF39 overexpressing (lv-PRPF39),
control (lv-vector), PRPF39 knockout (sh-PRPF39) and control
(sh-NC) HepG2 cells were treated with CIS and the IC50
values of CIS compared. (G) PRPF39 in miR-513b-5P mimics,
mimics-NC, miR-513b-5P inhibitor and inhibitor-NC groups. (H)
Potential binding sites of miR-513b-5P with PRPF39 3′UTR. (I) The
luciferase activity of WT PRPF39 3′UTR or mutant PRPF39 3′UTR of
two potential binding sites following transfection with miR-513b-5P
mimics in 293T cell lines. Data are presented as means ± SEM of at
least three independent experiments. CIS, cisplatin; miRNA,
microRNA; circRNA, circular RNA; NS, no significant difference; WT,
wild-type; MUT, mutant; UTR, untranslated region; sh, short
hairpin. |
PRPF39, as a predicted mRNA target of miR-513b-5p,
was significantly upregulated following TACE (Fig. S2C) and associated with cisplatin
resistance (15). The roles of
PRPF39 were determined in CIS sensitivity. Overexpressing and
knockout of PRPF39 was constructed in HepG2 cells (Fig. S3C). As shown in Fig. 5F, PRPF39 overexpression enhanced the
sensitivity of HepG2 cells to CIS. PRPF39 knockdown increased the
tolerance of HepG2 to CIS (Fig.
5F). The present study found that the miR-513b-5p inhibitor
increased PRPF39 expression, while the miR-513b-5p mimic decreased
PRPF39 expression (Fig. 5G).
According to MRE analysis, miR-513b-5p had two potential binding
sites in the 3′UTR of PRPF39 (Fig.
5H). Therefore, luciferase reporter plasmids containing the WT
and mutated 3′UTR of PRPF39 were designed to predict the binding
site. Following cotransfection of the miR-513b-5p mimic with the
PRPF39 3′UTR WT1 or PRPF39 3′UTR WT2, the luciferase activity was
significantly decreased. The mutant PRPF39 structure was unaffected
by miR-513b-5p (Fig. 5I).
The expression of miR-513b-5p and PRPF39 was
detected in HepG2 transplanted tumors. The present study
demonstrated that the expression of miR-513b-5p was decreased and
the expression of PRPF39 was increased in tumor tissues with
overexpression of circ-G004213 (Fig.
S3C and D). Meanwhile, the expression of miR-513b-5p in tumor
tissues was decreased and the expression of PRPF39 was increased in
tumor tissues with knockout of circ-G004213 (Fig. S3C and D). These results further
demonstrated that circ-G004213 regulated CIS sensitivity by
regulating miR-513b-5p/PRPR39.
Circ-G004213 is associated with the
prognosis of patients with LC following TACE
The circ-G004213 gene was significantly upregulated
following TACE treatment (Fig.
S2A), especially for case 1, 2 and 3, which demonstrated an
improved prognosis. To further verify the effect of circ-G004213 on
the prognosis of patients with LC, the clinicopathological
variables and circ-G004213 expression was analyzed in 50 patients
with LC following TACE. The results demonstrated that high
expression of circ-G004213 was negatively associated with advanced
TNM stage, BCLC stage or α-fetoprotein (AFP) and exhibited no
significant associations with other parameters, including age, sex,
HBV or Child-Pugh score (Table I).
Among the mRNAs that were negatively associated with the prognosis
of patients, the downstream mRNAs of circ-G004213, GINS2, ZIC2 and
CREB3L3 were significantly downregulated following TACE (Fig. S2D) and the mRNAs positively
associated with patient prognosis, PRPF39 and ZWINT, were
significantly upregulated following TACE (Fig. S2C). The above results suggested
that the expression of circ-G004213 in exosomes may be positively
associated with the prognosis of patients with LC following
TACE.
Discussion
Exosomes serve a crucial role in the carcinogenesis
of LC. In particular, small noncoding RNAs in exosomes could serve
as biomarkers for the prediction and diagnosis of LC (27). Exosomes from LC cells can promote
the occurrence and development of LC. For example, Exosomes derived
from HepG2 cell can reprogram biological behaviors of LO2 cells
(28) and exosomes derived from LC
patient serum and HepG2 cells can inhibit proliferation and
sorafenib chemoresistance in LC (29).
Recently, an increasing number of studies have
demonstrated that exosomal circRNAs serve a crucial role in the
prediction and diagnosis of LC (10–13).
Exosomal circPTGR1 from cells with high metastatic (LM3) can
promote LC metastasis via the miR449a-MET pathway (17). Although TACE is an important
treatment for LC, the alteration and effects of circRNAs in LC
following TACE remain to be elucidated. The alterations of the
expression profile of circRNAs in exosomes from patients with LC
following TACE were studied for the first time, to the best of the
authors' knowledge, by high-throughput whole transcriptome
sequencing and subsequent bioinformatic analysis in the present
study.
The present study identified 390 differentially
expressed circRNAs between the prior TACE and TACE1 groups and 489
differentially expressed circRNAs between the prior TACE and TACE2
groups. Based on these different circRNAs, GO analysis was
performed to annotate and determine the function of these circRNAs.
The present study demonstrated that they were associated with fatty
acid metabolism, receptor binding and membrane protein complexes.
The medium-chain fatty acid metabolic process is associated with
the metabolism, metastasis and prognosis of LC (30,31).
The insulin receptor is associated with LC cell viability and
proliferation (32). The nuclear
membrane protein complex promotes the early recurrence of
hepatocellular carcinoma in association with the Wnt/β-catenin
signaling pathway (33). KEGG
pathway analysis predicted that protein digestion and absorption
pathways were activated following TACE. This pathway is associated
with the anti-LC efficacy of DHA in combination with sorafenib
(34).
To identify the circRNAs that demonstrated
significant and consistent changes during TACE treatment, the
intersection of the circRNAs with significant changes for prior
TACE vs. TACE2 and TACE vs. TACE2 were selected. It was found that
circ-G004213 and circ_GABRG3 were upregulated during TACE
treatment. Cisplatin is a common chemotherapeutic agent in TACE.
Therefore, the sensitivity of cisplatin chemotherapy is an
important factor affecting TACE treatment. The expression of
circ-G004213 and circ_GABRG3 was detected in HepG2 and
cisplatin-resistant HepG2/CIS cells. The present study demonstrated
that circ-G004213 was expressed at low levels in HepG2/CIS cells
and downregulated in response to CIS induction. However, the role
and mechanism of circ_GABRG3 in TACE require further study. Further
analysis confirmed that circ-G004213 promoted the cisplatin
sensitivity of HepG2 cells and improved survival in
vivo.
Bioinformatic analysis has shown that some circRNAs
can act as miRNA sponges to regulate the expression levels of other
related RNAs via miRNA response elements. This mechanism is called
competitive endogenous RNA mechanism. miRNAs bind to mRNAs to
inhibit post-transcriptional gene expression and serve an important
role in regulating gene expression, the cell cycle and organismal
development (35). Therefore, it is
important to identify the interactions of circRNAs, miRNAs and
mRNAs. Based on the first five predicted miRNA targets and the
first five predicted mRNA targets of each miRNA, the
circRNA-miRNA-mRNA network was constructed. Among the predicted
miRNAs, miR-513b-5p was confirmed to be a candidate miRNA.
miR-513b-5p is associated with cell proliferation and migration in
ovarian cancer (20). Furthermore,
the present study confirmed that PRPF39, as a mRNA target of
miR-513b-5p, was significantly upregulated following TACE.
Knockdown of PRPF39 expression using siRNA resulted in a
significant increase in cisplatin resistance (21). Therefore, circ-G004213 regulates CIS
sensitivity via miR-513b-5p/PRPR39.
Studies demonstrate that exosomes from HCC cells can
promote the occurrence and development of LC (28,29).
However, the present study used HepG2 cells for the purpose of
elucidating the role of circ-G004213 in cisplatin chemotherapy and
further proved the interaction of circ-G004213/miR-513b-5p/PRPF39.
Therefore, it did not isolate exosomes from HepG2 cells for further
study. However, in a future study, will further analyze the role of
exosomal circRNAs source of HepG2 cells on cisplatin
chemotherapy.
The present study demonstrated that high expression
of circ-G004213 was negatively associated with advanced TNM stage,
BCLC stage or AFP and exhibited no significant associations with
other parameters, including age, sex, HBV or Child-Pugh score. TNM
stage, BCLC stage and AFP predict poor prognosis in patients with
LC (36–38). It was found that potential
downstream mRNAs of circ-G004213, GINS2, ZIC2 and CREB3L3 were
significantly downregulated and PRPF39 and ZWINT were significantly
upregulated following TACE. The upregulation and interrelated
expression of GINS subunits predicts poor prognosis in patients
with LC (39). ZIC2 could promote
tumor growth and metastasis via PAK4 in LC (40). ZIC2-dependent octamer-binding
transcription factor 4 activation drives the self-renewal of human
LC stem cells (41). CREBH, which
is activated by stress in the endoplasmic reticulum (ER), is an
ER-resident transmembrane basic leucine zipper (bZIP) transcription
factor that is specifically expressed in the liver and activated
CREBH may serve an important role in LC proliferation (42). Therefore, GINS2, ZIC2 and CREB3L3
were negatively associated with the prognosis of LC. Knockdown of
PRPF39 expression using siRNA results in a significant increase in
cisplatin resistance (21) and
cisplatin is the main chemotherapy drug used for TACE.
Overexpression of ZWINT predicts poor prognosis and promotes the
proliferation of LC by regulating cell cycle-related proteins
(43). Therefore, PRPF39 and ZWINT
were positively associated with the prognosis of LC and
circ-G004213 was positively associated with the prognosis of
LC.
In conclusion, the above results suggested that the
expression of circ-G004213 in exosomes could promote CIS
sensitivity via regulation of miR-513b-5p/PRPR39 and circ-G004213
was positively associated with the prognosis of patients with LC
following TACE. Circ-G004213 may be an indicator for predicting the
efficacy of TACE in patients with LC.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was supported by grants from the
National Natural Science Foundation of China (grant no. 81702446),
Department education of Sichuan Province (grant nos. 18ZA0151 and
18ZB0175), Department of Science and Technology of Sichuan Province
(grant no. C2018JY0654), Fund of Chengdu Medical College (grant no.
CDYXY007) and Special Research Fund of the First Affiliated
Hospital of Chengdu Medical College (grant no. CYFY2017YB09).
Availability of data and materials
The datasets generated and/or analyzed during the
current study are available in the Gene Expression Omnibus
repository at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE165183.
Authors' contributions
LQ was mainly involved in patient recruitment, TACE
treatment and clinical sample collection. ZZ was involved in
clinical data collection and analysis. CW performed the animal
experiment. XL was involved in clinic sample storage and
processing. TZ was involved in sequencing data analysis and drug
resistance experiment. JL mainly contributed to designing and
performing the present study and manuscript editing. LQ and JL were
responsible for the authenticity of the data in this article. All
authors reviewed and approved the final manuscript.
Ethics approval and consent to
participate
Informed consent was signed in writing by all
patients and the study protocol was approved by the ethics
committee of the First Affiliated Hospital of Chengdu Medical
College (approval no. 2017009). The animal experiments were
approved by the research ethics committee of Chengdu Medical
College (approval no. 2017CYA-021).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
LC
|
liver cancer
|
|
TACE
|
transarterial chemoembolization
|
|
circ-G004213
|
hsa-circRNA-G004213
|
|
mRNA
|
messenger RNA
|
|
miRNA
|
microRNA
|
|
lncRNA
|
long noncoding RNA
|
|
circRNA
|
circular RNA
|
|
prior TACE
|
prior to TACE treatment
|
|
TACE1
|
after the first TACE operation
|
|
TACE2
|
after the second TACE operation
|
|
TEM
|
transmission electron microscope
|
|
MNNA
|
Malvern Nano-Nano Analyzer
|
|
CIS
|
cisplatin
|
|
RT-qPCR
|
reverse transcription-quantitative
PCR
|
|
HBV
|
hepatitis virus B
|
|
AFP
|
alpha-fetoprotein
|
|
GO
|
Gene Ontology
|
|
BP
|
biological process
|
|
CC
|
cell component
|
|
MF
|
molecular function
|
|
KEGG
|
Kyoto Encyclopedia of Genes and
Genomes
|
|
NC
|
negative control
|
|
OE
|
overexpression
|
References
|
1
|
Lingiah VA, Niazi M, Olivo R, Paterno F,
Guarrera JV and Pyrsopoulos NT: Liver transplantation beyond milan
criteria. J Clin Transl Hepatol. 8:69–75. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Wu CC, Ho WL and Liu TJ: Hepatocellular
carcinoma with adjacent organ extension: The enhancement of
preoperative transcatheter arterial embolization and the results of
surgical resection. Surg Today. 24:882–888. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Grandhi MS, Kim AK, Ronnekleiv-Kelly SM,
Kamel IR, Ghasebeh MA and Pawlik TM: Hepatocellular carcinoma: From
diagnosis to treatment. Surg Oncol. 25:74–85. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Gu J, Zhang X, Cui R, Zhang J, Wang Z, Jia
Y, Miao R, Dong Y, Ma X, Fan H, et al: Prognostic predictors for
patients with hepatocellular carcinoma receiving adjuvant
transcatheter arterial chemoembolization. Eur J Gastroenterol
Hepatol. 31:836–844. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Loosen SH, Schulze-Hagen M, Leyh C, Benz
F, Vucur M, Kuhl C, Trautwein C, Tacke F, Bruners P, Roderburg C
and Luedde T: IL-6 and IL-8 serum levels predict tumor response and
overall survival after TACE for primary and secondary hepatic
malignancies. Int J Mol Sci. 19:17662018. View Article : Google Scholar
|
|
6
|
Daniele A, Divella R, Quaranta M, Mattioli
V, Casamassima P, Paradiso A, Garrisi VM, Gadaleta CD,
Gadaleta-Caldarola G, Savino E, et al: Clinical and prognostic role
of circulating MMP-2 and its inhibitor TIMP-2 in HCC patients prior
to and after trans-hepatic arterial chemo-embolization. Clin
Biochem. 47:184–190. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhou L, Lv T, Zhang Q, Zhu Q, Zhan P, Zhu
S, Zhang J and Song Y: The biology, function and clinical
implications of exosomes in lung cancer. Cancer Lett. 407:84–92.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Shi X, Wang B, Feng X, Xu Y, Lu K and Sun
M: circRNAs and exosomes: A mysterious frontier for human cancer.
Mol Ther Nucleic Acids. 19:384–392. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wan M, Ning B, Spiegel S, Lyon CJ and Hu
TY: Tumor-derived exosomes (TDEs): How to avoid the sting in the
tail? Med Res Rev. 40:385–412. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Han Q, Zhao H, Jiang Y, Yin C and Zhang J:
HCC-derived exosomes: Critical player and target for cancer immune
escape. Cells. 8:5582019. View Article : Google Scholar
|
|
11
|
Abudoureyimu M, Zhou H, Zhi Y, Wang T,
Feng B, Wang R and Chu X: Recent progress in the emerging role of
exosome in hepatocellular carcinoma. Cell Prolif. 52:e125412019.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Sarnow P and Sagan SM: Unraveling the
mysterious interactions between hepatitis C virus RNA and
liver-specific microRNA-122. Annu Rev Virol. 3:309–332. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Li B, Mao R, Liu C, Zhang W, Tang Y and
Guo Z: LncRNA FAL1 promotes cell proliferation and migration by
acting as a CeRNA of miR-1236 in hepatocellular carcinoma cells.
Life Sci. 197:122–129. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Liu J, Fan L, Yu H, Zhang J, He Y, Feng D,
Wang F, Li X, Liu Q, Li Y, et al: Endoplasmic reticulum stress
causes liver cancer cells to release exosomal miR-23a-3p and
up-regulate programmed death ligand 1 expression in macrophages.
Hepatology. 70:241–258. 2019.PubMed/NCBI
|
|
15
|
Huang XY, Huang ZL, Huang J, Xu B, Huang
XY, Xu YH, Zhou J and Tang ZY: Exosomal circRNA-100338 promotes
hepatocellular carcinoma metastasis via enhancing invasiveness and
angiogenesis. J Exp Clin Cancer Res. 39:202020. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Su Y, Lv X, Yin W, Zhou L, Hu Y, Zhou A
and Qi F: CircRNA Cdr1as functions as a competitive endogenous RNA
to promote hepatocellular carcinoma progression. Aging (Albany NY).
11:8182–8203. 2019.
|
|
17
|
Wang G, Liu W, Zou Y, Wang G, Deng Y, Luo
J, Zhang Y, Li H, Zhang Q, Yang Y and Chen G: Three isoforms of
exosomal circPTGR1 promote hepatocellular carcinoma metastasis via
the miR449a-MET pathway. EBioMedicine. 40:432–445. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhang H, Deng T, Ge S, Liu Y, Bai M, Zhu
K, Fan Q, Li J, Ning T, Tian F, et al: Exosome circRNA secreted
from adipocytes promotes the growth of hepatocellular carcinoma by
targeting deubiquitination-related USP7. Oncogene. 38:2844–2859.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wang X, Zhang X, Wang G, Wang L, Lin Y and
Sun F: Hsa-miR-513b-5p suppresses cell proliferation and promotes
P53 expression by targeting IRF2 in testicular embryonal carcinoma
cells. Gene. 626:344–353. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lin W, Ye H, You K and Chen L:
Up-regulation of circ_LARP4 suppresses cell proliferation and
migration in ovarian cancer by regulating miR-513b-5p/LARP4 axis.
Cancer Cell Int. 20:52020. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Stark AL, Delaney SM, Wheeler HE, Im HK
and Dolan ME: Functional consequences of PRPF39 on distant genes
and cisplatin sensitivity. Hum Mol Genet. 21:4348–4355. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
National Health and Family Planning
Commission, . Notice of general office of national health and
family planning commission on printing and issuing standards for
diagnosis and treatment of primary liver cancer (2017 edition).
http://www.nhfpc.gov.cn/yzygj/s7659/201706/80abf02a86c048fcd130e5e298f7aeee.shtmlJune
2–2017
|
|
23
|
Kim KM, Shim SG, Sinn DH, Song JE, Kim BS
and Kim HG: Child-Pugh, MELD, MELD-Na, and ALBI scores: Which liver
function models best predicts prognosis for HCC patient with
ascites? Scand J Gastroenterol. 55:951–957. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Li K, Gao B, Li J, Chen H, Li Y, Wei Y,
Gong D, Gao J, Zhang J, Tan W, et al: ZNF32 protects against
oxidative stress-induced apoptosis by modulation C1QBP
transcription. Oncotarget. 6:38107–38126. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li J, Ao J, Li K, Zhang J, Li Y, Zhang L,
Wei Y, Gong D, Gao J, Tan W, et al: ZNF32 contributes to the
induction of multidrug resistance by regulating TGF-β receptor 2
signaling in lung adenocarcinoma. Cell Death Dis. 7:e24282016.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wang H, Lu Z and Zhao X: Tumorigenesis,
diagnosis, and therapeutic potential of exosomes in liver cancer. J
Hematol Oncol. 12:1332019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
He X, Yu J, Xiong L, Liu Y, Fan L, Li Y,
Chen B, Chen J and Xu X: Exosomes derived from liver cancer cells
reprogram biological behaviors of LO2 cells by transferring
Linc-ROR. Gene. 719:1440442019. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wang G, Zhao W, Wang H, Qiu G, Jiang Z,
Wei G and Li X: Exosomal miR-744 inhibits proliferation and
sorafenib chemoresistance in hepatocellular carcinoma by targeting
PAX2. Med Sci Monit. 25:7209–7217. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Wakana H, Kono H, Fukushima H, Nakata Y,
Akazawa Y, Maruyama S, Hagio K, Fujii H and Ichikawa D: Effects of
medium-chain triglycerides administration in chemically-induced
carcinogenesis in mice. Anticancer Res. 39:6653–6660. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wang H, Lu J, Dolezal J, Kulkarni S, Zhang
W, Chen A, Gorka J, Mandel JA and Prochownik EV: Inhibition of
hepatocellular carcinoma by metabolic normalization. PLoS One.
14:e02181862019. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Sakurai Y, Kubota N, Takamoto I, Obata A,
Iwamoto M, Hayashi T, Aihara M, Kubota T, Nishihara H and Kadowaki
T: Role of insulin receptor substrates in the progression of
hepatocellular carcinoma. Sci Rep. 7:53872017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Chen J, Rajasekaran M, Xia H, Zhang X,
Kong SN, Sekar K, Seshachalam VP, Deivasigamani A, Goh BK, Ooi LL,
et al: The microtubule-associated protein PRC1 promotes early
recurrence of hepatocellular carcinoma in association with the
Wnt/β-catenin signalling pathway. Gut. 65:1522–1534. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Hou C, Guo D, Yu X, Wang S and Liu T:
TMT-based proteomics analysis of the anti-hepatocellular carcinoma
effect of combined dihydroartemisinin and sorafenib. Biomed
Pharmacother. 126:1098622020. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Cao D, Wang Y, Li D and Wang L:
Reconstruction and analysis of the differentially expressed
IncRNA-miRNA-mRNA network based on competitive endogenous RNA in
hepatocellular carcinoma. Crit Rev Eukaryot Gene Expr. 29:539–549.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhou ZR, Liu M, Lu HR, Li YF, Liang SX and
Zhang CY: Validation of different staging systems for
hepatocellular carcinoma in a cohort of 249 patients undergoing
radiotherapy. Oncotarget. 8:46523–46531. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
de Freitas LB, Longo L, Santos D,
Grivicich I and Álvares-da-Silva MR: Hepatocellular carcinoma
staging systems: Hong Kong liver cancer vs Barcelona clinic liver
cancer in a Western population. World J Hepatol. 11:678–688. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Bird TG, Dimitropoulou P, Turner RM, Jenks
SJ, Cusack P, Hey S, Blunsum A, Kelly S, Sturgeon C, Hayes PC and
Bird SM: Alpha-fetoprotein detection of hepatocellular carcinoma
leads to a standardized analysis of dynamic AFP to improve
screening based detection. PLoS One. 11:e01568012016. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Lian YF, Li SS, Huang YL, Wei H, Chen DM,
Wang JL and Huang YH: Up-regulated and interrelated expressions of
GINS subunits predict poor prognosis in hepatocellular carcinoma.
Biosci Rep. 38:BSR201811782018. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lu SX, Zhang CZ, Luo RZ, Wang CH, Liu LL,
Fu J, Zhang L, Wang H, Xie D and Yun JP: Zic2 promotes tumor growth
and metastasis via PAK4 in hepatocellular carcinoma. Cancer Lett.
402:71–80. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Zhu P, Wang Y, He L, Huang G, Du Y, Zhang
G, Yan X, Xia P, Ye B, Wang S, et al: ZIC2-dependent OCT4
activation drives self-renewal of human liver cancer stem cells. J
Clin Invest. 125:3795–3808. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Cho HK, Kim SY, Kyaw YY, Win AA, Koo SH,
Kim HH and Cheong J: HBx induces the proliferation of
hepatocellular carcinoma cells via AP1 over-expressed as a result
of ER stress. Biochem J. 466:115–121. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Ying H, Xu Z, Chen M, Zhou S, Liang X and
Cai X: Overexpression of Zwint predicts poor prognosis and promotes
the proliferation of hepatocellular carcinoma by regulating
cell-cycle-related proteins. Onco Targets Ther. 11:689–702. 2018.
View Article : Google Scholar : PubMed/NCBI
|